Submitted:
05 December 2024
Posted:
09 December 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
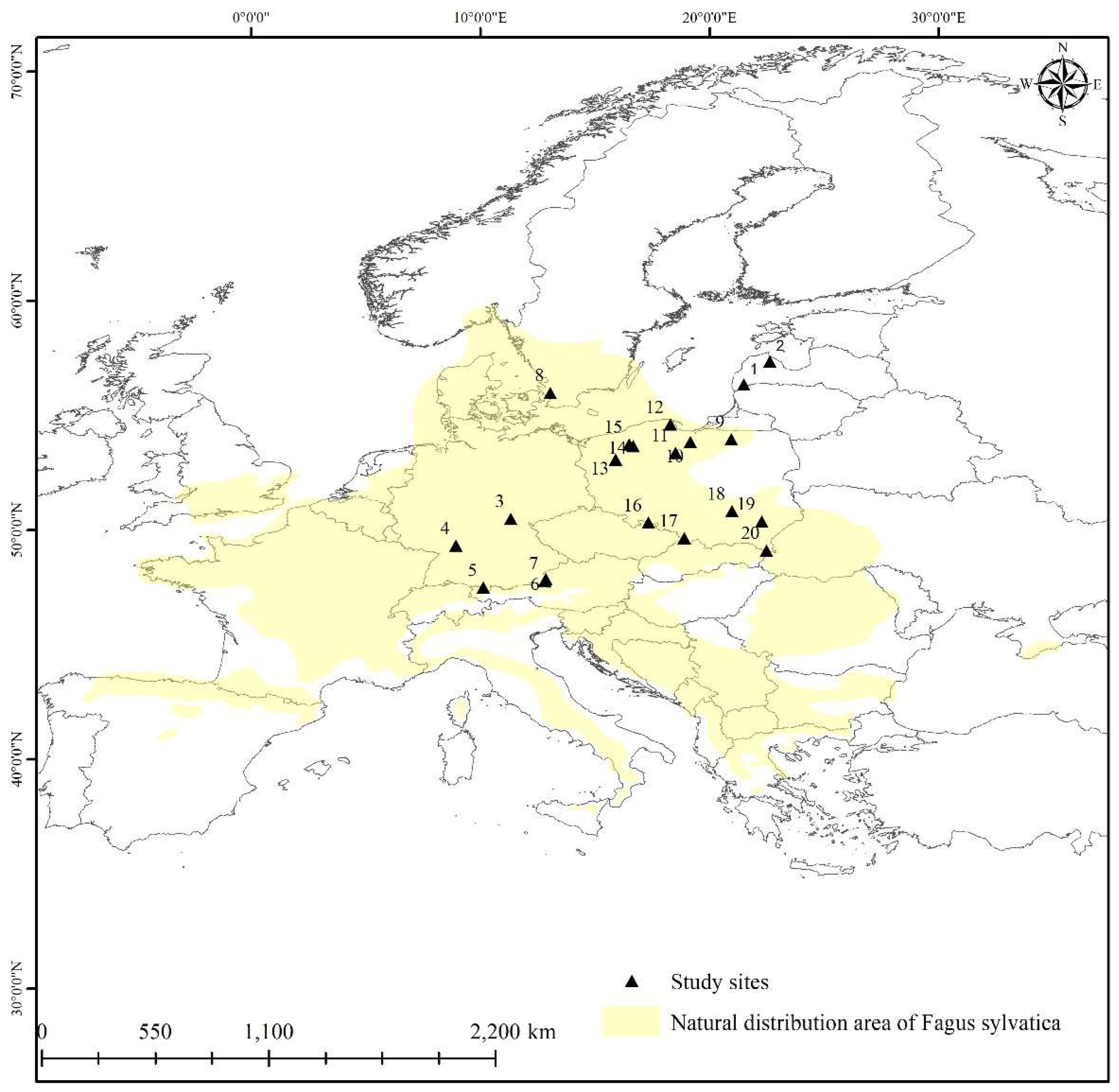
2.1. Characteristics of Latvian F.sylvatica stands and sample collection
2.2. DNA extraction and genotyping
2.3. Data analysis
3. Results
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Lenoir, J.; Gégout, J.C.; Marquet, P.A.; de Ruffray, P.; Brisse, H. A Significant Upward Shift in Plant Species Optimum Elevation During the 20th Century. Science. 2008, 320, 1768–1771. [CrossRef]
- Parmesan, C. Ecological and Evolutionary Responses to Recent Climate Change. Annu. Rev. Ecol. Evol. Syst. 2006, 37, 637–669. [CrossRef]
- Cortés, A.J.; Restrepo-Montoya, M.; Bedoya-Canas, L.E. Modern Strategies to Assess and Breed Forest Tree Adaptation to Changing Climate. Front. Plant Sci. 2020, 11. [CrossRef]
- Kreyling, J.; Buhk, C.; Backhaus, S.; Hallinger, M.; Huber, G.; Huber, L.; Jentsch, A.; Konnert, M.; Thiel, D.; Wilmking, M.; et al. Local adaptations to frost in marginal and central populations of the dominant forest tree Fagus sylvatica L. as affected by temperature and extreme drought in common garden experiments. Ecol. Evol. 2014, 4, 594–605. [CrossRef]
- Graudal, L.; Aravanopoulos, F.; Bennadji, Z.; Changtragoon, S.; Fady, B.; Kjær, E.D.; Loo, J.; Ramamonjisoa, L.; Vendramin, G.G. Global to local genetic diversity indicators of evolutionary potential in tree species within and outside forests. For. Ecol. Manage. 2014, 333, 35–51. [CrossRef]
- Williams, M.I.; Dumroese, R.K. Preparing for Climate Change: Forestry and Assisted Migration. J. For. 2013, 111, 287–297. [CrossRef]
- Pedlar, J.H.; McKenney, D.W.; Aubin, I.; Beardmore, T.; Beaulieu, J.; Iverson, L.; O’Neill, G.A.; Winder, R.S.; Ste-Marie, C. Placing Forestry in the Assisted Migration Debate. Bioscience 2012, 62, 835–842. [CrossRef]
