Submitted:
17 February 2025
Posted:
17 February 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Collection of Samples
2.2. Fungal Isolation and Identification
2.3. DNA Extraction
2.4. Metabarcoding Analysis
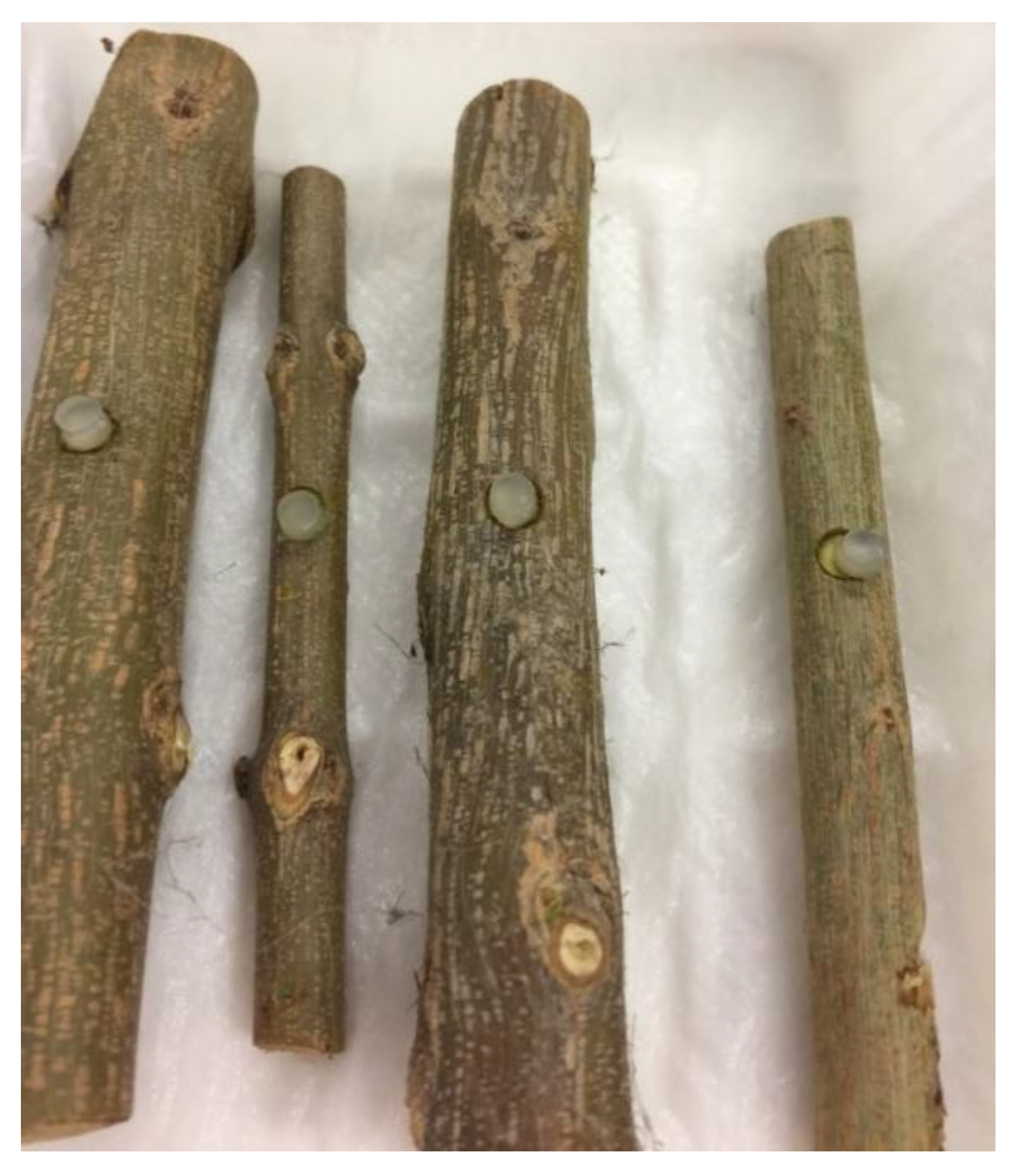
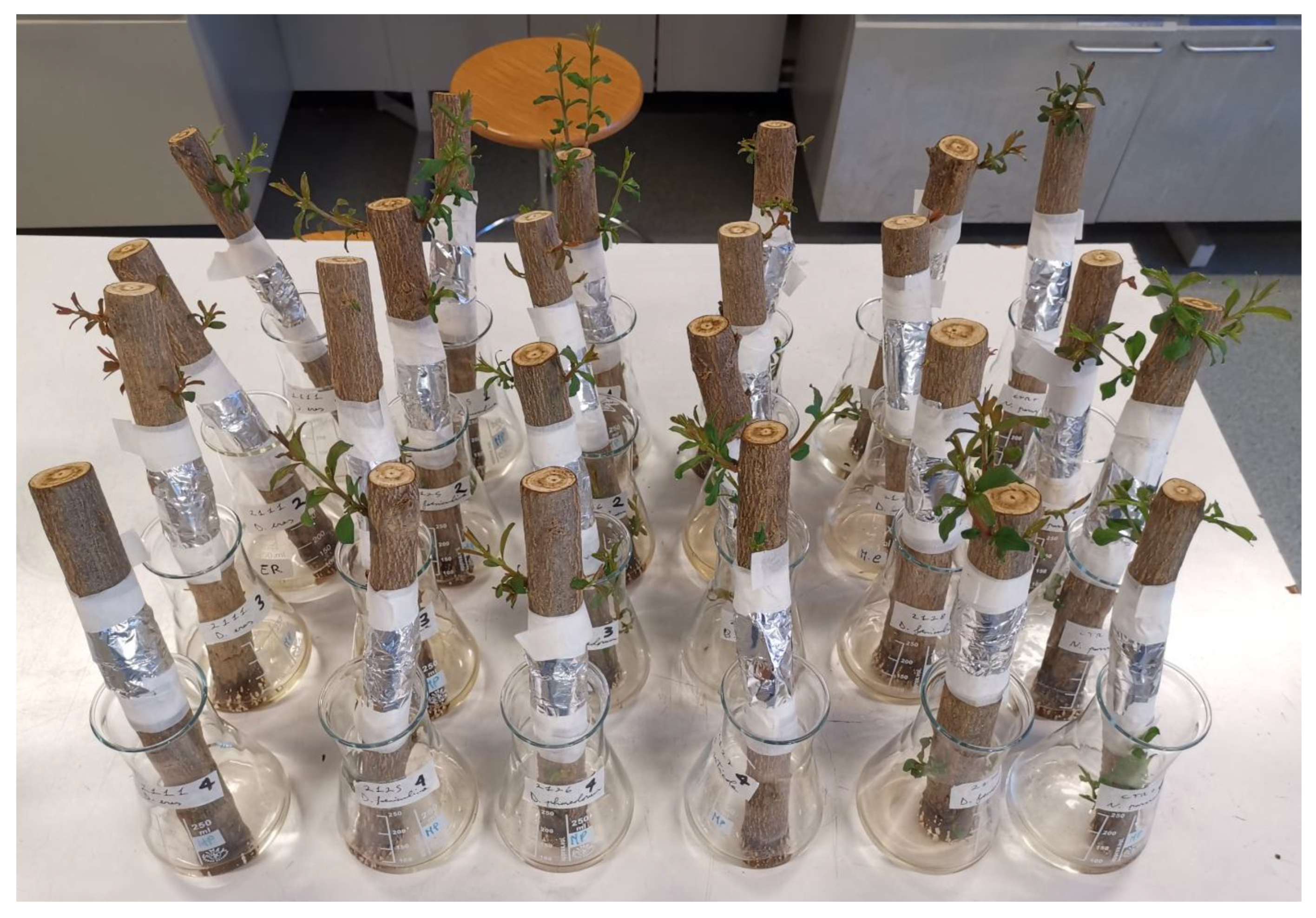
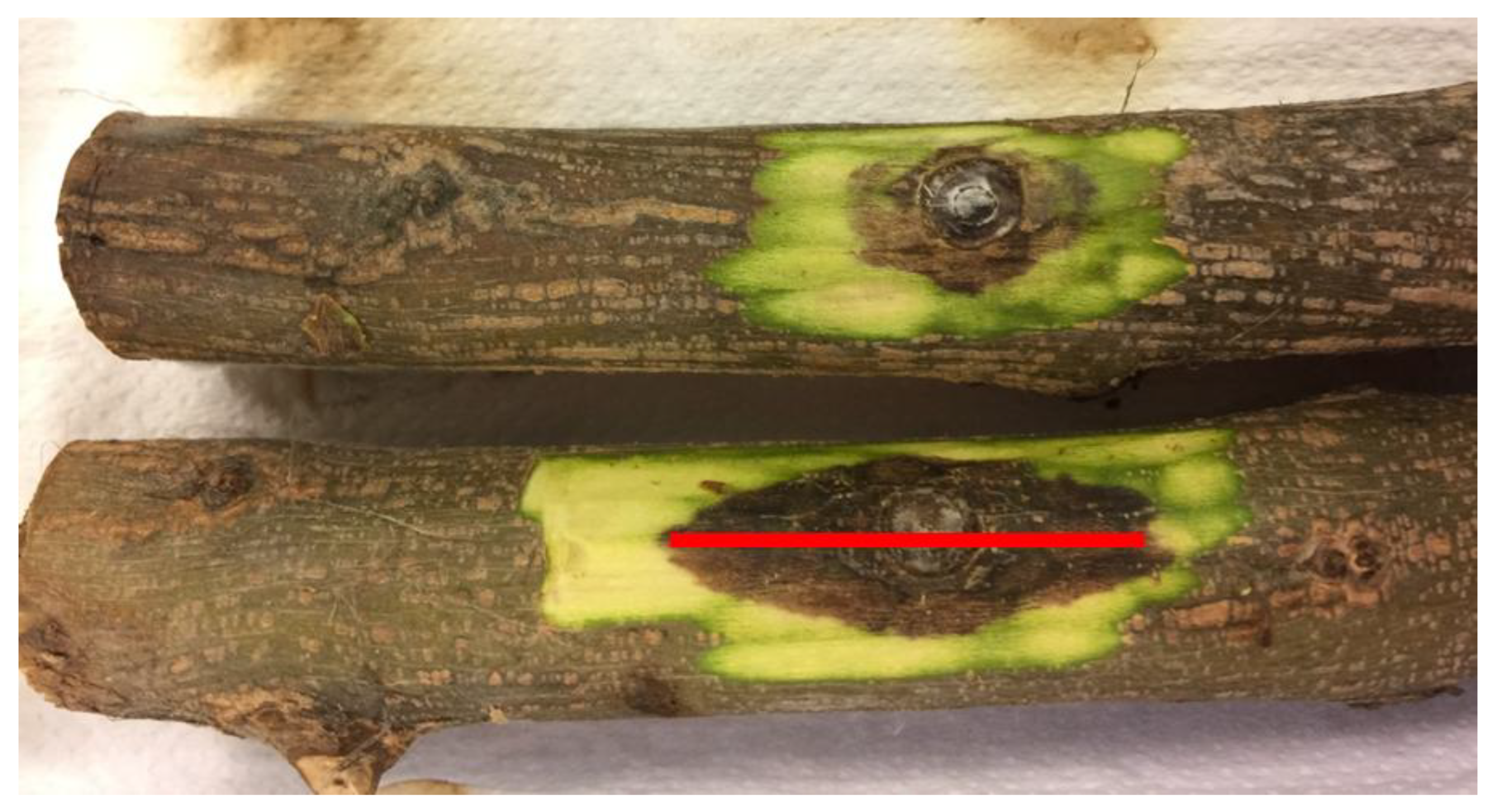
2.5. Pathogenicity Test
2.5. Statistical Analysis
