Submitted:
25 March 2025
Posted:
26 March 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. CRISPR/Cas9 Delivery Systems
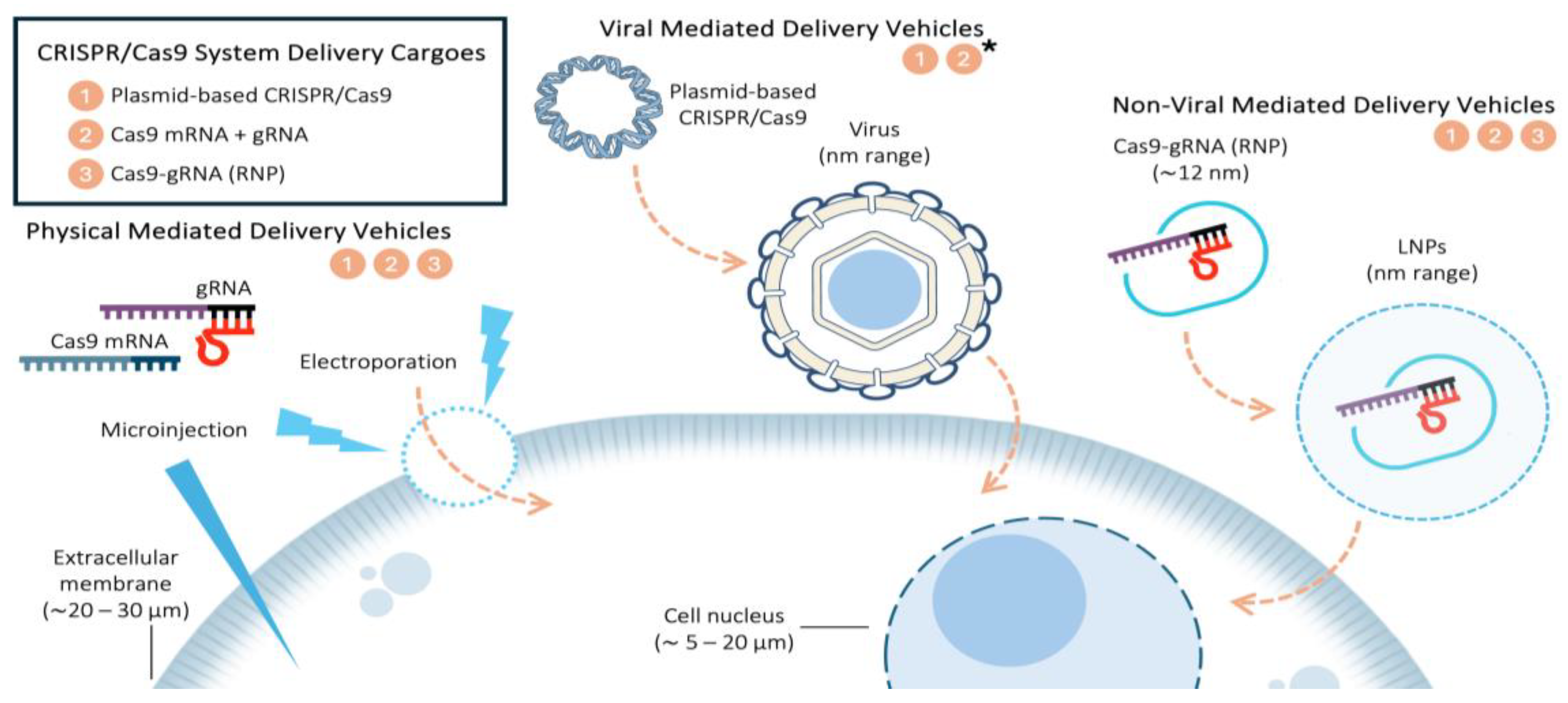
2.1. CRISPR/Cas9 System Delivery Cargoes
- Plasmid-based CRISPR/Cas9. This system is widely used due to its simplicity and low cost manipulation [14]. The moderate toxicity reported in certain cell lines could limit its application as the optimized lipid composition may be different [15]. In addition, both large Cas9 size and nuclear entry limit its genome editing efficiency [8,14]. gRNA can be encoded within the plasmid alongside Cas9 or introduced separately as a synthetic gRNA for more precise control over editing efficiency [16]. Viral vector cargoes might also included in this category, as they typically deliver DNA or RNA encoding both Cas9 and gRNA, rather than introducing them separately, leading to either, genome integration, transient or episomal expression [7].
