Submitted:
24 November 2024
Posted:
25 November 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
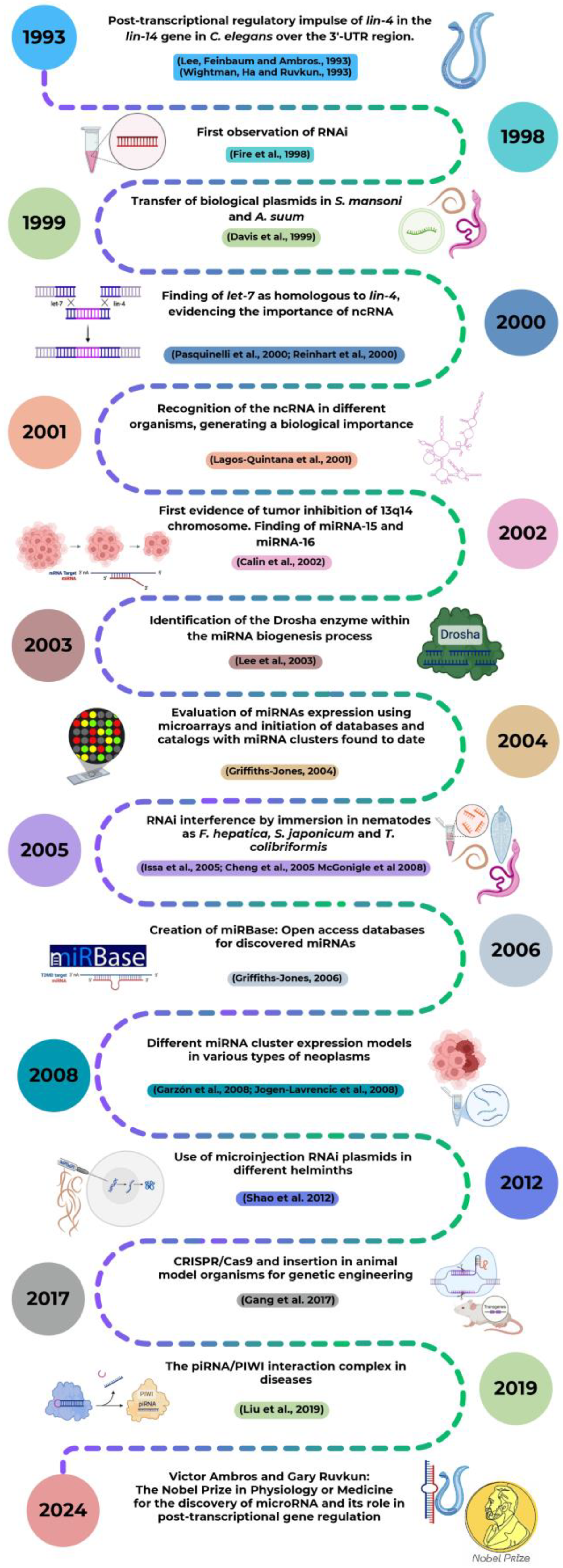
2. A Look at the Past in miRNAs: Discovery and Historical Landmarks
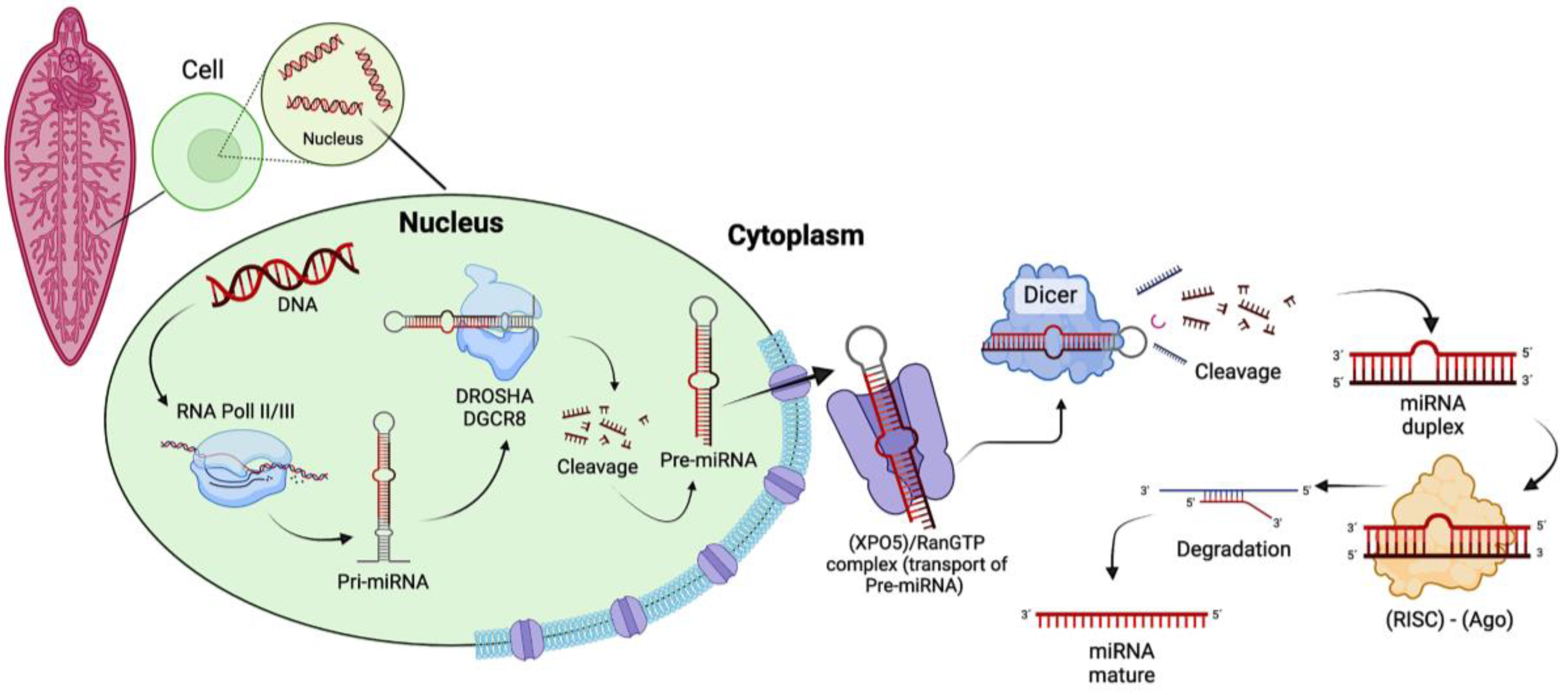
3. Location and Biogenesis of miRNAs