3. Results
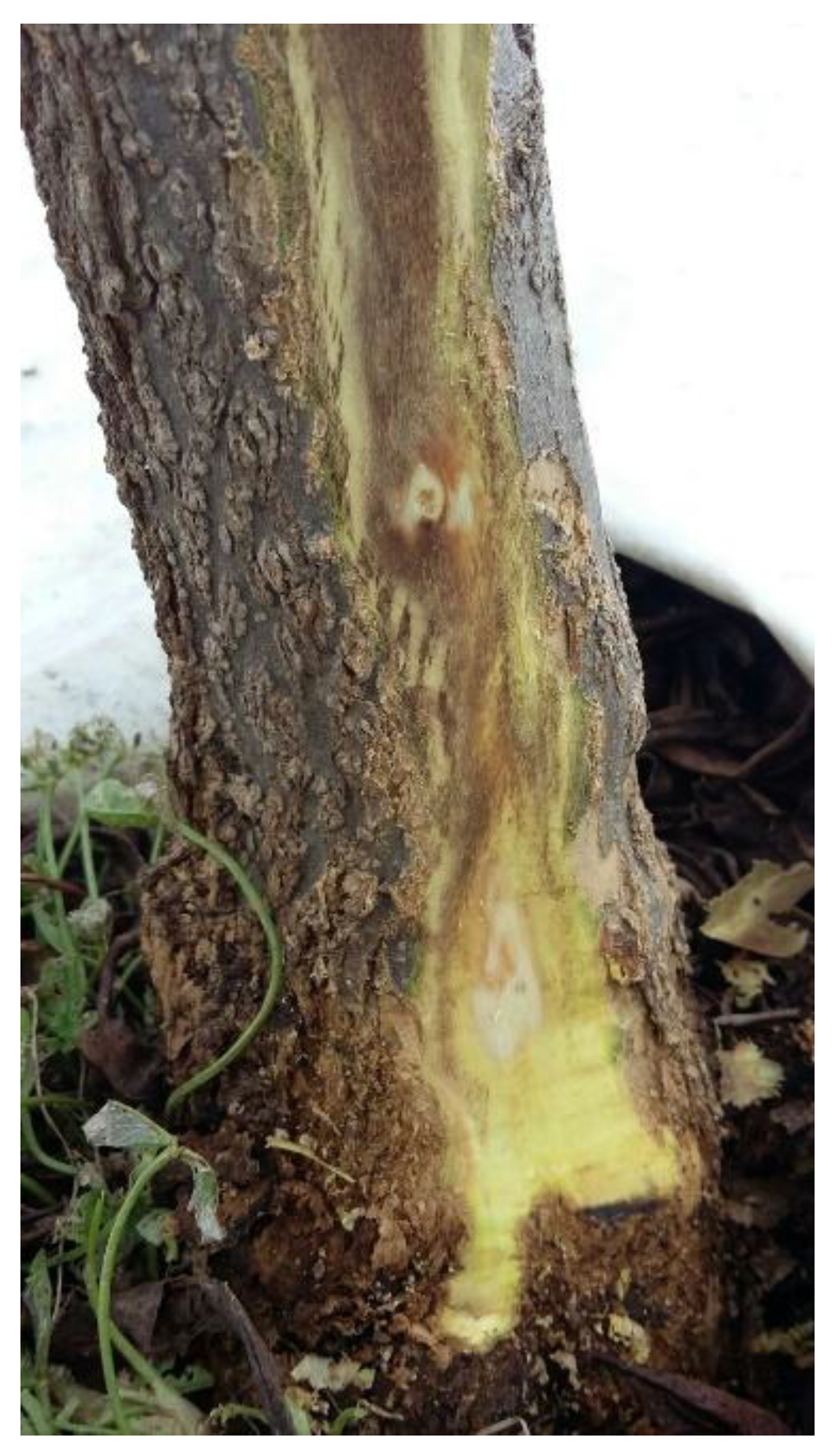
3.1. Phytosanitary Analysis of Pomegranate Plants Affected by Decay
3.2. Metabarcoding Analysis
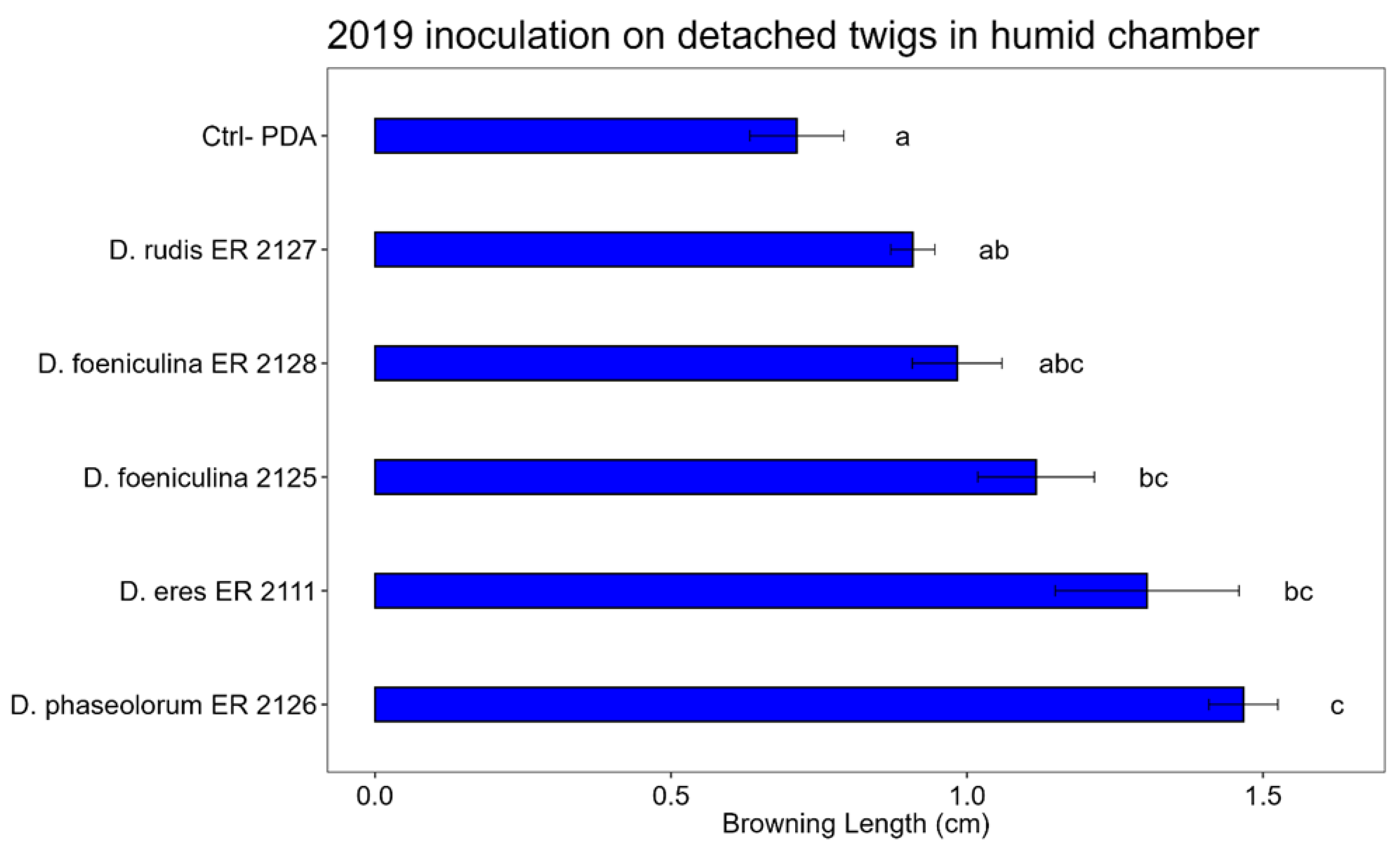
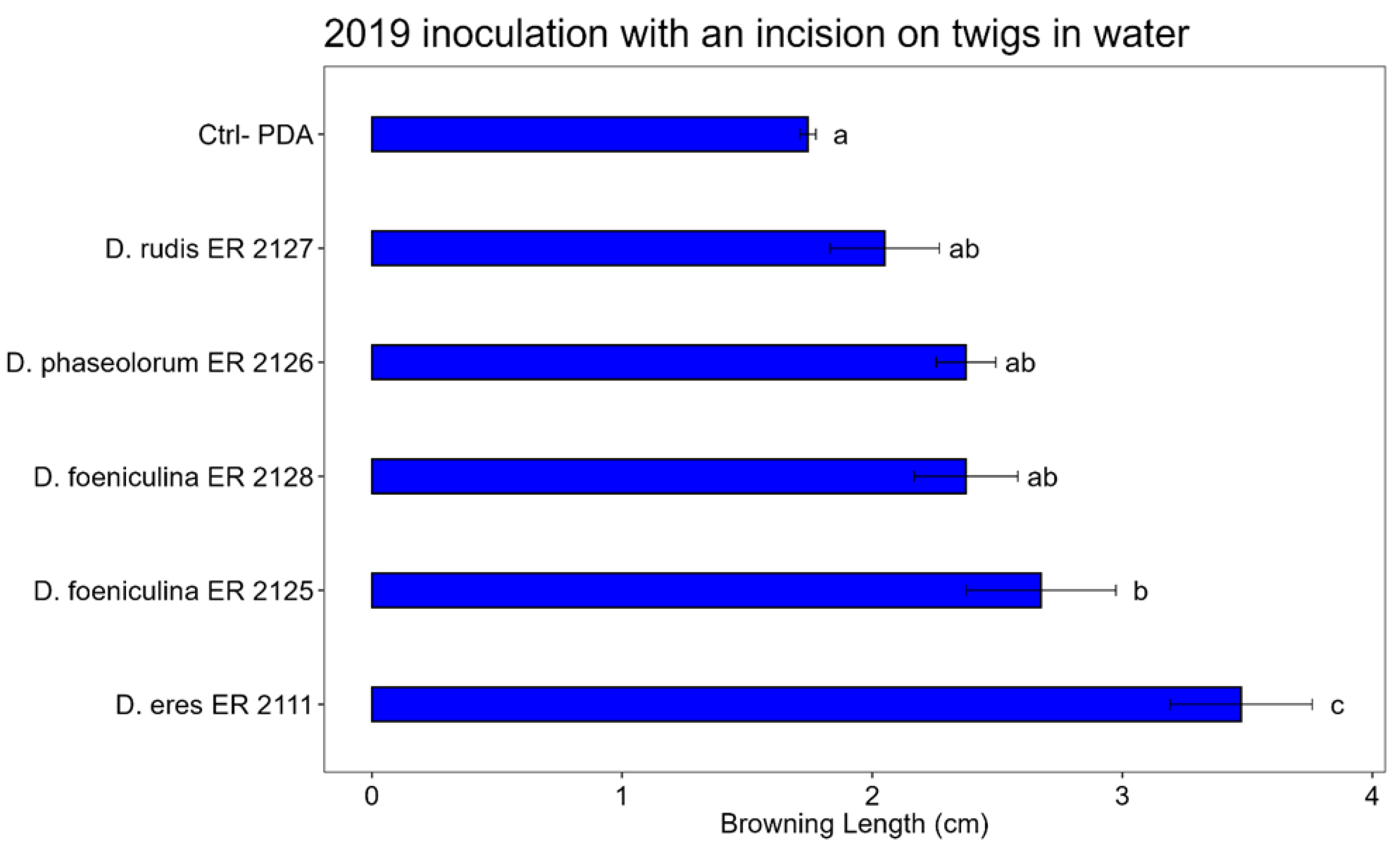
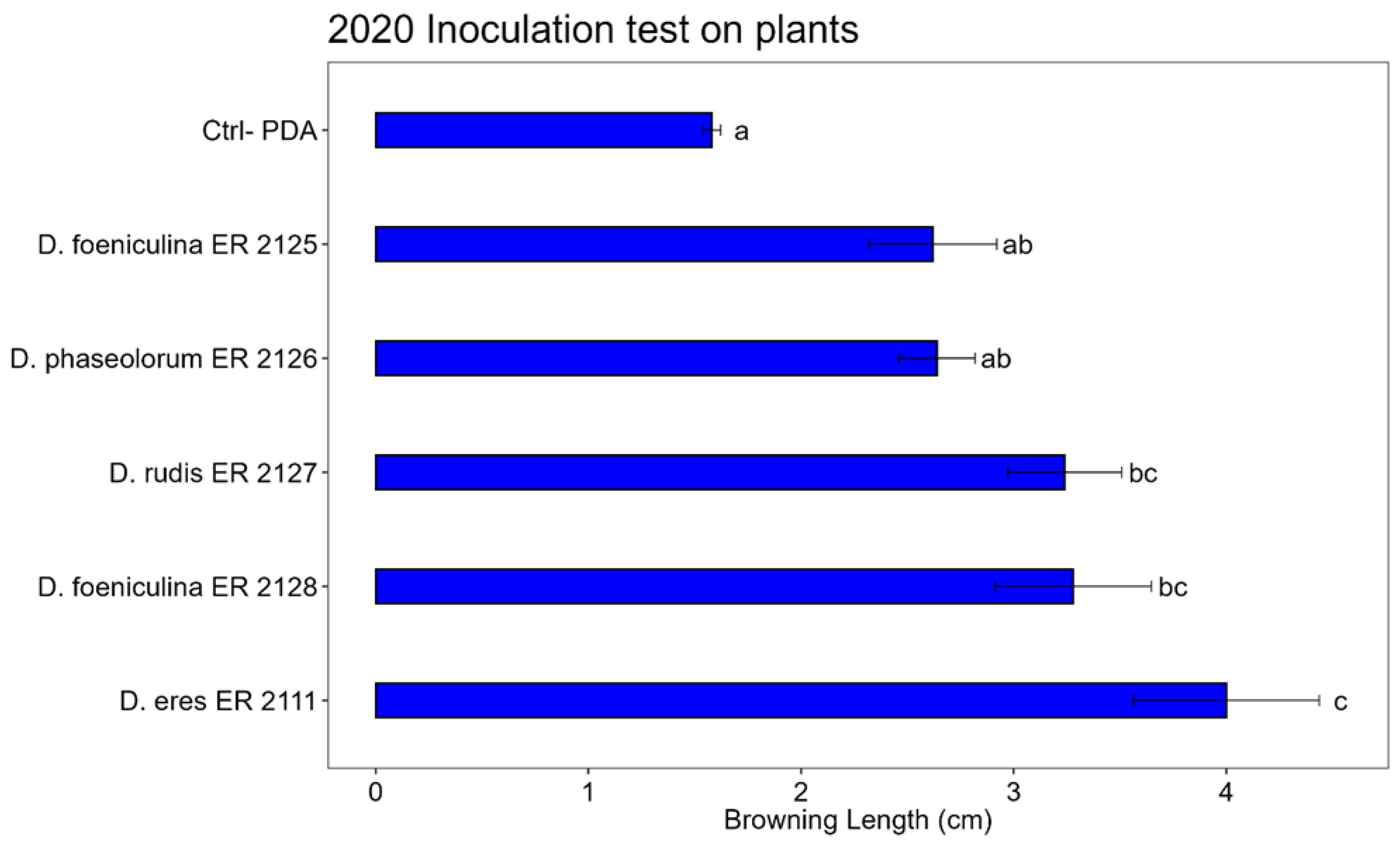
3.3. Pathogenicity Test
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cristofori, V.; Caruso, D.; Latini, G.; Dell’Agli, M.; Cammilli, C.; Rugini, E.; Bignami, C.; Muleo, R. Fruit quality of italian pomegranate (punica granatum l.) autochthonous varieties. European Food Research and Technology 2011, 232, 397–403. [Google Scholar] [CrossRef]
- Hajleh, M.N.A.; Al-Dujaili, E.A.S. In Anti-cancer activity of pomegranate and its biophenols; general review, 2016.
