Submitted:
18 December 2024
Posted:
19 December 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Samples
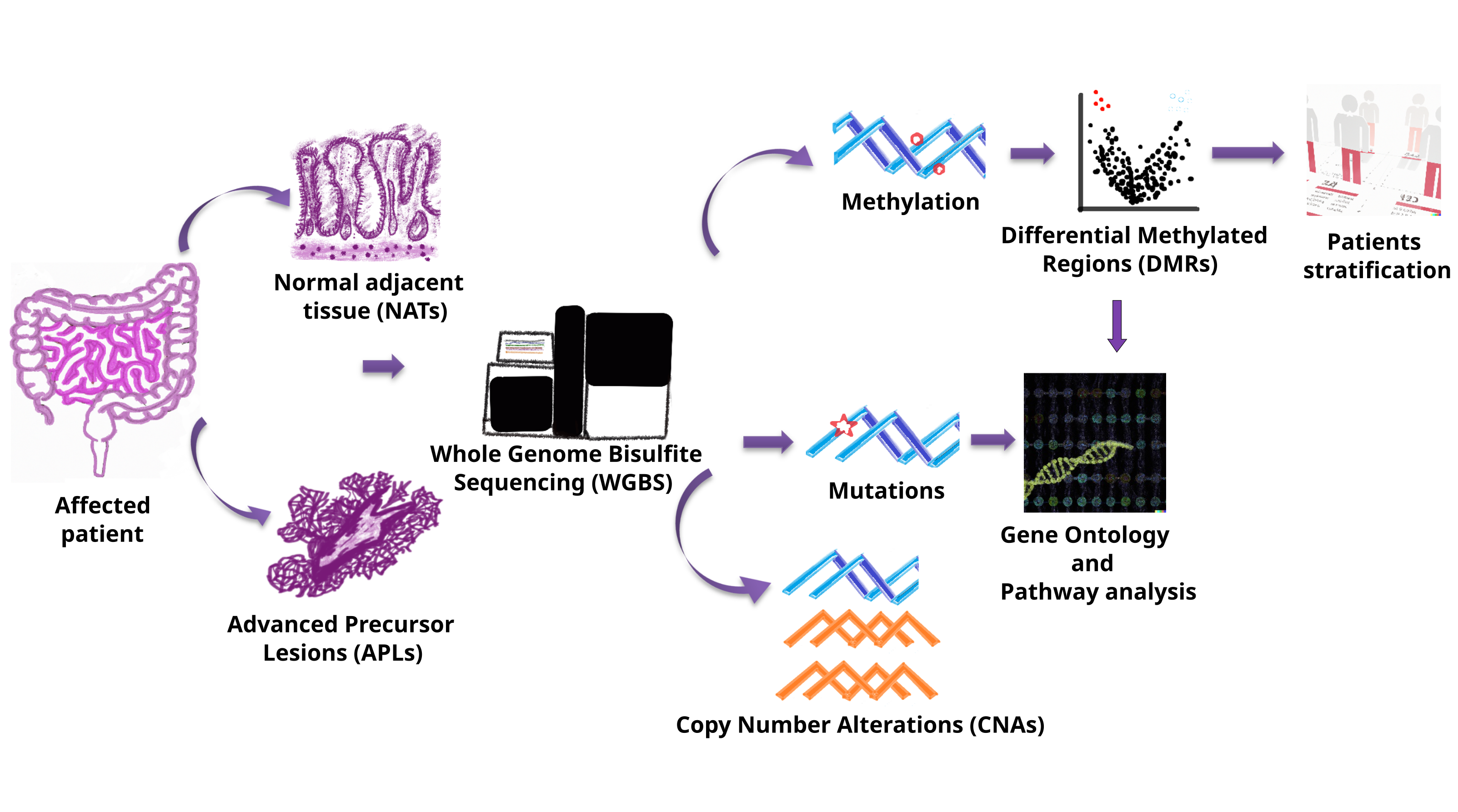
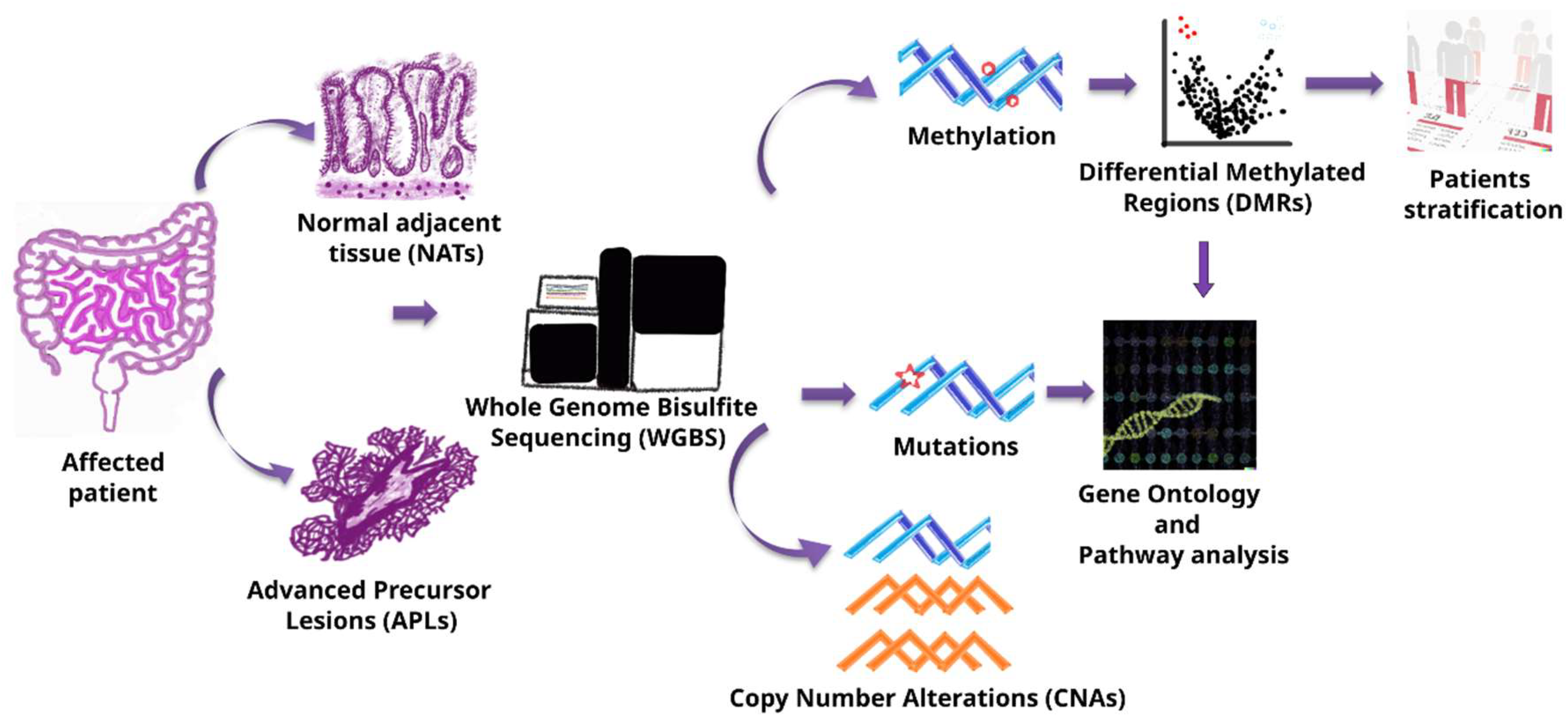
2.2. Sample Preparation and Whole-Genome Bisulfite Sequencing (WGBS)
2.3. Genomic Data and Methylation Processing
2.4. Analyses of Differentially Methylated Position/Region
2.5. Pathway Enrichment Analysis
2.6. Copy Number Alteration Detection
2.7. Somatic Mutation Detection
2.8. CpG Island Methylator Phenotype (CIMP)
2.9. Microsatellite Instability (MSI) Status Prediction
2.10. Principal Component Analysis (PCA)