- Cas9 mRNA coupled with gRNA. This method offers fast and low toxicity genome editing, making it ideal for sensitive cells. Liu et al. [17] clearly demonstrated biocompatibility and high genome editing efficacy using bioreducible LNPs by simultaneous delivery of Cas9 mRNA and gRNA. This system decreases off-target editing events, making it suitable for Cas9 transient expression [14]. The gRNA can either be co-transcribed within the Cas9 mRNA or delivered separately as an independent molecule to optimize stability and efficiency. Viral vectors cargoes could also be in this category when delivering RNA encoding Cas9 and gRNA, without genome integration.
- RNP complexes. RNP are composed by Cas9 protein and gRNA, and they offer the highest gene editing efficiency and specificity [16]. Wei et al. [18] demonstrated that lipid nanoparticles encapsulating RNP exhibit tissue-specific gene editing in mice lungs and liver. Moreover, this system also minimizes off-target effects and toxicity [14,18].
2.2. Types of Delivery Vehicles
2.2.1. Physical Mediated Delivery Vehicles
2.2.2. Viral Mediated Delivery Vehicles
2.2.3. Non-Viral Mediated Delivery Vehicles
3. Aggregation Behavior of Cas9
4. Encapsulation Efficiency of Cas9
| Delivery system | Cas9 encapsulation efficiency | Particle size (Hydrodynamic) | Gene editing efficiency/ Therapeutic outcome | Ref. |
|---|---|---|---|---|
| Exosomes (native) | ~1% (low stochastic loading) | Not specified | Poor delivery, not editing data | [60] |
| Cas9 conjugated to a 12 nm gold nanoparticle | ~45 Cas9 per particle (~6%) | ~23 ± 5 nm | Comparable to electroporation in reported assays | [62] |
| LNPs | Not reported numerically; varied with RNP ratio | ~100-200 nm (est.) | Up to 45.2% indels in IL-10 gene | [63] |
| LNPs | Not quantified | Not specified | ~3-3.5% HDR integration; 80 % restoration of cystic fibrosis transmembrane conductance regulator chloride channel function | [65] |
| Virus like particles | Not specified, but functional loading confirmed | ~100-200 nm | Knockout efficiency 70-90% in vitro and 60-70% in primary human T cells | [64] |
| Mini enveloped delivery vehicles | Not quantified | ~120-140 nm (est.) | Improved gene knockout in human T cells and reduced immunogenic content | [66] |
| Gold nanoparticle aggregates | Surface clustering observed (variable density) | >40nm in aggregated states | Decreased nuclear entry and editing when aggregation is not controlled | [62] |
5. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CRISPR | Clustered Regulatory Interspaced Short Palindromic Repeats |
| Cas9 | CRISPR-associated 9 |
| gRNA | Guide RNA |
| RNP | Ribonucleoprotein |