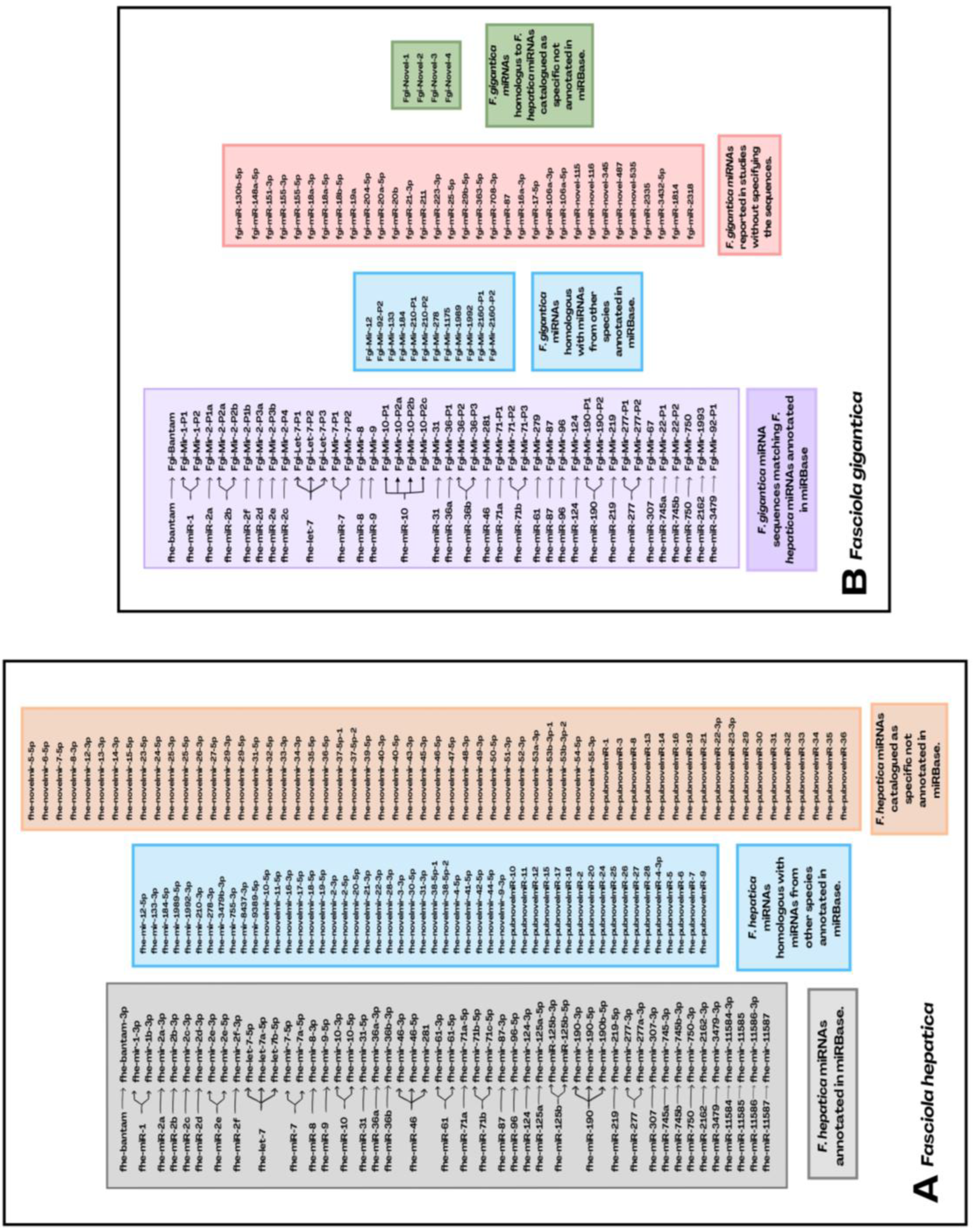
5. Unraveling the miRNome in Fasciolosis
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Available online:. Available online: https://www.who.int/teams/control-of-neglected-tropical-diseases (accessed on 16 September 2024).
