Submitted:
28 March 2025
Posted:
31 March 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Method
2.1. Data Acquisition
2.2. Data Preprocessing
2.3. Phenotype Simulation
2.4. Genome-Wide Association Study (GWAS)
2.5. Performance Metric Calculation
2.6. Visualization
2.7. Software and Code Availability
3. Results
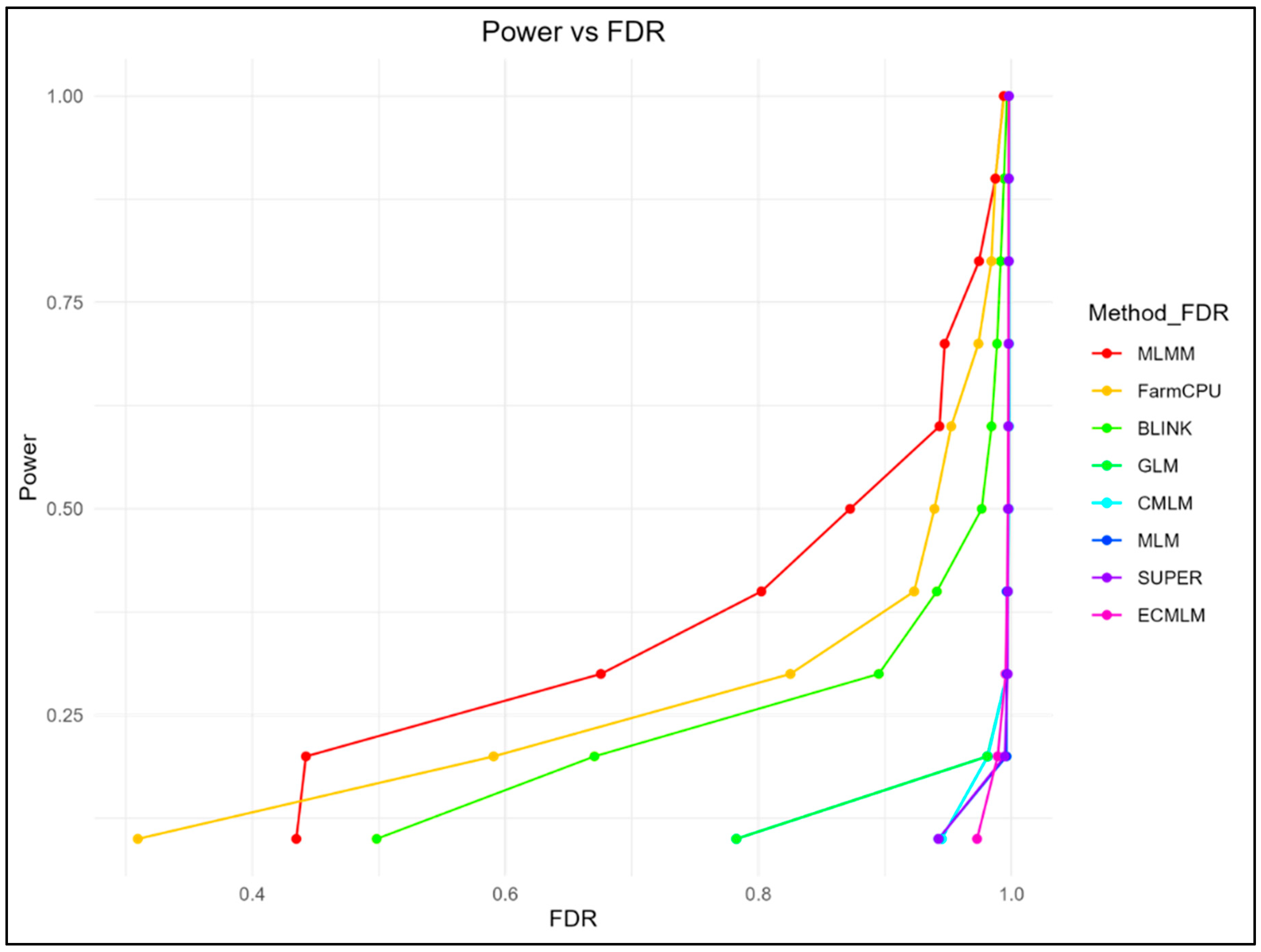
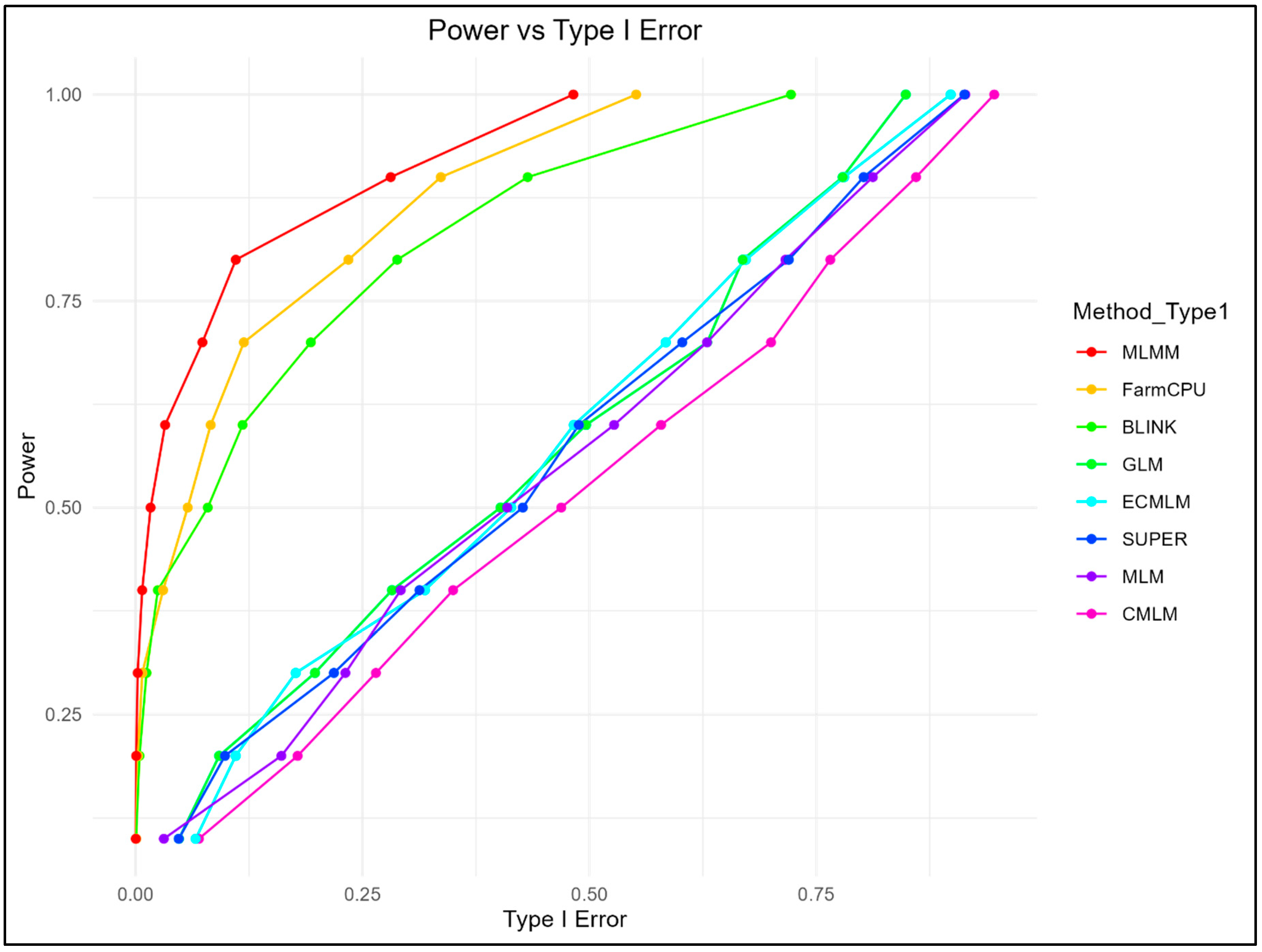
3.1. Power, FDR, and Type I Error
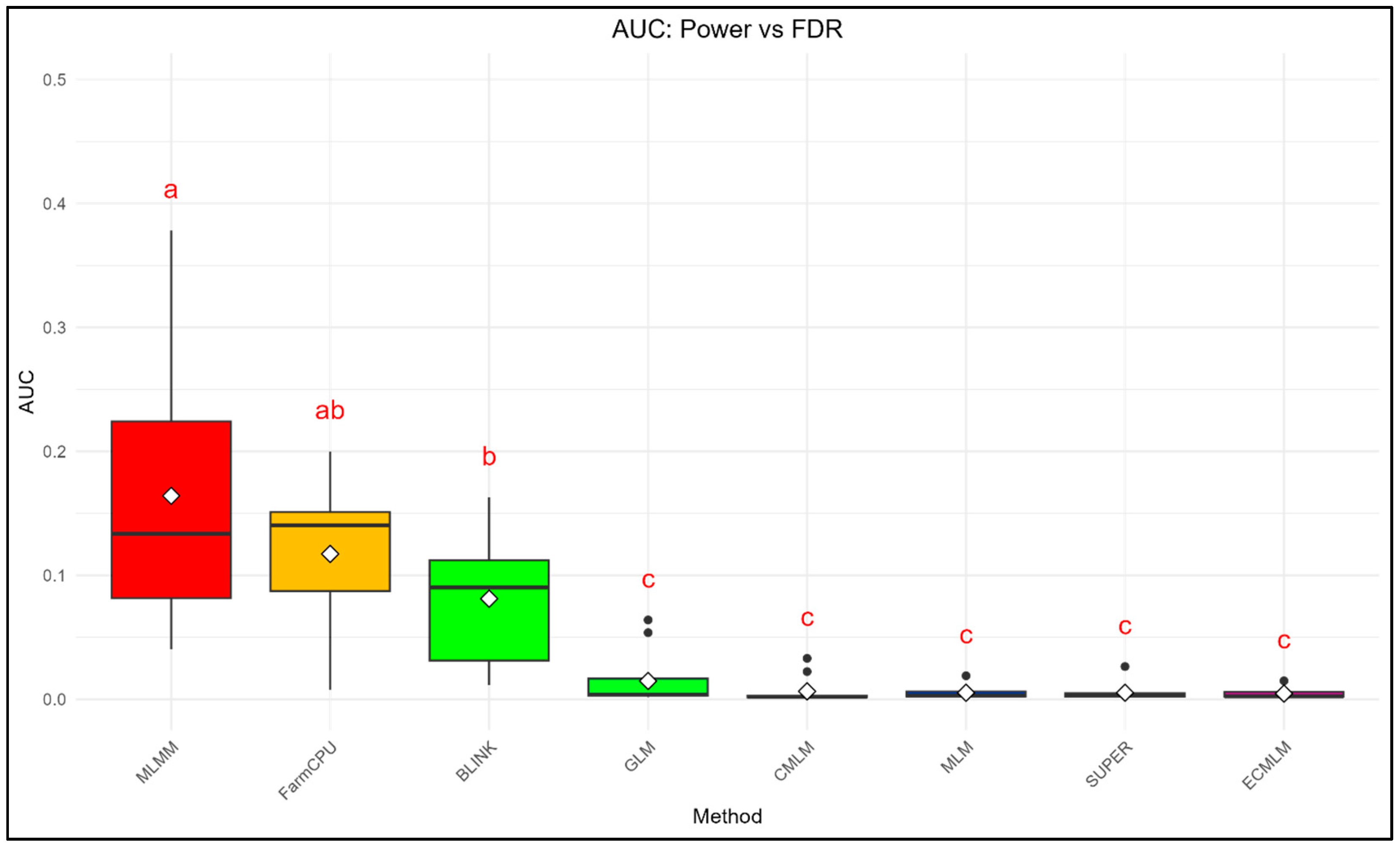
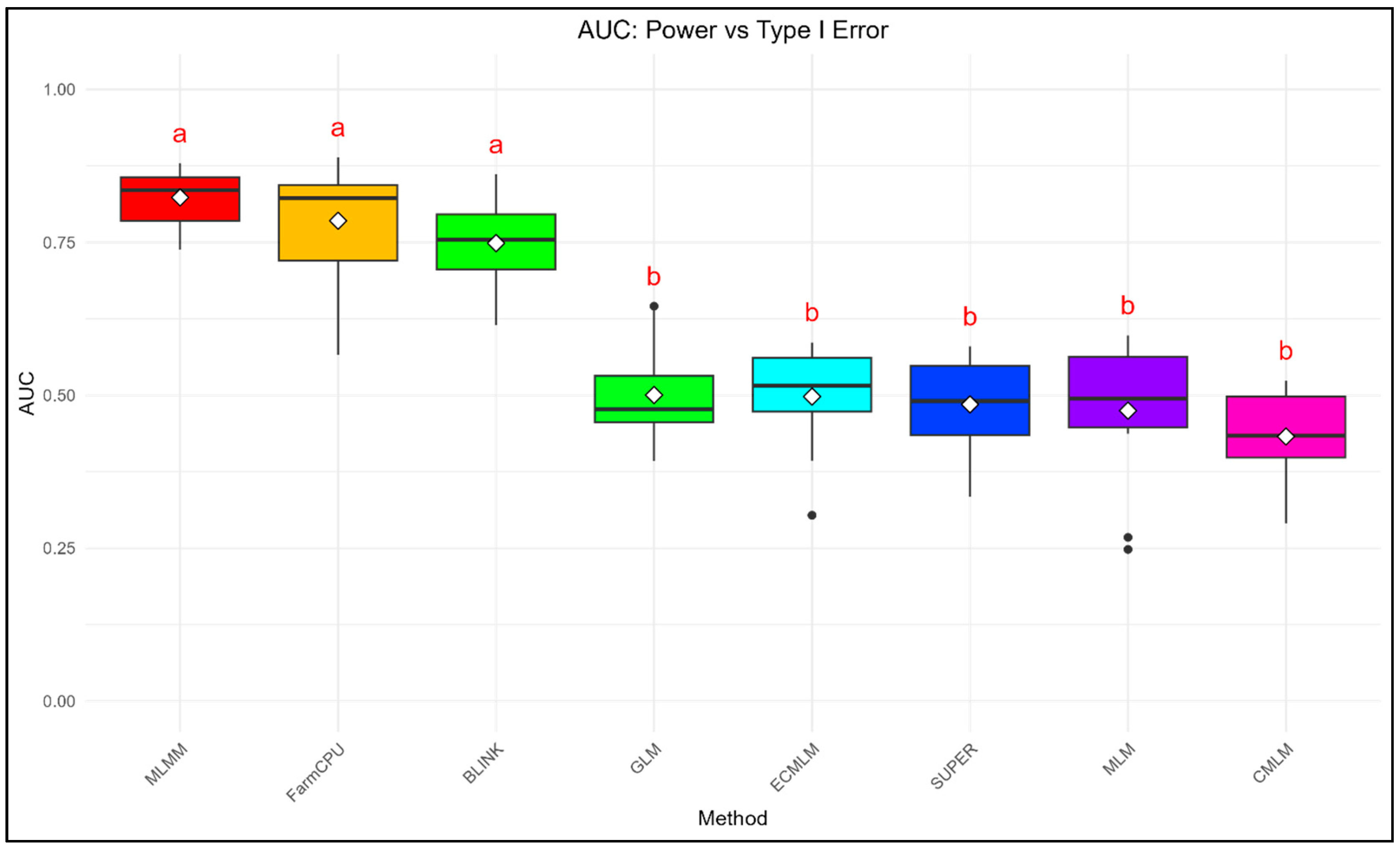
3.2. AUC Analysis
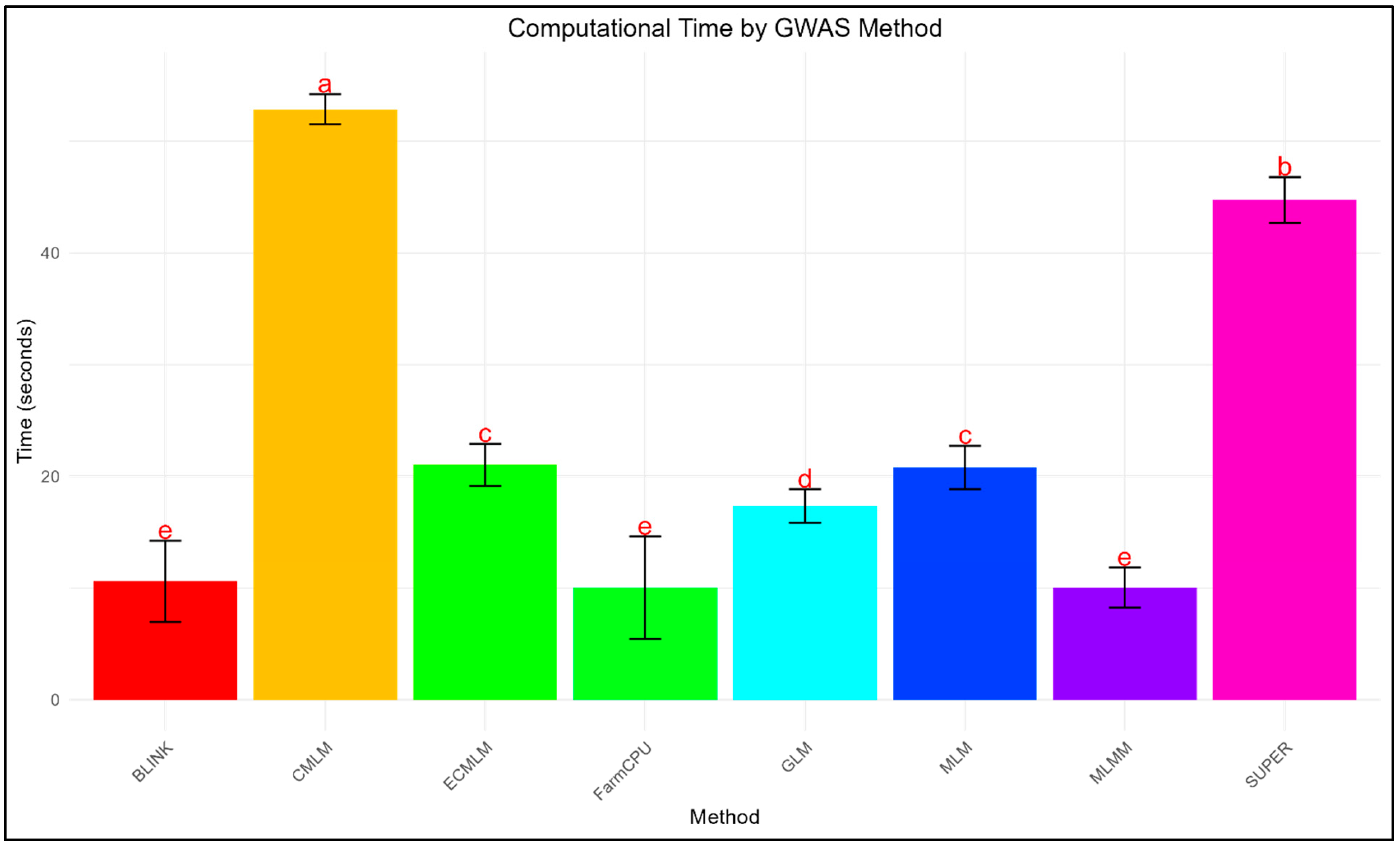
