Submitted:
02 January 2025
Posted:
03 January 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
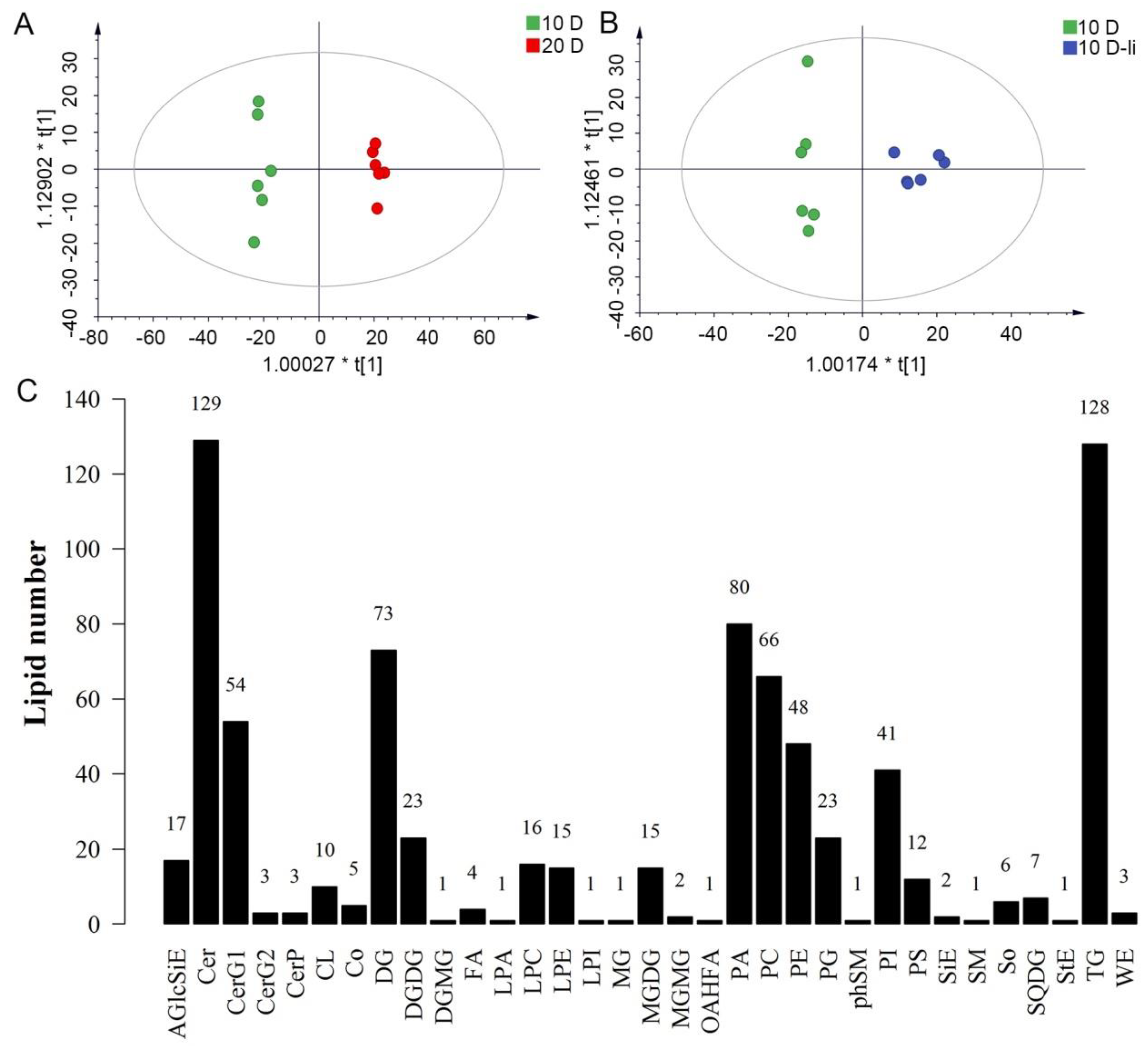
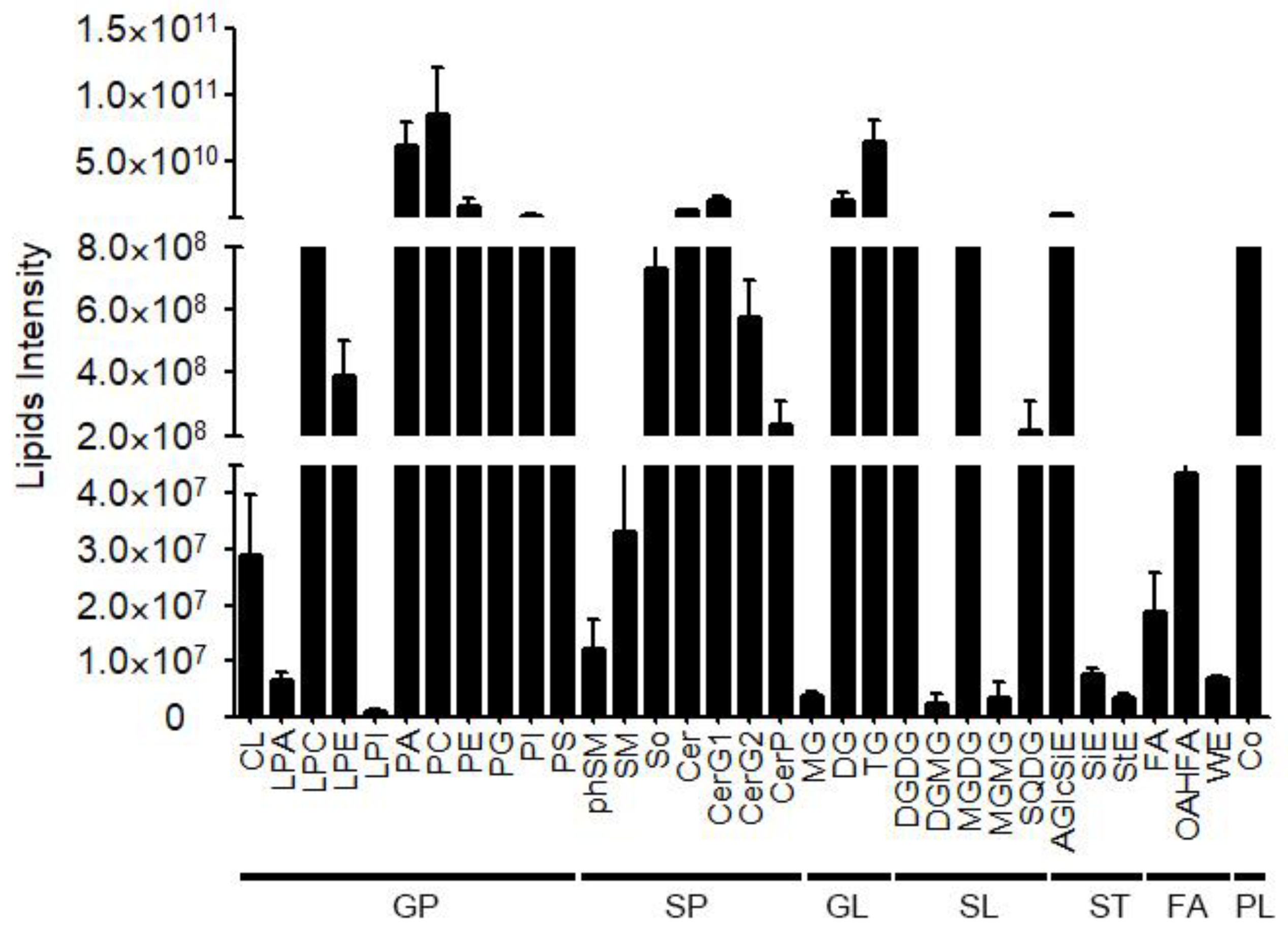
2.1. Untargeted Lipidomics Analysis in Fiber Cells
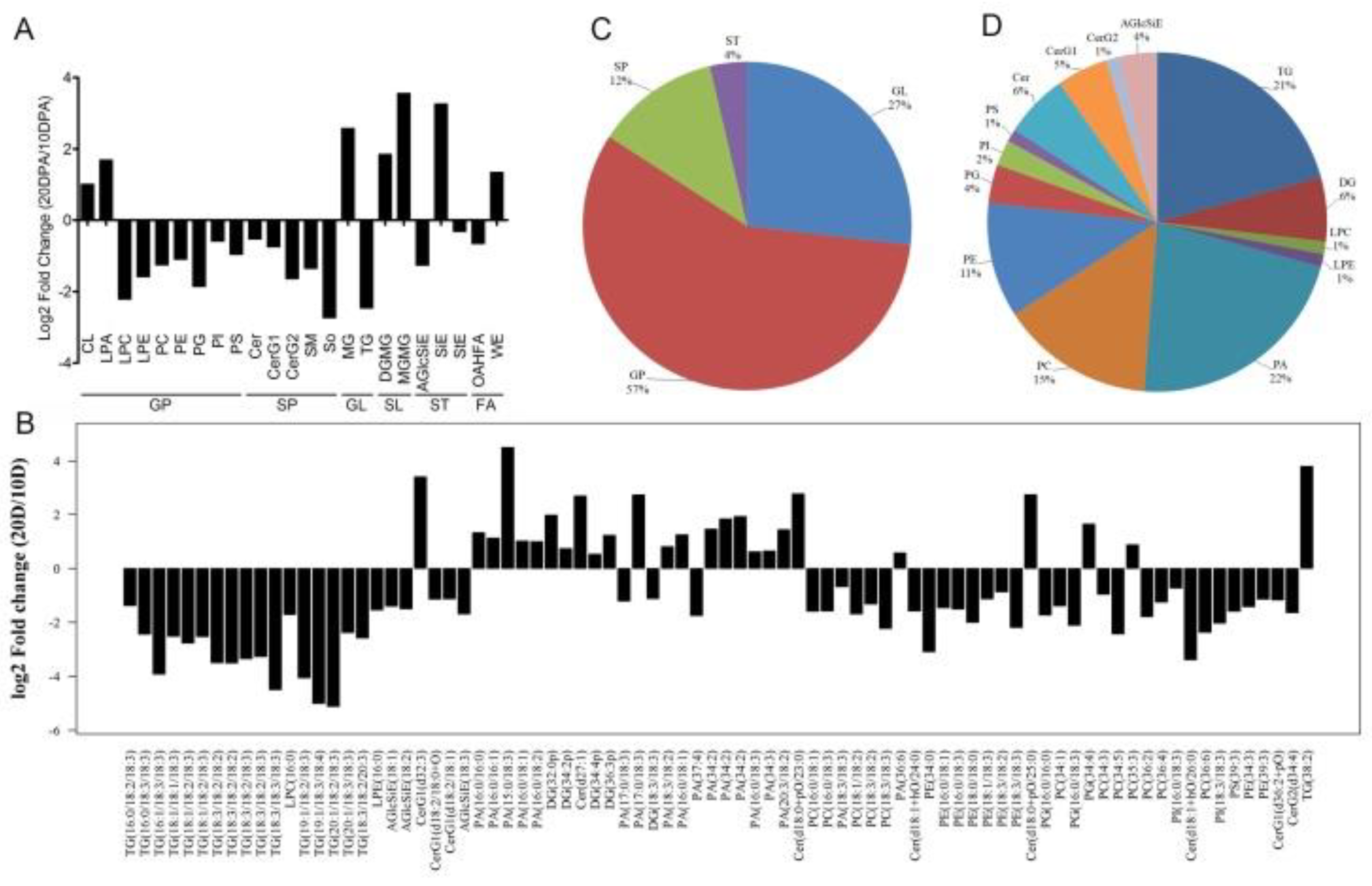
2.2. The Lipid Difference Between the Stage of Rapid Elongation and SCW Deposition of Fiber Cells
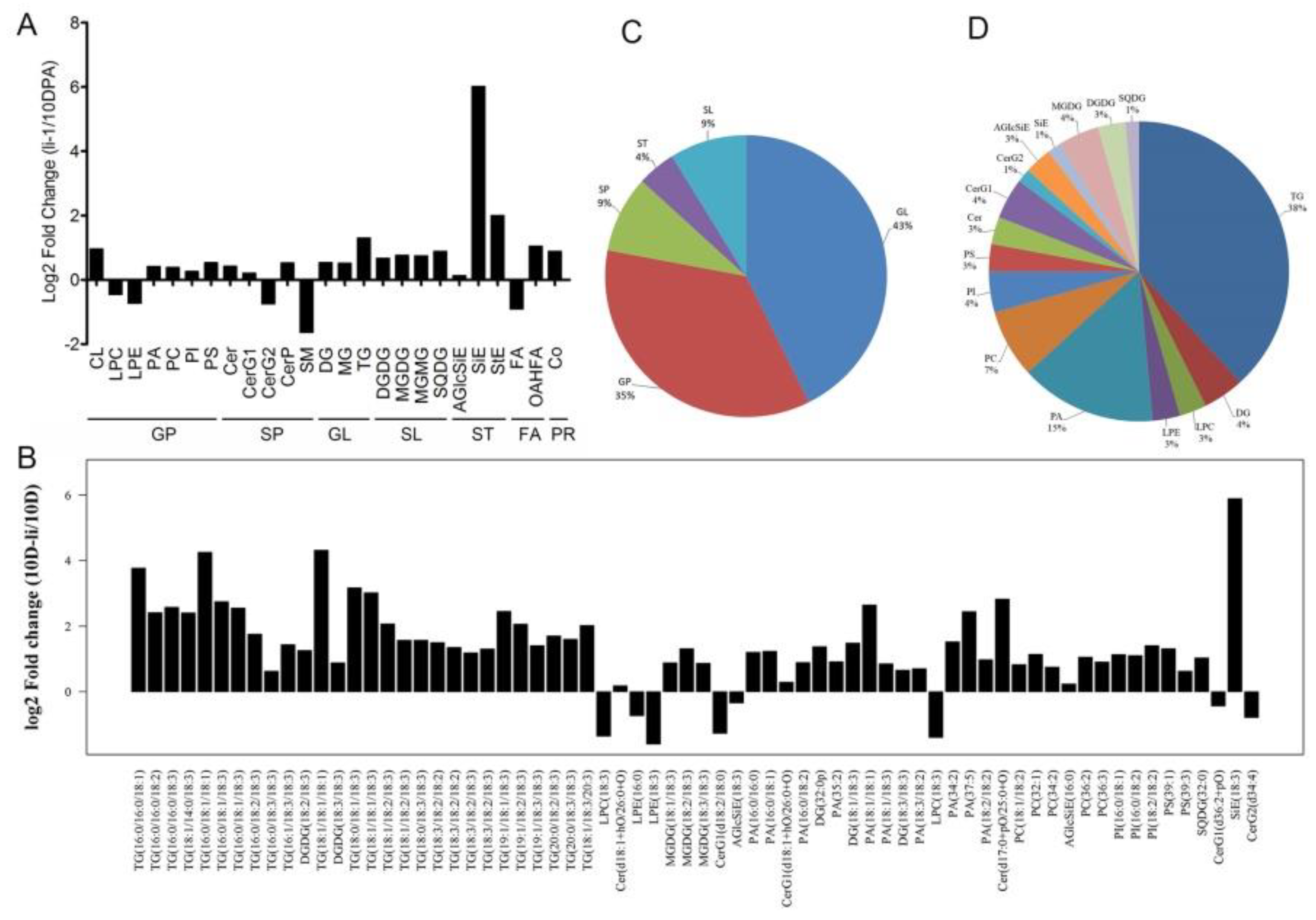
2.3. The Lipid Change of Fiber Cell of li-1 Mutant
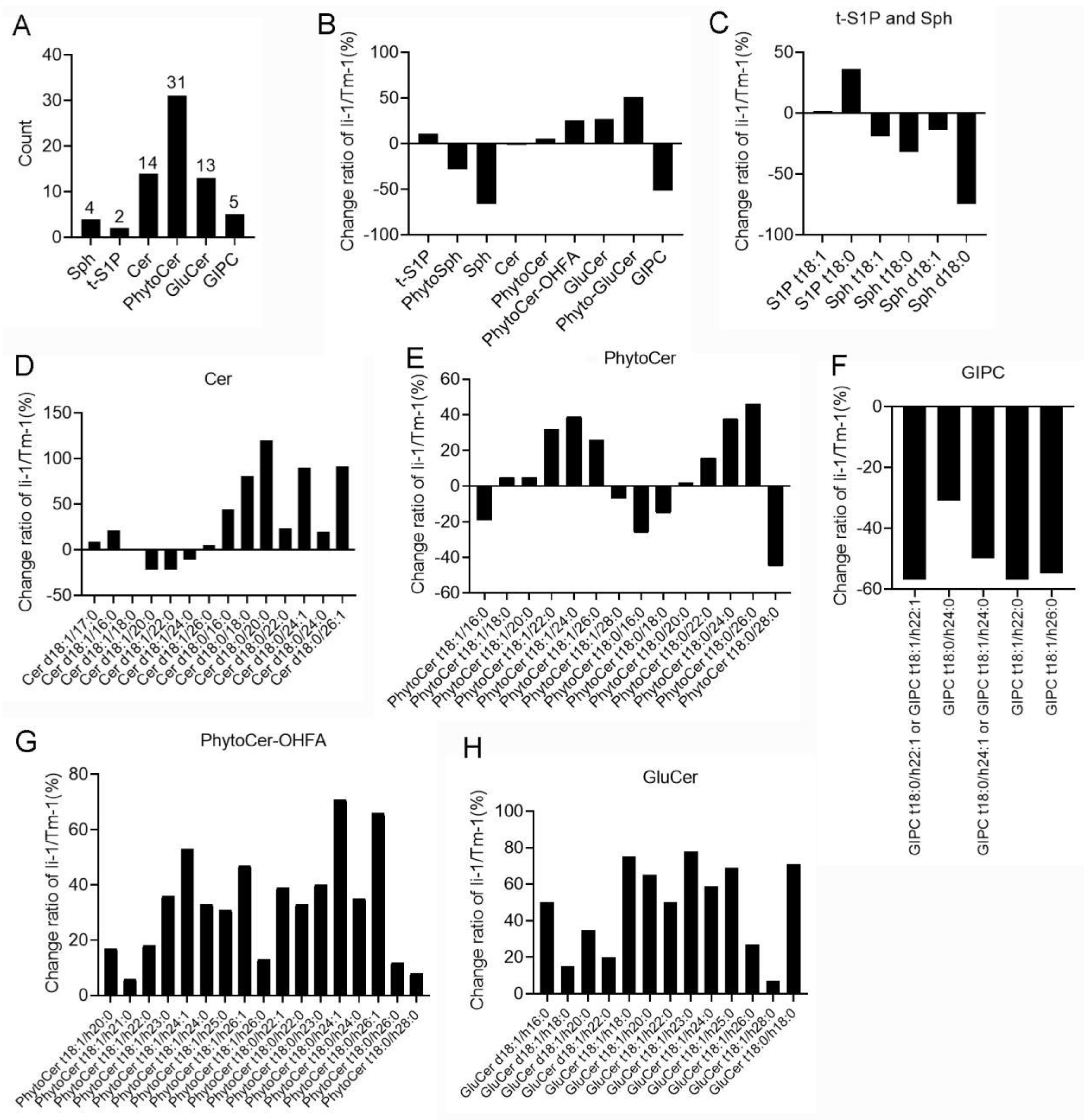
2.4. The Changes of Sphingolipids in li-1 Mutant Fiber Cells
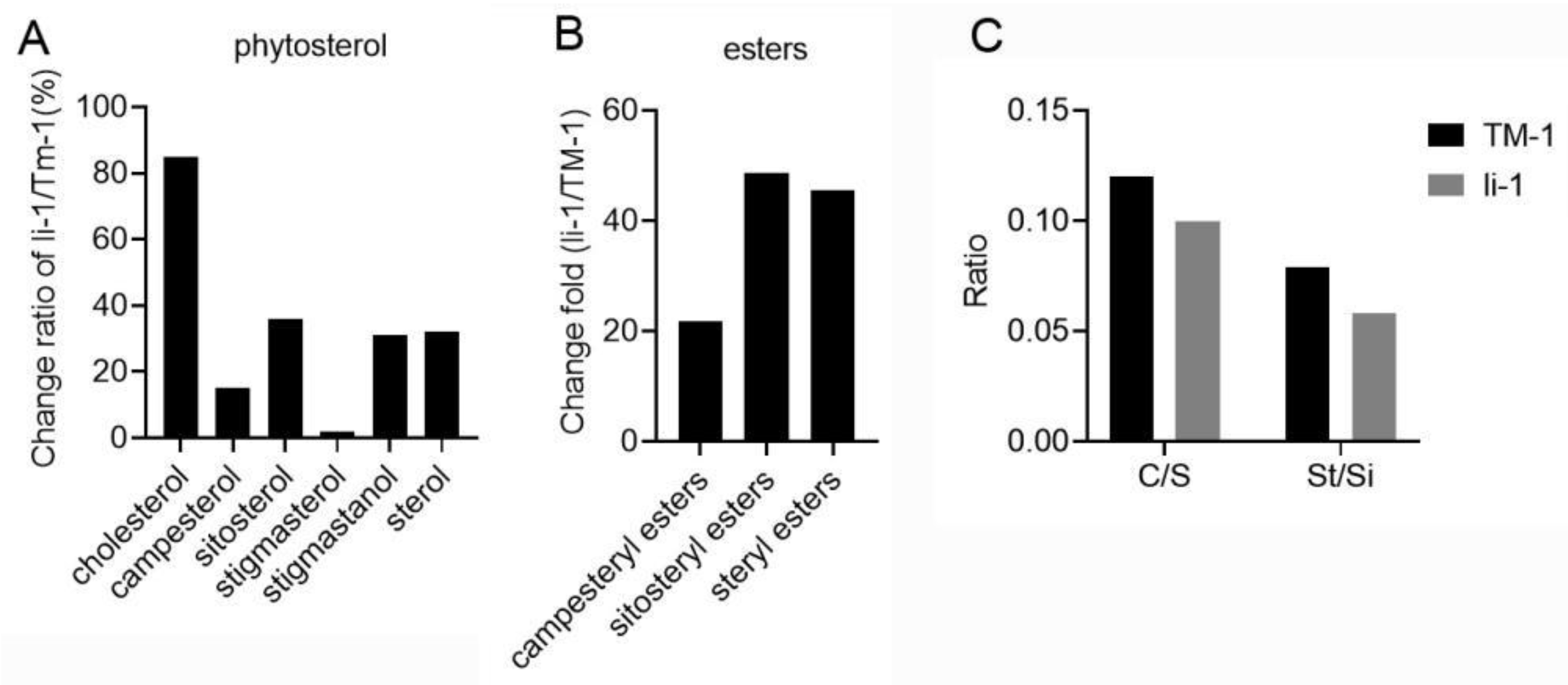
2.5. The Changes of Phytosterols in li-1 Mutant Fiber Cells
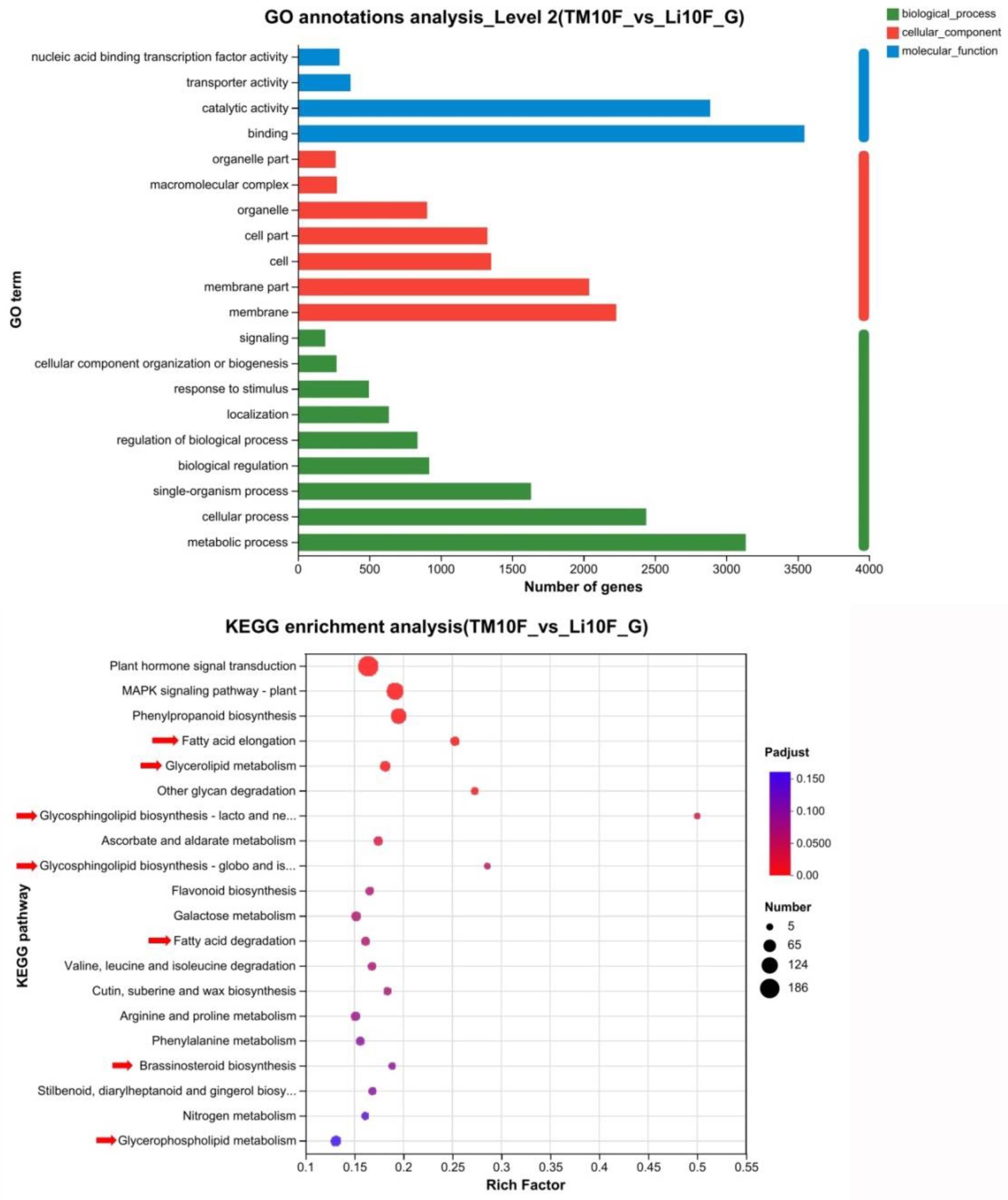
2.6. The Expression of Genes Involved in Lipid Metabolism Was Altered in li-1 Fiber Cells
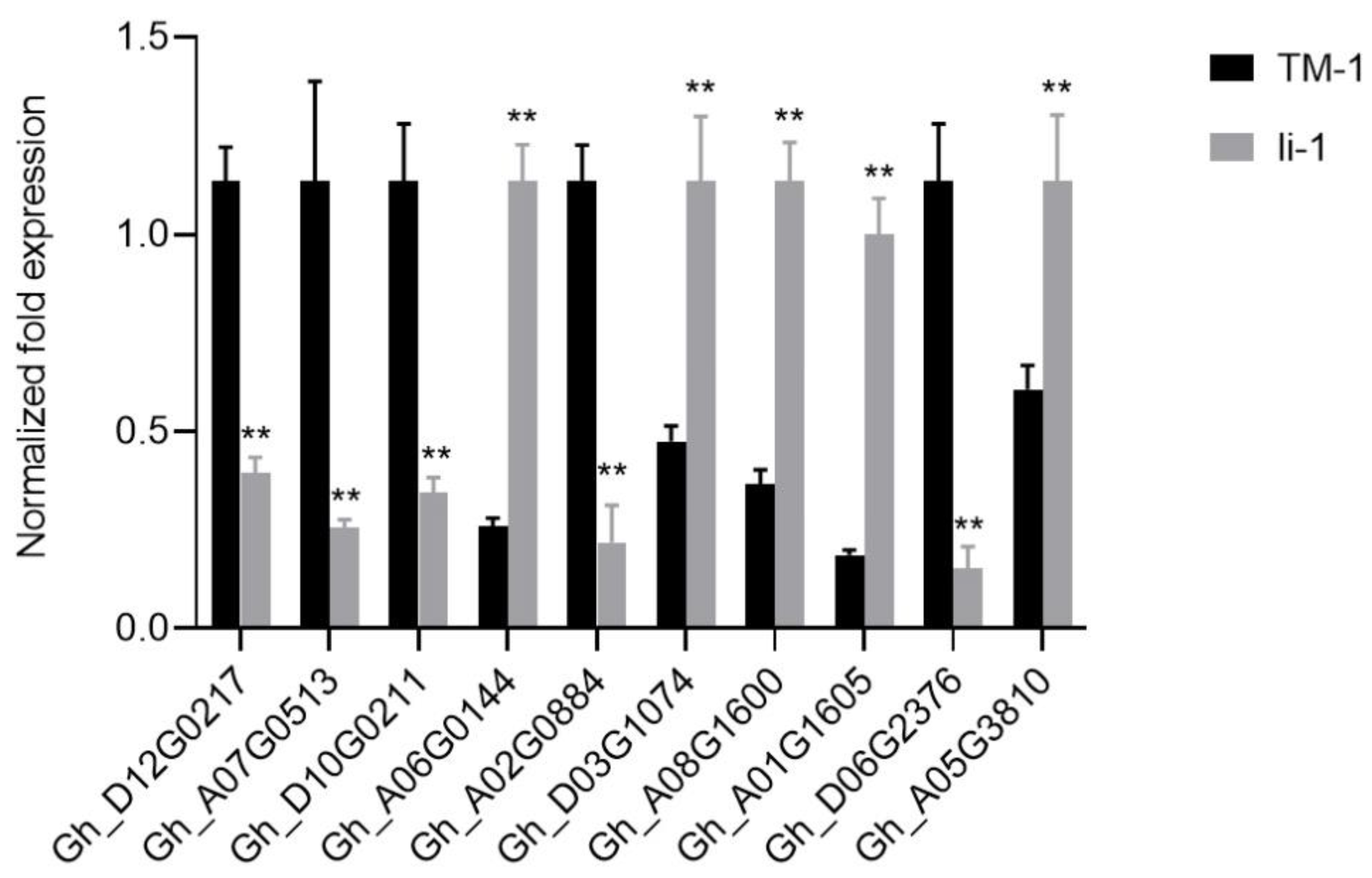
2.7. The Expression Levels of Key Genes in Lipid Metabolism Were Elevated in li-1 Fiber Cells
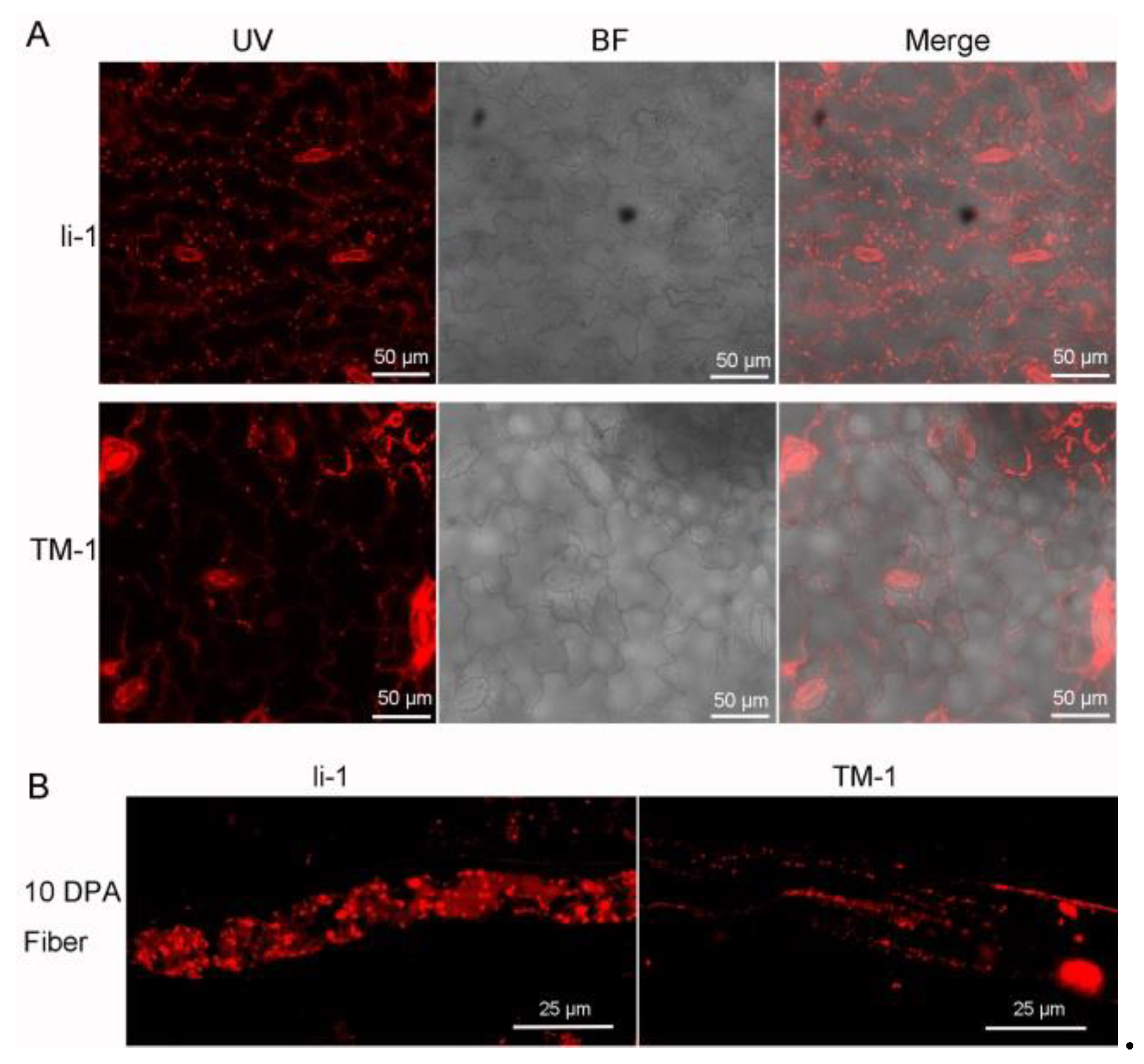
2.8. The Number of Oil Body Was Increased in li-1 Leaf and Fiber Cell
3. Discussion
3.1. The Role of Lipids in Fiber Elongation and SCW Deposition
3.2. The Lipid Metabolism Disruption in the li-1 Mutant Fibers
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. RNA Extraction and qRT-PCR Assay
4.3. Lipid Extraction and Lipidomics
4.4. The Nile Red Stain
4.5. Statistical Analysis
4.6. Bioinformatic Analysis
5. Conclusions
5.1. The Role of Lipids in Fiber Elongation and SCW Deposition
5.2. The Lipid Metabolism Disruption in the li-1 Mutant Fibers
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kim HJ, Triplett BA: Cotton fiber growth in planta and in vitro. Models for plant cell elongation and cell wall biogenesis. Plant physiology 2001, 127(4):1361-1366. [CrossRef]