- Cabada, M.M.; White Jr., C. A. New developments in epidemiology, diagnosis, and treatment of fascioliasis. Curr. Opin. Infect. Dis. 2012, 25, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Nyindo, M.; Lukambagire, A. Fascioliasis: An Ongoing Zoonotic Trematode Infection. Biomed. Res. Int. 2015, 2015, 786195. [Google Scholar] [CrossRef]
- Rufino-Moya, P.J.; Zafra, R.; Martínez-Moreno, A.; Buffoni, L.; Valderas, E.; Pérez, J.; Molina-Hernández, V.; Ruiz-Campillo, M.T.; Herrera-Torres, G. , Martínez-Moreno, F.J. Advancement in Diagnosis, Treatment, and Vaccines against Fasciola hepatica: A Comprehensive Review. Pathogens 2024, 13, 669. [Google Scholar] [CrossRef] [PubMed]
- Mehmood, K.; Zhang, H.; Sabir, A.J.; Abbas, R.Z.; Ijaz, M.; Durrani, A.Z.; Saleem, M.H.; Rehman, M.U. , Iqbal, M.K.; Wang, Y.; et al. A review on epidemiology, global prevalence and economical losses of fasciolosis in ruminants. Microb. Pathog. [CrossRef]
- Kamaludeen, J; Grahan-Borwn, J; Stephens, N. ; Miller, J.; Howell, A.; Beesley, N.J.; Hodgkinson J.; Learmount, J; Williams. D. Lack of efficacy of triclabendazole against Fasciola hepatica is present on sheep farms in three regions of England, and Wales. Vet. Rec. 2019, 184, 502. [Google Scholar] [CrossRef] [PubMed]
- Kelley, J.M.; Elliot, T.P.; Beddoe, T.; Anderson, G.; Skuce, P.; Spithill, T.W. Current Threat of Triclabendazole Resistance in Fasciola hepatica. Trends Parasitol. 2016, 32, 458–469. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.N.; Winters, A.; Milic, I.; Devitt, A.; Cookson, A.; Brophy, P.M.; Morphew, R.M. Evidence of sequestration of triclabendazole and associated metabolites by extracellular vesicles of Fasciola hepatica. Sci. Rep. 2020, 10, 13445. [Google Scholar] [CrossRef]
- Beesley, N.J.; Cwiklinski, K.; Allen, K.; Hoyle, R.C.; Spithill, T.W.; La Course, E.J.; Williams, D.J.L.; Paterson, S; Hodgkinson, J. E. A major locus confers triclabendazole resistance in Fasciola hepatica and shows dominant inheritance. PLoS Pathog. 2023, 19, e1011081. [Google Scholar] [CrossRef]
- Lalor, R; Cwiklinski, K; Davies Calvani, N. E.; Dorey, A.; Hamon, S.; López Corrales, J.; Dalton, J.P.; De Marco Verissimo, C. Pathogenicity and virulence of the liver flukes Fasciola hepatica and Fasciola gigantica that cause the zoonosis Fasciolosis. Virulence 2021, 12, 2839–2867. [Google Scholar] [CrossRef]
- Mas-Coma, S.; Valero, M.A.; Bargues, M.D. Human and Animal Fascioliasis: Origins and Worldwide Evolving Scenario. Clin. Microbiol. Rev. 2022, 35, e0008819. [Google Scholar] [CrossRef]
- Cwiklinski, K.; de la Torre-Escudero, E.; Trelis, M.; Bernal, D.; Dufresne, P.J.; Brennan, G.P.; O’Neill, S.; Tort, J.; Paterson, S.; Marcilla, A.; et al. The Extracellular Vesicles of the Helminth Pathogen, Fasciola hepatica: Biogenesis Pathways and Cargo Molecules Involved in Parasite Pathogenesis. Mol. Cell. Proteomics 2015, 14, 3258–3257. [Google Scholar] [CrossRef]
- Durey, C.; Maizels, R.M. Helminth extracellular vesicles: Interactions with the host immune system. Mol. Immunol. 2021, 137, 124–133. [Google Scholar] [CrossRef]
- Álvarez-Rojas, C.A.; Ansell, B.R.E.; Hall, R.S.; Gasser, R.B.; Young, N.D.; Jex, A.R.; Scheerlinck, J.P.Y. Transcriptional analysis identifies key genes involved in metabolism, fibrosis/tissue repair and the immune response against Fasciola hepatica in sheep liver. Parasites & Vectors 2015, 8, 124. [Google Scholar] [CrossRef]
- Young, N.D.; Hall, R.S.; Jex, A.E.; Cantacessi, C.; Gasser, R.B. Elucidating the transcriptome of Fasciola hepatica – A key to fundamental and biotechnological discoveries for a neglected parasite. Biotechnol. Adv. 2010, 28, 222–231. [Google Scholar] [CrossRef] [PubMed]
- Cwiklinski, K.; Dalton, J.P.; Dufresne, P.J.; La Course, J.; Williams, D.J.; Hodgkinson, J.; Paterson, S. The Fasciola hepatica genome: gene duplication and polymorphism reveals adaptation to the host environment and the capacity for rapid evolution. Genome Biol. [CrossRef]
- Cwiklinski, K.; Dalton, J.P. Advances in Fasciola hepatica research using ‘omics’ technologies. Int. J. Parasitol. 2018, 48, 321–331. [Google Scholar] [CrossRef]
- Ricafrente, A.; Cwiklinski, K.; Nguyen, H.; Dalton, J.P.; Tran, N.; Donnelly, S. Stage-specific miRNAs regulate gene expression associated with growth, development and parasite-host interaction during the intra-mammalian migration of the zoonotic helminth parasite Fasciola hepatica. BMC Genomics, 2022, 23, 419. [Google Scholar] [CrossRef] [PubMed]