3. Results
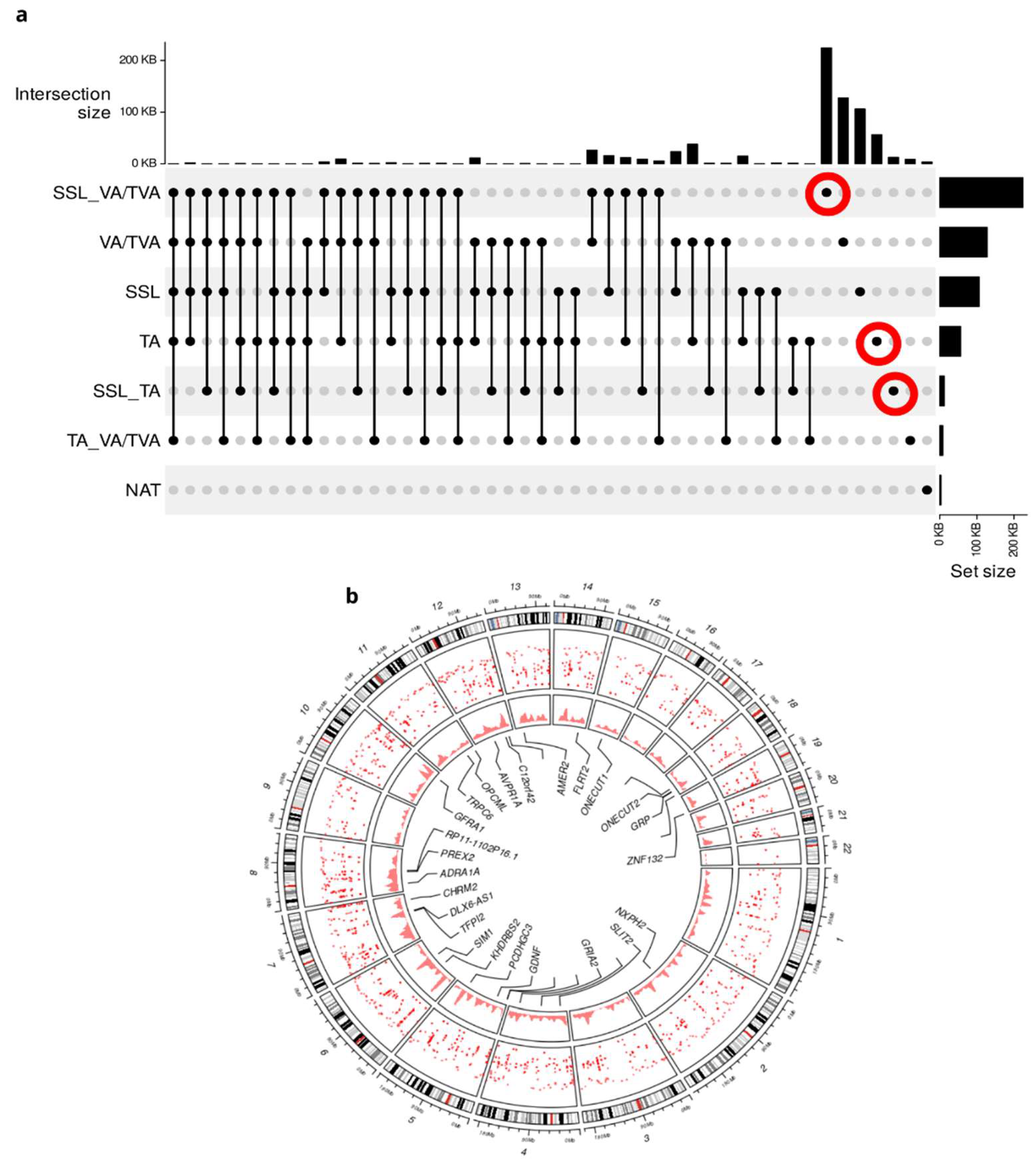
3.1. DMR Analysis Shows Unique Methylation Patterns Between APL’s Subtypes and CRC
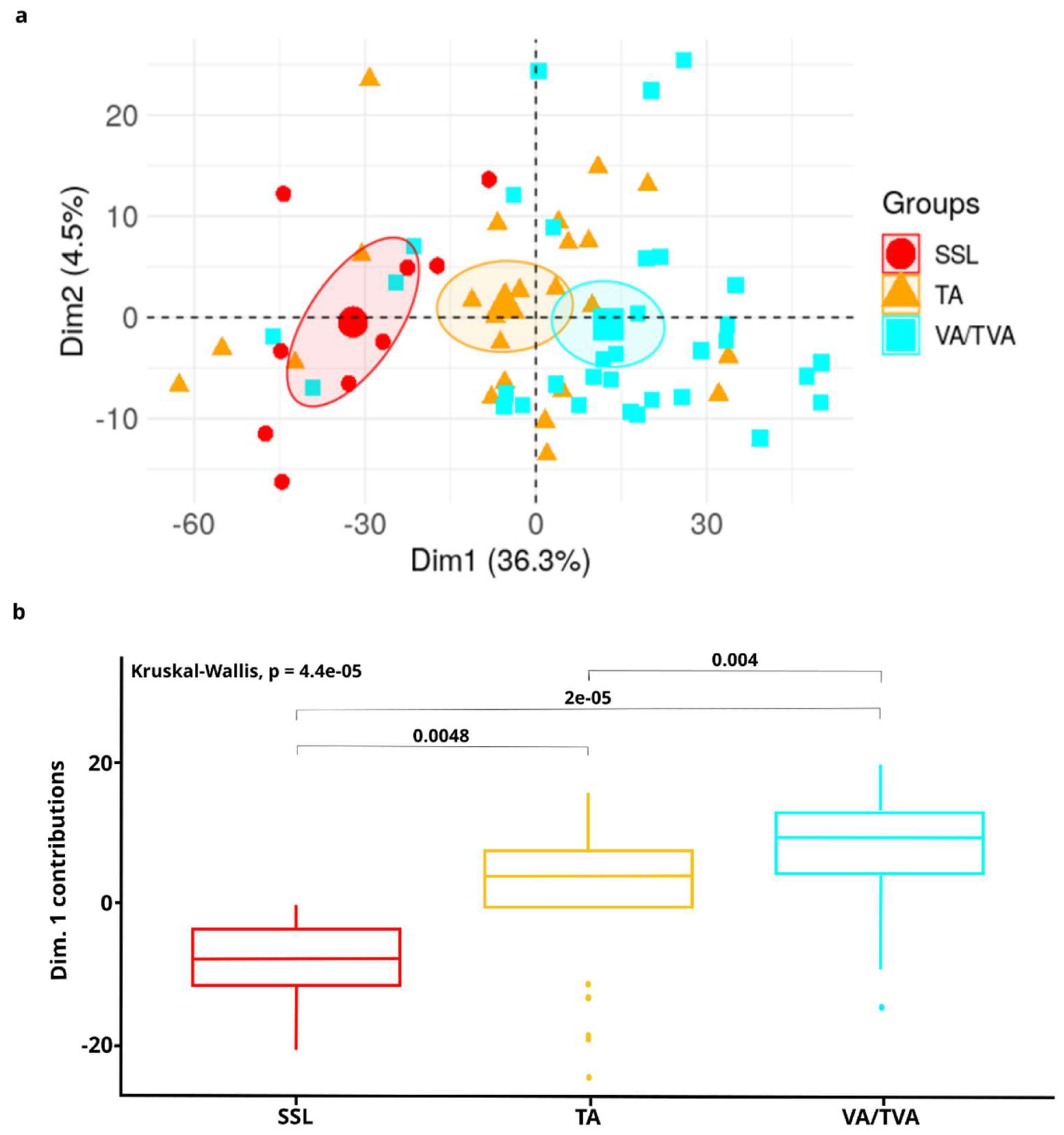
3.2. DMR Signature Enables Sample Stratification According to APL Subtype
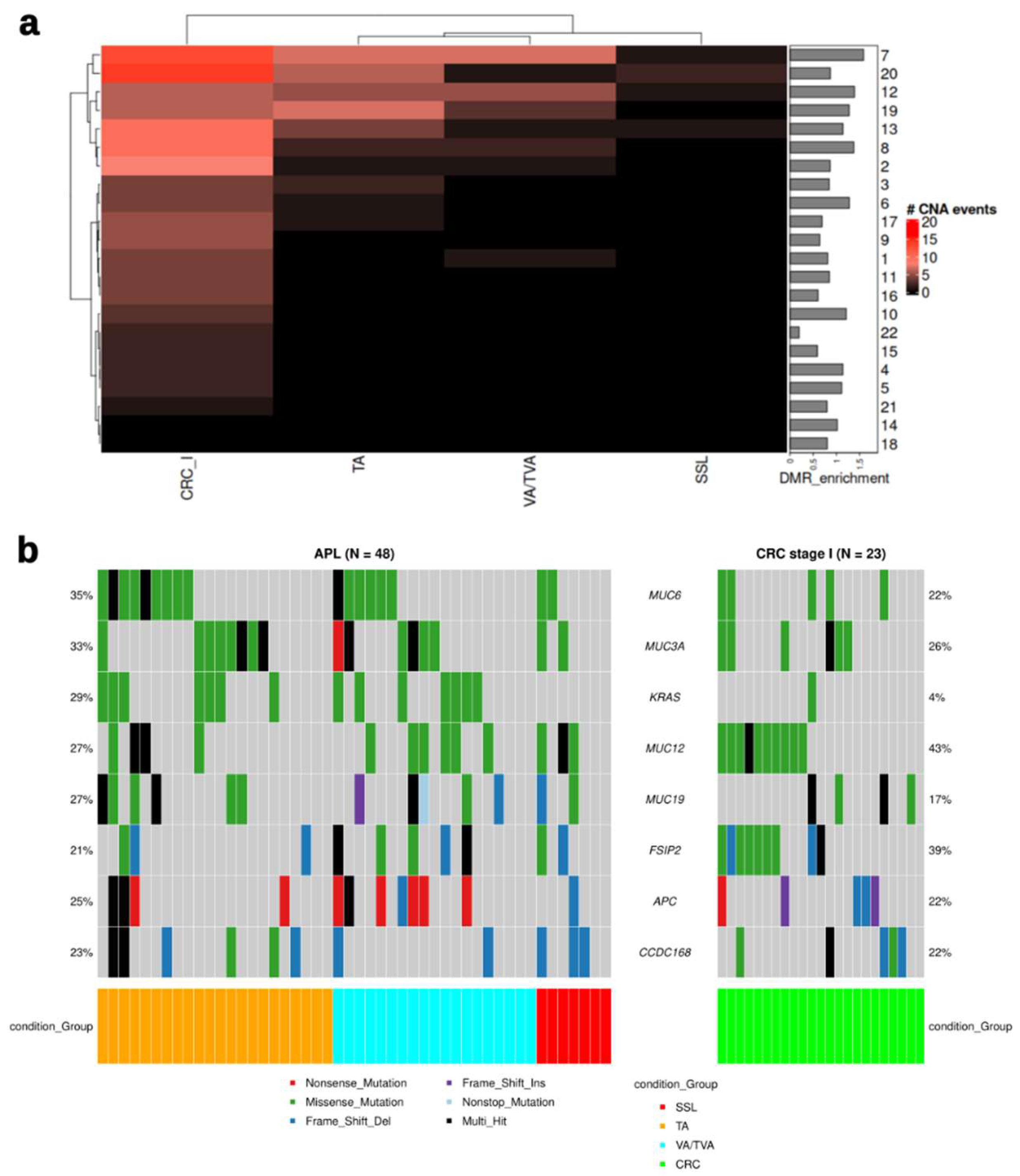
3.3. CNAs and Mutation Analysis Suggests Synergistic Mechanisms Complementary to Epigenetic Control in APLs
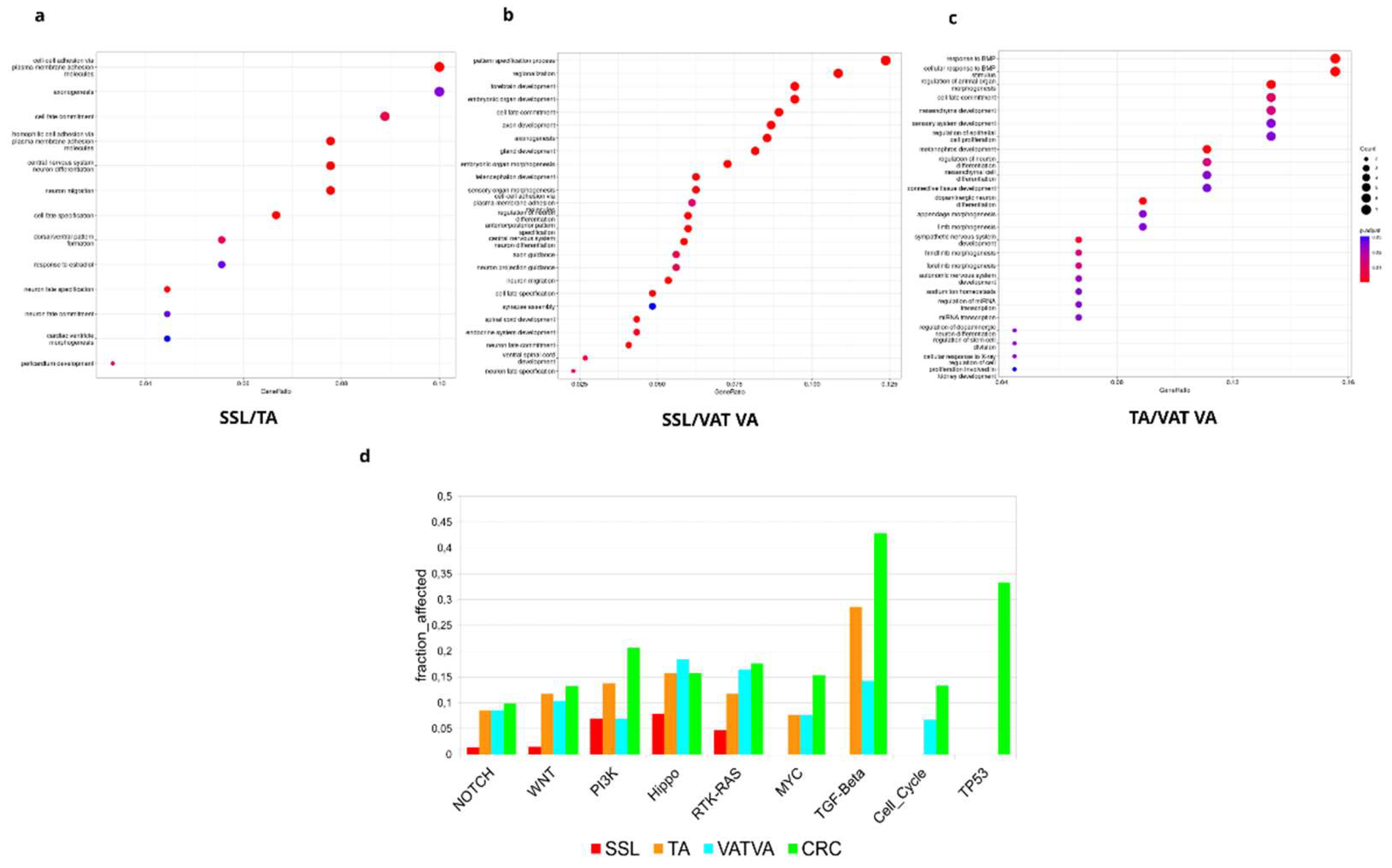
3.4. Pathway Enrichment Analysis Reveal Redundancy in the Circuitry of Transcriptional Regulation Cellular Fate and Proliferation Signaling in APL Progression