- Qin YM, Zhu YX: How cotton fibers elongate: a tale of linear cell-growth mode. Current opinion in plant biology 2011, 14(1):106-111. [CrossRef]
- Haigler CH, Betancur L, Stiff MR, Tuttle JR: Cotton fiber: a powerful single-cell model for cell wall and cellulose research. Frontiers in plant science 2012, 3. [CrossRef]
- Stiff MR, Haigler CH: Cotton fiber tips have diverse morphologies and show evidence of apical cell wall synthesis. Scientific reports 2016, 6. [CrossRef]
- Yang Z, Zhang C, Yang X, Liu K, Wu Z, Zhang X, Zheng W, Xun Q, Liu C, Lu L: PAG1, a cotton brassinosteroid catabolism gene, modulates fiber elongation. New Phytologist 2014, 203(2):437-448. [CrossRef]
- Kohel RJ, Quisenberry JE, Benedict CR: Fiber Elongation and Dry Weight Changes in Mutant Lines of Cotton1. Crop Sci 1974, 14(3):cropsci1974.0011183X001400030040x. [CrossRef]
- Karaca M, Saha S, Jenkins JN, Zipf A, Kohel R, Stelly DM: Simple sequence repeat (SSR) markers linked to the Ligon lintless (Li(1)) mutant in cotton. The Journal of heredity 2002, 93(3):221-224. [CrossRef]
- Kohel RJ, Benedict CR, Jividen GM: Incorporation of [14C]Glucose into Crystalline Cellulose in Aberrant Fibers of a Cotton Mutant. Crop Sci 1993, 33(5):cropsci1993.0011183X003300050032x. [CrossRef]
- Ohlrogge J, Browse J: Lipid biosynthesis. The Plant Cell 1995, 7(7):957-970. [CrossRef]
- Sanjaya, Miller R, Durrett TP, Kosma DK, Lydic TA, Muthan B, Koo AJK, Bukhman YV, Reid GE, Howe GA et al.: Altered Lipid Composition and Enhanced Nutritional Value of Arabidopsis Leaves following Introduction of an Algal Diacylglycerol Acyltransferase 2. Plant Cell 2013, 25(2):677-693. [CrossRef]
- Farmer EE, Weber H, Vollenweider S: Fatty acid signaling in Arabidopsis. Planta 1998, 206(2):167-174. [CrossRef]
