Submitted:
24 March 2025
Posted:
25 March 2025
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
Sample collecting and Fungal Isolates
DNA extraction and genome sequencing
Genome assembly and gene prediction
Chromosome alignments
Functional annotation
Phylogenetic analysis
Genome-wide Profiling of SIX gene effectors
Results and Discussion
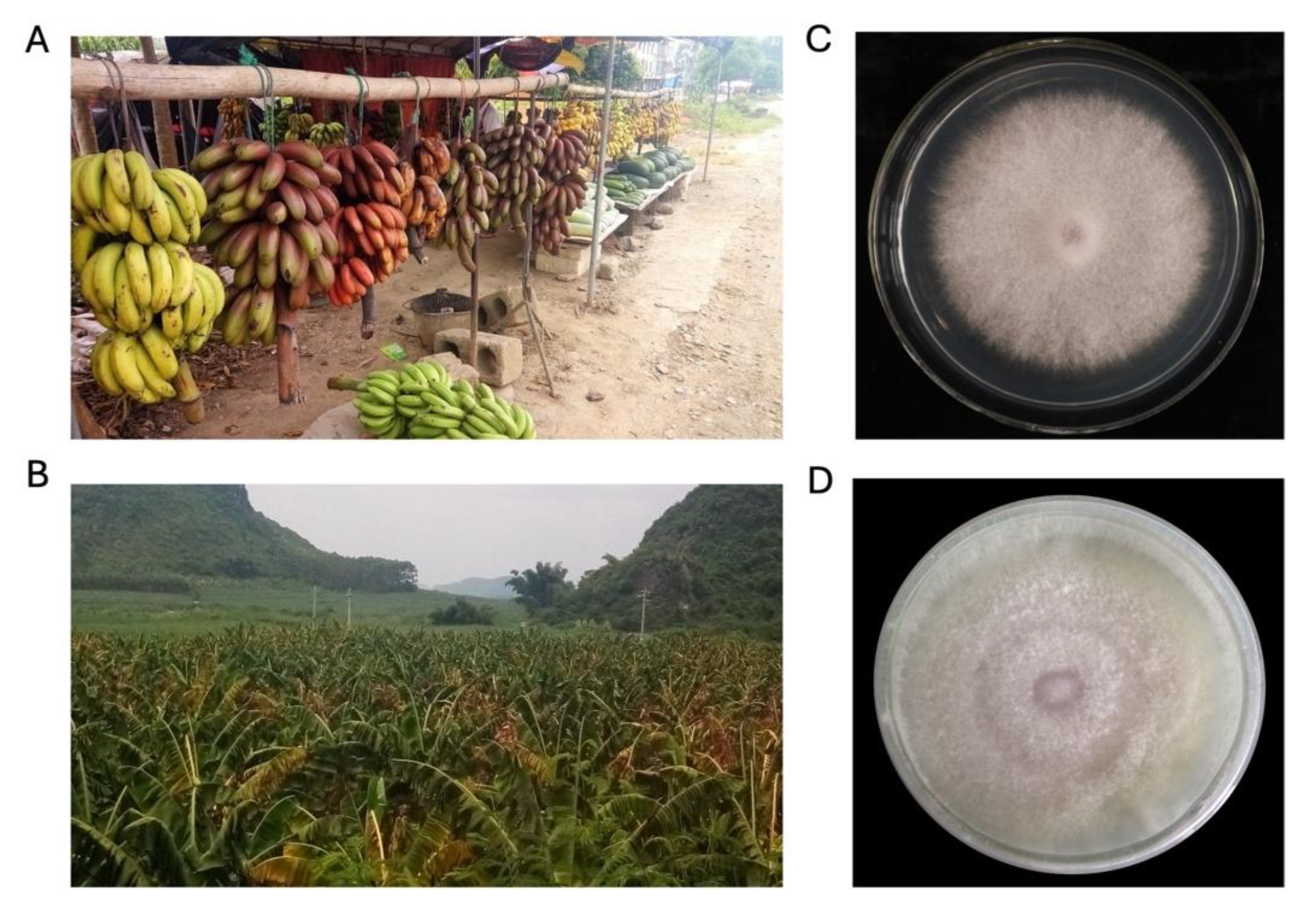



Pathogen isolation
Mitochondrial genomes
Nuclear genomes
Chromosome rearrangements
Functional Annotations
Phylogenetic analysis
Effector annotation
SIX gene profiles of Fusarium oxysporum f.sp. cubense genomes
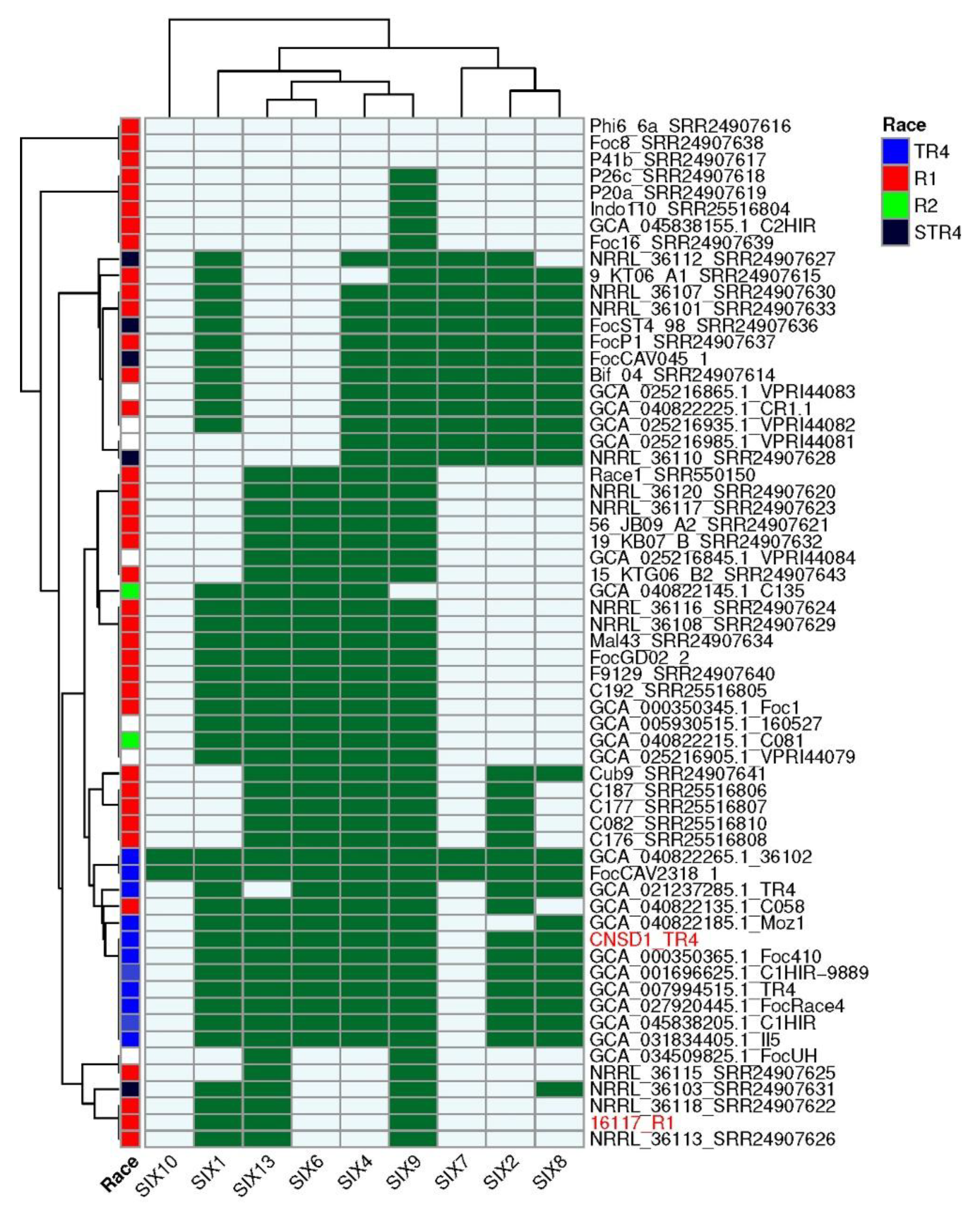
Conclusion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Declaration of Interest Statement
References
- Dita, M.; Barquero, M.; Heck, D.; Mizubuti, E.S.; Staver, C.P. Fusarium wilt of banana: current knowledge on epidemiology and research needs toward sustainable disease management. Front. Plant Sci. 2018, 9, 1468. [CrossRef]
- Ploetz, R.C. Fusarium wilt of banana. Phytopathology 2015, 105, 1512-1521. [CrossRef]
- Pegg, K.G.; Coates, L.M.; O’Neill, W.T.; Turner, D.W. The epidemiology of Fusarium wilt of banana. Front. Plant Sci. 2019, 10, 1395. [CrossRef]