- Arora, N.; Tripathi, S.; Singh, A.K.; Mondal, P.; Mishra, A; Prasad, A. Micromanagement of Immune System: Role of miRNAs in Helminthic Infections. Front. Microbiol. 2017, 8, 586. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, S.; Ricafrente, A.; Cwiklinski, K.; Sais, D.; Dalton, J.P.; Tran, N.; Donnelly, S. Exploring the utility of circulating miRNAs as diagnostic biomarkers of fasciolosis. Sci. Rep. 2024, 14, 7431. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Ghildiyal, M.; Zamore, P.D. Small silencing RNAs: an expanding universe. Nat. Rev. Genet. 2009, 10, 94–108. [Google Scholar] [CrossRef]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef]
- Anglicheau, D.; Muthukumar, T.; Suthanthiran, M. MicroRNAs: small RNAs with big effects. Transplantation 2010, 90, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Chandra, S.; Vimal, D.; Sharma, D.; Rai, V.; Gupta, S.C.; Chowdhuri, D.K. Role of miRNAs in development and disease: Lessons learnt from small organisms. Life Sci. 2017, 185, 8–14. [Google Scholar] [CrossRef]
- Almeida, M.I.; Reins, R.M.; Calin, G.A. MicroRNA history: Discovery, recent applications, and next frontiers. Mutat. Res. 2011, 717, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Pirela, M; Andrade-Alviárez, D. ; Medina, L.; Castillo, C., Liempi, A., Guerrero-Muñoz, J, Ortega, Y, Maya, J.D, Rojas, V, Quiñones, W., Eds.; et al. MicroRNAs: master regulators in host-parasitic protist intertactions. Open. Biol. 2022, 12, 210395. [Google Scholar] [CrossRef]
- Rojas-Pirela, M.; Andrade-Alviárez, D.; Quiñones, W.; Rojas, M.V.; Castillo, C.; Liempi, A.; Medina, L.; Guerrero-Muñoz, J.; Fernández-Moya, A.; Ortega, Y.A. , et al. microRNAs: Critical Players during Helminth Infections. Microorganisms 2022, 11, 61. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 2004, 32, 109–111. [Google Scholar] [CrossRef]
- Kozomara, A.; Griffiths-Jones, S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2011, 39, 152–157. [Google Scholar] [CrossRef]
- Backes, C.; Fehlmann, T.; Kern, F.; Kehl, T.; Lenhof, H.; Meese, E.; Keller, A. miRCarta: a central repository for collecting miRNA candidates. Nucleic Acids Res. 2018, 46, D160–D167. [Google Scholar] [CrossRef]
- Bourdon, C.; Bardou, P.; Aujean, E.; Le Guillou, S.; Tosser-Klopp, G.; Le Provost, F. RumimiR: a detailed microRNA database focused on ruminant species. Database (Oxford). 2019, 2019, baz099. [Google Scholar] [CrossRef]
- Fromm, B.; Høye, E.; Domanska, D.; Zhong, X.; Aparicio-Puerta, E.; Ovchinnikov, V.; Umu, S.U.; Chabot, P.J.; Kang, W.; Aslanzadeh, M.; et al. MirGeneDB 2.1: toward a complete sampling of all major animal phyla. Nucleic Acids Res. 2021, 50, D204–D210. [Google Scholar] [CrossRef]
- Xu, M.; Ai, L.; Fu, J.; Nisbet, A.J.; Liu, Q.; Chen, M.; Zhou, D.; Zhu, X. Comparative characterization of microRNAs from the liver flukes Fasciola gigantica and F. hepatica. PLoS One 2012, 7, e53387. [Google Scholar] [CrossRef]
- Tran, N.; Ricafrente, A.; To, J.; Lund, M; Marques, T. M.; Gama-Carvalho, M.; Cwiklinski, K.; Dalton, J.P.; Donnely, S. Fasciola hepatica hijacks host macrophages miRNA machinery to modulate early innate immune responses. Sci. Rep. 2021, 11, 6712. [Google Scholar] [CrossRef] [PubMed]
- Britton, C.; Winter, A.D.; Marks, N.D.; Gu, H.; McNeilly, T.N.; Gillan, V.; Devaney, E. Application of small RNA technology for improved control of parasitic helminths. Vet. Parasitol. 2015, 212, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Ricafrente, A.; Nguyen, H.; Tran, N.; Donnelly, S. An Evaluation of the Fasciola hepatica miRnome Predicts a Targeted Regulation of Mammalian Innate Immune Responses. Front. Immunol. 2021, 11, 608686. [Google Scholar] [CrossRef] [PubMed]
- Fontenla, S.; Langleib, M.; de la Torre-Escudero, E.; Domínguez, M.F.; Robinson, M.W.; Tort, J. Role of Fasciola hepatica Small RNAs in the Interaction With the Mammalian Host. Front. Cell. Infect. Microbiol. 2022, 11, 812141. [Google Scholar] [CrossRef]
- Karapetyan, A.R.; Buiting, C.; Kuiper, R.A.; Coolen, M.W. Regulatory Roles for Long ncRNA and mRNA. Cancers 2013, 5, 462–490. [Google Scholar] [CrossRef]
- Ruvkun, G.; Giusto, J. The Caenorhabditis elegans heterochromic gene lin-14 encodes a nuclear protein that forms a temporal developmental switch. Nature 1989, 338, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementary to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Davis, R.E.; Parra, A.; LoVerde, P.T.; Ribeiro, E.; Glorioso, G.; Hodgson, S. Transient expression of DNA and RNA in parasitic helminths by using particle bombardment. Proc. Natl. Acad. Sci. U S A. 1999, 96, 8687–8692. [Google Scholar] [CrossRef]