- Liu G-J, Xiao G-H, Liu N-J, Liu D, Chen P-S, Qin Y-M, Zhu Y-X: Targeted Lipidomics Studies Reveal that Linolenic Acid Promotes Cotton Fiber Elongation by Activating Phosphatidylinositol and Phosphatidylinositol Monophosphate Biosynthesis. Mol Plant 2015, 8(6):911-921. [CrossRef]
- Ji SJ, Lu YC, Feng JX, Wei G, Li J, Shi YH, Fu Q, Liu D, Luo JC, Zhu YX: Isolation and analyses of genes preferentially expressed during early cotton fiber development by subtractive PCR and cDNA array. Nucleic acids research 2003, 31(10):2534-2543. [CrossRef]
- Shi YH, Zhu SW, Mao XZ, Feng JX, Qin YM, Zhang L, Cheng J, Wei LP, Wang ZY, Zhu YX: Transcriptome profiling, molecular biological, and physiological studies reveal a major role for ethylene in cotton fiber cell elongation. Plant Cell 2006, 18(3):651-664. [CrossRef]
- Yang SS, Cheung F, Lee JJ, Ha M, Wei NE, Sze SH, Stelly DM, Thaxton P, Triplett B, Town CD et al.: Accumulation of genome-specific transcripts, transcription factors and phytohormonal regulators during early stages of fiber cell development in allotetraploid cotton. Plant Journal 2006, 47(5):761-775. [CrossRef]
- Gou JY, Wang LJ, Chen SP, Hu WL, Chen XY: Gene expression and metabolite profiles of cotton fiber during cell elongation and secondary cell wall synthesis. Cell Res 2007, 17(5):422-434. [CrossRef]
- Qin YM, Hu CY, Pang Y, Kastaniotis AJ, Hiltunen JK, Zhu YX: Saturated very-long-chain fatty acids promote cotton fiber and Arabidopsis cell elongation by activating ethylene biosynthesis. Plant Cell 2007, 19(11):3692-3704. [CrossRef]
- Qin YM, Pujol FM, Hu CY, Feng JX, Kastaniotis AJ, Hiltunen JK, Zhu YX: Genetic and biochemical studies in yeast reveal that the cotton fibre-specific GhCER6 gene functions in fatty acid elongation. Journal of Experimental Botany 2007, 58(3):473-481. [CrossRef]
- Zhu L, Dou L, Shang H, Li H, Yu J, Xiao G: GhPIPLC2D promotes cotton fiber elongation by enhancing ethylene biosynthesis. iScience 2021, 24(3):102199. [CrossRef]
