Submitted:
16 December 2024
Posted:
18 December 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Samples, DNA Extraction and Sequencing
2.2. Genome Assembly and Annotation
2.3. Codon Usage Analysis
2.4. Ka/Ks Value Analyses
2.5. Phylogenetic Analysis TimeTree Estimation
3. Results
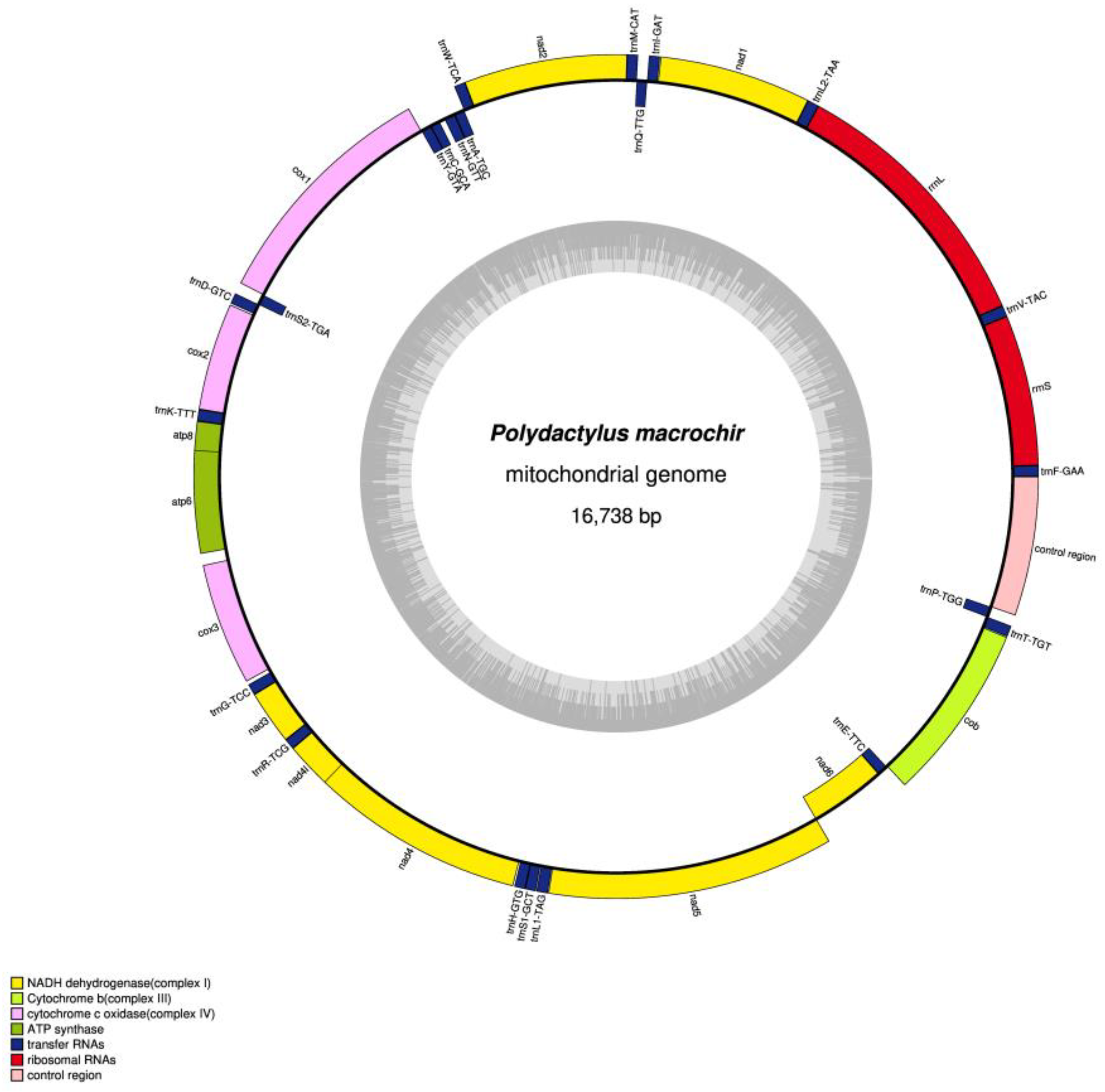
3.1. Mitogenomic Structure and Organization
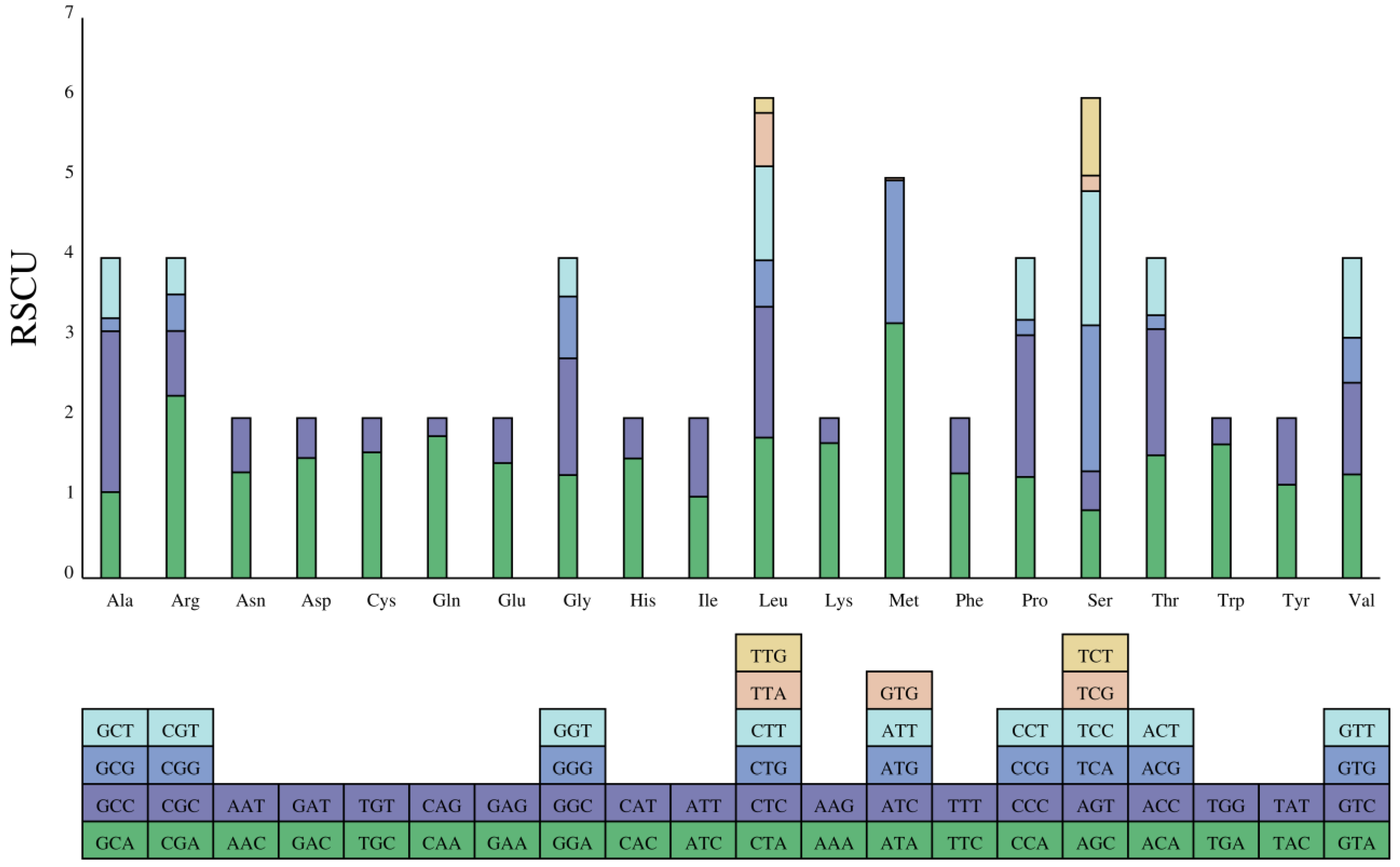
3.2. Protein-Coding Genes and Codon Usage
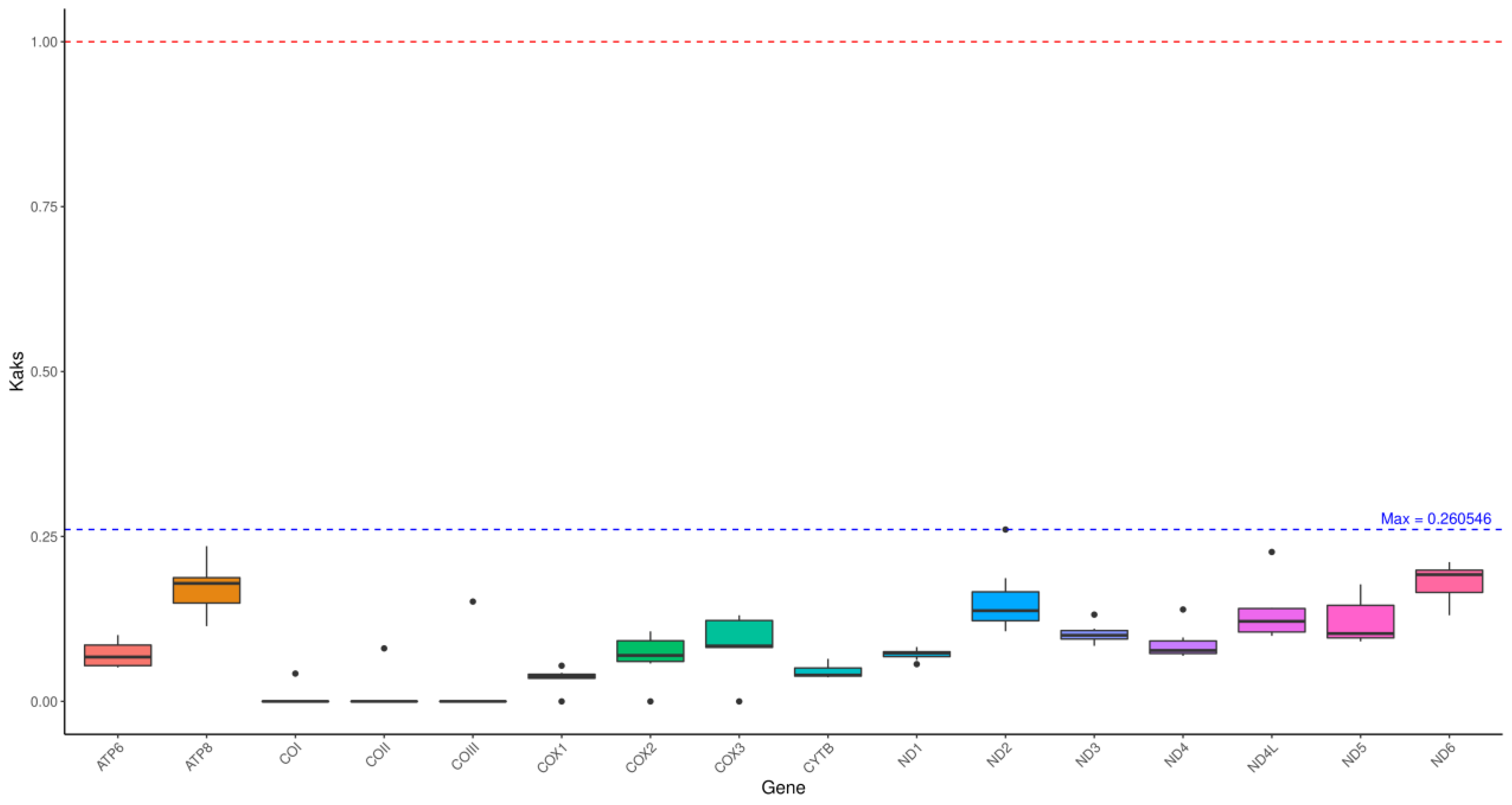
3.3. Ka/Ks Value Analyses
3.4. Phylogenetic Analysis and TimeTree Estimation