- Ste-Marie, C. Adapting Sustainable Forest Management to Climate Change: A Review of Assisted Tree Migration and its Potential Role in Adapting Sustainable Forest Management to Climate Change. 2014, 28.
- Magri, D. Patterns of post-glacial spread and the extent of glacial refugia of European beech (Fagus sylvatica). J. Biogeogr. 2008, 35, 450–463. [CrossRef]
- Merzeau, D.; Comps, B.; Thiébaut, B.; Cuguen, J.; Letouzey, J. Genetic structure of natural stands of Fagus sylvatica L. (beech). Heredity (Edinb). 1994, 72, 269–277. [CrossRef]
- Oddou-Muratorio, S.; Bontemps, A.; Klein, E.K.; Chybicki, I.; Vendramin, G.G.; Suyama, Y. Comparison of direct and indirect genetic methods for estimating seed and pollen dispersal in Fagus sylvatica and Fagus crenata. For. Ecol. Manage. 2010, 259, 2151–2159. [CrossRef]
- von Wühlisch, G. Technical Guidelines for genetic conservation and use for European beech (Fagus sylvatica).; Bioversity International: Rome, 2008; ISBN 978-92-9043-787-1.
- Leuschner, C.; Meier, I.C.; Hertel, D. On the niche breadth of Fagus sylvatica: Soil nutrient status in 50 Central European beech stands on a broad range of bedrock types. Ann. For. Sci. 2006, 63, 355–368. [CrossRef]
- Matisone, I.; Jaunslaviete, I.; Adamovičs, A.; Matisons, R.; Jansons, Ā. Response of underplanted European beech to shelterwood thinning at the northeasternmost forpost plantation in Europe. New For. 2024, 55, 1267–1281. [CrossRef]
- Pretzsch, H.; del Río, M.; Ammer, C.; Avdagic, A.; Barbeito, I.; Bielak, K.; Brazaitis, G.; Coll, L.; Dirnberger, G.; Drössler, L.; et al. Growth and yield of mixed versus pure stands of Scots pine (Pinus sylvestris L.) and European beech (Fagus sylvatica L.) analysed along a productivity gradient through Europe. Eur. J. For. Res. 2015, 134, 927–947. [CrossRef]
- Peters, R. Beech Forests; Springer Netherlands: Dordrecht, 1997; ISBN 978-90-481-4824-0.
