Submitted:
14 November 2024
Posted:
18 November 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results


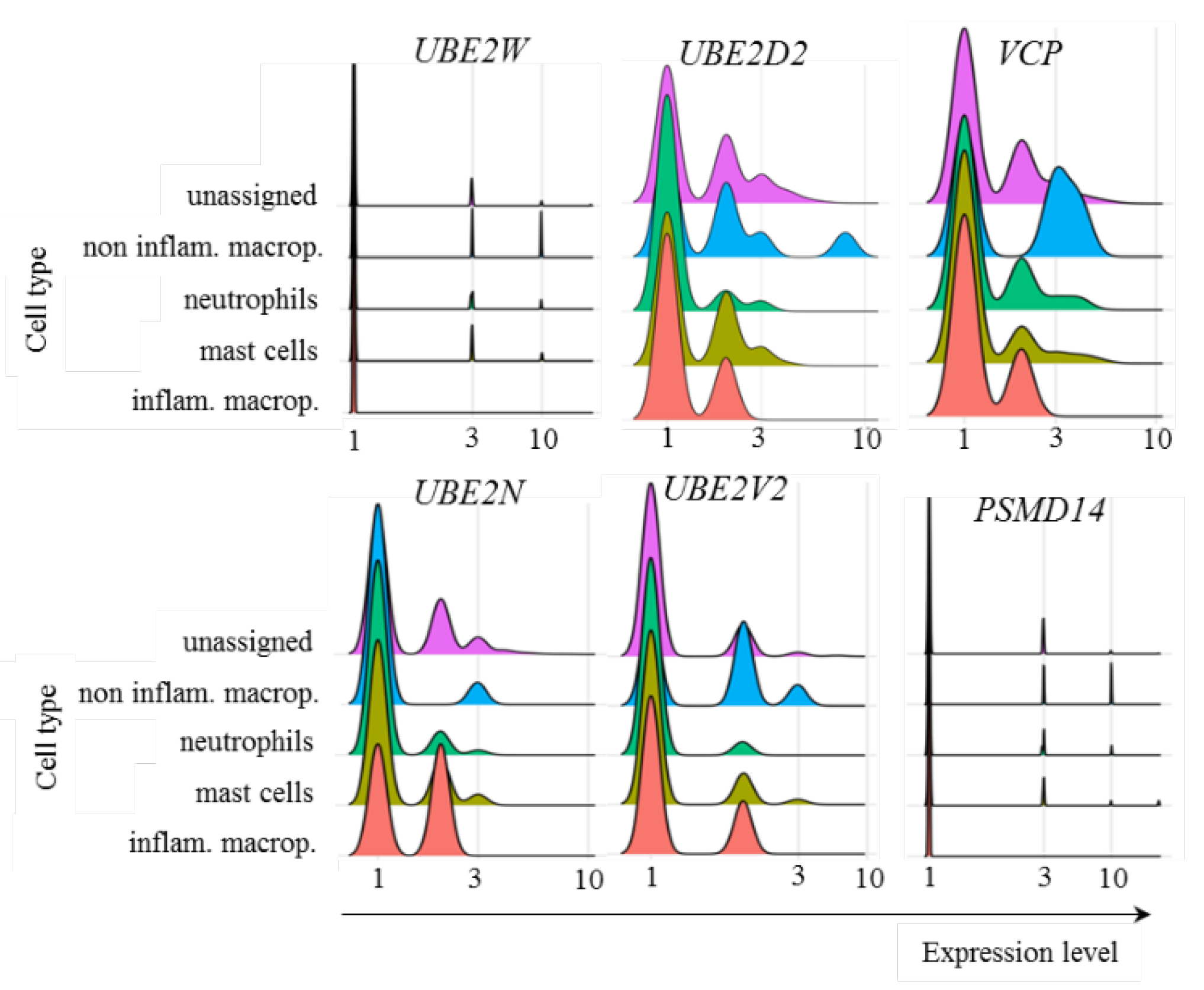
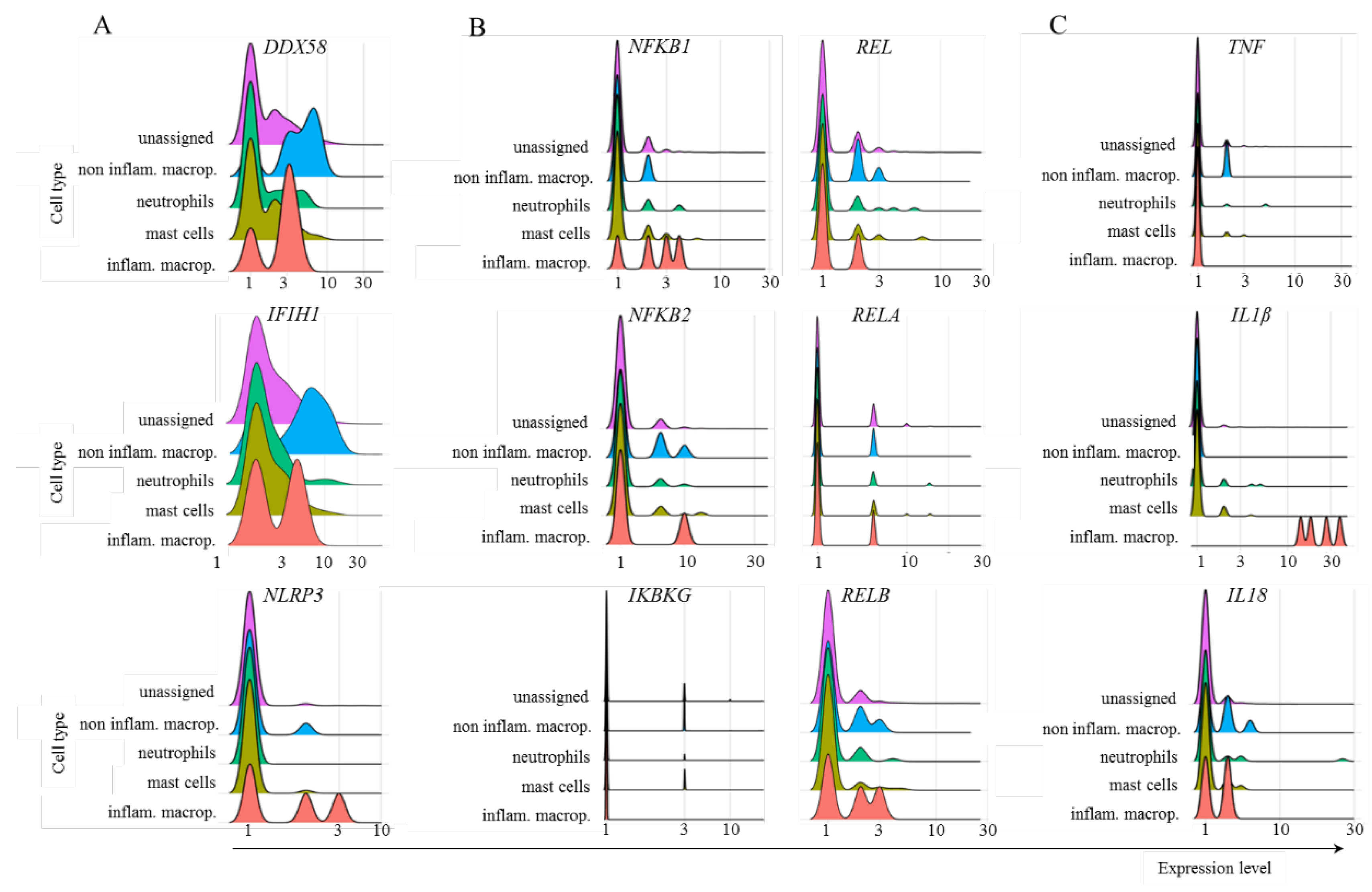
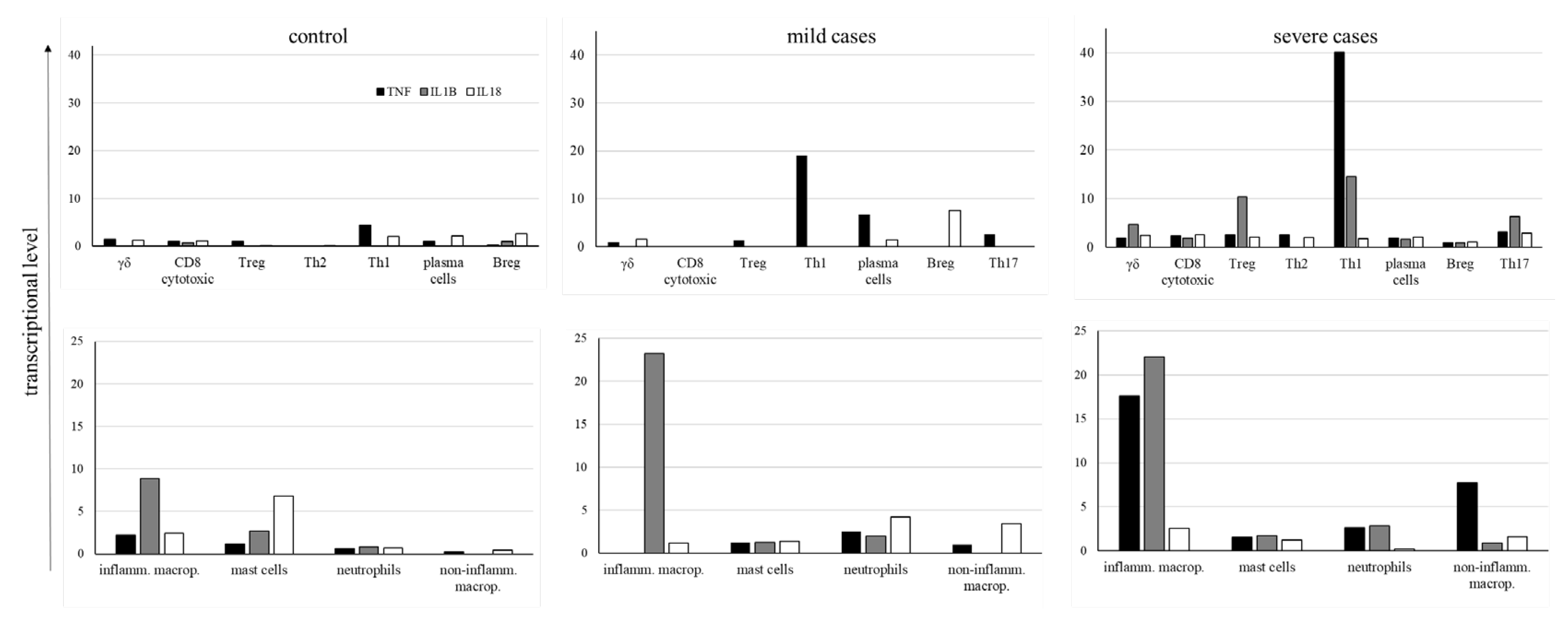
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data availability statement
Conflicts of Interest
References
- M.E.S. El Keshky, S.S. Basyouni, and A.M. Al Sabban, (2020) Getting Through COVID-19: The Pandemic’s Impact on the Psychology of Sustainability, Quality of Life, and the Global Economy - A Systematic Review. Front Psychol 11 585897.