- Pasquinelli, A.E.; Reinhart, B.J.; Slack, F.; Martindale, M.Q.; Kuroda, M.I.; Maller, B.; Hayward, D.C.; Ball, E.E.; Degnan, B.; Müller, P.; et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 2000, 408, 86–89. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature, 2000, 403, 901–906. [Google Scholar] [CrossRef] [PubMed]
- Lagos-Quintana, M.; Rauhut, R.; Lendeckel, W.; Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 2001, 294, 853–858. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci U S A. 2002, 99, 15524–15529. [Google Scholar] [CrossRef]
- Garzón, R.; Volinia, S.; Liu, C.; Fernandez-Cymering, C.; Palumbo, T.; Pichiorri, F.; Fabri, M.; Coombes, K.; Alder, H.; Nakamura, T. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 2008, 111, 3183–3189. [Google Scholar] [CrossRef]
- Jogen-Lavrencic, M.; Sun, S.M.; Dijkstra, M.K.; Valk, P.J.M.; Löwenberg, B. MicroRNA expression profiling in relation to the genetic heterogeneity of acute myeloid leukemia. Blood 2008, 111, 5078–5085. [Google Scholar] [CrossRef]
- Lee, Y.; Ahn, C.; Han, J.; Choi, H.; Kim, J.; Yim, J.; Lee, J.; Provost, P.; Radmark, O.; Kim, S.; Kim, V.N. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003, 425, 415–419. [Google Scholar] [CrossRef]
- Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 2004, 32, D109–D111. [Google Scholar] [CrossRef]
- Griffiths-Jones, S.; Grocock, R.J.; van Dongen, S.; Bateman, A.; Enright, A.J. miRbase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef]
- Issa, Z.; Grant, W.N.; Stasiuk, S.; Shoemaker, C.B. Development of methods for RNA interference in the sheep gastrointestinal parasite, Trichostrongylus colubriformis. Int. J. Parasitol. 2005, 35, 935–940. [Google Scholar] [CrossRef]
- Cheng, G.; Lin, J.; Jin, Y.; Fu, Z.; Jin, Y.; Zhou, Y.; Cai, Y. Dose-dependent inhibition of gynecophoral canal protein gene expression in vitro in the schistosome (Schistosoma japonicum) by RNA interference. Acta Biochim. Biophys. Sin. (Shangai) 2005, 37, 386–390. [Google Scholar] [CrossRef] [PubMed]
- McGonigle, L.; Mousley, A.; Marks, N.J.; Brennan, G.P.; Dalton, J.P.; Spithill, T.W.; Day, T.A.; Maule, A.G. The silencing of cysteine proteases in Fasciola hepatica newly excysted juveniles using RNA interference reduces gut penetration. Int. J. Parasitol. 2008, 38, 149–155. [Google Scholar] [CrossRef]
- Shao, H.; Li, X.; Nolan, T.J.; Massey Jr, H.C.; Pearce, E.J.; Lok, J.B. Transposon-mediated chromosomal integration of transgenes in the parasitic nematode Strongyloides ratti and establishment of stable transgenic lines. PLoS Pathog. 2012, 8, e1002871. [Google Scholar] [CrossRef] [PubMed]
- Gang, S.S.; Castelletto, M.L.; Bryant, A.S.; Yang, E.; Mancuso, N.; Lopez, J.B.; Pellegrini, M.; Hallem, E.A. Targeted mutagenesis in a human-parasitic nematode. PLoS Pathog. 2017, 13, e1006675. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Dou, M.; Song, X.; Dong, Y.; Liu, S.; Liu, H.; Tao, J.; Li, W.; Yin, X.; Xu, W. The emerging role of the piRNA/piwi complex in cancer. Mol. Cancer 2019, 18, 123. [Google Scholar] [CrossRef]
- Available online:. Available online: https://www.nobelprize.org/prizes/medicine/2024/press-release/ (accessed on 4 November 2024).
- Weiss, C.N.; Ito, K. A. Macro View of MicroRNAs: The Discovery of MicroRNAs and Their Role in Hematopoiesis and Hematologic Disease. Int. Rev. Cell Mol. Biol. 2017, 334, 99–175. [Google Scholar] [CrossRef]
- Quinzo, M.J.; Perteguer, M.J.; Brindley, P.J.; Loukas, A.; Sotillo, J. Transgenesis in parasitic helminths: a brief history and prospects for the future. Parasit Vectors 2022, 15, 110. [Google Scholar] [CrossRef]
- Liu, B.; Shyr, Y.; Cai, J.; Liu, Q. Interplay between miRNAs and host genes and their role in cancer. Brief Func. Genomics 2019, 18, 155–266. [Google Scholar] [CrossRef]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Treiber, T.; Treiber, N.; Meister, G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell Biol. 2019, 20, 5–20. [Google Scholar] [CrossRef]
- Denli, A.M.; Tops, B.B.J.; Plasterk, R.H.A.; Ketting, R.F.; Hannon, G.J. Processing of primary microRNAs by the Microprocessor complex. Nature. [CrossRef]
- Saliminejad, K.; Khorshid, H.R.K.; Fard, S.S.; Ghaffari, S.H. An overview of microRNAs: Biology, functions, therapeutics, and analysis methods. J. Cell Physiol. 2019, 234, 5451–5465. [Google Scholar] [CrossRef] [PubMed]