- Gailis, A.; Šmaukstelis, E. Oak and beech resources in Latvia. In First EUFORGEN Meeting on Social Broadleaves, Bordeaux, France, 23–25 October 1997; Turok, J., Kremer, A., De Vries, S.M.G., Eds.; International Plant Genetic Resources Institute: Rome, 1998; pp. 121–126.
- Matisons, R.; Šņepsts, G.; Puriņa, L.; Donis, J.; Jansons, Ā. Dominant height growth of European beech at the northeasternmost stands in Europe. Silva Fenn. 2018, 52. [CrossRef]
- 2023; European Commission Guidelines on closer-to-nature forest management; Publications Office of the European Union, 2023;
- Dreimanis, A. Dižskābaržu audžu ražība Šķēdes mežu novadā [Productivity of European beech in Šķede forest district]. LLU Raksti 2006, 16, 97–100.
- Puriņa, L.; Neimane, U.; Džeriņa, B.; Jansons, Ā. Eiropas dižskābarža (Fagus sylvatica L.) atjaunošanos ietekmējošie faktori [Factors affecting regeneration of European beech (Fagus sylvatica L.)]. Mežzinātne 2013, 27, 67–76.
- Jansons, Ā.; Matisons, R.; Puriņa, L.; Neimane, U.; Jansons, J. Relationships between climatic variables and Tree-Ring width of European beech and European larch growing outside of their natural distribution area. Silva Fenn. 2015, 49, 1–8. [CrossRef]
- Augustaitis, A.; Kliučius, A.; Marozas, V.; Pilkauskas, M.; Augustaitiene, I.; Vitas, A.; Staszewski, T.; Jansons, A.; Dreimanis, A. Sensitivity of European beech trees to unfavorable environmental factors on the edge and outside of their distribution range in northeastern Europe. iForest - Biogeosciences For. 2016, 9, 259–269. [CrossRef]
- Bolte, A.; Czajkowski, T.; Kompa, T. The north-eastern distribution range of European beech - A review. Forestry 2007, 80, 413–429. [CrossRef]
- Buras, A.; Menzel, A. Projecting Tree Species Composition Changes of European Forests for 2061–2090 Under RCP 4.5 and RCP 8.5 Scenarios. Front. Plant Sci. 2019, 9. [CrossRef]
- Bolte, A.; Hilbrig, L.; Grundmann, B.; Kampf, F.; Brunet, J.; Roloff, A. Climate change impacts on stand structure and competitive interactions in a southern Swedish spruce–beech forest. Eur. J. For. Res. 2010, 129, 261–276. [CrossRef]
- Gallois, A.; Audran, J.C.; Burrus, M. Assessment of genetic relationships and population discrimination among Fagus sylvatica L. by RAPD. Theor. Appl. Genet. 1998, 97, 211–219. [CrossRef]
- Pastorelli, R.; Smulders, M.J.M.; Van’t Westende, W.P.C.; Vosman, B.; Giannini, R.; Vettori, C.; Vendramin, G.G. Characterization of microsatellite markers in Fagus sylvatica L. and Fagus orientalis Lipsky. Mol. Ecol. Notes 2003, 3, 76–78.
