Submitted:
05 February 2025
Posted:
06 February 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
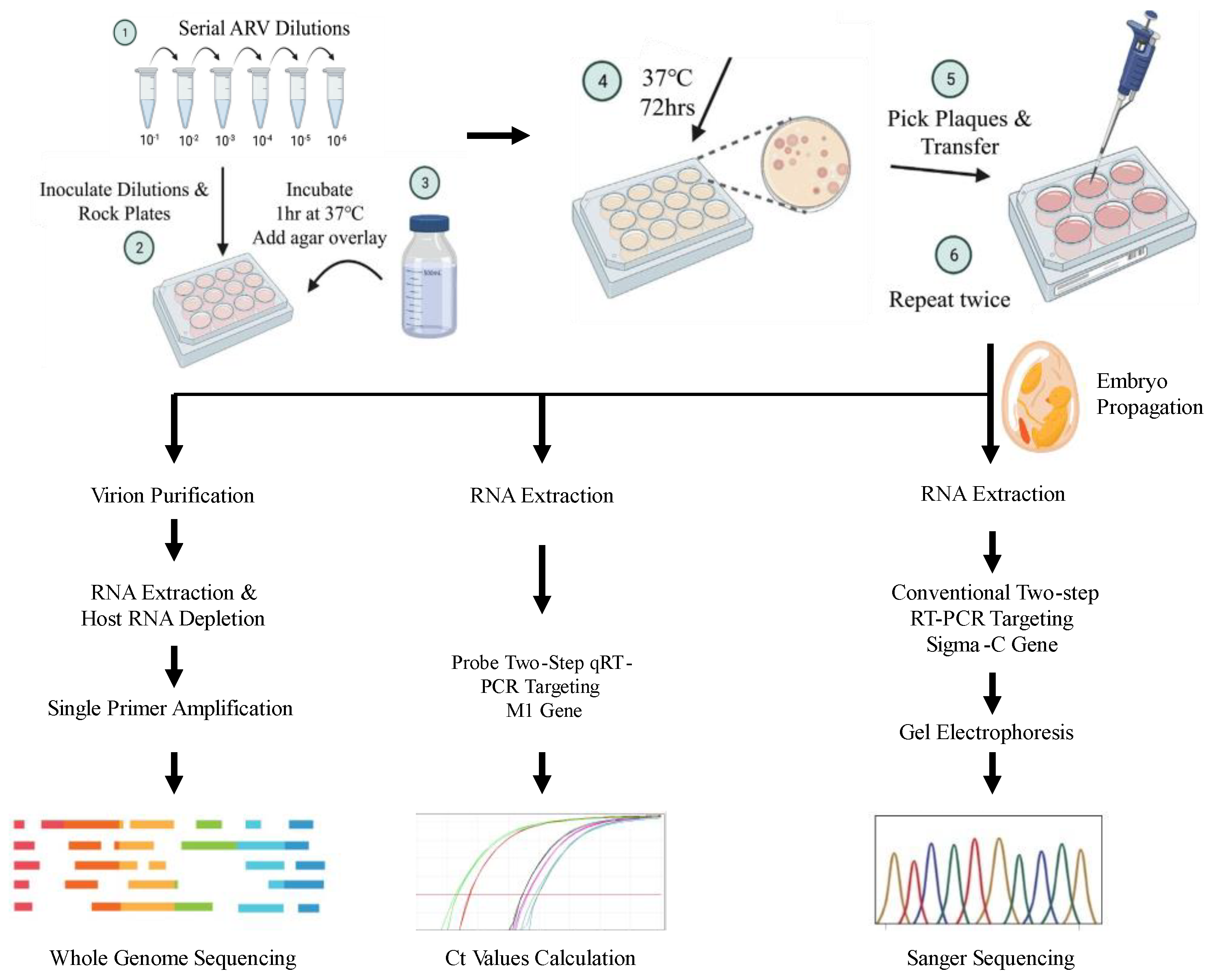
2. Materials and Methods
Clinical History of AL Isolate
Chicken Embryo Liver Cell Culture
Plaque Purification of AL Isolate
Preparation of Viral Stocks in Chicken Embryos
Viral RNA Detection and Quantification
Sanger Sequencing and Data Analysis
Whole Genome Sequencing: Library Preparation
Whole Genome Sequencing: Data Analysis
3. Results
qPCR Targeting ARV M1 Gene Quantified RNA of Both AL and S1133 Isolates
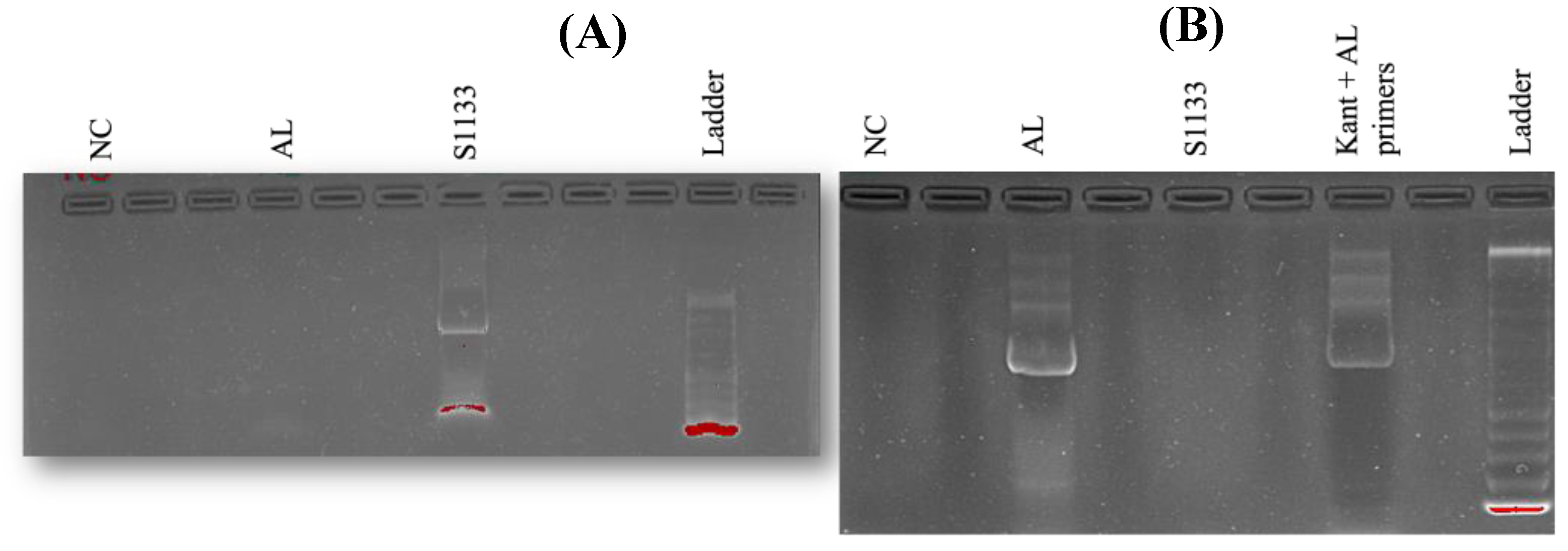
PCR Using Kant Primer Amplified S1133 and Sanger Sequencing Confirmed Identity with S1133
Whole Genome Sequencing: Read Quality
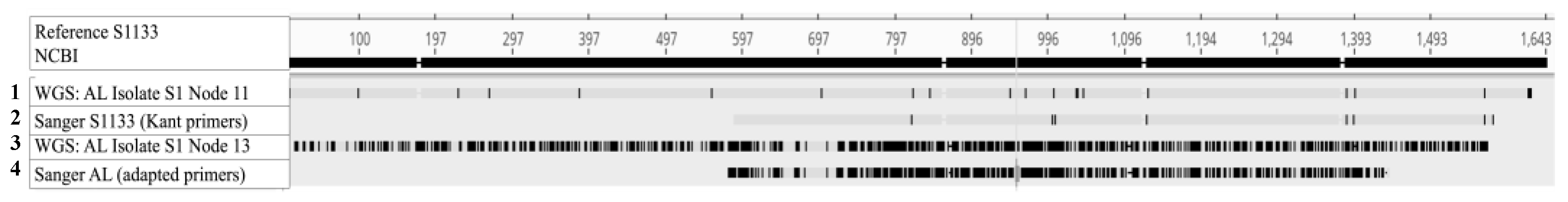
WGS: Contigs Similar and Divergent from the Reference S1133 Were Found
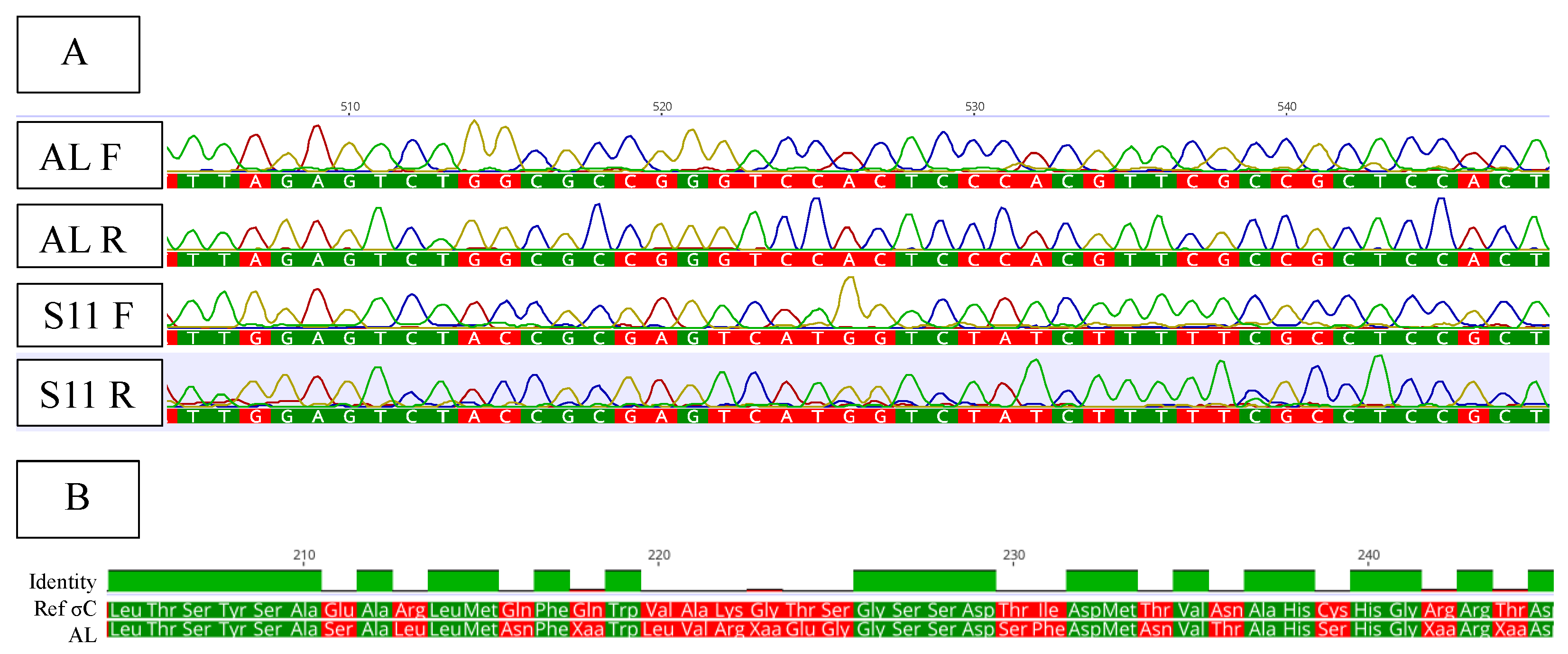
Primers Designed Based on WGS Amplified σC Gene of AL Isolate
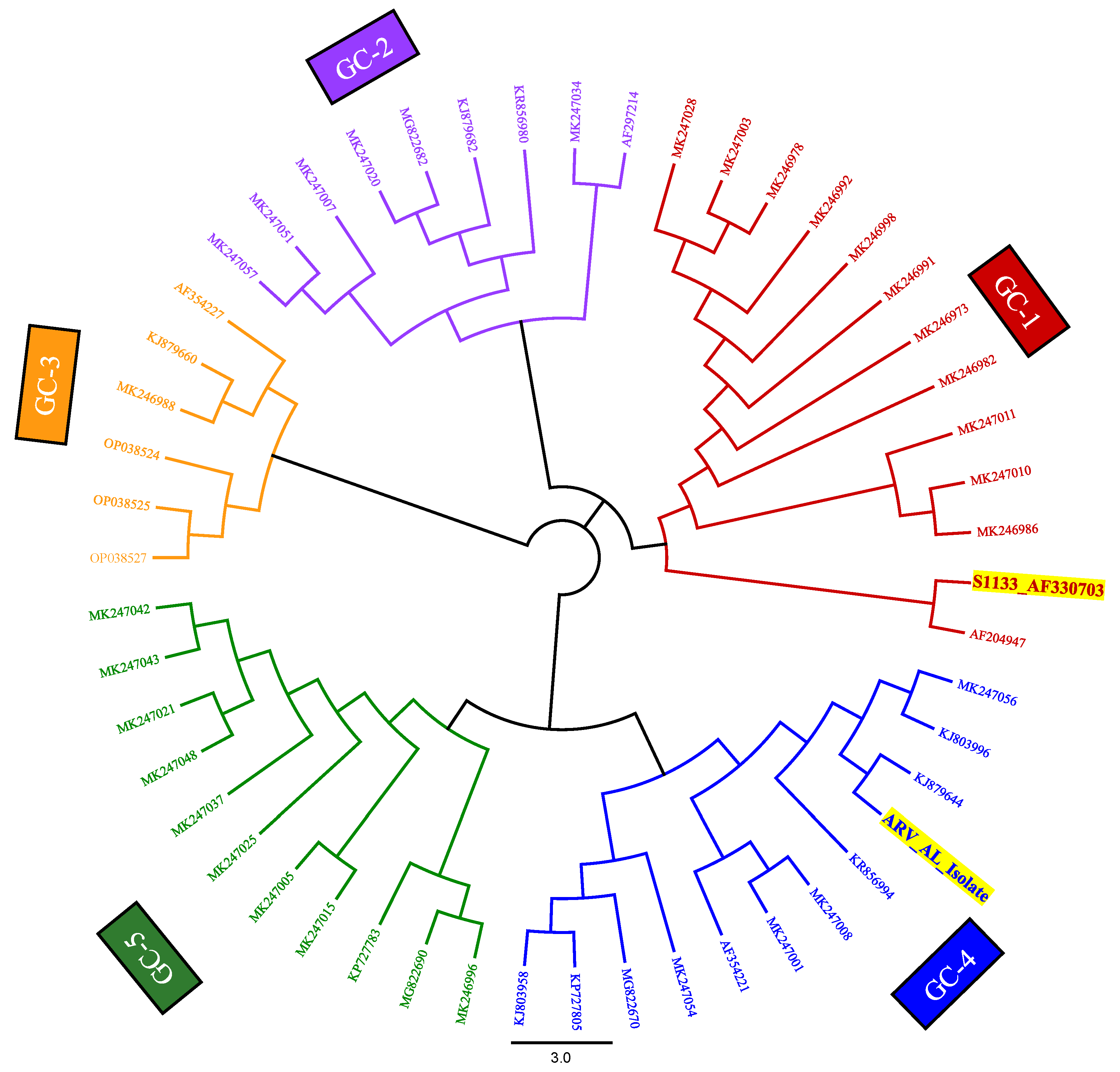
Phylogenetic Analysis Confirmed a Mixture of Divergent Sequences
4. Discussion
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kovács, E.; Varga-Kugler, R.; Mató, T.; Homonnay, Z.; Tatár-Kis, T.; Farkas, S.; Kiss, I.; Bányai, K.; Palya, V. Identification of the Main Genetic Clusters of Avian Reoviruses from a Global Strain Collection. Front. Vet. Sci. 2023, 9, 1094761. [Google Scholar] [CrossRef] [PubMed]