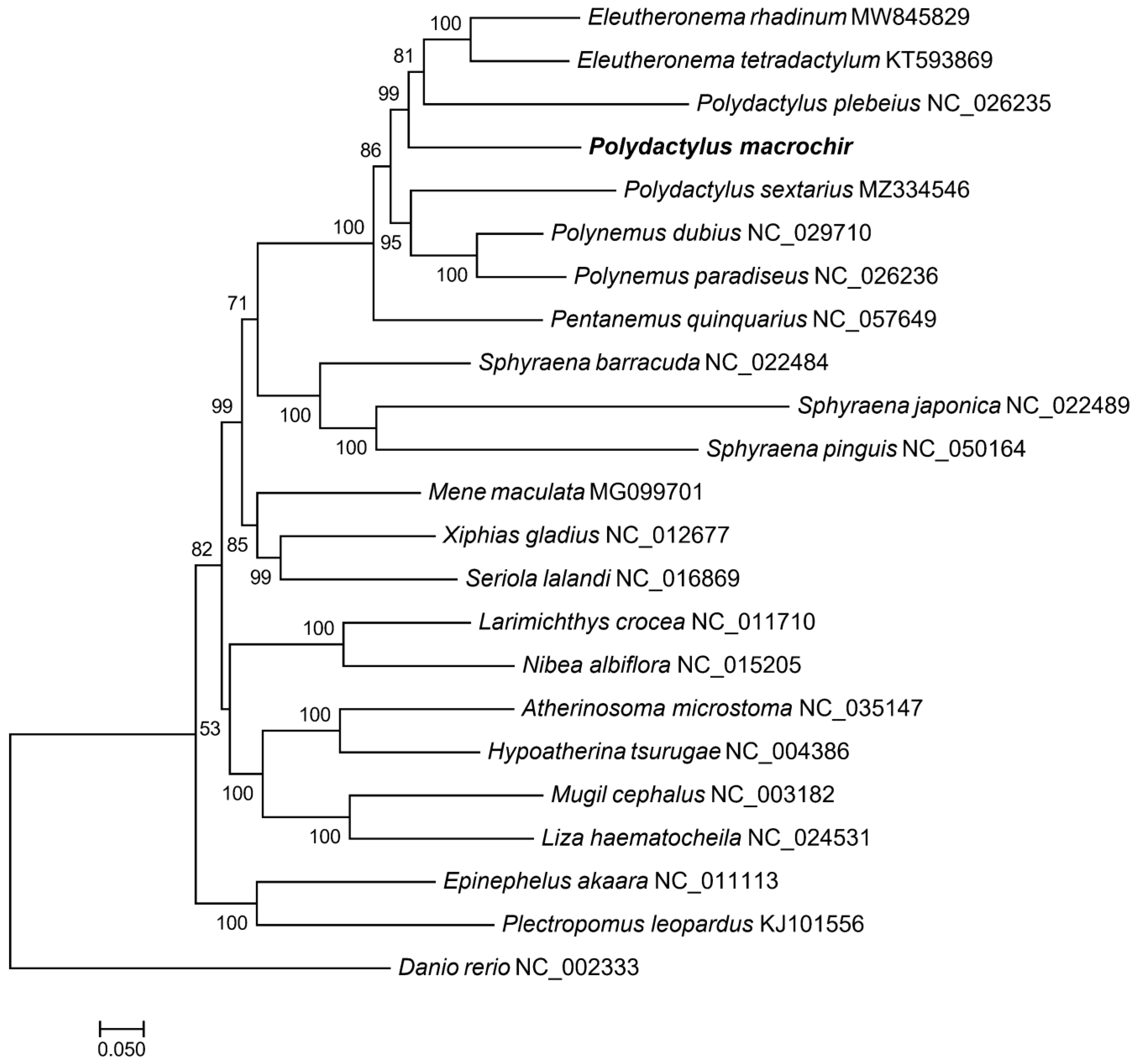

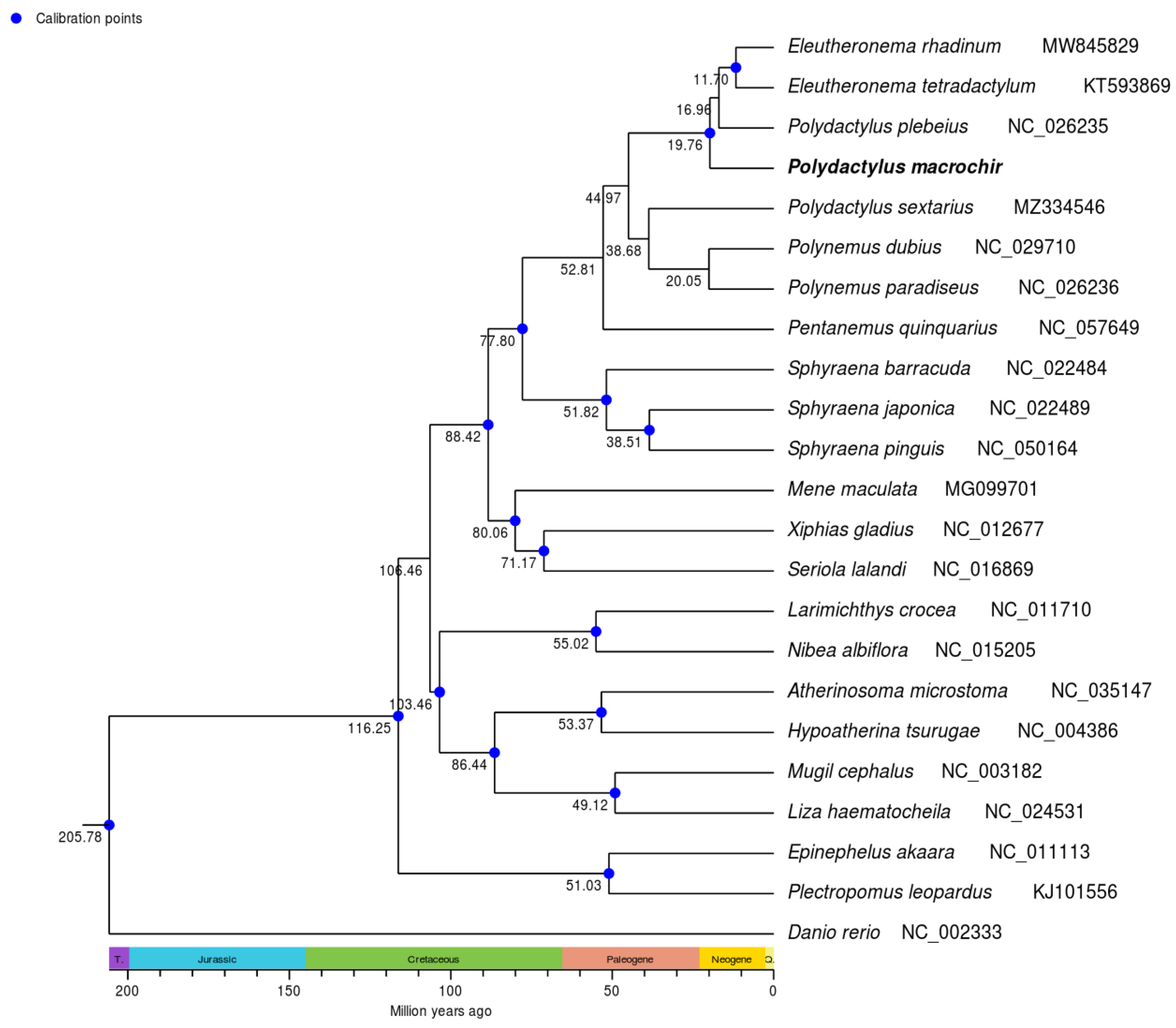
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Motomura: H. Treadfns of the world (Family Polynemidae): An annotated and illustrated catalogue of polynemid species known to date (Food & Agriculture Org, Rome, 2004).
- Presti, P.; Johnson, G.D.; Datovo, A. Anatomy and evolution of the pectoral filaments of threadfins (Polynemidae). Sci Rep. 2020, 10, 17751. [CrossRef]
- Fricke R, Eschmeyer W, Fong J. Eschmeyer’s catalog of fishes: genera/species by family/subfamily. San Francisco: California Academy of Sciences, 2022.
- Gosline WA. Systematic position and relationships of the percesocine fishes. Pac. Sci. 1962, 16, 207-217.
- Rosen DE. The relationships and taxonomic position of the halfbeaks, killifishes, silversides, and their relatives. Bull. Am. Mus. Nat. Hist. 1964, 127, 217-268.