3.5. DMR Phenotype in APLs Is Not CIMP and MSI Related
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023 Jan 12;73(1):17–48.
- Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024 May 4;74(3):229–63.
- Fornasarig M, Magris R, De Re V, Bidoli E, Canzonieri V, Maiero S, et al. Molecular and Pathological Features of Gastric Cancer in Lynch Syndrome and Familial Adenomatous Polyposis. Int J Mol Sci. 2018 Jun 6;19(6):1682. [CrossRef]
- Sawicki T, Ruszkowska M, Danielewicz A, Niedźwiedzka E, Arłukowicz T, Przybyłowicz KE. A Review of Colorectal Cancer in Terms of Epidemiology, Risk Factors, Development, Symptoms and Diagnosis. Cancers (Basel). 2021 Apr 22;13(9):2025. [CrossRef]
- Sinicrope FA. Increasing Incidence of Early-Onset Colorectal Cancer. New England Journal of Medicine. 2022 Apr 21;386(16):1547–58. [CrossRef]
- Cho KR, Vogelstein B. Genetic alterations in the adenoma--carcinoma sequence. Cancer. 1992 Sep 15;70(6 Suppl):1727–31.
- Tan C, Du X. KRAS mutation testing in metastatic colorectal cancer. World J Gastroenterol. 2012 Oct 7;18(37):5171–80.
- Song Y, Huang J, Wang K, Li Y. To Identify Adenomatous Polyposis Coli Gene Mutation as a