- French, D. Incidence and Economic Impact of Reovirus in the Poultry Industries in the United States. Avian Dis. 2022, 66, 432–434. [Google Scholar] [CrossRef] [PubMed]
- Spandidos, D. and; Graham, A.F. Physical and Chemical Characterization of an Avian Reovirus. J. Virol. 1976, 19, 968–976. [Google Scholar] [CrossRef] [PubMed]
- Ayalew, L.E.; Ahmed, K.A.; Mekuria, Z.H.; Lockerbie, B.; Popowich, S.; Tikoo, S.K.; Ojkic, D.; Gomis, S. The Dynamics of Molecular Evolution of Emerging Avian Reoviruses through Accumulation of Point Mutations and Genetic Re-Assortment. Virus Evol. 2020, 6, veaa025. [Google Scholar] [CrossRef]
- Kant, A.; Balk, F.; Born, L.; van Roozelaar, D.; Heijmans, J.; Gielkens, A.; ter Huurne, A. Classification of Dutch and German Avian Reoviruses by Sequencing the Sigma C Protein. Vet. Res. 2003, 34, 203–212. [Google Scholar] [CrossRef]
- Goldenberg, D.; Pasmanik-Chor, M.; Pirak, M.; Kass, N.; Lublin, A.; Yeheskel, A.; Heller, D.; Pitcovski, J. Genetic and Antigenic Characterization of Sigma C Protein from Avian Reovirus. Avian Pathol 2010, 39, 189–199. [Google Scholar] [CrossRef]