- Johnson, D.G Percomorph phylogeny: Progress and problems. Bulletin of Marine Science. 1993, 52, 3-28.
- Kang, S.; Imamura, H.; Kawai, T. Morphological evidence supporting the monophyly of the family Polynemidae (Teleostei: Perciformes) and its sister relationship with Sciaenidae. Ichthyol. Res. 2017, 65, 29-41. [CrossRef]
- Presti P, Johnson GD, Datovo A. Facial and gill musculature of polynemid fishes, with notes on their possible relationships with sciaenids (Percomorphacea: Perciformes). J Morphol. 2020, 281, 662-675. [CrossRef]
- Mirande, J.M. Combined phylogeny of ray-ffnned fishes (Actinopterygii) and the use of morphological characters in large-scale analyses. Cladistics. 2017, 33, 333-350.
- Sanciangco, M.D.; Carpenter, K.E.; Betancur-R, R. Phylogenetic placement of enigmatic percomorph families (Teleostei: Percomorphaceae). Molecular Phylogenetics and Evolution. 2015, 94, 565-576. [CrossRef]
- Betancur-R, R.; Broughton, R.E.; Wiley, E.O.; Carpenter, K.; López, J.A.; Li, C.; Holcroft, N.I.; Arcila, D.; Sanciangco, M.; Cureton, Ii.J.C.; Zhang, F.; Buser, T.; Campbell, M.A.; Ballesteros, J.A.; Roa-Varon, A.; Willis, S.; Borden, W.C.; Rowley, T.; Reneau, P.C.; Hough, D.J.; Lu, G.; Grande, T.; Arratia, G.; Ortí, G. The tree of life and a new classification of bony fishes. PLoS Currents. 2013, 5. [CrossRef]
- Hughes, L.C.; Ortí, G.; Huang, Y.; Sun, Y.; Baldwin, C.C.; Thompson, A.W.; Arcila, D.; Betancur-R, R.; Li, C.; Becker, L.; Bellora, N.; Zhao, X.; Li, X.; Wang, M.; Fang, C.; Xie, B.; Zhou, Z.; Huang, H.; Chen, S.; Venkatesh, B.; Shi, Q. Comprehensive phylogeny of rayffnned ffshes (Actinopterygii) based on transcriptomic and genomic data. Proc. Natl. Acad. Sci. U.S.A. 2018, 115, 6249-6254.
- Girard, M.G.; Davis, M.P.; Smith, W.L. The phylogeny of Carangiform fishes: morphological and genomic investigations of a new fish clade. Copeia. 2020, 108, 265-298. [CrossRef]
- Zhong, L.Q.; Wang, M.H.; Li, D.M.; Tang, S.K.; Chen, X.H. Mitochondrial genome of Eleutheronema rhadinum with an additional non-coding region and novel insights into the phylogenetics. Front. Mar. Sci. 2021, 8, 746598. [CrossRef]
- Taanman, J.W. The mitochondrial genome: structure, transcription, translation and replication. Biochim. Biophys. Acta Bioenerg. 1999, 1410, 103-123. [CrossRef]
- Moritz, C.; Dowling, T. E.; Brown, W.M. Evolution of animal mitochondrial DNA: relevance for population biology and systematics. Annu. Rev. Ecol. Syst. 1987, 18, 269–292. [CrossRef]
- Wang, C.; Ye, P.; Liu, M.; Zhang, Y.; Feng, H.; Liu, J.; Zhou, H.; Wang, J.; Chen, X. Comparative Analysis of Four Complete Mitochondrial Genomes of Epinephelidae (Perciformes). Genes (Basel). 2022, 13, 660. [CrossRef]
- Gao, J.; Li, C.; Yu, D.; Wang, T.; Lin, L.; Xiao, Y.; Wu, P.; Liu, Y. Comparative mitogenome analyses uncover mitogenome features and phylogenetic implications of the parrotfishes (Perciformes: Scaridae). Biology (Basel). 2023, 12, 410. [CrossRef]
- Yang, L.M.; Xue, J.F.; Zhao, X.M.; Ding, K.; Liu, Z.W.; Wang, Z.S.; Chen, J.B.; Huang, Y.K. Mitochondrial genome characteristics reveal evolution of Acanthopsetta nadeshnyi (Jordan and Starks, 1904) and phylogenetic relationships. Genes (Basel). 2024, 15, 893.