- Wanjie SW, Welti R, Moreau RA, Chapman KD: Identification and quantification of glycerolipids in cotton fibers: Reconciliation with metabolic pathway predictions from DNA databases. Lipids 2005, 40(8):773-785. [CrossRef]
- Chen M, Cahoon EB, Saucedo-García M, Plasencia J, Gavilanes-Ruíz M: Plant Sphingolipids: Structure, Synthesis and Function. In: Lipids in Photosynthesis: Essential and Regulatory Functions. Edited by Wada H, Murata N. Dordrecht: Springer Netherlands; 2009: 77-115.
- Lynch DV, Dunn TM: An introduction to plant sphingolipids and a review of recent advances in understanding their metabolism and function. New Phytol 2004, 161(3):677-702. [CrossRef]
- Markham JE, Jaworski JG: Rapid measurement of sphingolipids from Arabidopsis thaliana by reversed-phase high-performance liquid chromatography coupled to electrospray ionization tandem mass spectrometry. Rapid Commun Mass Sp 2007, 21(7):1304-1314. [CrossRef]
- Xu F, Suo X, Li F, Bao C, He S, Huang L, Luo M: Membrane lipid raft organization during cotton fiber development. Journal of Cotton Research 2020, 3(1):13. [CrossRef]
- Xu F, Chen Q, Huang L, Luo M: Advances about the Roles of Membranes in Cotton Fiber Development. 2021. [CrossRef]
- Wang L, Liu C, Liu Y, Luo M: Fumonisin B1-Induced Changes in Cotton Fiber Elongation Revealed by Sphingolipidomics and Proteomics. Biomolecules 2020, 10(9). [CrossRef]
- Chen Q, Xu F, Wang L, Suo XD, Wang QL, Meng Q, Huang L, Ma CX, Li GM, Luo M: Sphingolipid Profile during Cotton Fiber Growth Revealed That a Phytoceramide Containing Hydroxylated and Saturated VLCFA Is Important for Fiber Cell Elongation. Biomolecules 2021, 11(9). [CrossRef]