- Narvaez, S.A.; Harrell, T.L.; Oluwayinka, O.L.T.D.; Sellers, H.S.; Khalid, Z.; Hauck, R.; Chowdhury, E.U.; Conrad, S.J. Optimizing the Conditions for Whole-Genome Sequencing of Avian Reoviruses. Viruses 2023, 15, 1938. [Google Scholar] [CrossRef]
- Nour, I.; Alvarez-Narvaez, S.; Harrell, T.L.; Conrad, S.J.; Mohanty, S.K. Whole Genomic Constellation of Avian Reovirus Strains Isolated from Broilers with Arthritis in North Carolina, USA. Viruses 2023, 15. [Google Scholar] [CrossRef]
- Egaña-Labrin, S.; Hauck, R.; Figueroa, A.; Stoute, S.; Shivaprasad, H.L.; Crispo, M.; Corsiglia, C.; Zhou, H.; Kern, C.; Crossley, B.; et al. Genotypic Characterization of Emerging Avian Reovirus Genetic Variants in California. Sci. Rep. 2019, 9, 9351. [Google Scholar] [CrossRef]
- Rafique, S.; Rashid, F.; Wei, Y.; Zeng, T.T.; Xie, L.J.; Xie, Z.X. Avian Orthoreoviruses: A Systematic Review of Their Distribution, Dissemination Patterns, and Genotypic Clustering. Viruses 2024, 16, 1056. [Google Scholar] [CrossRef]
- Meanger, J.; Wickramasinghe, R.; Enriquez, C.E.; Robertson, M.D.; Wilcox, G.E. Type-specific Antigenicity of Avian Reoviruses. Avian Pathol. 1995, 24, 121–134. [Google Scholar] [CrossRef] [PubMed]
- Goldenberg, D.; Lublin, A.; Rosenbluth, E.; Heller, E.D.; Pitcovski, J. Differentiating Infected from Vaccinated Animals, and among Virulent Prototypes of Reovirus. J. Virol. Methods 2011, 177, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.H.; Lee, L.H.; Shih, W.L.; Hu, Y.C.; Liu, H.J. Baculovirus Surface Display of σC and σB Proteins of Avian Reovirus and Immunogenicity of the Displayed Proteins in a Mouse Model. Vaccine 2008, 26, 6361–6367. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.-H.; Hsu, A.-P.; Shien, J.-H.; Chang, T.-J.; Liao, J.-W.; Chen, J.-R.; Lin, C.-F.; Hsu, W.-L. Avian Reovirus Sigma C Enhances the Mucosal and Systemic Immune Responses Elicited by Antigen-Conjugated Lactic Acid Bacteria. Vaccine 2012, 30, 5019–5029. [Google Scholar] [CrossRef]
- Goldenberg, D.; Lublin, A.; Rosenbluth, E.; Heller, E.D.; Pitcovski, J. Optimized Polypeptide for a Subunit Vaccine against Avian Reovirus. Vaccine 2016, 34, 3178–3183. [Google Scholar] [CrossRef]
- Saikia, D.P.; Yadav, K.; Pathak, D.C.; Ramamurthy, N.; D’Silva, A.L.; Marriappan, A.K.; Ramakrishnan, S.; Vakharia, V.N.; Chellappa, M.M.; Dey, S. Recombinant Newcastle Disease Virus (NDV) Expressing Sigma C Protein of Avian Reovirus (ARV) Protects against Both ARV and NDV in Chickens. Pathogens 2019, 8. [Google Scholar] [CrossRef]
- Clark, F.D.; Ni, Y.; Collisson, E.W.; Kemp, M.C. Characterization of Avian Reovirus Strain-Specific Polymorphisms. Avian Dis. 1990, 34, 304–314. [Google Scholar] [CrossRef]