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M, Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; Pyshkin, A.V.; Sirotkin, A.V.; Vyahhi, N.; Tesler, G.; Alekseyev, M.A.; Pevzner, P.A. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012, 19, 455-77.
- Bernt, M.; Donath, A.; Jühling, F.; Externbrink, F.; Florentz, C.; Fritzsch, G.; Pütz, J.; Middendorf, M.; Stadler, P.F. MITOS: Improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 2013, 69, 313-319. [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2019, 20, 1160-1166. [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A Toolkit incorporating γ-series methods and sliding window strategies. Genom. Proteom. Bioinform. 2010, 8, 77-80.
- Montaña-Lozano, P.; Moreno-Carmona, M.; Ochoa-Capera, M.; Medina, N.S.; Boore, J.L.; Prada, C.F. Comparative genomic analysis of vertebrate mitochondrial reveals a differential of rearrangements rate between taxonomic class. Sci. Rep. 2022, 12, 5479. [CrossRef]
- Ke Z, Zhou K, Hou M, Luo H, Li Z, Pan X, Zhou J, Jing T, Ye H. Characterization of the Complete Mitochondrial Genome of the Elongate Loach and Its Phylogenetic Implications in Cobitidae. Animals (Basel). 2023, 13, 3841. [CrossRef]
- Gun, L.; Yumiao, R.; Haixian, P.; Liang, Z. Comprehensive analysis and comparison on the codon usage pattern of whole Mycobacterium tuberculosis coding genome from different area. Biomed. Res. Int. 2018, 2018, 3574976. [CrossRef]
- Kundu S, Kang HE, Kim AR, Lee SR, Kim EB, Amin MHF, Andriyono S, Kim HW, Kang K. Mitogenomic Characterization and Phylogenetic Placement of African Hind, Cephalopholis taeniops: Shedding Light on the Evolution of Groupers (Serranidae: Epinephelinae). Int. J. Mol. Sci. 2024, 25, 1822.
- Yang, Z.H.; Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 2000, 17, 32-43. [CrossRef]
| Order | Family | Genus | Species | Size (bp) | Accession No. |
|---|---|---|---|---|---|
| Perciformes | Polynemidae | Eleutheronema | Eleutheronema rhadinum | 16718 | MW845829.1 |
| Eleutheronema tetradactylum | 16491 | KT593869.1 | |||
| Pentanemus | Pentanemus quinquarius | 16708 | NC_057649.1 | ||
| Polydactylus | Polydactylus sextarius | 16841 | MZ334546.1 | ||
| Polydactylus plebeius | 16765 | NC_026235.1 | |||
| Polynemus | Polynemus paradiseus | 16710 | NC_026236.1 | ||
| Polynemus dubius | 16555 | NC_029710.1 | |||
| Sphyraenidae | Sphyraena | Sphyraena pinguis | 16620 | NC_050164.1 | |
| Sphyraena barracuda |
16707 | NC_022484.1 | |||
| Sphyraena japonica | 16760 | NC_022489.1 | |||
| Xiphiidae | Xiphias | Xiphias gladius | 16520 | NC_012677.1 | |
| Carangidae | Seriola | Seriola lalandi | 16532 | NC_016869.1 | |
| Menidae | Mene | Mene maculata | 16733 | MG099701.1 | |
| Sciaenidae | Larimichthys | Larimichthys crocea | 16466 | NC_011710.1 | |
| Nibea | Nibea albiflora | 16499 | NC_015205.1 | ||
| Atheriniformes | Atherinidae | Atherinosoma | Atherinosoma microstoma | 16573 | NC_035147.1 |