| LNPs | Lipid Nanoparticles |
| pDNA | Plasmid DNA |
| mRNA | Messenger RNA |
| iPSCs | Induced Pluripotent Stem Cells |
| AVs | Adenoviruses |
| AAVs | Adeno-associated viruses |
| DOPE | 1,2-dioleoyl-sn-glycero-3-phosphorylethanolamine |
| PEG | Polyethyleneglycol |
| CDK11 | Cyclin-Dependent Kinase 11 |
| PEI | Polyethylenimine |
| ZIFs | Zeolitic Imidazolate Frameworks |
| DLS | Dynamic Light Scattering |
| FCS | Fluorescence Correlation Spectroscopy |
References
- Sternberg, S.H.; Redding, S.; Jinek, M.; Greene, E.C.; Doudna, J.A. DNA Interrogation by the CRISPR RNA-Guided Endonuclease Cas9. Nature 2014, 507, 62–67. [CrossRef]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science 2012, 337, 816–821. [CrossRef]
- Zhang, L.; Wang, P.; Feng, Q.; Wang, N.; Chen, Z.; Huang, Y.; Zheng, W.; Jiang, X. Lipid Nanoparticle-Mediated Efficient Delivery of CRISPR/Cas9 for Tumor Therapy. NPG Asia Mater 2017, 9, e441–e441. [CrossRef]
- Du, Y.; Liu, Y.; Hu, J.; Peng, X.; Liu, Z. CRISPR/Cas9 Systems: Delivery Technologies and Biomedical Applications. Asian Journal of Pharmaceutical Sciences 2023, 18, 100854. [CrossRef]
- Sinclair, F.; Begum, A.A.; Dai, C.C.; Toth, I.; Moyle, P.M. Recent Advances in the Delivery and Applications of Nonviral CRISPR/Cas9 Gene Editing. Drug Deliv. and Transl. Res. 2023, 13, 1500–1519. [CrossRef]
- Kazemian, P.; Yu, S.-Y.; Thomson, S.B.; Birkenshaw, A.; Leavitt, B.R.; Ross, C.J.D. Lipid-Nanoparticle-Based Delivery of CRISPR/Cas9 Genome-Editing Components. Mol. Pharmaceutics 2022, 19, 1669–1686. [CrossRef]
- Naso, M.F.; Tomkowicz, B.; Perry, W.L.; Strohl, W.R. Adeno-Associated Virus (AAV) as a Vector for Gene Therapy. BioDrugs 2017, 31, 317–334. [CrossRef]
- Duan, L.; Ouyang, K.; Xu, X.; Xu, L.; Wen, C.; Zhou, X.; Qin, Z.; Xu, Z.; Sun, W.; Liang, Y. Nanoparticle Delivery of CRISPR/Cas9 for Genome Editing. Front Genet 2021, 12, 673286. [CrossRef]
- Nguyen, D.N.; Roth, T.L.; Li, P.J.; Chen, P.A.; Apathy, R.; Mamedov, M.R.; Vo, L.T.; Tobin, V.R.; Goodman, D.; Shifrut, E.; et al. Polymer-Stabilized Cas9 Nanoparticles and Modified Repair Templates Increase Genome Editing Efficiency. Nat Biotechnol 2020, 38, 44–49. [CrossRef]
- den Engelsman, J.; Garidel, P.; Smulders, R.; Koll, H.; Smith, B.; Bassarab, S.; Seidl, A.; Hainzl, O.; Jiskoot, W. Strategies for the Assessment of Protein Aggregates in Pharmaceutical Biotech Product Development. Pharm Res 2011, 28, 920–933. [CrossRef]
- Pukala, T.L. Mass Spectrometric Insights into Protein Aggregation. Essays Biochem 2023, 67, 243–253. [CrossRef]