- Vasserman, Y.; Eliahoo, D.; Hemsani, E.; Kass, N.; Ayali, G.; Pokamunski, S.; Pitcovski, J. The Influence of Reovirus Sigma C Protein Diversity on Vaccination Efficiency. Avian Dis. 2004, 48, 271–278. [Google Scholar] [CrossRef]
- Dawe, W.H.; Kapczynski, D.R.; Linnemann, E.G.; Gauthiersloan, V.R.; Sellers, H.S. Analysis of the Immune Response and Identification of Antibody Epitopes against the Sigma C Protein of Avian Orthoreovirus Following Immunization with Live or Inactivated Vaccines. Avian Dis. 2023, 66, 465–475. [Google Scholar] [CrossRef]
- Egaña-Labrin, S.; Gallardo, R.A. Concomitant Virus Detection on Avian Reovirus Isolates. In Proceedings of the Proceedings of Western Poultry Disease Conference.
- Tran, A.; Berard, A.; Coombs, K.M. Avian Reoviruses: Propagation, Quantification, and Storage. In Current Protocols in Microbiology; Coico, R., McBride, A., Quarles, J.M., Stevenson, B., Taylor, R.K., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2009; ISBN 978-0-471-72925-9. [Google Scholar]
- Rekik, M.R.; Silim, A.; Elazhary, M.A.S.Y. Characteristics and Analysis of Electropherotypes of Avian Reovirus Field Isolates. Spec. Issue: Adv. Vet. Virol. 1990, 23, 273–281. [Google Scholar] [CrossRef]
- Wu, W.Y.; Shien, J.H.; Lee, L.H.; Shieh, H.K. Analysis of the Double-Stranded RNA Genome Segments among Avian Reovirus Field Isolates. J. Virol. Methods 1994, 48, 119–122. [Google Scholar] [CrossRef] [PubMed]