| Hypoatherina | Hypoatherina tsurugae | 16566 | NC_004386.1 | ||
| Serranidae | Epinephelus | Epinephelus akaara | 16795 | NC_011113.1 | |
| Plectropomus | Plectropomus leopardus | 16754 | KJ101556.1 | ||
| Mugiliformes | Mugilidae | Mugil | Mugil cephalus | 16685 | NC_003182.1 |
| Planiliza | Liza haematocheila | 16822 | NC_024531.1 | ||
| Cypriniformes | Danionidae | Danio | Danio rerio | 16596 | NC_002333.2 |
| Position | codon | |||||||
|---|---|---|---|---|---|---|---|---|
| Gene | stand | From | To | size | Intergenic Length | start | stop | |
| trnF | H | 1 | 70 | 70 | 0 | |||
| rrnS | H | 71 | 1037 | 967 | 0 | |||
| trnV | H | 1038 | 1110 | 73 | 0 | |||
| rrnL | H | 1111 | 2853 | 1743 | 0 | |||
| trnL2 | H | 2854 | 2927 | 74 | 0 | |||
| nad1 | H | 2928 | 3902 | 975 | 0 | ATG | TAA | |
| trnI | H | 3907 | 3976 | 70 | 4 | |||
| trnQ | L | 3976 | 4046 | 71 | -1 | |||
| trnM | H | 4046 | 4114 | 69 | -1 | |||
| nad2 | H | 4115 | 5160 | 1046 | 0 | ATG | TA- | |
| trnW | H | 5161 | 5231 | 71 | 0 | |||
| trnA | L | 5233 | 5301 | 69 | 1 | |||
| trnN | L | 5303 | 5375 | 73 | 1 | |||
| trnC | L | 5414 | 5479 | 66 | 38 | |||
| trnY | L | 5480 | 5549 | 70 | 0 | |||
| cox1 | H | 5551 | 7101 | 1551 | 1 | GTG | TAA | |
| trnS2 | L | 7102 | 7173 | 72 | 0 | |||
| trnD | H | 7174 | 7242 | 69 | 0 | |||
| cox2 | H | 7252 | 7942 | 691 | 9 | ATG | T-- | |
| trnK | H | 7943 | 8016 | 74 | 0 | |||
| atp8 | H | 8018 | 8206 | 189 | 1 | ATG | TAA | |
| atp6 | H | 8176 | 8859 | 684 | -31 | ATG | TAA | |
| cox3 | H | 8933 | 9718 | 786 | 73 | ATG | TAA | |
| trnG | H | 9742 | 9813 | 72 | 23 | |||
| nad3 | H | 9814 | 10162 | 349 | 0 | ATG | T-- | |
| trnR | H | 10163 | 10231 | 69 | 0 | |||
| nad4l | H | 10232 | 10528 | 297 | 0 | ATG | TAA | |
| nad4 | H | 10522 | 11896 | 1375 | -7 | ATG | T-- | |
| trnH | H | 11906 | 11974 | 69 | 9 | |||
| trnS1 | H | 11975 | 12043 | 69 | 0 | |||
| trnL1 | H | 12048 | 12120 | 73 | 4 | |||
| nad5 | H | 12124 | 13962 | 1839 | 3 | ATG | TAA | |
| nad6 | L | 13959 | 14480 | 522 | -4 | ATG | TAG | |
| trnE | L | 14482 | 14551 | 70 | 1 | |||
| cob | H | 14556 | 15698 | 1143 | 4 | ATG | TAA | |
| trnT | H | 15703 | 15774 | 72 | 4 | |||
| trnP | L | 15774 | 15845 | 72 | -1 | |||
| D-loop | H | 15846 | 16738 | 893 | 0 | |||
| Polydactylus_macrochir | Size(bp) | A% | T% | G% | C% | A+T% | G+C% | AT-skew | GC-skew |
|---|---|---|---|---|---|---|---|---|---|
| Mitogenome | 16738 | 28.4 | 25.75 | 16.36 | 29.49 | 54.15 | 45.85 | 0.049 | -0.286 |
| PCGs | 11447 | 25.6 | 27.71 | 15.65 | 31.03 | 53.32 | 46.68 | -0.039 | -0.329 |
| tRNAs | 1557 | 27.49 | 26.59 | 23.83 | 22.09 | 54.08 | 45.92 | 0.017 | 0.038 |
| rRNAs | 2710 | 31.51 | 22.77 | 20.92 | 24.8 | 54.28 | 45.72 | 0.161 | -0.085 |
| Dloop | 893 | 35.72 | 31.02 | 14.22 | 19.04 | 66.74 | 33.26 | 0.07 | -0.145 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