- Finn, J.D.; Smith, A.R.; Patel, M.C.; Shaw, L.; Youniss, M.R.; van Heteren, J.; Dirstine, T.; Ciullo, C.; Lescarbeau, R.; Seitzer, J.; et al. A Single Administration of CRISPR/Cas9 Lipid Nanoparticles Achieves Robust and Persistent In Vivo Genome Editing. Cell Rep 2018, 22, 2227–2235. [CrossRef]
- Zhou, S.; Li, Y.; Wu, Q.; Gong, C. Nanotechnology-Based CRISPR/Cas9 Delivery System for Genome Editing in Cancer Treatment. MedComm – Biomaterials and Applications 2024, 3, e70. [CrossRef]
- Li, L.; Hu, S.; Chen, X. Non-Viral Delivery Systems for CRISPR/Cas9-Based Genome Editing: Challenges and Opportunities. Biomaterials 2018, 171, 207–218. [CrossRef]
- Kulkarni, J.A.; Myhre, J.L.; Chen, S.; Tam, Y.Y.C.; Danescu, A.; Richman, J.M.; Cullis, P.R. Design of Lipid Nanoparticles for in Vitro and in Vivo Delivery of Plasmid DNA. Nanomedicine 2017, 13, 1377–1387. [CrossRef]
- Zuris, J.A.; Thompson, D.B.; Shu, Y.; Guilinger, J.P.; Bessen, J.L.; Hu, J.H.; Maeder, M.L.; Joung, J.K.; Chen, Z.-Y.; Liu, D.R. Cationic Lipid-Mediated Delivery of Proteins Enables Efficient Protein-Based Genome Editing in Vitro and in Vivo. Nat Biotechnol 2015, 33, 73–80. [CrossRef]
- Liu, J.; Chang, J.; Jiang, Y.; Meng, X.; Sun, T.; Mao, L.; Xu, Q.; Wang, M. Fast and Efficient CRISPR/Cas9 Genome Editing In Vivo Enabled by Bioreducible Lipid and Messenger RNA Nanoparticles. Adv Mater 2019, 31, e1902575. [CrossRef]
- Wei, T.; Cheng, Q.; Min, Y.-L.; Olson, E.N.; Siegwart, D.J. Systemic Nanoparticle Delivery of CRISPR-Cas9 Ribonucleoproteins for Effective Tissue Specific Genome Editing. Nat Commun 2020, 11, 3232. [CrossRef]
- Lotfi, M.; Morshedi Rad, D.; Mashhadi, S.S.; Ashouri, A.; Mojarrad, M.; Mozaffari-Jovin, S.; Farrokhi, S.; Hashemi, M.; Lotfi, M.; Ebrahimi Warkiani, M.; et al. Recent Advances in CRISPR/Cas9 Delivery Approaches for Therapeutic Gene Editing of Stem Cells. Stem Cell Rev and Rep 2023, 19, 2576–2596. [CrossRef]
- Chen, S.; Jiao, Y.; Pan, F.; Guan, Z.; Cheng, S.H.; Sun, D. Knock-In of a Large Reporter Gene via the High-Throughput Microinjection of the CRISPR/Cas9 System. IEEE Trans Biomed Eng 2022, 69, 2524–2532. [CrossRef]
- Hashimoto, M.; Takemoto, T. Electroporation Enables the Efficient mRNA Delivery into the Mouse Zygotes and Facilitates CRISPR/Cas9-Based Genome Editing. Sci Rep 2015, 5, 11315. [CrossRef]
- Moffa, J.C.; Bland, I.N.; Tooley, J.R.; Kalyanaraman, V.; Heitmeier, M.; Creed, M.C.; Copits, B.A. Cell Specific Single Viral Vector CRISPR/Cas9 Editing and Genetically Encoded Tool Delivery in the Central and Peripheral Nervous Systems. bioRxiv 2023, 2023.10.10.561249. [CrossRef]
- Waehler, R.; Russell, S.J.; Curiel, D.T. Engineering Targeted Viral Vectors for Gene Therapy. Nat Rev Genet 2007, 8, 573–587. [CrossRef]
- Li, C.; Samulski, R.J. Engineering Adeno-Associated Virus Vectors for Gene Therapy. Nat Rev Genet 2020, 21, 255–272. [CrossRef]