Predictive Marker of Endometrial Cancer Immunotherapy. Front Cell Dev Biol. 2022 Jul 22;10. [CrossRef]
- Leslie A, Carey FA, Pratt NR, Steele RJC. The colorectal adenoma-carcinoma sequence. Br J Surg. 2002 Jul;89(7):845–60. [CrossRef]
- Ternet C, Kiel C. Signaling pathways in intestinal homeostasis and colorectal cancer: KRAS at centre stage. Cell Commun Signal. 2021 Mar 10;19(1):31. [CrossRef]
- Naini B V, Odze RD. Advanced precancerous lesions (APL) in the colonic mucosa. Best Pract Res Clin Gastroenterol. 2013 Apr;27(2):235–56.
- Rupa JD, de Bruïne AP, Gerbers AJ, Leers MPG, Nap M, Kessels AGH, et al. Simultaneous detection of apoptosis and proliferation in colorectal carcinoma by multiparameter flow cytometry allows separation of high and low-turnover tumors with distinct clinical outcome. Cancer. 2003 May 15;97(10):2404–11. [CrossRef]
- Schimek V, Strasser K, Beer A, Göber S, Walterskirchen N, Brostjan C, et al. Tumour cell apoptosis modulates the colorectal cancer immune microenvironment via interleukin-8-dependent neutrophil recruitment. Cell Death Dis. 2022 Feb 4;13(2):113. [CrossRef]
- Taherian M, Lotfollahzadeh S, Daneshpajouhnejad P, Arora K. Tubular Adenoma. 2024.