- Guneratne, J.R.M.; Jones, R.C.; Georgiou, K. Some Observations on the Isolation and Cultivation of Avian Reoviruses. Avian Pathol. 1982, 11, 453–462. [Google Scholar] [CrossRef] [PubMed]
- van Loon, A.A.W.M.; Koopman, H.C.; Kosman, W.; Mumczur, J.; Szeleszczuk, O.; Karpinska, E.; Kosowska, G.; Lütticken, D. Virology: Isolation of a New Serotype of Avian Reovirus Associated with Malabsorption Syndrome in Chickens. Vet. Q 2001, 23, 129–133. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Lu, H.G. Whole Genome Alignment Based One-Step Real-Time RT-PCR for Universal Detection of Avian Orthoreoviruses of Chicken, Pheasant and Turkey Origins. Infect., Genet. Evol. 2016, 39, 120–126. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: A Multiple Sequence Alignment Method with Reduced Time and Space Complexity. BMC Bioinf. 2004, 5, 113. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Tamura, K.; Nei, M. Estimation of the Number of Nucleotide Substitutions in the Control Region of Mitochondrial DNA in Humans and Chimpanzees. Mol. Biol. Evol. 1993, 10, 512–526. [Google Scholar] [CrossRef]
- Alvarez-Narvaez, S.; Harrell, T.L.; Conrad, S.J. Protocol: Optimized Conditions for Whole Genome Sequencing of Avian Reoviruses 2024.