3.3. Computational Efficiency
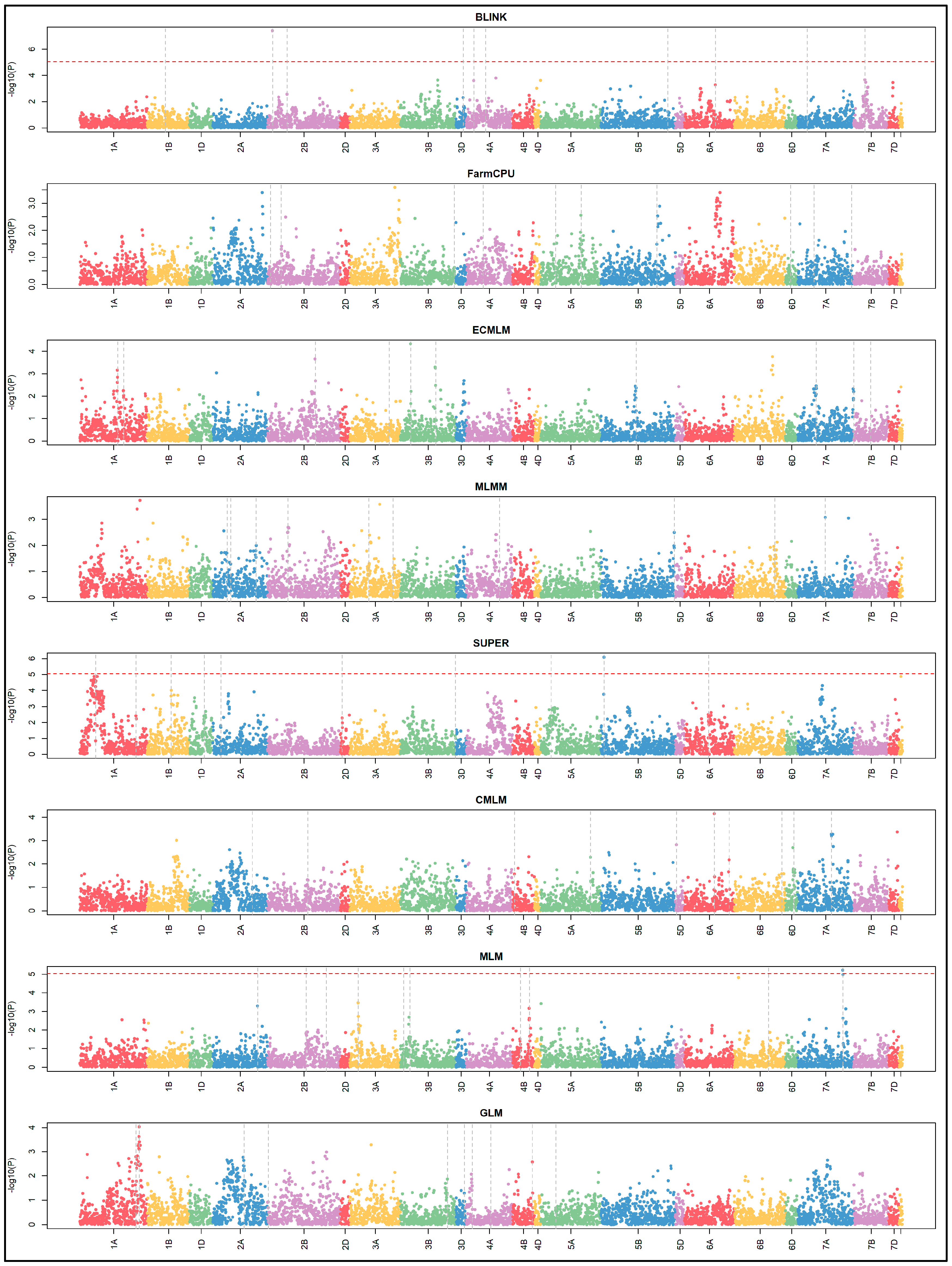
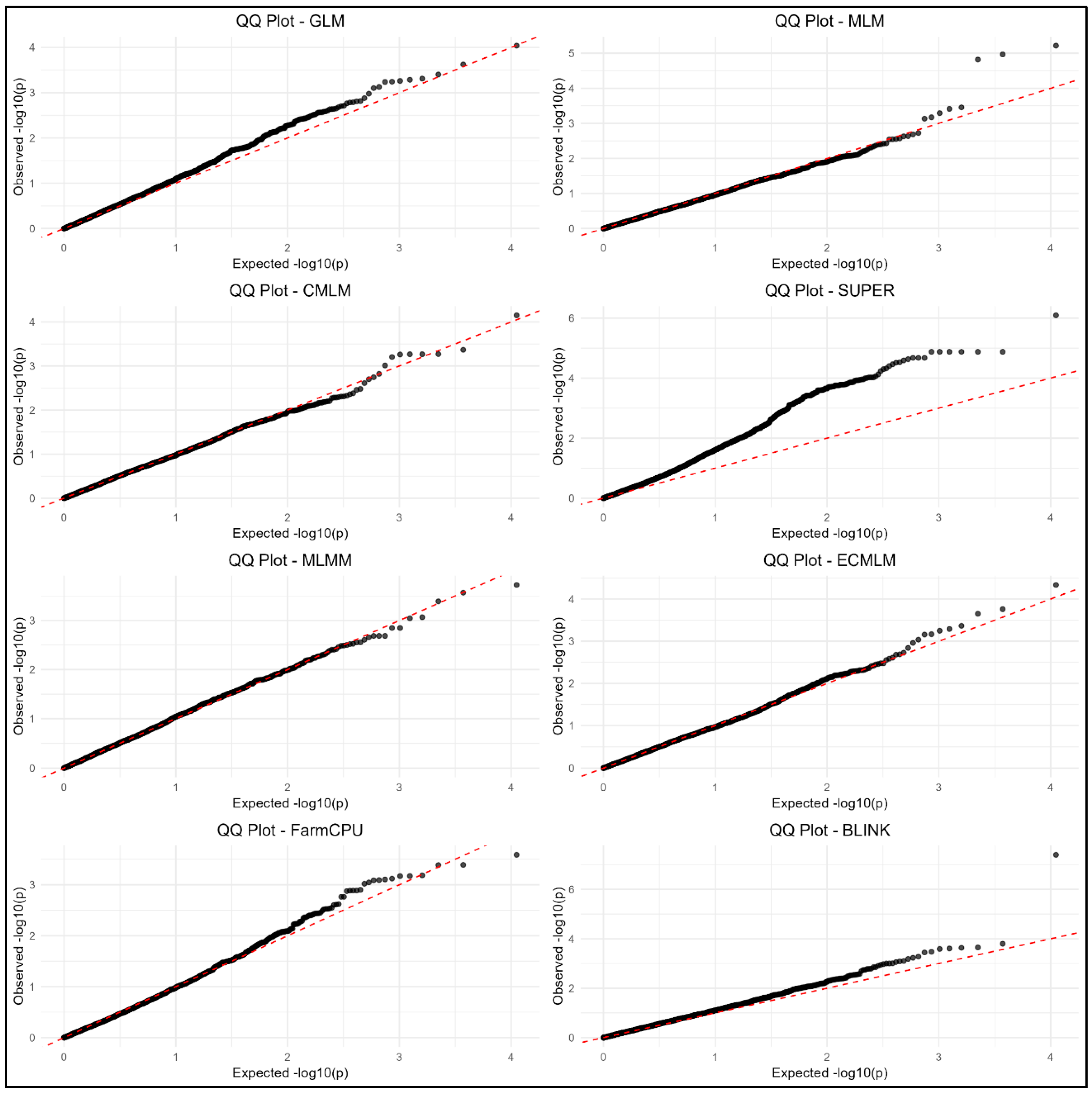
3.4. Manhattan and QQ Plots
4. Discussion
5. Conclusions
6. Acknowledgement
References
- Cebeci, Z., Bayraktar, M., & Gökçe, G. (2023). Comparison of the statistical methods for genome-wide association studies on simulated quantitative traits of domesticated goats (Capra hircus L.). Small Ruminant Research, 227, 107053. [CrossRef]
- da Silva Linge, C., Cai, L., Fu, W., Clark, J., Worthington, M., Rawandoozi, Z., ... & Gasic, K. (2022). Corrigendum: Multi-Locus