- Follenzi, A.; Sabatino, G.; Lombardo, A.; Boccaccio, C.; Naldini, L. Efficient Gene Delivery and Targeted Expression to Hepatocytes In Vivo by Improved Lentiviral Vectors. Human Gene Therapy 2002, 13, 243–260. [CrossRef]
- Wang, M.; Zuris, J.A.; Meng, F.; Rees, H.; Sun, S.; Deng, P.; Han, Y.; Gao, X.; Pouli, D.; Wu, Q.; et al. Efficient Delivery of Genome-Editing Proteins Using Bioreducible Lipid Nanoparticles. Proc. Natl. Acad. Sci. U.S.A. 2016, 113, 2868–2873. [CrossRef]
- Lee, K.; Conboy, M.; Park, H.M.; Jiang, F.; Kim, H.J.; Dewitt, M.A.; Mackley, V.A.; Chang, K.; Rao, A.; Skinner, C.; et al. Nanoparticle Delivery of Cas9 Ribonucleoprotein and Donor DNA in Vivo Induces Homology-Directed DNA Repair. Nat Biomed Eng 2017, 1, 889–901. [CrossRef]
- Alsaiari, S.K.; Patil, S.; Alyami, M.; Alamoudi, K.O.; Aleisa, F.A.; Merzaban, J.S.; Li, M.; Khashab, N.M. Endosomal Escape and Delivery of CRISPR/Cas9 Genome Editing Machinery Enabled by Nanoscale Zeolitic Imidazolate Framework. J. Am. Chem. Soc. 2018, 140, 143–146. [CrossRef]
- Averina, O.A.; Permyakov, O.A.; Grigorieva, O.O.; Starshin, A.S.; Mazur, A.M.; Prokhortchouk, E.B.; Dontsova, O.A.; Sergiev, P.V. Comparative Analysis of Genome Editors Efficiency on a Model of Mice Zygotes Microinjection. Int J Mol Sci 2021, 22, 10221. [CrossRef]
- Chen, L.-N.; Fan, X.-Y.; Liu, Y.-T.; Chen, S.-Q.; Xie, F.-Y.; Zeng, L.; Wen, J.; Li, J.; Ma, J.-Y.; Ou, X.-H.; et al. High-Survival Rate After Microinjection of Mouse Oocytes and Early Embryos With mRNA by Combining a Tip Pipette and Piezoelectric-Assisted Micromanipulator. Front Cell Dev Biol 2021, 9, 735971. [CrossRef]
- Niola, F.; Dagnæs-Hansen, F.; Frödin, M. In Vivo Editing of the Adult Mouse Liver Using CRISPR/Cas9 and Hydrodynamic Tail Vein Injection. Methods Mol Biol 2019, 1961, 329–341. [CrossRef]
- Kanefuji, T.; Yokoo, T.; Suda, T.; Abe, H.; Kamimura, K.; Liu, D. Hemodynamics of a Hydrodynamic Injection. Molecular Therapy - Methods & Clinical Development 2014, 1, 14029. [CrossRef]
- Goodwin, T.; Huang, L. Nonviral Vectors: We Have Come a Long Way. Adv Genet 2014, 88, 1–12. [CrossRef]
- Wang, S.-W.; Gao, C.; Zheng, Y.-M.; Yi, L.; Lu, J.-C.; Huang, X.-Y.; Cai, J.-B.; Zhang, P.-F.; Cui, Y.-H.; Ke, A.-W. Current Applications and Future Perspective of CRISPR/Cas9 Gene Editing in Cancer. Mol Cancer 2022, 21, 57. [CrossRef]
- Breyer, B.; Jiang, W.; Cheng, H.; Haydon, R.; Zhou, L.; Feng, T.; He, T.-C. Development and Use of Viral Vectors for Gene Transfer: Lessons from Their Applications in Gene Therapy. Viral vectors for gene therapy.
- Walther, W.; Stein, U. Viral Vectors for Gene Transfer. 2000.