- Dornblaser D, Young S, Shaukat A. Colon polyps: updates in classification and management. Curr Opin Gastroenterol. 2024 Jan;40(1):14–20. [CrossRef]
- Tornillo L, Lehmann FS, Garofoli A, Paradiso V, Ng CKY, Piscuoglio S. The Genomic Landscape of Serrated Lesion of the Colorectum: Similarities and Differences With Tubular and Tubulovillous Adenomas. Front Oncol. 2021 Oct 12;11. [CrossRef]
- Boregowda U, Umapathy C, Echavarria J, Saligram S. Risk of Metachronous Neoplasia with High-Risk Adenoma and Synchronous Sessile Serrated Adenoma: A Systematic Review and Meta-Analysis. Diagnostics. 2023 Apr 27;13(9):1569. [CrossRef]
- Kambara T, Simms LA, Whitehall VLJ, Spring KJ, Wynter CVA, Walsh MD, et al. BRAF mutation is associated with DNA methylation in serrated polyps and cancers of the colorectum. Gut. 2004 Aug;53(8):1137–44. [CrossRef]
- Nazemalhosseini Mojarad E, Kuppen PJ, Aghdaei HA, Zali MR. The CpG island methylator phenotype (CIMP) in colorectal cancer. Gastroenterol Hepatol Bed Bench. 2013;6(3):120–8.
- Boland CR, Goel A. Microsatellite Instability in Colorectal Cancer. Gastroenterology. 2010 May;138(6):2073-2087.e3.
- Tamura K, Kaneda M, Futagawa M, Takeshita M, Kim S, Nakama M, et al. Genetic and genomic basis of the mismatch repair system involved in Lynch syndrome. Int J Clin Oncol. 2019 Sep;24(9):999–1011.
- Kakar S, Deng G, Cun L, Sahai V, Kim YS. CpG island methylation is frequently present in tubulovillous and villous adenomas and correlates with size, site, and villous component. Hum Pathol. 2008 Jan;39(1):30–6. [CrossRef]
- Hong SN. Genetic and epigenetic alterations of colorectal cancer. Intest Res. 2018;16(3):327. [CrossRef]
- Molinari C, Marisi G, Passardi A, Matteucci L, De Maio G, Ulivi P. Heterogeneity in Colorectal Cancer: A Challenge for Personalized Medicine? Int J Mol Sci. 2018 Nov 23;19(12).
- Estécio MRH, Issa JPJ. Dissecting DNA hypermethylation in cancer. FEBS Lett. 2011 Jul 7;585(13):2078–86. [CrossRef]
- Baylin SB, Jones PA. A decade of exploring the cancer epigenome — biological and translational implications. Nat Rev Cancer. 2011 Oct 23;11(10):726–34. [CrossRef]
- Andrews S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Babraham, UK: Babraham Institute; 2023.
- Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011 May 2;17(1):10. [CrossRef]
- Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011 Jun 1;27(11):1571–2. [CrossRef]
- Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10(3):R25. [CrossRef]
- Du J, Yuan Z, Ma Z, Song J, Xie X, Chen Y. KEGG-PATH: Kyoto encyclopedia of genes and genomes-based pathway analysis using a path analysis model. Mol Biosyst. 2014 Jul 29;10(9):2441–7. [CrossRef]
- Gu Z, Hübschmann D. rGREAT : an R/bioconductor package for functional enrichment on genomic regions. Bioinformatics. 2023 Jan 1;39(1).
- Xu S, Hu E, Cai Y, Xie Z, Luo X, Zhan L, et al. Using clusterProfiler to characterize multiomics data. Nat Protoc. 2024 Nov 17;19(11):3292–320. [CrossRef]
- Yu G. Gene Ontology Semantic Similarity Analysis Using GOSemSim. In: Stem Cell Transcriptional Networks. 2020. p. 207–15.
- Adalsteinsson VA, Ha G, Freeman SS, Choudhury AD, Stover DG, Parsons HA, et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat Commun. 2017 Nov 6;8(1):1324. [CrossRef]
- Lai D, Shah S. HMMcopy: Copy number prediction with correction for GC and mappability bias for HTS data. 2024.