Genome-Wide Association Studies Reveal Fruit Quality Hotspots in Peach Genome. Frontiers in Plant Science, 13, 879112. [CrossRef]
- Huang, M., Liu, X., Zhou, Y., Summers, R. M., & Zhang, Z. (2019). BLINK: a package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience, 8(2), giy154. [CrossRef]
- Kaler, A. S., Gillman, J. D., Beissinger, T., & Purcell, L. C. (2020). Comparing different statistical models and multiple testing corrections for association mapping in soybean and maize. Frontiers in plant science, 10, 1794. [CrossRef]
- Lipka, A. E., Tian, F., Wang, Q., Peiffer, J., Li, M., Bradbury, P. J., ... & Zhang, Z. (2012). GAPIT: genome association and prediction integrated tool. Bioinformatics, 28(18), 2397-2399. [CrossRef]
- Li, M., Zhang, Y. W., Zhang, Z. C., Xiang, Y., Liu, M. H., Zhou, Y. H., ... & Zhang, Y. M. (2022). A compressed variance component mixed model for detecting QTNs and QTN-by-environment and QTN-by-QTN interactions in genome-wide association studies. Molecular Plant, 15(4), 630-650. [CrossRef]
- Liu, X., Huang, M., Fan, B., Buckler, E. S., & Zhang, Z. (2016). Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS genetics, 12(2), e1005767. [CrossRef]