- Wang Q, Meng Q, Xu F, Chen Q, Ma C, Huang L, Li G, Luo M: Comparative Metabolomics Analysis Reveals Sterols and Sphingolipids Play a Role in Cotton Fiber Cell Initiation. Int J Mol Sci 2021, 22(21). [CrossRef]
- Li G, Wang Q, Meng Q, Wang G, Xu F, Chen Q, Liu F, Hu Y, Luo M: Overexpression of a ceramide synthase gene,GhCS1, inhibits fiber cell initiation and elongation by promoting the synthesis of ceramides containing dihydroxy LCB and VLCFA. Front Plant Sci 2022, 13. [CrossRef]
- Meng Q, Wang Q, Xu F, Chen Q, Ma C, Huang L, Li G, Liu F, Luo M: Down-regulating a fiber-specific KCR like gene GhKCRL1 suppressed fiber elongation through blocking the synthesis of sphingolipids in fiber cell. Industrial Crops and Products 2022, 186:115290. [CrossRef]
- Zhang J, Meng Q, Wang Q, Zhang H, Tian H, Wang T, Xu F, Yan X, Luo M: Cotton sphingosine kinase GhLCBK1 participates in fiber cell elongation by affecting sphingosine-1-phophate and auxin synthesis. 2024(1879-0003 (Electronic)). [CrossRef]
- Tartaglio V, Rennie EA, Cahoon R, Wang G, Baidoo E, Mortimer JC, Cahoon EB, Scheller HV: Glycosylation of inositol phosphorylceramide sphingolipids is required for normal growth and reproduction in Arabidopsis. The Plant Journal 2017, 89(2):278-290. [CrossRef]
- Olzmann JA, Carvalho P: Dynamics and functions of lipid droplets. Nat Rev Mol Cell Bio 2019, 20(3):137-155. [CrossRef]
- Chapman KD, Dyer JM, Mullen RT: Biogenesis and functions of lipid droplets in plants: Thematic Review Series: Lipid Droplet Synthesis and Metabolism: from Yeast to Man. J Lipid Res 2012, 53(2):215-226. [CrossRef]
- Tang K, Liu JY: Molecular characterization of GhPLD alpha 1 and its relationship with se(c)ondary cell wall thickening in cotton fibers. Acta Bioch Bioph Sin 2017, 49(1):33-43. [CrossRef]