- Fanali, C.; Belluomo, M.G.; Cirilli, M.; Cristofori, V.; Zecchini, M.; Cacciola, F.; Russo, M.; Muleo, R.; Dugo, L. Antioxidant activity evaluation and hplc-photodiode array/ms polyphenols analysis of pomegranate juice from selected italian cultivars: A comparative study. Electrophoresis 2016, 37, 1947–1955. [Google Scholar] [CrossRef] [PubMed]
- Reddy, M.K.; Gupta, S.K.; Jacob, M.R.; Khan, S.I.; Ferreira, D. Antioxidant, antimalarial and antimicrobial activities of tannin-rich fractions, ellagitannins and phenolic acids from punica granatum l. Planta medica 2007, 73, 461–467. [Google Scholar] [CrossRef]
- Giménez-Bastida, J.A.; Ávila-Gálvez, M.Á.; Espín, J.C.; González-Sarrías, A. Evidence for health properties of pomegranate juices and extracts beyond nutrition: A critical systematic review of human studies. Trends in Food Science & Technology 2021, 114, 410–423. [Google Scholar] [CrossRef]
- Pollastro, S.; Dongiovanni, C.; Gerin, D.; Pollastro, P.; Fumarola, G.; De Miccolis Angelini, R.M.; Faretra, F. First report of coniella granati as a causal agent of pomegranate crown rot in southern italy. Plant Disease 2016, 100, 1498. [Google Scholar] [CrossRef]
- Mirabolfathy, M.; Groenewald, J.Z.; Crous, P.W. First report of pilidiella granati causing dieback and fruit rot of pomegranate (punica granatum) in iran. Plant Dis 2012, 96, 461. [Google Scholar] [CrossRef]
- Levy, E.; Elkind, G.; Ben-Arie, R.; Ben-Ze’ev, I.S. First report of coniella granati causing pomegranate fruit rot in israel. Phytoparasitica 2011, 39, 403–405. [Google Scholar] [CrossRef]
- Çeliker, N.M.; Uysal, A.; Çetinel, B.; Poyraz, D. Crown rot on pomegranate caused by coniella granati in turkey. Australasian Plant Disease Notes 2012, 7, 161–162. [Google Scholar] [CrossRef]
- Tziros, G.T.; Tzavella-Klonari, K. Pomegranate fruit rot caused by coniella granati confirmed in greece. Plant Pathology 2008, 57, 783. [Google Scholar] [CrossRef]
- Palou, L.; Guardado, A.; Montesinos-Herrero, C. First report of penicillium spp. And pilidiella granati causing postharvest fruit rot of pomegranate in spain. New Disease Reports 2010, 22, 21. [Google Scholar] [CrossRef]
- Palavouzis, S.C.; Tzamos, S.; Paplomatas, E.; Thomidis, T. First report of neofusicoccum parvum causing shoot blight of pomegranate in northern greece. New Disease Reports 2015, 32, 10. [Google Scholar] [CrossRef]
- KC, A.N.; Vallad, G.E. First report of neofusicoccum parvum causing shoot blight and stem cankers on pomegranate in florida. Plant Disease 2016, 100, 1783. [Google Scholar] [CrossRef]
- Riccioni, L.; Valente, M.T.; Giambattista, G.d. First report of neofusicoccum parvum causing shoot blight and plant decay on pomegranate in tarquinia, italy. Journal of Plant Pathology 2017, 99, 294. [Google Scholar]
- Molina-Gayosso, E.; Silva-Rojas, H.V.; García-Morales, S.; Avila-Quezada, G. First report of black spots on avocado fruit caused by neofusicoccum parvum in mexico. Plant Dis 2012, 96, 287. [Google Scholar] [CrossRef]
- Phillips, A.J.L.; Alves, A.; Abdollahzadeh, J.; Slippers, B.; Wingfield, M.J.; Groenewald, J.Z.; Crous, P.W. The botryosphaeriaceae: Genera and species known from culture. Studies in Mycology 2013, 76, 51–167. [Google Scholar] [CrossRef]
- Úrbez-Torres, J.R.; Gubler, W.D. Susceptibility of grapevine pruning wounds to infection by lasiodiplodia theobromae and neofusicoccum parvum. Plant Pathology 2011, 60, 261–270. [Google Scholar] [CrossRef]
- Lamichhane, J.R.; Venturi, V. Synergisms between microbial pathogens in plant disease complexes: A growing trend. Frontiers in Plant Science 2015, 6. [Google Scholar] [CrossRef]
- Schmidt, P.-A.; Bálint, M.; Greshake, B.; Bandow, C.; Römbke, J.; Schmitt, I. Illumina metabarcoding of a soil fungal community. Soil Biology and Biochemistry 2013, 65, 128–132. [Google Scholar] [CrossRef]
- Abdelfattah, A.; Li Destri Nicosia, M.G.; Cacciola, S.O.; Droby, S.; Schena, L. Metabarcoding analysis of fungal diversity in the phyllosphere and carposphere of olive (olea europaea). PLoS ONE 2015, 10, e0131069. [Google Scholar] [CrossRef]
- Abdelfattah, A.; Cacciola, S.O.; Mosca, S.; Zappia, R.; Schena, L. Analysis of the fungal diversity in citrus leaves with greasy spot disease symptoms. Microbial ecology 2017, 73, 739–749. [Google Scholar] [CrossRef]
- Morales-Cruz, A.; Figueroa-Balderas, R.; García, J.F.; Tran, E.; Rolshausen, P.E.; Baumgartner, K.; Cantu, D. Profiling grapevine trunk pathogens in planta: A case for community-targeted DNA metabarcoding. BMC Microbiology 2018, 18, 214. [Google Scholar] [CrossRef] [PubMed]
- Del Frari, G.; Gobbi, A.; Aggerbeck, M.R.; Oliveira, H.; Hansen, L.H.; Ferreira, R.B. Characterization of the wood mycobiome of vitis vinifera in a vineyard affected by esca. Spatial distribution of fungal communities and their putative relation with leaf symptoms. Frontiers in Plant Science 2019, 10. [Google Scholar] [CrossRef]
- Kõljalg, U.; Larsson, K.-H.; Abarenkov, K.; Nilsson, R.H.; Alexander, I.J.; Eberhardt, U.; Erland, S.; Høiland, K.; Kjøller, R.; Larsson, E.; et al. Unite: A database providing web-based methods for the molecular identification of ectomycorrhizal fungi. New Phytologist 2005, 166, 1063–1068. [Google Scholar] [CrossRef] [PubMed]
- Meiklejohn, K.A.; Damaso, N.; Robertson, J.M. Assessment of bold and genbank – their accuracy and reliability for the identification of biological materials. PLoS ONE 2019, 14, e0217084. [Google Scholar] [CrossRef] [PubMed]
- van der Loos, L.M.; Nijland, R. Biases in bulk: DNA metabarcoding of marine communities and the methodology involved. Molecular ecology 2021, 30, 3270–3288. [Google Scholar] [CrossRef]
- Blaalid, R.; Kumar, S.; Nilsson, R.H.; Abarenkov, K.; Kirk, P.M.; Kauserud, H. Its1 versus its2 as DNA metabarcodes for fungi. Molecular Ecology Resources 2013, 13, 218–224. [Google Scholar] [CrossRef]
- Mbareche, H.; Veillette, M.; Bilodeau, G.; Duchaine, C. Comparison of the performance of its1 and its2 as barcodes in amplicon-based sequencing of bioaerosols. PeerJ 2020, 8, e8523. [Google Scholar] [CrossRef]
- Yang, R.-H.; Su, J.-H.; Shang, J.-J.; Wu, Y.-Y.; Li, Y.; Bao, D.-P.; Yao, Y.-J. Evaluation of the ribosomal DNA internal transcribed spacer (its), specifically its1 and its2, for the analysis of fungal diversity by deep sequencing. PLoS ONE 2018, 13, e0206428. [Google Scholar] [CrossRef]
- Napoli, R.; Paolanti, M.; Di Ferdinando, S. Atlante dei suoli del lazio. ARSIAL Regione Lazio. ISBN 978-88-904841-2-4 2019.