- Parris, D.J.; Kariithi, H.; Suarez, D.L. Non-Target RNA Depletion Strategy to Improve Sensitivity of next-Generation Sequencing for the Detection of RNA Viruses in Poultry. J. Vet. Diagn. Invest. 2022, 34, 638–645. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data 2010.
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and Accurate Long-Read Alignment with Burrows–Wheeler Transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve Years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative Genomics Viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.R.; Schultz, M.B.; Zobel, J.; Holt, K.E. Bandage: Interactive Visualization of de Novo Genome Assemblies. Bioinformatics 2015, 31, 3350–3352. [Google Scholar] [CrossRef]
- Murata, H.; Macauley, J.; Lewis, A.M.; Peden, K. Plaque Purification as a Method to Mitigate the Risk of Adventitious-Agent Contamination in Influenza Vaccine Virus Seeds. Vaccine 2011, 29, 3155–3161. [Google Scholar] [CrossRef]
- Markis, M. In Vivo and in Vitro Characterization of Virus Isolates Involved in Viral Enteritis/Enteropathy, Also Known as Runting Stunting Syndrome (Rss), in Broiler Chickens. Ph.D., University of Delaware, 2013.
- Shapouri, M.R.S.; Arella, M.; Silim, A. Evidence for the Multimeric Nature and Cell Binding Ability of Avian Reovirus Σ3 Protein. J. Gen. Virol. 1996, 77, 1203–1210. [Google Scholar] [CrossRef]
- Rekik, M.R.; Silim, A.; Bernier, G. Serological and Pathogenic Characterization of Avian Reoviruses Isolated in Quebec. Avian Pathol. 1991, 20, 607–617. [Google Scholar] [CrossRef]
- Aguilera, E.R.; Erickson, A.K.; Jesudhasan, P.R.; Robinson, C.M.; Pfeiffer, J.K. Plaques Formed by Mutagenized Viral Populations Have Elevated Coinfection Frequencies. mBio 2017, 8, e02020–16. [Google Scholar] [CrossRef]
- Ni, Y.W.; Kemp, M.C. A Comparative Study of Avian Reovirus Pathogenicity: Virus Spread and Replication and Induction of Lesions. Avian Dis. 1995, 39, 554–566. [Google Scholar] [CrossRef]
- Deshmukh, D.R.; Dutta, S.K.; Pomeroy, B.S. Avian Reoviruses V. Studies of Ultrastructural Morphology by Electron Microscopy. Avian Dis. 1971, 15, 588–595. [Google Scholar] [CrossRef]
- Gouvea, V.S.; Schnitzer, T.J. Polymorphism of the Migration of Double-Stranded RNA Genome Segments of Avian Reoviruses. J. Virol. 1982, 43, 465–471. [Google Scholar] [CrossRef] [PubMed]
- Masaji Mase; Makiko Gotou; Daisuke Inoue; Tsuneyuki Masuda; Satoko Watanabe; Hiroshi Iseki Genetic Analysis of Avian Reovirus Isolated from Chickens in Japan. Avian Dis. 2021, 65, 346–350. [CrossRef]