- Deng S, Wei T, Tan K, Hu M, Li F, Zhai Y, Ye S, Xiao Y, Hou L, Pei Y et al.: Phytosterol content and the campesterol:sitosterol ratio influence cotton fiber development: role of phytosterols in cell elongation. Sci China Life Sci 2016, 59(2):183-193. [CrossRef]
- Niu Q, Tan KL, Zang ZL, Xiao ZY, Chen KJ, Hu MY, Luo M: Modification of phytosterol composition influences cotton fiber cell elongation and secondary cell wall deposition. Bmc Plant Biology 2019, 19. [CrossRef]
- Zhang Z, Ruan Y-L, Zhou N, Wang F, Guan X, Fang L, Shang X, Guo W, Zhu S, Zhang T: Suppressing a Putative Sterol Carrier Gene Reduces Plasmodesmal Permeability and Activates Sucrose Transporter Genes during Cotton Fiber Elongation. The Plant Cell 2017, 29(8):2027-2046. [CrossRef]
- Liu F, Wei T, Wang Q, Li G, Meng Q, Huang L, Cheng X, Yan X, Hu Y, Xu F et al.: GhSMO2-2 is regulated by brassinosteroid signal and involved in cotton fiber elongation via influencing phytosterol and sphingolipid biosynthesis. Industrial Crops and Products 2023, 205:117527. [CrossRef]
- Ferrer A, Altabella T, Arro M, Boronat A: Emerging roles for conjugated sterols in plants. Progress in lipid research 2017, 67:27-37. [CrossRef]
- Valitova YN, Kotlova ER, Novikov AV, Shavarda AL, Artemenko KA, Zubarev RA, Minibayeva FV: Binding of sterols affects membrane functioning and sphingolipid composition in wheat roots. Biochemistry (Moscow) 2010, 75(5):554-561. [CrossRef]
- Mongrand S, Morel J, Laroche J, Claverol S, Carde JP, Hartmann MA, Bonneu M, Simon-Plas F, Lessire R, Bessoule JJ: Lipid rafts in higher plant cells - Purification and characterization of triton X-100-insoluble microdomains from tobacco plasma membrane. J Biol Chem 2004, 279(35):36277-36286. [CrossRef]
- Lefebvre B, Furt F, Hartmann MA, Michaelson LV, Carde JP, Sargueil-Boiron F, Rossignol M, Napier JA, Cullimore J, Bessoule JJ et al.: Characterization of lipid rafts from Medicago truncatula root plasma membranes: A proteomic study reveals the presence of a raft-associated redox system. Plant Physiol 2007, 144(1):402-418. [CrossRef]