- Fratianni, S.; Acquaotta, F. The climate of italy. In Landscapes and landforms of italy, Soldati, M.; Marchetti, M., Eds. Springer International Publishing: Cham, 2017; pp 29-38.
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. 38 - amplification and direct sequencing of fungal ribosomal rna genes for phylogenetics. In Pcr protocols, Innis, M.A.; Gelfand, D.H.; Sninsky, J.J.; White, T.J., Eds. Academic Press: San Diego, 1990; pp 315-322.
- Cenis, J.L. Rapid extraction of fungal DNA for pcr amplification. Nucleic Acids Research 1992, 20, 2380. [Google Scholar] [CrossRef]
- Aragona, M.; Minio, A.; Ferrarini, A.; Valente, M.T.; Bagnaresi, P.; Orrù, L.; Tononi, P.; Zamperin, G.; Infantino, A.; Valè, G.; et al. De novo genome assembly of the soil-borne fungus and tomato pathogen pyrenochaeta lycopersici. BMC Genomics 2014, 15, 313. [Google Scholar] [CrossRef]
- Miller, K.E.; Hopkins, K.; Inward, D.J.G.; Vogler, A.P. Metabarcoding of fungal communities associated with bark beetles. Ecology and Evolution 2016, 6, 1590–1600. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Uparse: Highly accurate otu sequences from microbial amplicon reads. Nature Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- Abarenkov, K.Z., Allan; Piirmann, Timo; Pöhönen, Raivo; Ivanov, Filipp; Nilsson, R. Henrik; Kõljalg, Urmas. Unite usearch/utax release for eukaryotes. Version 04.04.2024. UNITE Community. 2024.
- Manetti, G.; Brunetti, A.; Lumia, V.; Sciarroni, L.; Marangi, P.; Cristella, N.; Faggioli, F.; Reverberi, M.; Scortichini, M.; Pilotti, M. Identification and characterization of neofusicoccum stellenboschiana in branch and twig dieback-affected olive trees in italy and comparative pathogenicity with n. Mediterraneum. Journal of Fungi 2023, 9, 292. [Google Scholar] [CrossRef] [PubMed]
- Moncini, L.; Pachetti, G.; Pollastro, S. Il cancro del fusto minaccia il melograno italiano. 2017.
- Graniti, A.; Surico, G.; Mugnai, L. Esca of grapevine: A disease complex or a complex of diseases? Phytopathologia Mediterranea 2000, 39, 16–20. [Google Scholar]
- Hennion, B.; Lecomte, P.; Larignon, P.; Baudry, A.; Durpaire, M.P.; Mouillon, M.; Tailleur, J.L.; Dupuy, O. First observations of a wood decay (esca-loke disease) on kiwifruit in france. Acta Horticulturae 2003, 409–413. [Google Scholar] [CrossRef]
- Savian, F.; Ginaldi, F.; Musetti, R.; Sandrin, N.; Tarquini, G.; Pagliari, L.; Firrao, G.; Martini, M.; Ermacora, P. Studies on the aetiology of kiwifruit decline: Interaction between soil-borne pathogens and waterlogging. Plant and Soil 2020, 456, 113–128. [Google Scholar] [CrossRef]