- Bilela, S.; Dounavi, A.; Fussi, B.; Konnert, M.; Holst, J.; Mayer, H.; Rennenberg, H.; Simon, J. Natural regeneration of Fagus sylvatica L. adapts with maturation to warmer and drier microclimatic conditions. For. Ecol. Manage. 2012, 275, 60–67. [CrossRef]
- Lefèvre, S.; Wagner, S.; Petit, R.J.; De Lafontaine, G. Multiplexed microsatellite markers for genetic studies of beech. Mol. Ecol. Resour. 2012, 12, 484–491. [CrossRef]
- Stefanini, C.; Csilléry, K.; Ulaszewski, B.; Burczyk, J.; Schaepman, M.E.; Schuman, M.C. A novel synthesis of two decades of microsatellite studies on European beech reveals decreasing genetic diversity from glacial refugia. Tree Genet. Genomes 2023, 19, 3. [CrossRef]
- Kempf, M.; Konnert, M. Distribution of genetic diversity in Fagus sylvatica at the north-eastern edge of the natural range. Silva Fenn. 2016, 50, 1–17. [CrossRef]
- Demesure, B.; Comps, B.; Petit, R.J. Chloroplast DNA Phylogeography of the Common Beech (Fagus sylvatica L.) in Europe. Evolution (N. Y). 1996, 50, 2515. [CrossRef]
- Vettori, C.; Vendramin, G.G.; Anzidei, M.; Pastorelli, R.; Paffetti, D.; Giannini, R. Geographic distribution of chloroplast variation in Italian populations of beech (Fagus sylvatica L.). Theor. Appl. Genet. 2004, 109, 1–9. [CrossRef]
- Ramesh, P.; Mallikarjuna, G.; Sameena, S.; Kumar, A.; Gurulakshmi, K.; Reddy, B.V.; Reddy, P.C.O.; Sekhar, A.C. Advancements in molecular marker technologies and their applications in diversity studies. J. Biosci. 2020, 45, 123. [CrossRef]
- Buiteveld, J.; Vendramin, G.G.; Leonardi, S.; Kamer, K.; Geburek, T. Genetic diversity and differentiation in European beech (Fagus sylvatica L.) stands varying in management history. For. Ecol. Manage. 2007, 247, 98–106. [CrossRef]
- Westergren, M.; Bozic, G.; Ferreira, A.; Kraigher, H. Insignificant effect of management using irregular shelterwood system on the genetic diversity of European beech (Fagus sylvatica L.): A case study of managed stand and old growth forest in Slovenia. For. Ecol. Manage. 2015, 335, 51–59. [CrossRef]
- Frank, A.; Pluess, A.R.; Howe, G.T.; Sperisen, C.; Heiri, C. Quantitative genetic differentiation and phenotypic plasticity of European beech in a heterogeneous landscape: Indications for past climate adaptation. Perspect. Plant Ecol. Evol. Syst. 2017, 26, 1–13. [CrossRef]
- Vornam, B.; Decarli, N.; Gailing, O. Spatial Distribution of Genetic Variation in a Natural Beech Stand (Fagus sylvaticaL.) Based on Microsatellite Markers. Conserv. Genet. 2004, 5, 561–570. [CrossRef]