- Islands, C.; Province, M. Vegetative compatibility among races of Fusarium oxysporum f. sp. cubense. Plant Dis. 1988, 72, 325-328.
- Fourie, G.; Steenkamp, E.T.; Ploetz, R.C.; Gordon, T.; Viljoen, A. Current status of the taxonomic position of Fusarium oxysporum formae specialis cubense within the Fusarium oxysporum complex. Infect. Genet. Evol. 2011, 11, 533-542. [CrossRef]
- Mostert, D.; Molina, A.B.; Daniells, J.; Fourie, G.; Hermanto, C.; Chao, C.-P.; Fabregar, E.; Sinohin, V.G.; Masdek, N.; Thangavelu, R. The distribution and host range of the banana Fusarium wilt fungus, Fusarium oxysporum f. sp. cubense, in Asia. PLoS One 2017, 12, e0181630. [CrossRef]
- Munhoz, T.; Vargas, J.; Teixeira, L.; Staver, C.; Dita, M. Fusarium Tropical Race 4 in Latin America and the Caribbean: status and global research advances towards disease management. Front. Plant Sci. 2024, 15. [CrossRef]
- van Westerhoven, A.C.; Meijer, H.J.; Seidl, M.F.; Kema, G.H. Uncontained spread of Fusarium wilt of banana threatens African food security. PLoS Pathog. 2022, 18, e1010769. [CrossRef]