- Guasconi, L.; Serradell, M.C.; Garro, A.P.; Iaconelli, L.; Masih, D.T. C-type lectins on macrophages participate in the immunomodulatory response to Fasciola hepatica products. Immunology 2011, 133, 386–396. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Campillo, M.T.; Molina-Hernández, V.; Pérez, J.; Pacheco, I.L.; Pérez, R.; Escamilla, A.; Martínez-Moreno, F.J.; Martínez-Moreno, A.; Zafra, R. Study of peritoneal macrophage immunophenotype in sheep experimentally infected with Fasciola hepatica. Vet. Parasitol. 2018, 257, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Dalton, J.P.; Robinson, M.W.; Mulcahy, G.; O’Neill, S.M.; Donnelly, S. Immunomodulatory molecules of Fasciola hepatica: candidates for both vaccine and immunotherapeutic development. Vet. Parasitol. 2013, 195, 272–285. [Google Scholar] [CrossRef]
- Santiago-Figueroa, O.; Espino, A.M. Fasciola hepatica Fatty Acid Binding Protein Induces the Alternative Activation of Human Macrophages. Infect. Immun. 2014, 82, 5005–5012. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Campillo, M.T.; Barrero-Torres, D.M. , Abril, N.; Pérez, J.; Zafra, R.; Buffoni, L.; Martínez-Moreno, Á.; Martínez-Moreno, F.J.; Molina-Hernández, V. Fasciola hepatica primoinfections and reinfections in sheep drive distinct Th1/Th2/Treg immune responses in liver and hepatic lymph node at early and late stages. Vet. Res. 2023, 54, 2. [Google Scholar] [CrossRef]
- Zheng, Y.; Cai, X.; Bradley, J.E. microRNAs in parasites and parasite infection. RNA Biol. 2013, 10, 371–379. [Google Scholar] [CrossRef]
- Cucher, M.A.; Ancarola, M.E.; Kamenetzky, L. The challenging world of extracellular RNAs of helminth parasites. Mol. Immunol. 2021, 234, 150–160. [Google Scholar] [CrossRef]
- Alizadeh, Z.; Mahami-Oskouei, M.; Spotin, A.; Kazemi, T.; Ahmadpour, E.; Cai, P.; Shanehbandi, D.; Shekari, N. Parasite-derived microRNAs in plasma as novel promising biomarkers for the early detection of hydatid cyst infection and post-surgery follow-up. Acta Trop. 2020, 202, 105255. [Google Scholar] [CrossRef]
- Britton, C.; Laing, R.; Devaney, E. Small RNAs in parasitic nematodes – forms and functions. Parasitology 2020, 147, 855–864. [Google Scholar] [CrossRef]
- Fromm, B.; Trelis, M.; Hackenberg, M.; Cantalapiedra, F.; Bernal, D.; Marcilla, A. The revised microRNA complement of Fasciola hepatica reveals a plethora of overlooked microRNAs and evidence for enrichment of immune-regulatory microRNAs in extracellular vesicles. Int. J. Parasitol. 2015, 45, 697–702. [Google Scholar] [CrossRef]
- Murphy, A.J.; Guyre, P.M.; Pioli, P.A. Estradiol suppresses NF-kappa B activation through coordinated regulation of let-7a and miR-125b in primary human macrophages. J. Immunol. 2010, 184, 5029–5037. [Google Scholar] [CrossRef] [PubMed]
- Duroux-Richard, I.; Roubert, C.; Ammari, M.; Présumey, J.; Grün, J.R.; Häupl, T.; Grützkau, A.; Lecellier, C.; Boitez, V.; Codogno, P.; et al. miR-125b controls monocyte adaptation to inflammation through mitochondrial metabolism and dynamics. Blood 2016, 128, 3125–3136. [Google Scholar] [CrossRef] [PubMed]
- Chaudhuri, A.A.; So, A.Y.; Sinha, N.; Gibson, W.S.J.; Taganov, K.D.; O’Connell, R.M.; Baltimore, D. MicroRNA-125b potentiates macrophage activation. J. Immunol. 2011, 187, 5062–5068. [Google Scholar] [CrossRef]
- Fontenla, S.; Dell’Oca, N.; Smircich, P.; Tort, J.F.; Siles-Lucas, M. The miRnome of Fasciola hepatica juveniles endorses the existence of a reduced set of highly divergent micro RNAs in parasitic flatworms. Int. J. Parasitol. 2015, 45, 901–913. [Google Scholar] [CrossRef] [PubMed]
- Hoy, A.M.; Lundie, R.J.; Ivens, A.; Quintana, J.F.; Nausch, N.; Forster, T.; Jones, F.; Kabatereine, N.B. Parasite-Derived MicroRNAs in Host Serum As Novel Biomarkers of Helminth Infection. PLoS Negl. Trop. Dis. 2014, 8, e2701. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Ambros, V.R. Caenorhabditis elegans microRNAs of the let-7 family act in innate immune response circuits and confer robust developmental timing against pathogen stress. Proc. Natl. Acad. Sci. U S A 2015, 112, E2366–E2375. [Google Scholar] [CrossRef]
- Stark, A.; Brennecke, J.; Russell, R.B.; Cohen, S.M. Identification of Drosophila MicroRNA targets. PLoS Biol. 2003, 1, E60. [Google Scholar] [CrossRef]
- Rubio, M.; Belles, X. Subtle roles of microRNAs let-7, miR-100 and miR-125 on wing morphogenesis in hemimetabolan metamorphosis. J. Insect. Physiol. 2013, 59, 1089–1094. [Google Scholar] [CrossRef]
- Sun, X.; Jiao, X.; Pestell, T.G.; Fan, C.; Qin, S.; Mirabelli, E.; Ren, H.; Pestell, R.G. MicroRNAs and cancer stem cells: The sword and the shield. Oncogene 2014, 33, 4967–4977. [Google Scholar] [CrossRef]
- Banerjee, S.; Xie, N.; Cui, H.; Tan, Z.; Yang, S.; Icyuz, M.; Abraham, E.; Liu, G. MicroRNA let-7c regulates macrophage polarization. J. Immunol. 2013, 190, 6542–6549. [Google Scholar] [CrossRef] [PubMed]