- Prencipe, S.; Savian, F.; Nari, L.; Ermacora, P.; Spadaro, D.; Martini, M. First report of phytopythium vexans causing decline syndrome of actinidia deliciosa ‘hayward’ in italy. Plant Disease 2020, 104, 2032. [Google Scholar] [CrossRef]
- Donati, I.; Cellini, A.; Sangiorgio, D.; Caldera, E.; Sorrenti, G.; Spinelli, F. Pathogens associated to kiwifruit vine decline in italy. Agriculture 2020, 10, 119. [Google Scholar] [CrossRef]
- Tacconi, G.; Paltrinieri, S.; Mejia, J.F.; Fuentealba, S.P.; Bertaccini, A.; Tosi, L.; Giacopini, A.; Mazzucchi, U.; Favaron, F.; Sella, L.; et al. Vine decline in kiwifruit: Climate change and effect on waterlogging and phytophthora in north italy. 2015, 93–97.
- Spigaglia, P.; Barbanti, F.; Marocchi, F.; Mastroleo, M.; Baretta, M.; Ferrante, P.; Caboni, E.; Lucioli, S.; Scortichini, M. Clostridium bifermentans and c. Subterminale are associated with kiwifruit vine decline, known as moria, in italy. Plant Pathology 2020, 69, 765–774. [Google Scholar] [CrossRef]
- Chandra, R.; Babu, K.D.; Jadhav, V.T.; Silva, J.T.d. In Origin, history and domestication of pomegranate, 2010.
- Mello, A.; Lumini, E.; Napoli, C.; Bianciotto, V.; Bonfante, P. Arbuscular mycorrhizal fungal diversity in the tuber melanosporum brûlé. Fungal Biology 2015, 119, 518–527. [Google Scholar] [CrossRef]
- Bazzicalupo, A.L.; Bálint, M.; Schmitt, I. Comparison of its1 and its2 rdna in 454 sequencing of hyperdiverse fungal communities. Fungal Ecology 2013, 6, 102–109. [Google Scholar] [CrossRef]
- Monard, C.; Gantner, S.; Stenlid, J. Utilizing its1 and its2 to study environmental fungal diversity using pyrosequencing. FEMS Microbiology Ecology 2013, 84, 165–175. [Google Scholar] [CrossRef]
- Orgiazzi, A.; Bianciotto, V.; Bonfante, P.; Daghino, S.; Ghignone, S.; Lazzari, A.; Lumini, E.; Mello, A.; Napoli, C.; Perotto, S.; et al. 454 pyrosequencing analysis of fungal assemblages from geographically distant, disparate soils reveals spatial patterning and a core mycobiome. Diversity 2013, 5, 73–98. [Google Scholar] [CrossRef]
- Nilsson, R.H.; Ryberg, M.; Abarenkov, K.; Sjökvist, E.; Kristiansson, E. The its region as a target for characterization of fungal communities using emerging sequencing technologies. FEMS Microbiology Letters 2009, 296, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Bellemain, E.; Davey, M.L.; Kauserud, H.; Epp, L.S.; Boessenkool, S.; Coissac, E.; Geml, J.; Edwards, M.; Willerslev, E.; Gussarova, G.; et al. Fungal palaeodiversity revealed using high-throughput metabarcoding of ancient DNA from arctic permafrost. Environ Microbiol 2013, 15, 1176–1189. [Google Scholar] [CrossRef]