- Sun, J.; Zhang, J.; Fang, H.; Peng, L.; Wei, S.; Li, C.; Zheng, S.; Lu, J. Comparative transcriptome analysis reveals resistance-related genes and pathways in Musa acuminata banana 'Guijiao 9' in response to Fusarium wilt. Plant Physiol.Biochem. 2019, 141, 83-94. [CrossRef]
- Cheng, H.; Concepcion, G.T.; Feng, X.; Zhang, H.; Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 2021, 18, 170-175. [CrossRef]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072-1075. [CrossRef]
- Manni, M.; Berkeley, M.R.; Seppey, M.; Zdobnov, E.M. BUSCO: Assessing genomic data quality and beyond. Curr. Protoc. 2021, 1, e323. [CrossRef]
- Flynn, J.M.; Hubley, R.; Goubert, C.; Rosen, J.; Clark, A.G.; Feschotte, C.; Smit, A.F. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Nat. Acad. Sci. 2020, 117, 9451-9457. [CrossRef]
- Majoros, W.H.; Pertea, M.; Salzberg, S.L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 2004, 20, 2878-2879. [CrossRef]
- Stanke, M.; Morgenstern, B. AUGUSTUS: a web server for gene prediction in eukaryotes that allows user-defined constraints. Nucleic Acids Res. 2005, 33, W465-W467. DOI:org/10.1093/nar/gki458.