- Farnoushi, Y.; Heller, D.; Lublin, A. Genetic Characterization of Newly Emerging Avian Reovirus Variants in Chickens with Viral Arthritis/Tenosynovitis in Israel. Virology 2024, 589, 109908. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.-J.; Giambrone, J.J. Amplification, Cloning and Sequencing of the σC-Encoded Gene of Avian Reovirus. J. Virol. Methods 1997, 63, 203–208. [Google Scholar] [CrossRef] [PubMed]
- Wickramasinghe, R.; Meanger, J.; Enriquez, C.E.; Wilcox, G.E. Avian Reovirus Proteins Associated with Neutralization of Virus Infectivity. Virology 1993, 194, 688–696. [Google Scholar] [CrossRef]
- Kim, S.-W.; Choi, Y.-R.; Park, J.-Y.; Wei, B.; Shang, K.; Zhang, J.-F.; Jang, H.-K.; Cha, S.-Y.; Kang, M. Isolation and Genomic Characterization of Avian Reovirus from Wild Birds in South Korea. Front. Vet. Sci. 2022, 9. [Google Scholar] [CrossRef]
- De Carli, S.; Wolf, J.M.; Gräf, T.; Lehmann, F.K.M.; Fonseca, A.S.K.; Canal, C.W.; Lunge, V.R.; Ikuta, N. Genotypic Characterization and Molecular Evolution of Avian Reovirus in Poultry Flocks from Brazil. Avian Pathol. 2020, 49, 611–620. [Google Scholar] [CrossRef]
- Hsu, H.W.; Su, H.Y.; Huang, P.H.; Lee, L.H.; Liu, H.J. Sequence and Phylogenetic Analysis of P10- and P17-Encoding Genes of Avian Reovirus. Avian Dis. 2005, 49, 36–42. [Google Scholar] [CrossRef]
- Teng, L.; Xie, Z.; Xie, L.; Liu, J.; Pang, Y.; Deng, X.; Xie, Z.; Fan, Q.; Luo, S.; Feng, J.; et al. Sequencing and Phylogenetic Analysis of an Avian Reovirus Genome. Virus Genes 2014, 48, 381–386. [Google Scholar] [CrossRef]
- Egaña-Labrin, S.; Jerry, C.; Roh, H.J.; Silva, A.P. da; Corsiglia, C.; Crossley, B.; Rejmanek, D.; Gallardo, R.A. Avian Reoviruses of the Same Genotype Induce Different Pathology in Chickens. Avian Dis. 2021, 65, 529–539. [Google Scholar] [CrossRef]

| Technique | Primers | S1133 Isolate | Plaque-purified AL isolate |
|---|---|---|---|
| Conventional PCR targeting σC gene | Kant et. al, 2003 | + | - |
| Sanger sequencing | Kant et. al, 2003 | + | - |
| Conventional PCR targeting σC gene | Designed de novo for AL based on WGS | - | + |
| Sanger sequencing | Designed de novo for AL based on WGS | - | + |
| Quantitative PCR targeting M1 gene | Tang and Lu, 2016 | + | + |
| Whole genome sequencing | Single primer amplification approach | S1133-like gene segments only | Both AL & S1133 gene segments |
| Genome segment | Pairwise identity of AL isolate contigs to S1133 | ||
| Contig 1 | Contig 2 | Contig 3 | |
| L1 | 91.2 | - | - |
| L2 | 88.4 | 88.2 | - |
| L3 | 96.2 | - | - |
| M1 | 93.5 | 94.4 | 93.2 |
| M2 | 83.3 | 83.2 | - |
| M3 | 89.5 | 89.4 | 87.9 |
| S1 | 55.8 | 90.9 | 93.3 |
| S2 | 93.2 | 93.7 | - |
| S3 | 97.6 | - | - |
| S4 | 90 | 89.1 | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