- Nunn A, Can SN, Otto C, Fasold M, Díez Rodríguez B, Fernández-Pozo N, et al. EpiDiverse Toolkit: a pipeline suite for the analysis of bisulfite sequencing data in ecological plant epigenetics. NAR Genom Bioinform. 2021 Oct 4;3(4). [CrossRef]
- Koboldt DC, Chen K, Wylie T, Larson DE, McLellan MD, Mardis ER, et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics. 2009 Sep 1;25(17):2283–5. [CrossRef]
- Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012 Mar;22(3):568–76. [CrossRef]
- Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly (Austin). 2012 Apr 27;6(2):80–92. [CrossRef]
- Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018 Nov;28(11):1747–56. [CrossRef]
- Wilkerson MD, Hayes DN. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010 Jun 15;26(12):1572–3. [CrossRef]
- Santamarina-García M, Brea-Iglesias J, Bramsen JB, Fuentes-Losada M, Caneiro-Gómez FJ, Vázquez-Bueno JÁ, et al. MSIMEP: Predicting microsatellite instability from microarray DNA methylation tumor profiles. iScience. 2023 Mar 17;26(3):106127. [CrossRef]
- Yamanoi K, Arai E, Tian Y, Takahashi Y, Miyata S, Sasaki H, et al. Epigenetic clustering of gastric carcinomas based on DNA methylation profiles at the precancerous stage: its correlation with tumor aggressiveness and patient outcome. Carcinogenesis. 2015 May;36(5):509–20. [CrossRef]
- Vincent A, Omura N, Hong SM, Jaffe A, Eshleman J, Goggins M. Genome-Wide Analysis of Promoter Methylation Associated with Gene Expression Profile in Pancreatic Adenocarcinoma. Clinical Cancer Research. 2011 Jul 1;17(13):4341–54.
- Negrini S, Gorgoulis VG, Halazonetis TD. Genomic instability — an evolving hallmark of cancer. Nat Rev Mol Cell Biol. 2010 Mar;11(3):220–8. [CrossRef]
- Druliner BR, Wang P, Bae T, Baheti S, Slettedahl S, Mahoney D, et al. Molecular characterization of colorectal adenomas with and without malignancy reveals distinguishing genome, transcriptome and methylome alterations. Sci Rep. 2018 Feb 16;8(1):3161. [CrossRef]
- Bomme L, Bardi G, Pandis N, Fenger C, Kronborg O, Heim S. Clonal karyotypic abnormalities in colorectal adenomas: Clues to the early genetic events in the adenoma-carcinoma sequence. Genes Chromosomes Cancer. 1994 Jul 14;10(3):190–6.
- Bollen Y, Stelloo E, van Leenen P, van den Bos M, Ponsioen B, Lu B, et al. Reconstructing single-cell karyotype alterations in colorectal cancer identifies punctuated and gradual diversification patterns. Nat Genet. 2021 Aug;53(8):1187–95. [CrossRef]
- Cox KE, Liu S, Lwin TM, Hoffman RM, Batra SK, Bouvet M. The Mucin Family of Proteins: Candidates as Potential Biomarkers for Colon Cancer. Cancers (Basel). 2023 Feb 27;15(5). [CrossRef]
- Lei H, Tao K. Somatic mutations in colorectal cancer are associated with the epigenetic modifications. J Cell Mol Med. 2020 Oct;24(20):11828–36. [CrossRef]
| APL | APL (validation) | CRC stage I | |
|---|---|---|---|
| n | 48 | 26 | 23 |
|
Age median (IQR), years (range) |
68.5 (14.75) (39-86) |
68.5 (19.75) (23-84) |
68 (13) (37-81) |
| Gender Female/Male, n | 17/31 | 16/10 | 14/9 |
|
Histology SSL, n TA, n VA/TVA, n |
7 22 19 |
7 4 15 |
| Comparison | DMRs | Hyper DMRs | Hypo DMRs |
|---|---|---|---|
| SSL vs SSL NAT | 6317 | 1271 | 5046 |
| TA vs TA NAT | 36297 | 861 | 35436 |
| VA/TVA vs VA/TVA NAT | 65794 | 1807 | 63987 |
| SSL vs TA | 1299 | 137 | 1162 |
| SSL vs VA/TVA | 11955 | 2096 | 9859 |
| TA vs VA/TVA | 651 | 91 | 560 |
| SSL NAT vs TA NAT | 4 | ||
| SSL NAT vs VA/TVA NAT | 30 | ||
| TA NAT vs VA/TVA NAT | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