- Ter-Hovhannisyan, V.; Lomsadze, A.; Chernoff, Y.O.; Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 2008, 18, 1979-1990. [CrossRef]
- Slater, G.S.C.; Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 2005, 6, 31. [CrossRef]
- Haas, B.J.; Salzberg, S.L.; Zhu, W.; Pertea, M.; Allen, J.E.; Orvis, J.; White, O.; Buell, C.R.; Wortman, J.R. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 2008, 9, 1-22. [CrossRef]
- Uliano-Silva, M.; Ferreira, J.G.R.N.; Krasheninnikova, K.; Blaxter, M.; Mieszkowska, N.; Hall, N.; Holland, P.; Durbin, R.; Richards, T.; Kersey, P.; et al. MitoHiFi: a python pipeline for mitochondrial genome assembly from PacBio high fidelity reads. BMC Bioinformatics 2023, 24, 288. [CrossRef]
- Community, T.G. The Galaxy platform for accessible, reproducible, and collaborative data analyses: 2024 update. Nucleic Acids Res. 2024, 52, W83-W94. [CrossRef]
- Brown, M.R.; Manuel Gonzalez de La Rosa, P.; Blaxter, M. tidk: a toolkit to rapidly identify telomeric repeats from genomic datasets. Bioinformatics 2025, 41. [CrossRef]
- Marçais, G.; Delcher, A.L.; Phillippy, A.M.; Coston, R.; Salzberg, S.L.; Zimin, A. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 2018, 14, e1005944. [CrossRef]
- Blake, J.D.; Cohen, F.E. Pairwise sequence alignment below the twilight zone. J. Mol. Biol. 2001, 307, 721-735. [CrossRef]
- Finn, R.D.; Attwood, T.K.; Babbitt, P.C.; Bateman, A.; Bork, P.; Bridge, A.J.; Chang, H.-Y.; Dosztányi, Z.; El-Gebali, S.; Fraser, M.; et al. InterPro in 2017—beyond protein family and domain annotations. Nucleic Acids Res. 2016, 45, D190-D199. [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59-60. [CrossRef]
- Almagro Armenteros, J.J.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 2019, 37, 420-423. [CrossRef]
- Emanuelsson, O.; Brunak, S.; von Heijne, G.; Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2007, 2, 953-971. [CrossRef]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 2001, 305, 567-580. [CrossRef]
- Sperschneider, J.; Dodds, P.N. EffectorP 3.0: prediction of apoplastic and cytoplasmic effectors in fungi and oomycetes. Mol. Plant Microbe Interact. 2022, 35, 146-156. [CrossRef]
- van Westerhoven, A.C.; Aguilera-Galvez, C.; Nakasato-Tagami, G.; Shi-Kunne, X.; Martinez de la Parte, E.; Chavarro-Carrero, E.; Meijer, H.J.G.; Feurtey, A.; Maryani, N.; Ordóñez, N.; et al. Segmental duplications drive the evolution of accessory regions in a major crop pathogen. New Phytol. 2024, 242, 610-625. [CrossRef]
- Zhang, Y.; Liu, S.; Mostert, D.; Yu, H.; Zhuo, M.; Li, G.; Zuo, C.; Haridas, S.; Webster, K.; Li, M.; et al. Virulence of banana wilt-causing fungal pathogen Fusarium oxysporum tropical race 4 is mediated by nitric oxide biosynthesis and accessory genes. Nat. Microbiol. 2024, 9, 2232-2243. [CrossRef]
- Kaliapan, K.; Mazlin, S.N.A.; Chua, K.O.; Rejab, N.A.; Mohd-Yusuf, Y. Secreted in Xylem (SIX) genes in Fusarium oxysporum f. sp. cubense (Foc) unravels the potential biomarkers for early detection of Fusarium wilt disease. Arch. Microbiol. 2024, 206, 271. [CrossRef]
- Gardiner, D.P.; Aitken, E.A.B.; Le, D.P.; Smith, L.J.; Chen, A. De novo long-read assembly and annotation for genomes of two cotton-associated Fusarium oxysporum isolates (Submitted). 2025.