- Boulan, L.; Martín, D.; Milán, M. Bantam miRNA promotes systemic growth by connecting insulin signaling and ecdysone production. Curr. Biol. 2013, 23, 473–478. [Google Scholar] [CrossRef]
- Xue, X.; Sun, J.; Zhang, Q. Wanf, Z.; Huang, Y.; Pan, W. Identification and characterization of novel microRNAs from Schistosoma japonicum. PLoS One 2008, 3, e4034. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Zhu, L.; Wang, J.; Qiu, L.; Chen, Y.; Davis, R.E.; Cheng, G. Schistosoma japonicum extracellular vesicle miRNA cargo regulates host macrophage functions facilitating parasitism. PLoS Pathog. 2019, 15, e1007817. [Google Scholar] [CrossRef]
- Lopes-Junior, E.; Bertevello, C.; Silveira, G.; Guedes, C.; Rodrigues, G.; Ribeiro, V.; Amaral, M.; Kanamura, C.; Pinto, P.; Krüger, R.; et al. Human tumor necrosis factor alpha affects the egg-laying dynamics and glucose metabolism of Schistosoma mansoni adult worms in vitro. Parasit. Vectors 2022, 15, 176. [Google Scholar] [CrossRef]
- Buck, A.H.; Coakley, G.; Simbari, F.; McSorley, H.J.; Quintana, J.F.; Le Bihan, T.; Kumar, S.; Abreu-Goodger, C.; Lear, M.; Harcus, Y.; et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 2014, 5, 5488. [Google Scholar] [CrossRef]
- Wu, J.; Liao, Y.; Li, D.; Zhu, Z.; Zhang, L.; Wu, Z.; He, P.; Wang, L. Extracellular vesicles derived from Trichinella Spiralis larvae promote the polarization of macrophages to M2b type and inhibit the activation of fibroblasts. Front. Immunol. 2022, 13, 974322. [Google Scholar] [CrossRef] [PubMed]
- Fromm, B.; Ovchinnikov, V.; Høye, E.; Bernal, D.; Hackenberg, M.; Marcilla, A. On the presence and immunoregulatory functions of extracellular microRNAs in the trematode Fasciola hepatica. Parasite Immunol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Montaner, S.; Galiano, A.; Trelis, M.; Martin-Jaular, L.; del Portillo, H.A.; Bernal, D.; Marcilla, A. The Role of Extracellular Vesicles in Modulating the Host Immune Response during Parasitic Infections. Front. Immunol. 2014, 5, 433. [Google Scholar] [CrossRef]
- Guo, X.; Guo, A. Profiling circulating microRNAs in serum of Fasciola gigantica-infected buffalo. Mol. Biochem. Parasitol. 2019, 232, 111201. [Google Scholar] [CrossRef]
- Ricciardi, A.; Bennuru, S.; Tariq, S.; Kaur, S.; Wu, W.; Elkahloun, A.G.; Arakelyan, A.; Shaik, J.; Dorward, D.W.; Nutman, T.B.; Semnani, R.T. Extracellular vesicles released from the filarial parasite Brugia malayi downregulate the host m TOR pathway. PLoS Negl. Trop. Dis. 2021, 15, e0008884. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Guo, X.; He, W.; Shao, Z.; Zhang, X.; Yang, J.; Shen, Y.; Luo, X.; Cao, J. Effects of Echinococcus multilocularis miR-71 mimics on murine macrophage RAW264.7 cells. Int. Immunopharmacol. 2016, 34, 259–262. [Google Scholar] [CrossRef] [PubMed]
- Robb, E.; McCammick, E.M.; Wells, D.; McVeigh, P.; Gardiner, E.; Armstrong, R.; McCusker, P.; Mousley, A.; Clarke, N.; Marks, N.J.; et al. Transcriptomic analysis supports a role for the nervous system in regulating growth and development of Fasciola hepatica juveniles. PLoS Negl. Trop. Dis. 2022, 16, e0010854. [Google Scholar] [CrossRef]
- Shi, W.; He, J.; Mei, X.; Lu, K.; Zeng, Z.; Zhang, Y.; Sheng, Z.; Elsheikha, H.M.; Huang, W.; Zhu, X. Dysregulation of hepatic microRNA expression in C57BL/6 mice affected by excretory-secretory products of Fasciola gigantica. PLoS Negl. Trop. Dis. 2020, 14, e0008951. [Google Scholar] [CrossRef]
- Saravanan, S.; Thirugnanasambantham, K.; Hanieh, H.; Karikalan, K.; Sekar, D.; Rajagopalan, R.; Islam, V.I.H. miRNA-24 and miRNA-466I-5p controls inflammation in rat hepatocytes. Cell. Mol. Immunol. 2014, 12, 113–115. [Google Scholar] [CrossRef]
- El-Guendy, N.M.; Helwa, R.; El-Halawany, M.S.; Ali, S.A.R.; Aly, M.T.; Alieldin, N.H.; Fouad, S.A.H.; Saeid, H.; Abdel-Wahab, A.A. The Liver MicroRNA Expression Profiles Associated With Chronic Hepatitis C Virus (HCV) Genotype-4 Infection: A Preliminary Study. Hepat. Mon. 2016, 16, e33881. [Google Scholar] [CrossRef] [PubMed]
- Guidi, R.; Wedeles, C.J.; Wilson, M.S. ncRNAs in Type-2 Immunity. Noncoding RNA 2020, 6, 10. [Google Scholar] [CrossRef]
- Ovchinnikov, V.Y. Kashina, E.V.; Mordninov, V.A.; Fromm, B. EV-transported microRNAs of Schistosoma mansoni and Fasciola hepatica: Potential targets in definitive hosts. Infect. Genet. Evol. 2020, 85, 104528. [Google Scholar] [CrossRef]
- Kimura, K.; Hohjoh, H.; Fukuoka, M.; Sato, W.; Oki, S.; Tomi, C.; Yamaguchi, H.; Kondo, Takahashi, R. ; Yamamura, T. Circulating exosomes suppress the induction of regulatory T cells via let-7i in multiple sclerosis. Nat. Commun, 2018, 9, 17. [Google Scholar] [CrossRef]