- Nicolas Morales, Jean-Luc Jannink, Clay L. Birkett, David J Waring, Lukas A Mueller, et. al. (2022) Breedbase: a digital ecosystem for modern plant breeding, G3 Genes|Genomes|Genetics,12(7), jkac078, . [CrossRef]
- Price, A. L., Patterson, N. J., Plenge, R. M., Weinblatt, M. E., Shadick, N. A., & Reich, D. (2006). Principal components analysis corrects for stratification in genome-wide association studies. Nature genetics, 38(8), 904-909. [CrossRef]
- R Core Team. R: A language and environment for statistical computing.(2023) R foundation for statistical computing, Vienna, Austria. https://www.R-project.org/.
- Sandhu, K. S., Burke, A. B., Merrick, L. F., Pumphrey, M. O., & Carter, A. H. (2024). Comparing performances of different statistical models and multiple threshold methods in a nested association mapping population of wheat. Frontiers in Plant Science, 15, 1460353. [CrossRef]
- Segura, V., Vilhjálmsson, B. J., Platt, A., Korte, A., Seren, Ü., Long, Q., & Nordborg, M. (2012). An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nature genetics, 44(7), 825-830. [CrossRef]
- Tam, V., et al. (2019). Benefits and limitations of genome-wide association studies. Nature Reviews Genetics, 20(8), 467-484. [CrossRef]
- Visscher, P. M., et al. (2017). 10 Years of GWAS Discovery: Biology, Function, and Translation. American Journal of Human Genetics, 101(1), 5-22. [CrossRef]
- Wang, Q., Tian, F., Pan, Y., Buckler, E. S., & Zhang, Z. (2014). A SUPER powerful method for genome wide association study. PloS one, 9(9), e107684. [CrossRef]
- Wen, Y. J., Zhang, H., Zhang, J., Feng, J. Y., Huang, B., Dunwell, J. M., ... & Wu, R. (2016). A fast multi-locus random-SNP-effect EMMA for genome-wide association studies. bioRxiv, 077404. [CrossRef]
- Yu, J., Pressoir, G., Briggs, W. H., Vroh Bi, I., Yamasaki, M., Doebley, J. F., ... & Buckler, E. S. (2006). A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nature genetics, 38(2), 203-208. [CrossRef]
- Zhang, Z., Ersoz, E., Lai, C. Q., Todhunter, R. J., Tiwari, H. K., Gore, M. A., ... & Buckler, E. S. (2010). Mixed linear model approach adapted for genome-wide association studies. Nature genetics, 42(4), 355-360. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