- Li, X.; Le, Y.; Zhang, Z.; Nian, X.; Liu, B.; Yang, X. Viral Vector-Based Gene Therapy. IJMS 2023, 24, 7736. [CrossRef]
- Warrington, K.H.; Herzog, R.W. Treatment of Human Disease by Adeno-Associated Viral Gene Transfer. Hum Genet 2006, 119, 571–603. [CrossRef]
- Schauber, C.A.; Tuerk, M.J.; Pacheco, C.D.; Escarpe, P.A.; Veres, G. Lentiviral Vectors Pseudotyped with Baculovirus Gp64 Efficiently Transduce Mouse Cells in Vivo and Show Tropism Restriction against Hematopoietic Cell Types in Vitro. Gene Ther 2004, 11, 266–275. [CrossRef]
- Coroadinha, A.S. Host Cell Restriction Factors Blocking Efficient Vector Transduction: Challenges in Lentiviral and Adeno-Associated Vector Based Gene Therapies. Cells 2023, 12, 732. [CrossRef]
- Xie, L.; Han, Y.; Liu, Y.; Zhou, Y.; Yu, J.; Von Brunn, A.; Lei, J. Viral Vector-based Cancer Treatment and Current Clinical Applications. MedComm – Oncology 2023, 2, e55. [CrossRef]
- Nasimuzzaman, M. Viral Vectors for Gene Therapy of Genetic Diseases: Challenges and Prospects. JHVRV 2014, 2. [CrossRef]
- Miao, C.H.; Nakai, H.; Thompson, A.R.; Storm, T.A.; Chiu, W.; Snyder, R.O.; Kay, M.A. Nonrandom Transduction of Recombinant Adeno-Associated Virus Vectors in Mouse Hepatocytes In Vivo: Cell Cycling Does Not Influence Hepatocyte Transduction. J Virol 2000, 74, 3793–3803. [CrossRef]
- Kao, C.-Y.; Pan, Y.-C.; Hsiao, Y.-H.; Lim, S.-K.; Cheng, T.-W.; Huang, S.-W.; Wu, S.M.-Y.; Sun, C.-P.; Tao, M.-H.; Mou, K.Y. Improvement of Gene Delivery by Minimal Bacteriophage Particles. ACS Nano 2023, 17, 14532–14544. [CrossRef]
- Bezeljak, U. Cancer Gene Therapy Goes Viral: Viral Vector Platforms Come of Age. Radiology and Oncology 2022, 56, 1–13. [CrossRef]
- Dudek, A.M.; Porteus, M.H. Answered and Unanswered Questions in Early-Stage Viral Vector Transduction Biology and Innate Primary Cell Toxicity for Ex-Vivo Gene Editing. Front. Immunol. 2021, 12, 660302. [CrossRef]
- Mendonça, M.C.P.; Kont, A.; Kowalski, P.S.; O’Driscoll, C.M. Design of Lipid-Based Nanoparticles for Delivery of Therapeutic Nucleic Acids. Drug Discovery Today 2023, 28, 103505. [CrossRef]
- Kim, M.; Hwang, Y.; Lim, S.; Jang, H.-K.; Kim, H.-O. Advances in Nanoparticles as Non-Viral Vectors for Efficient Delivery of CRISPR/Cas9. Pharmaceutics 2024, 16, 1197. [CrossRef]
- Miteva, M.; Kirkbride, K.C.; Kilchrist, K.V.; Werfel, T.A.; Li, H.; Nelson, C.E.; Gupta, M.K.; Giorgio, T.D.; Duvall, C.L. Tuning PEGylation of Mixed Micelles to Overcome Intracellular and Systemic siRNA Delivery Barriers. Biomaterials 2015, 38, 97–107. [CrossRef]
- Guzman Gonzalez, V.; Grunenberger, A.; Nicoud, O.; Czuba, E.; Vollaire, J.; Josserand, V.; Le Guével, X.; Desai, N.; Coll, J.-L.; Divita, G.; et al. Enhanced CRISPR-Cas9 RNA System Delivery Using Cell Penetrating Peptides-Based Nanoparticles for Efficient in Vitro and in Vivo Applications. J Control Release 2024, 376, 1160–1175. [CrossRef]