- Hu, R.; Zhang, X.; Ma, Q.; Elsheikha, H.M.; Ehsan, M.; Zhao, Q.; Fromm, B.; Zhu, X. Differential expression of microRNAs and t RNA fragments mediate the adaptation of the liver fluke Fasciola gigantica to its intermediate snail and definitive mammalian hosts. Int. J. Parasitol. 2021, 51, 405–414. [Google Scholar] [CrossRef]
- Wang, S.; Chen, D.; He, J.; Zheng, W.; Tian, A.; Zhao, G.; Elsheikha, H.M.; Zhu, X. Fasciola gigantica-Derived Excretory-Secretory Products Alter the Expression of mRNAs, miRNAs, lncRNAs, and circRNAs Involved in the Immune Response and Metabolism in Goat Peripheral Blood Mononuclear Cells. Front. Immunol. 2021, 12, 653755. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-López, C.M.; Trelis, M.; Jara, L.; Cantalapiedra, F.; Marcilla, A.; Bernal, D. Diversity of extracellular vesicles from different developmental stages of Fasciola hepatica. Int. J. Parasitol. 2020, 50, 663–669. [Google Scholar] [CrossRef] [PubMed]
- Herron, C.M.; O’Connor, A.; Robb, E.; McCammick, E.; Hill, C.; Marks, N.; Robinson, M.W.; Maula, A.G.; McVeigh, P. Developmental Regulation and Functional Prediction of microRNAs in an Expanded Fasciola hepatica miRNome. Front. Cell Infect. Microbiol. 2022, 12, 811123. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V.; Bartel, B.; Bartel, D.P.; Burge, C.B.; Carrington, J.C.; Chen, X.; Dreyfus, G.; Eddy, S.R.; Griffiths-Jones, S.; Marshall, M.; et al. A uniform system for microRNA annotation. RNA 2003, 9, 277–279. [Google Scholar] [CrossRef]
- Cai, P.; Hou, N.; Piao, X.; Liu, S.; Liu, H.; Yang, F.; Wang, J.; Jin, Q.; Wang, H.; Chen, Q. Profiles of small non-coding RNAs in Schistosoma japonicum during development, PLoS Negl. Trop. Dis. 2011, 5, e1256. [Google Scholar] [CrossRef]
- Macchiaroli, N.; Cucher, M.; Kamenetzky, L.; Yones, C.; Bugnon, L.; Berriman, M.; Olson, P.D.; Rosenzvit, M.C. Identification and expression profiling of microRNAs in Hymenolepis. Int. J. Parasitol. 2019, 49, 2111–2223. [Google Scholar] [CrossRef]
- McVeigh, P.; McCammick, E.; Robb, E.; Brophy, P.; Morphew, R.M.; Marks, N.J.; Maule, A.G. Discovery of long non-coding RNAs in the liver fluke, Fasciola hepatica. PLoS Negl. Trop. Dis. 2023, 17, e0011663. [Google Scholar] [CrossRef]
- Ulitsky, I. Interactions between short and long noncoding RNAs. FEBS Lett. 2018, 592, 2874–2883. [Google Scholar] [CrossRef]
- Zhang, M.; He, P.; Bian, Z. Long Noncoding RNAs in Neurodegenerative Diseases: Pathogenesis and Potential Implications as Clinical Biomarkers. Front. Mol. Neurosci. 2021, 14, 685143. [Google Scholar] [CrossRef]
- He, M.; Wan, J.; Zeng, H.; Tang, Y. , He, M. miR-133a-5p suppresses gastric cancer through TCF4 down-regulation. J. Gastrointest. Oncol. 2021, 12, 1007–1019. [Google Scholar] [CrossRef]
- Shandilya, U.K.; Sharma, A.; Naylor, D.; Canovas, A.; Mallard, B.; Karrow, N.A. Expression Profile of miRNA from High, Middle and Low Stress-Responding Sheep during Bacterial Endotoxin Challenge. Animals 2023, 13, 508. [Google Scholar] [CrossRef]
- Bąska, P.; Majewska, A.; Zygner, W.; Długosz, E.; Wiśniewski, M. Fasciola hepatica Excretory-Secretory Products (Fh-ES) Either Do Not Affect miRNA Expression Profile in THP-1 Macrophages or the Changes Are Undetectable by a Microarray Technique. Pathogens 2024, 13, 854. [Google Scholar] [CrossRef]
- Batan Pumeda, S. Comparison Between Next-Generation Sequencing and Microarrays for miRNA Expression in Cancer Samples. Natl. Acad. Sci. Lett. 2023. [Google Scholar] [CrossRef]
- Sais, D.; Chowdhury, S.; Dalton, J.P.; Tran, N.; Donnelly, S. Both host and parasite non-coding RNAs co-ordinate the regularion of macrophage gene expression to reduce pro-inflammatory immune responses and promote tissue repair pathways during infection with Fasciola hepatica. RNA Biol. 2024, 21, 62–77. [Google Scholar] [CrossRef]
- Alhallaf, R.; Agha, Z.; Miller, C.M.; et al. The NLRP3 inflammasome suppresses protective immunity to gastrointestinal helminth infection. Cell Rep. 2018, 23, 1085–1098. [Google Scholar] [CrossRef]
- Guan, F.; Jiang, W.; Bai, Y.; et al. Purinergic P2X7 receptor mediates the elimination of Trichinella spiralis by activating nf-κB/NLRP3/IL-1β pathway in macrophages. Infect Immun. 2021, 89, e00683–20. [Google Scholar] [CrossRef]
- De la Torre-Escudero, E.; Gerlach, J.Q.; Bennet, A.P.S.; Cwiklinski, K.; Jewhurst, H.L.; Huson, K.M.; Joshi, L.; Kilcoyne, M.; O’Neill, S.; Dalton, J.P.; Robinson, M.W. Surface molecules of extracellular vesicles secreted by the helminth pathogen Fasciola hepatica direct their internalisation by host cells. PLoS Negl. Trop. Dis. 2019, 13, e0007087. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).