- Badotti, F.; de Oliveira, F.S.; Garcia, C.F.; Vaz, A.B.M.; Fonseca, P.L.C.; Nahum, L.A.; Oliveira, G.; Góes-Neto, A. Effectiveness of its and sub-regions as DNA barcode markers for the identification of basidiomycota (fungi). BMC Microbiology 2017, 17, 42. [Google Scholar] [CrossRef]
- Tedersoo, L.; Anslan, S.; Bahram, M.; Põlme, S.; Riit, T.; Liiv, I.; Kõljalg, U.; Kisand, V.; Nilsson, H.; Hildebrand, F.; et al. Shotgun metagenomes and multiple primer pair-barcode combinations of amplicons reveal biases in metabarcoding analyses of fungi. MycoKeys 2015, 10. [Google Scholar] [CrossRef]
- Tedersoo, L.; Lindahl, B. Fungal identification biases in microbiome projects. Environmental Microbiology Reports 2016, 8, 774–779. [Google Scholar] [CrossRef]
- Martin, K.J.; Rygiewicz, P.T. Fungal-specific pcr primers developed for analysis of the its region of environmental DNA extracts. BMC Microbiology 2005, 5, 28. [Google Scholar] [CrossRef]
- Langarica-Fuentes, A.; Zafar, U.; Heyworth, A.; Brown, T.; Fox, G.; Robson, G.D. Fungal succession in an in-vessel composting system characterized using 454 pyrosequencing. FEMS Microbiology Ecology 2014, 88, 296–308. [Google Scholar] [CrossRef]
- U’Ren, J.M.; Riddle, J.M.; Monacell, J.T.; Carbone, I.; Miadlikowska, J.; Arnold, A.E. Tissue storage and primer selection influence pyrosequencing-based inferences of diversity and community composition of endolichenic and endophytic fungi. Molecular Ecology Resources 2014, 14, 1032–1048. [Google Scholar] [CrossRef]
- Cuadros-Orellana, S.; Leite, L.R.; Smith, A.; Medeiros, J.D.; Badotti, F.; Fonseca, P.L.; Vaz, A.B.; Oliveira, G.; Góes-Neto, A. Assessment of fungal diversity in the environment using metagenomics: A decade in review. Fungal Genomics & Biology 2013, 3, 1. [Google Scholar]
- Bálint, M.; Schmidt, P.-A.; Sharma, R.; Thines, M.; Schmitt, I. An illumina metabarcoding pipeline for fungi. Ecology and Evolution 2014, 4, 2642–2653. [Google Scholar] [CrossRef] [PubMed]
- Gweon, H.S.; Oliver, A.; Taylor, J.; Booth, T.; Gibbs, M.; Read, D.S.; Griffiths, R.I.; Schonrogge, K. Pipits: An automated pipeline for analyses of fungal internal transcribed spacer sequences from the illumina sequencing platform. Methods in ecology and evolution 2015, 6, 973–980. [Google Scholar] [CrossRef] [PubMed]
- Hibbett, D.; Abarenkov, K.; Kõljalg, U.; Öpik, M.; Chai, B.; Cole, J.; Wang, Q.; Crous, P.; Robert, V.; Helgason, T.; et al. Sequence-based classification and identification of fungi. Mycologia 2016, 108, 1049–1068. [Google Scholar] [CrossRef]
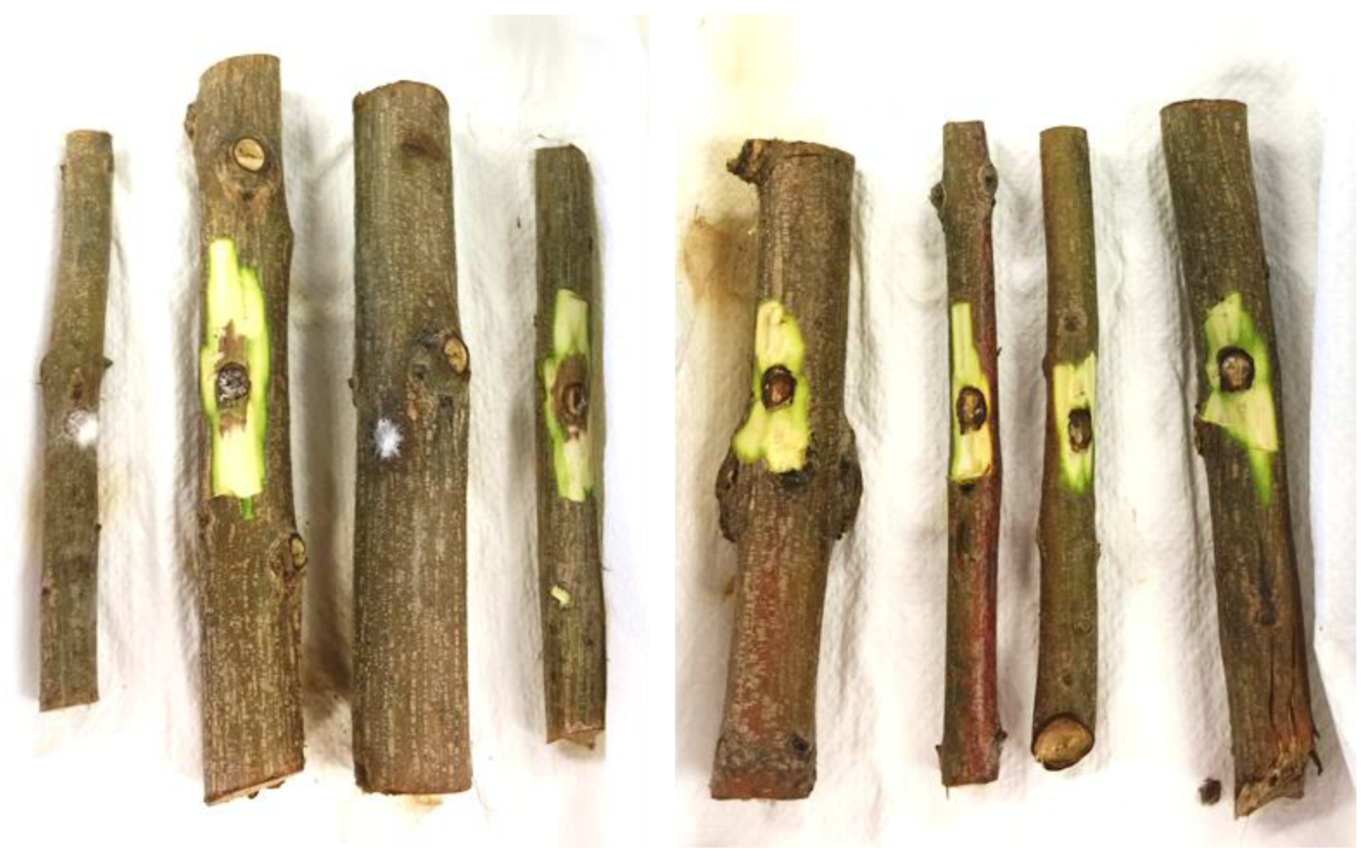
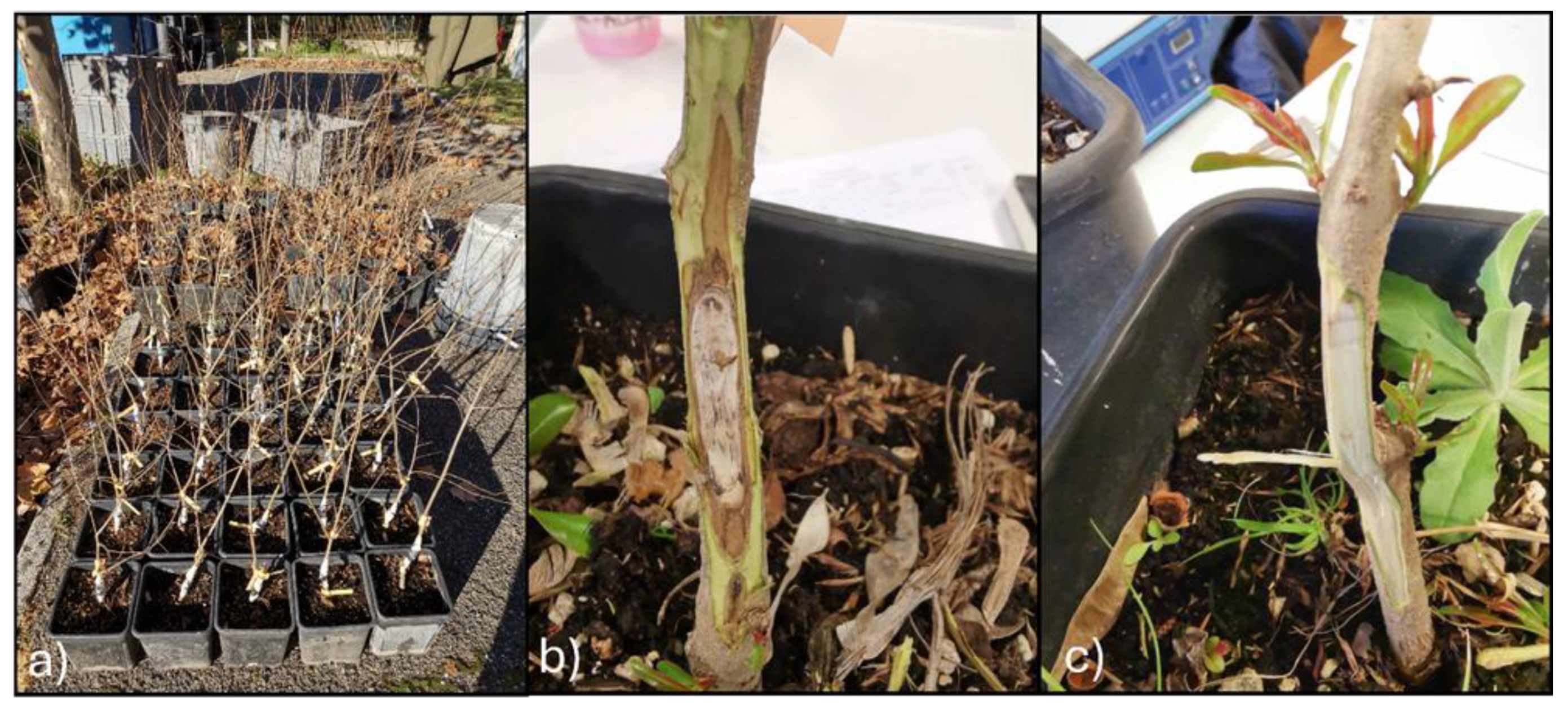

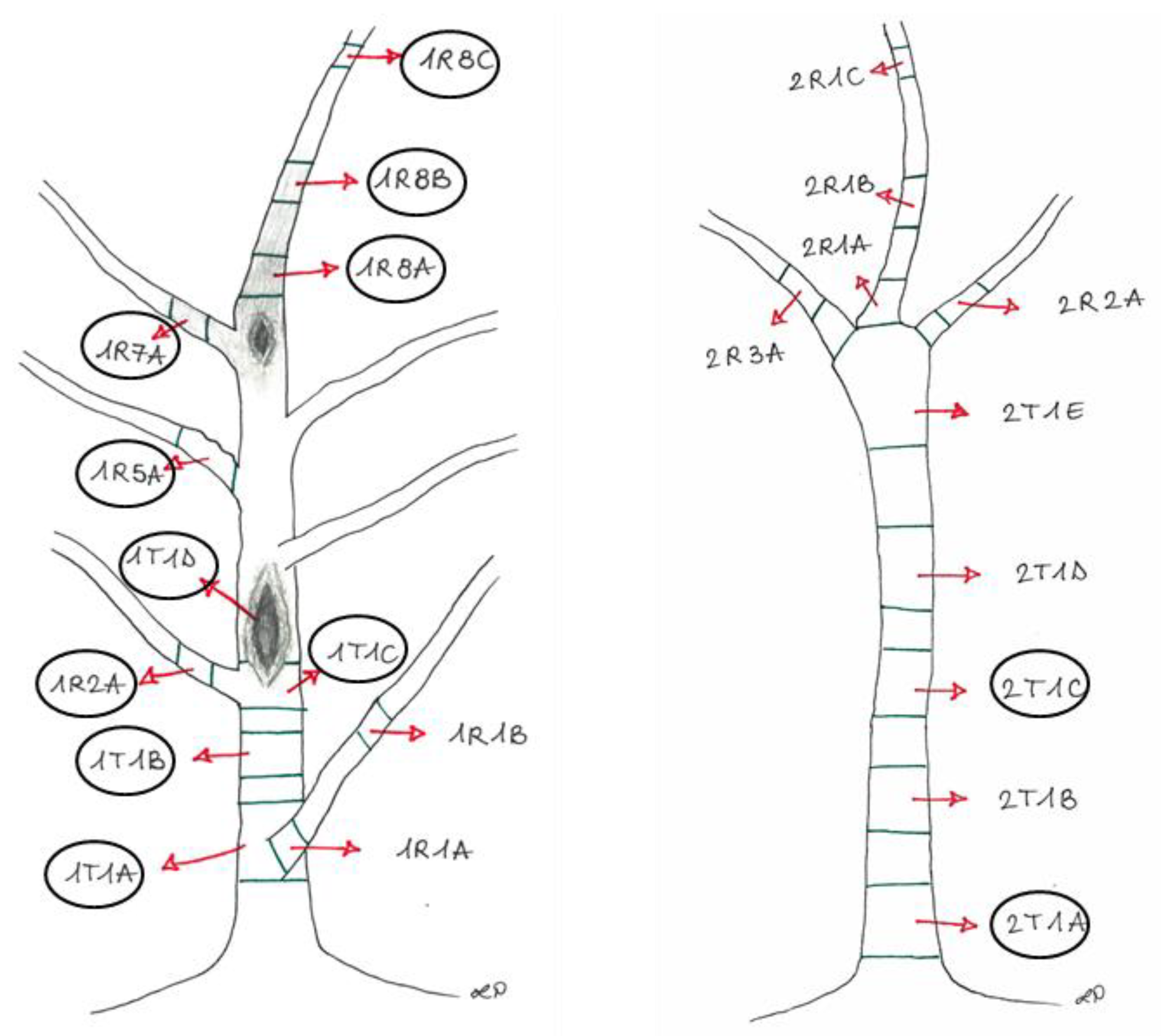



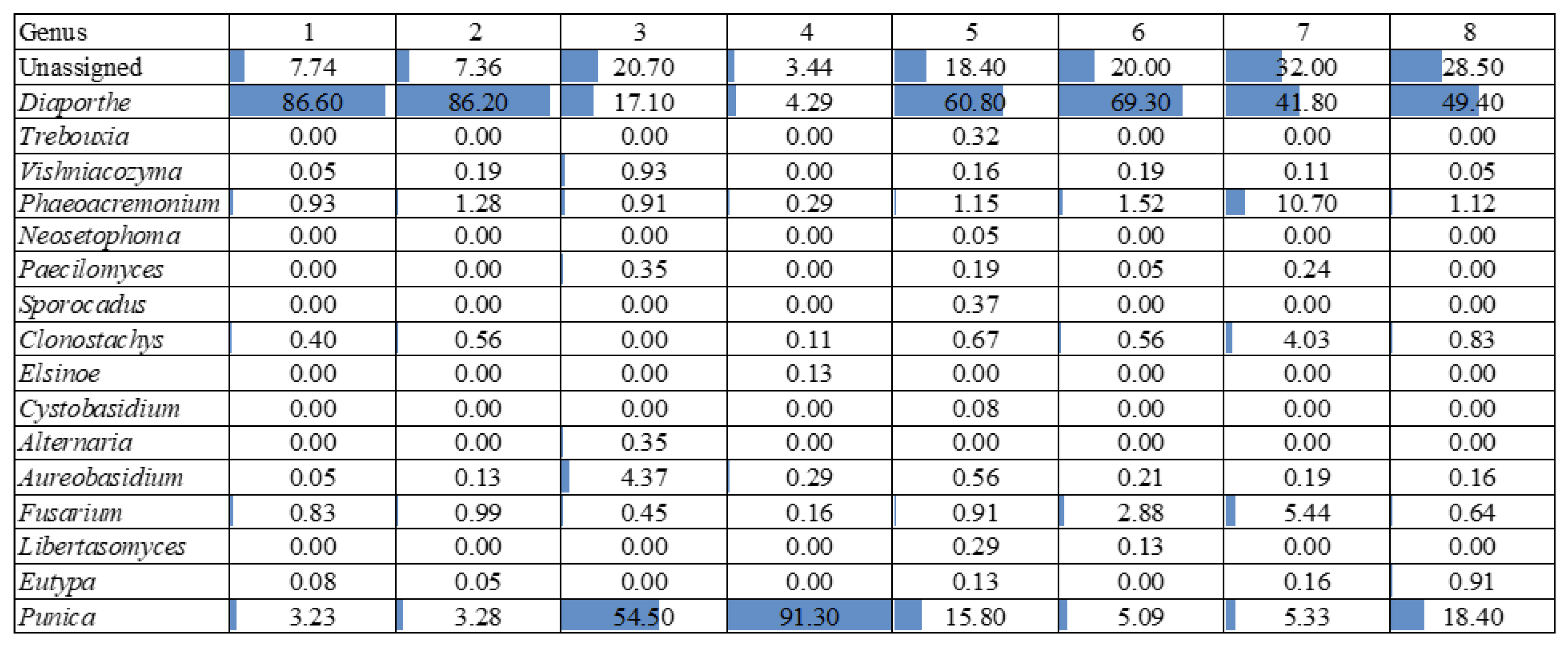
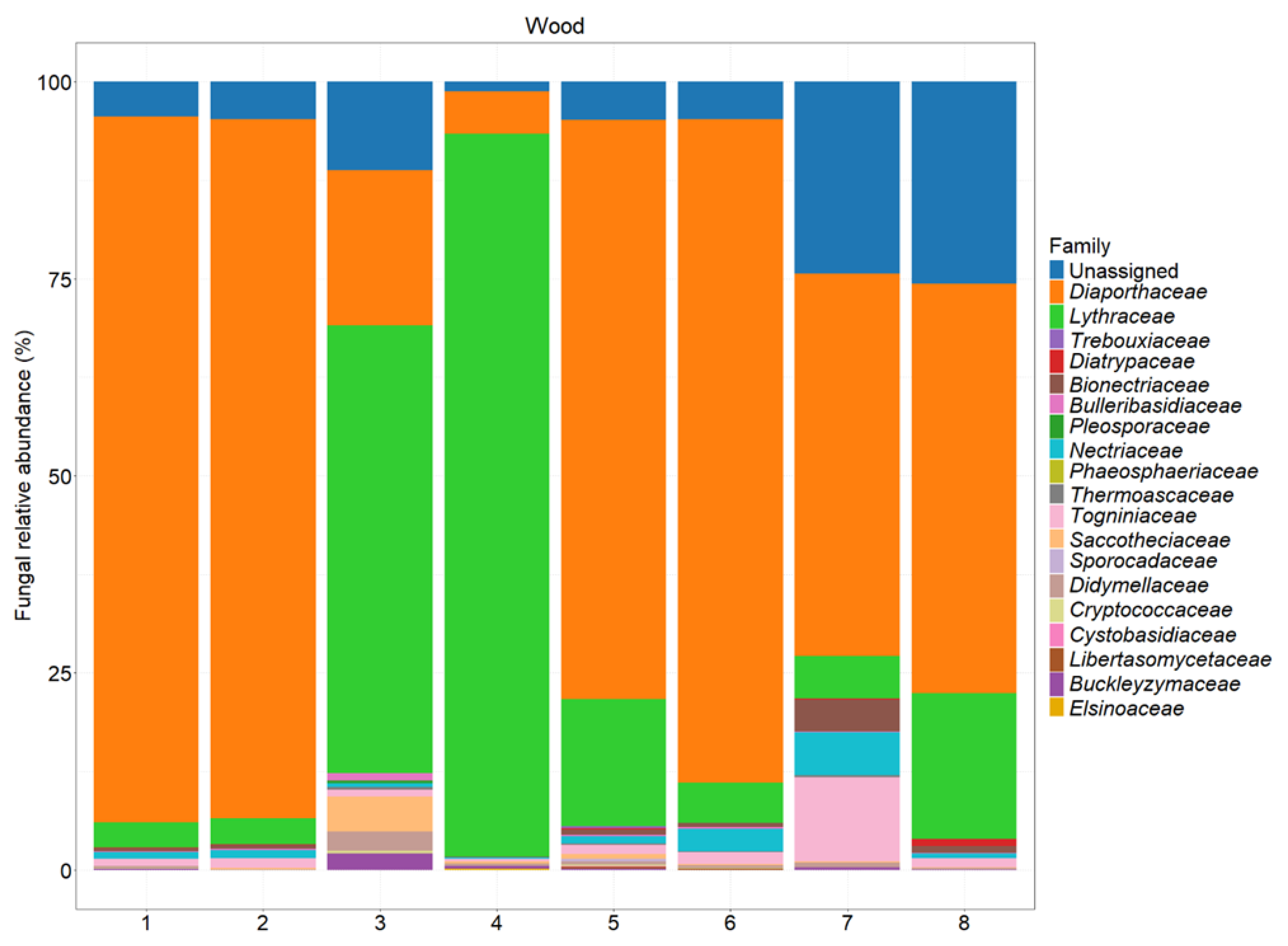