- Liu, B.-Y.; He, X.-Y.; Zhuo, R.-X.; Cheng, S.-X. Tumor Targeted Genome Editing Mediated by a Multi-Functional Gene Vector for Regulating Cell Behaviors. Journal of Controlled Release 2018, 291, 90–98. [CrossRef]
- Sun, W.; Ji, W.; Hall, J.M.; Hu, Q.; Wang, C.; Beisel, C.L.; Gu, Z. Self-Assembled DNA Nanoclews for the Efficient Delivery of CRISPR-Cas9 for Genome Editing. Angew Chem Int Ed Engl 2015, 54, 12029–12033. [CrossRef]
- Yang, X.-C.; Samanta, B.; Agasti, S.S.; Jeong, Y.; Zhu, Z.-J.; Rana, S.; Miranda, O.R.; Rotello, V.M. Drug Delivery Using Nanoparticle-Stabilized Nanocapsules. Angew Chem Int Ed Engl 2011, 50, 477–481. [CrossRef]
- Alyami, M.Z.; Alsaiari, S.K.; Li, Y.; Qutub, S.S.; Aleisa, F.A.; Sougrat, R.; Merzaban, J.S.; Khashab, N.M. Cell-Type-Specific CRISPR/Cas9 Delivery by Biomimetic Metal Organic Frameworks. J. Am. Chem. Soc. 2020, 142, 1715–1720. [CrossRef]
- Liang, Y.; Xu, X.; Xu, L.; Iqbal, Z.; Ouyang, K.; Zhang, H.; Wen, C.; Duan, L.; Xia, J. Chondrocyte-Specific Genomic Editing Enabled by Hybrid Exosomes for Osteoarthritis Treatment. Theranostics 2022, 12, 4866–4878. [CrossRef]
- Novo, M.; Pérez-González, C.; Freire, S.; Al-Soufi, W. Early Aggregation of Amyloid-β(1–42) Studied by Fluorescence Correlation Spectroscopy. In Protein Aggregation; Cieplak, A.S., Ed.; Methods in Molecular Biology; Springer US: New York, NY, 2023; Vol. 2551, pp. 1–14 ISBN 978-1-07-162596-5.
- Novo, M.; Freire, S.; Al-Soufi, W. Critical Aggregation Concentration for the Formation of Early Amyloid-β (1-42) Oligomers. Sci Rep 2018, 8, 1783. [CrossRef]
- Manzano, I.; Taylor, N.; Csordas, M.; Vezeau, G.E.; Salis, H.M.; Zydney, A.L. Purification of Cas9—RNA Complexes by Ultrafiltration. Biotechnol. Prog. 2021, 37. [CrossRef]
- Camperi, J.; Moshref, M.; Dai, L.; Lee, H.Y. Physicochemical and Functional Characterization of Differential CRISPR-Cas9 Ribonucleoprotein Complexes. Anal. Chem. 2022, 94, 1432–1440. [CrossRef]
- Ponomareva, N.I.; Brezgin, S.A.; Kostyusheva, A.P.; Slatinskaya, O.V.; Bayurova, E.O.; Gordeychuk, I.V.; Maksimov, G.V.; Sokolova, D.V.; Babaeva, G.; Khan, I.I.; et al. Stochastic Packaging of Cas Proteins into Exosomes. Mol Biol 2024, 58, 147–156. [CrossRef]
- Li, Q.; Zhang, Z.; Wu, X.; Zhao, Y.; Liu, Y. Cascade-Responsive Nanoparticles for Efficient CRISPR/Cas9-Based Glioblastoma Gene Therapy. ACS Appl. Mater. Interfaces 2025, 17, 4480–4489. [CrossRef]
- Konstantinidou, S.; Lindstaedt, A.; Schmidt, T.J.N.; Nocilla, F.; Maltinti, G.; Rocco, M.A.; Landi, E.; Carli, A.D.; Crucitta, S.; Lai, M.; et al. A Transfection-Free Approach of Gene Editing via a Gold-Based Nanoformulation of the Cas9 Protein 2024.