- Kramer, K.; Buiteveld, J.; Forstreuter, M.; Geburek, T.; Leonardi, S.; Menozzi, P.; Povillon, F.; Schelhaas, M.J.; Teissier du Cros, E.; Vendramin, G.G.; et al. Bridging the gap between ecophysiological and genetic knowledge to assess the adaptive potential of European beech. Ecol. Modell. 2008, 216, 333–353. [CrossRef]
- Rajendra, K.C.; Seifert, S.; Prinz, K.; Gailing, O.; Finkeldey, R. Subtle human impacts on neutral genetic diversity and spatial patterns of genetic variation in European beech (Fagus sylvatica). For. Ecol. Manage. 2014, 319, 138–149. [CrossRef]
- Kembrytė, R.; Danusevičius, D.; Baliuckas, V.; Buchovska, J. Phenology Is Associated with Genetic and Stem Morphotype Variation in European Beech (Fagus sylvatica L.) Stands. Forests 2022, 13, 664. [CrossRef]
- Matisons, R.; Puriņa, L.; Adamovičs, A.; Robalte, L.; Jansons, Ā. European beech in its northeasternmost stands in Europe: Varying climate-growth relationships among generations and diameter classes. Dendrochronologia 2017, 45, 123–131. [CrossRef]
- Jansone, D.; Matisons, R.; Kārše, V.; Bāders, E.; Kaupe, D.; Jansons, Ā. Structural Heterogeneity of European Beech (Fagus sylvatica L.) Stands at Its Northernmost Limits. Sustainability 2023, 15, 14681. [CrossRef]
- Kembrytė, R.; Danusevičius, D.; Buchovska, J.; Baliuckas, V.; Kavaliauskas, D.; Fussi, B.; Kempf, M. DNA-based tracking of historical introductions of forest trees: the case of European beech (Fagus sylvatica L.) in Lithuania. Eur. J. For. Res. 2021, 140, 435–449. [CrossRef]
- Vranckx, G.; Jacquemyn, H.; Mergeay, J.; Cox, K.; Kint, V.; Muys, B.; Honnay, O. Transmission of genetic variation from the adult generation to naturally established seedling cohorts in small forest stands of pedunculate oak (Quercus robur L.). For. Ecol. Manage. 2014, 312, 19–27. [CrossRef]
- Porebski, S.; Bailey, L.G.; Baum, B.R. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Report. 1997. [CrossRef]
- Van Oosterhout, C.; Hutchinson, W.F.; Wills, D.P.M.; Shipley, P. MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 2004, 4, 535–538. [CrossRef]
- Ritland, K. Estimators for pairwise relatedness and individual inbreeding coefficients. Genet. Res. 1996, 67, 175–185. [CrossRef]
- Peakall, R.; Smouse, P.E. GenALEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 2012, 28, 2537–2539. [CrossRef]
- Kalinowski, S.T.; Taper, M.L.; Marshall, T.C. Revising how the computer program cervus accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 2007, 16, 1099–1106. [CrossRef]