- Borner GHH, Sherrier DJ, Weimar T, Michaelson LV, Hawkins ND, MacAskill A, Napier JA, Beale MH, Lilley KS, Dupree P: Analysis of detergent-resistant membranes in Arabidopsis. Evidence for plasma membrane lipid rafts. Plant Physiol 2005, 137(1):104-116. [CrossRef]
- Zäuner S, Ternes P, Warnecke D: Biosynthesis of sphingolipids in plants (and some of their functions). Adv Exp Med Biol 2010, 688:249-263. [CrossRef]
- Ali U, Li H, Wang X, Guo L: Emerging Roles of Sphingolipid Signaling in Plant Response to Biotic and Abiotic Stresses. Mol Plant 2018, 11(11):1328-1343. [CrossRef]
- Hannun YA, Luberto C: Ceramide in the eukaryotic stress response. Trends Cell Biol 2000, 10(2):73-80. [CrossRef]
- Scheller HV, Moore WM, Fang L, Chan C, Ishikawa T, Ebert B, Rautengarten C, Kawai-Yamada M, Heazlewood JL, Mortimer JC: Sphingolipid glycosylation and its role in membrane organization and plant-microbe interactions. Glycobiology 2018, 28(12):1020-1020.
- Liang WH, Fang L, Xiang D, Hu Y, Feng H, Chang LJ, Zhang TZ: Transcriptome Analysis of Short Fiber Mutant Ligon lintless-1 (Li-1) Reveals Critical Genes and Key Pathways in Cotton Fiber Elongation and Leaf Development. PloS one 2015, 10(11). [CrossRef]
- Bolton JJ, Soliman KM, Wilkins TA, Jenkins JN: Aberrant Expression of Critical Genes during Secondary Cell Wall Biogenesis in a Cotton Mutant, Ligon Lintless-1 (Li-1). Comparative and functional genomics 2009, 2009:659301. [CrossRef]
- Gilbert MK, Turley RB, Kim HJ, Li P, Thyssen G, Tang YH, Delhom CD, Naoumkina M, Fang DD: Transcript profiling by microarray and marker analysis of the short cotton (Gossypium hirsutum L.) fiber mutant Ligon lintless-1 (Li-1). BMC genomics 2013, 14. [CrossRef]