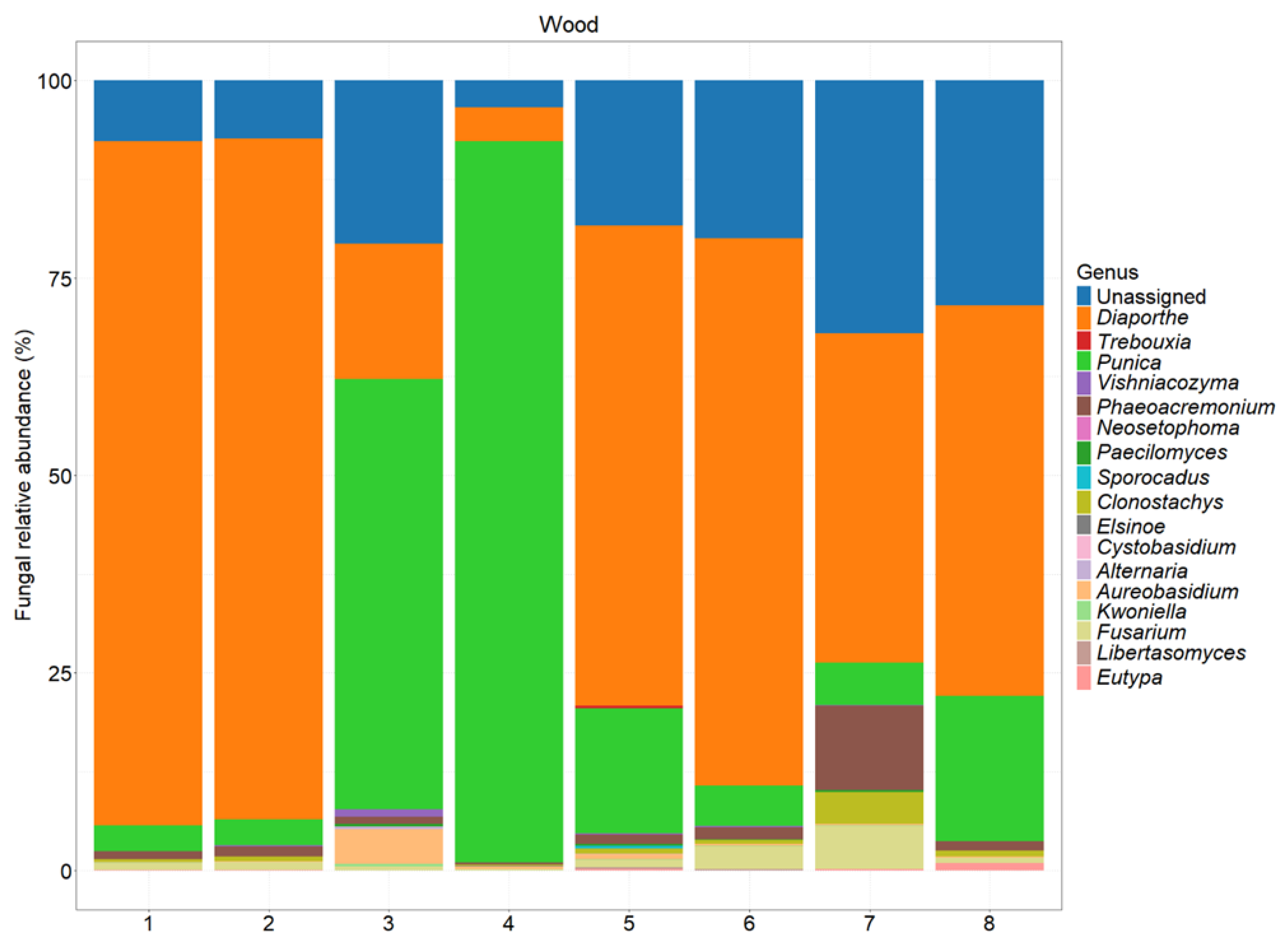


| Experimental Units | Section | Phenotypic Features |
|---|---|---|
| 1 | 1R8A+1R8B | Brown tissues above the canker from the apical portion (Plant 1) |
| 2 | 1R7A | Brown tissue from the lateral branch close to the canker (Plant 1) |
| 3 | 1R2A+1R8C | Asymptomatic tissues from branches (Plant 1) |
| 4 | 1R5A | Asymptomatic tissue (showing bark roughness) from the lateral branch (Plant 1) |
| 5 | 1T1A | Brown tissue from the basal area of the trunk, corresponding to the collar (Plant 1) |
| 6 | 1T1B | Brown tissue below the canker from the trunk (Plant 1) |
| 7 | 1T1C+1T1D | Brown tissues into the canker area (Plant 1) |
| 8 | 2T1A+2T1C | Brown tissues from the trunk (Plant 2) |
| Strain CREA-DC Collection Code | Species | Isolation incidence (% of the total fungal colonies) | Pavona | Sermoneta | GenBank Accession Number | |||
|---|---|---|---|---|---|---|---|---|
| July 2017 | October 2017 | March 2018* | September 2018 | February 2019 | ||||
| ER2123 | N. parvum | 27.6 | Absent | Present | Absent | Present | Absent | MW020287 |
| ER2111 | D. eres | 20.7 | Present | Absent | Present | Absent | Absent | MW020285 |
| ER2125, ER2128 | D. foeniculina | 17.2 | Absent | Present | Present | Absent | Absent | MW020288, MW032268 |
| ER2126 | D. phaseolorum | 3.4 | Absent | Present | Absent | Absent | Absent | MW020289 |
| ER2127 | D. rudis | 3.4 | Absent | Present | Present | Absent | Absent | MW032267 |
| ER2145 | Cytospora acaciae | 3.4 | Absent | Absent | Absent | Present | Absent | / |
| / | Eutypa lata | 0.5 | Absent | Absent | Present | Absent | Absent | / |
| ER2130 | Gliocladium spp. | 24.1 | Absent | Present | Present | Present | Present | / |
| / | Phoma glomerata | 5.3 | Absent | Absent | Present | Absent | Present | / |
| / | Neopestalotiopsis clavispora | 1 | Absent | Absent | Present | Absent | Absent | / |
| / | Fusarium spp. | 18.6 | Absent | Absent | Present | Absent | Absent | / |
| EFFECT | SS | DF | MS | F | ProbF |
|---|---|---|---|---|---|
| Isolate | 1.49215 | 5 | 0.29843 | 13.64701997 | 3.93174E-05 |
| Residual | 0.328016667 | 15 | 0.021867778 | ||
| Total | 1.94685 | 23 | 0.084645652 |
| EFFECT | SS | DF | MS | F | ProbF |
|---|---|---|---|---|---|
| Isolate | 8.419153439 | 5 | 1.683831 | 14.04785 | 4.26E-06 |
| Residual | 2.517142857 | 21 | 0.119864 | ||
| Total | 10.9362963 | 26 | 0.420627 |
| EFFECT | SS | DF | MS | F | ProbF |
|---|---|---|---|---|---|
| Isolate | 16.79066667 | 5 | 3.358133 | 9.631358 | 3.84E-05 |
| Residual | 8.368 | 24 | 0.348667 | ||
| Total | 25.15866667 | 29 | 0.86754 | ||
| C.V. (%): 20,4082959922632 | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