- Jones, O.R.; Wang, J. COLONY: A program for parentage and sibship inference from multilocus genotype data. Mol. Ecol. Resour. 2010, 10, 551–555. [CrossRef]
- Porras-Hurtado, L.; Ruiz, Y.; Santos, C.; Phillips, C.; Carracedo, Á.; Lareu, M. V. An overview of STRUCTURE: applications, parameter settings, and supporting software. Front. Genet. 2013, 4. [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [CrossRef]
- Li, Y.; Liu, J. StructureSelector : A web-based software to select and visualize the optimal number of clusters using multiple methods. Mol. Ecol. Resour. 2018, 18, 176–177. [CrossRef]
- Sandurska, E.; Ulaszewski, B.; Meyza, K.; Sztupecka, E.; Burczyk, J. Factors determining fine-scale spatial genetic structure within coexisting populations of common beech (Fagus sylvatica L.), pedunculate oak (Quercus robur L.), and sessile oak (Q. petraea (Matt.) Liebl.). Ann. For. Sci. 2024, 81, 3. [CrossRef]
- Merzeau, D.; Comps, B.; Thiebaut, B.; Letouzey, J. Estimation of Fagus sylvatica L mating system parameters in natural populations. Ann. des Sci. For. 1994, 51, 163–173. [CrossRef]
- Balloux, F. Heterozygote excess in small populations and the heterozygote-excess effective population size. Evolution (N. Y). 2004, 58, 1891–1900. [CrossRef]
- Leuschner, C. Drought response of European beech (Fagus sylvatica L.)—A review. Perspect. Plant Ecol. Evol. Syst. 2020, 47, 125576. [CrossRef]
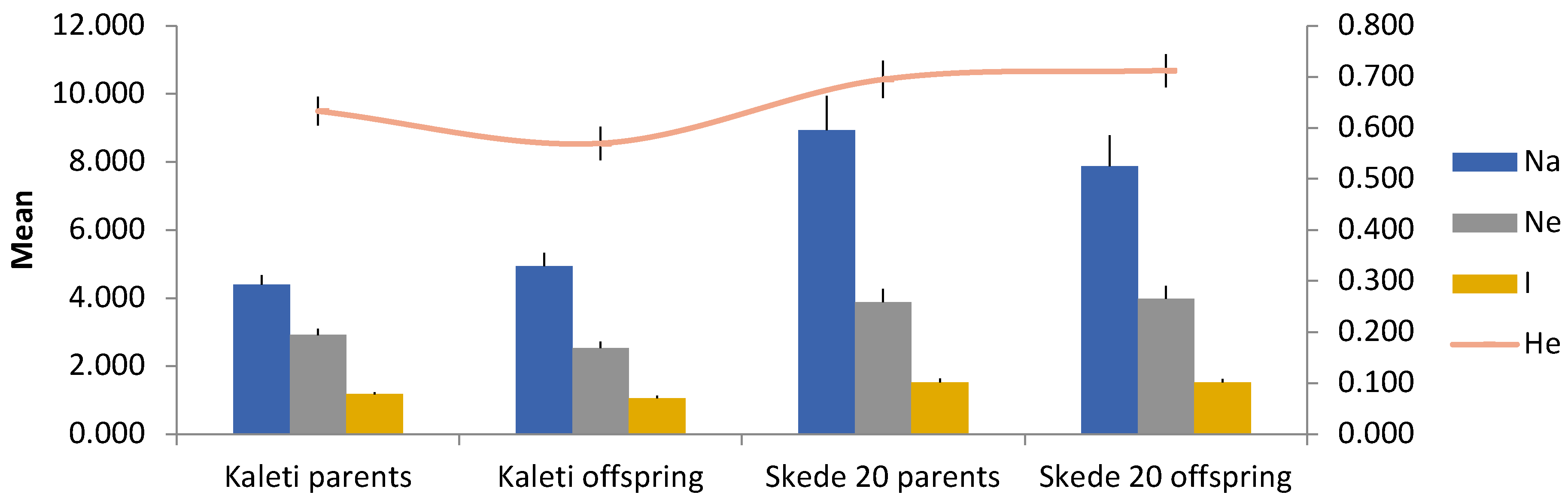

| Pop | Na (SE) | Ne (SE) | I (SE) | Ho (SE) | He (SE) | F (SE) |
|---|---|---|---|---|---|---|
| Kaleti parents | 4.400 (0.273) | 2.912 (0.184) | 1.172 (0.059) | 0.697 (0.041) | 0.633 (0.028) | -0.100 (0.036) |
| Kaleti offspring | 4.933 (0.396) | 2.524 (0.197) | 1.059 (0.065) | 0.588 (0.035) | 0.569 (0.033) | -0.040 (0.031) |
| Skede 20 parents | 8.933 (1.007) | 3.881 (0.389) | 1.516 (0.114) | 0.672 (0.037) | 0.695 (0.036) | 0.030 (0.027) |
| Skede 20 offspring | 7.867 (0.920) | 3.983 (0.378) | 1.523 (0.104) | 0.718 (0.040) | 0.712 (0.032) | -0.012 (0.038) |
| Stand | Na (SE) | Ne (SE) | I (SE) | Ho (SE) | He (SE) | F (SE) |
|---|---|---|---|---|---|---|
| Skede_20 | 8.500 (0.976) | 3.722 (0.382) | 1.474 (0.114) | 0.669 (0.040) | 0.685 (0.037) | 0.021 (0.027) |
| Skede_21 | 7.286 (0.952) | 3.713 (0.386) | 1.456 (0.124) | 0.649 (0.043) | 0.676 (0.044) | 0.031 (0.029) |
| Skede_23 | 7.000 (0.864) | 3.553 (0.413) | 1.396 (0.125) | 0.618 (0.047) | 0.658 (0.045) | 0.058 (0.03) |
| Kaleti | 4.286 (0.266) | 2.956 (0.192) | 1.177 (0.063) | 0.707 (0.042) | 0.638 (0.030) | -0.107 (0.038) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