- Mullen TD, Hannun YA, Obeid LM: Ceramide synthases at the centre of sphingolipid metabolism and biology. The Biochemical journal 2012, 441(3):789-802. [CrossRef]
- Ternes P, Feussner K, Werner S, Lerche J, Iven T, Heilmann I, Riezman H, Feussner I: Disruption of the ceramide synthase LOH1 causes spontaneous cell death in Arabidopsis thaliana. New Phytol 2011, 192(4):841-854. [CrossRef]
- Worgall TS: Sphingolipids: major regulators of lipid metabolism. Curr Opin Clin Nutr Metab Care 2007, 10(2):149-155. [CrossRef]
- Shimada TL, Shimada T, Okazaki Y, Higashi Y, Saito K, Kuwata K, Oyama K, Kato M, Ueda H, Nakano A et al.: HIGH STEROL ESTER 1 is a key factor in plant sterol homeostasis. Nat Plants 2019, 5(11):1154-1166. [CrossRef]
- Deevska GM, Nikolova-Karakashian MN: The expanding role of sphingolipids in lipid droplet biogenesis. Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids 2017, 1862(10, Part B):1155-1165. [CrossRef]
- Guzha A, Whitehead P, Ischebeck T, Chapman KD: Lipid Droplets: Packing Hydrophobic Molecules Within the Aqueous Cytoplasm. Annual Review of Plant Biology 2023, 74(Volume 74, 2023):195-223. [CrossRef]
- Krahmer N, Hilger M, Kory N, Wilfling F, Stoehr G, Mann M, Farese RV, Jr., Walther TC: Protein correlation profiles identify lipid droplet proteins with high confidence. Mol Cell Proteomics 2013, 12(5):1115-1126. [CrossRef]
- Ischebeck T, Krawczyk HE, Mullen RT, Dyer JM, Chapman KD: Lipid droplets in plants and algae: Distribution, formation, turnover and function. Semin Cell Dev Biol 2020, 108:82-93. [CrossRef]
- Qin Z, Wang T, Zhao Y, Ma C, Shao Q: Molecular Machinery of Lipid Droplet Degradation and Turnover in Plants. Int J Mol Sci 2023, 24(22):16039. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