- Im, S.H.; Jang, M.; Park, J.-H.; Chung, H.J. Finely Tuned Ionizable Lipid Nanoparticles for CRISPR/Cas9 Ribonucleoprotein Delivery and Gene Editing. J Nanobiotechnol 2024, 22, 175. [CrossRef]
- Borovikova, S.E.; Shepelev, M.V.; Mazurov, D.V.; Kruglova, N.A. Efficient Genome Editing Using ‘NanoMEDIC’ AsCas12a-VLPs Produced with Pol II-Transcribed crRNA. IJMS 2024, 25, 12768. [CrossRef]
- Foley, R.A.; Ayoub, P.G.; Sinha, V.; Juett, C.; Sanoyca, A.; Duggan, E.C.; Lathrop, L.E.; Bhatt, P.; Coote, K.; Illek, B.; et al. Lipid Nanoparticles for the Delivery of CRISPR/Cas9 Machinery to Enable Site-Specific Integration of CFTR and Mutation-Agnostic Disease Rescue 2025.
- Ngo, W.; Peukes, J.T.; Baldwin, A.; Xue, Z.W.; Hwang, S.; Stickels, R.R.; Lin, Z.; Satpathy, A.T.; Wells, J.A.; Schekman, R.; et al. Mechanism-Guided Engineering of a Minimal Biological Particle for Genome Editing.
- Schubert, M.S.; Thommandru, B.; Woodley, J.; Turk, R.; Yan, S.; Kurgan, G.; McNeill, M.S.; Rettig, G.R. Optimized Design Parameters for CRISPR Cas9 and Cas12a Homology-Directed Repair. Sci Rep 2021, 11, 19482. [CrossRef]
- Chenouard, V.; Leray, I.; Tesson, L.; Remy, S.; Allan, A.; Archer, D.; Caulder, A.; Fortun, A.; Bernardeau, K.; Cherifi, Y.; et al. Excess of Guide RNA Reduces Knockin Efficiency and Drastically Increases On-Target Large Deletions. iScience 2023, 26, 106399. [CrossRef]
- Palermo, G.; Miao, Y.; Walker, R.C.; Jinek, M.; McCammon, J.A. CRISPR-Cas9 Conformational Activation as Elucidated from Enhanced Molecular Simulations. Proc Natl Acad Sci U S A 2017, 114, 7260–7265. [CrossRef]
- Shibata, M.; Nishimasu, H.; Kodera, N.; Hirano, S.; Ando, T.; Uchihashi, T.; Nureki, O. Real-Space and Real-Time Dynamics of CRISPR-Cas9 Visualized by High-Speed Atomic Force Microscopy. Nat Commun 2017, 8, 1430. [CrossRef]
| Delivery vehicle | Details | Editing efficiency | Ref. |
|---|---|---|---|
| Physical | Microinjection to insert green fluorescent protein in HepG2 cells | Around 40% | [20] |
| Electroporation in mouse zygotes | Highly efficient genome editing | [21] | |
| Viral | AAVs to specifically edit cell lines in mice nervous system | Efficient gene editing | [22] |
| Engineered AAVs with capsid modifications to enhance transduction | Improved efficiency in muscle and neural tissues | [23,24] | |
| Lentiviral vectors optimized for stable genome editing in hematopoietic and liver cells | High efficiency, long-term expression | [25] | |
| Non-viral | Bioreducible LNPs | Up to 90% in cultured cells and up to 80% in vivo | [17] |
| LNPs to deliver RNP into cells and edit tissues such as muscle, brain, liver and lungs | High editing efficient | [18] | |
| Bioreducible LNPs with negative charge in mammalian cells and rodent brain | Approximately 70% | [26] | |
| Nanoparticles with polyglutamic acid in different types of T cells | Knock-in efficiency up to >50% | [9] | |
| Gold nanoparticles to correct mutations of Duchenne muscular dystrophy in different cell types | 5.4% of the gene edited to wild type | [27] | |
| Nanoscale ZIFs | 37% reduction in gene expression | [28] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