- Keller, O.; Kollmar, M.; Stanke, M.; Waack, S. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 2011, 27, 757-763. [CrossRef]
- Stanke, M.; Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 2003, 19, 215-225.
- Shen, W.; Sipos, B.; Zhao, L. SeqKit2: A Swiss army knife for sequence and alignment processing. iMeta 2024, e191. [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.i.; Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059-3066. [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972-1973. [CrossRef]
- Edler, D.; Klein, J.; Antonelli, A.; Silvestro, D. raxmlGUI 2.0: a graphical interface and toolkit for phylogenetic analyses using RAxML. Methods Ecol. Evol. 2021, 12, 373-377. [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 2024. [CrossRef]
- Brenes Guallar, M.A.; Fokkens, L.; Rep, M.; Berke, L.; van Dam, P. Fusarium oxysporum effector clustering version 2: An updated pipeline to infer host range. Front. Plant Sci. 2022, 13. [CrossRef]
- van Dam, P.; Fokkens, L.; Schmidt, S.M.; Linmans, J.H.; Kistler, H.C.; Ma, L.J.; Rep, M. Effector profiles distinguish formae speciales of Fusarium oxysporum. Environ. Microbiol. 2016, 18, 4087-4102. [CrossRef]
- Brankovics, B.; van Dam, P.; Rep, M.; de Hoog, G.S.; J. van der Lee, T.A.; Waalwijk, C.; van Diepeningen, A.D. Mitochondrial genomes reveal recombination in the presumed asexual Fusarium oxysporum species complex. BMC Genomics 2017, 18, 735. [CrossRef]
- Cantarel, B.L.; Coutinho, P.M.; Rancurel, C.; Bernard, T.; Lombard, V.; Henrissat, B. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009, 37, D233-238. [CrossRef]
- Ordonez, N.; Seidl, M.F.; Waalwijk, C.; Drenth, A.; Kilian, A.; Thomma, B.P.H.J.; Ploetz, R.C.; Kema, G.H.J. Worse comes to worst: bananas and Panama disease—when plant and pathogen clones meet. PLoS Pathog. 2015, 11, e1005197. [CrossRef]
- Czislowski, E.; Fraser-Smith, S.; Zander, M.; O'Neill, W.T.; Meldrum, R.A.; Tran-Nguyen, L.T.; Batley, J.; Aitken, E.A. Investigation of the diversity of effector genes in the banana pathogen, Fusarium oxysporum f. sp. cubense, reveals evidence of horizontal gene transfer. Mol. Plant pathol. 2018, 19, 1155-1171. [CrossRef]
- Gardiner, D.M.; McDonald, M.C.; Covarelli, L.; Solomon, P.S.; Rusu, A.G.; Marshall, M.; Kazan, K.; Chakraborty, S.; McDonald, B.A.; Manners, J.M. Comparative Pathogenomics reveals horizontally acquired novel virulence genes in fungi infecting cereal hosts. PLoS Pathog. 2012, 8, e1002952. [CrossRef]
| Statistics | 16117 (Race 1) | CNSD1 (TR4) |
|---|---|---|
| Assembly | ||
| Total sequence data (Gbp) | 8.46 | 10.5 |
| Coverage (fold) | 170 | 223 |
| Assembly size (bp) | 51,695,064 | 49,684,144 |
| No. of contigs | 92 | 77 |
| Largest contig (bp) | 5,598,307 | 6,666,412 |
| N50 contig length (bp) | 4,227,447 | 4,512,489 |
| Contig L50 | 6 | 5 |
| Contig L90 | 14 | 11 |
| GC content (%) | 47.87 | 47.72 |
| BUSCO coverage (%) | 98.4 | 98.4 |
| Total no. of BUSCOs | 1706 | 1706 |
| No. of duplicated BUSCOs | 9 | 6 |
| No. of fragmented BUSCOs | 8 | 8 |
| No. of missing BUSCOs | 19 | 20 |
| Gene models | ||
| Total no. of genes | 15,943 | 15,247 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