- L. Yao, L. Lu, and W. Ma, (2022) Immunopathological changes, complications, sequelae and immunological memory in COVID-19 patients. Heliyon 8 e09302. [CrossRef]
- M. Daeron, (2014) Fc receptors as adaptive immunoreceptors. Curr Top Microbiol Immunol 382 131-64.
- L.L. Lu, T.J. Suscovich, S.M. Fortune, and G. Alter, (2018) Beyond binding: antibody effector functions in infectious diseases. Nat Rev Immunol 18 46-61. [CrossRef]
- M. Daeron, S. Jaeger, L. Du Pasquier, and E. Vivier, (2008) Immunoreceptor tyrosine-based inhibition motifs: a quest in the past and future. Immunol Rev 224 11-43. [CrossRef]
- M. Raghavan, and P.J. Bjorkman, (1996) Fc receptors and their interactions with immunoglobulins. Annu Rev Cell Dev Biol 12 181-220. [CrossRef]
- M. Pyzik, K.M.K. Sand, J.J. Hubbard, J.T. Andersen, I. Sandlie, and R.S. Blumberg, (2019) The Neonatal Fc Receptor (FcRn): A Misnomer? Front Immunol 10 1540.
- F.L. van de Veerdonk, E. Giamarellos-Bourboulis, P. Pickkers, L. Derde, H. Leavis, R. van Crevel, J.J. Engel, W.J. Wiersinga, A.P.J. Vlaar, M. Shankar-Hari, T. van der Poll, M. Bonten, D.C. Angus, J.W.M. van der Meer, and M.G. Netea, (2022) A guide to immunotherapy for COVID-19. Nat Med 28 39-50. [CrossRef]
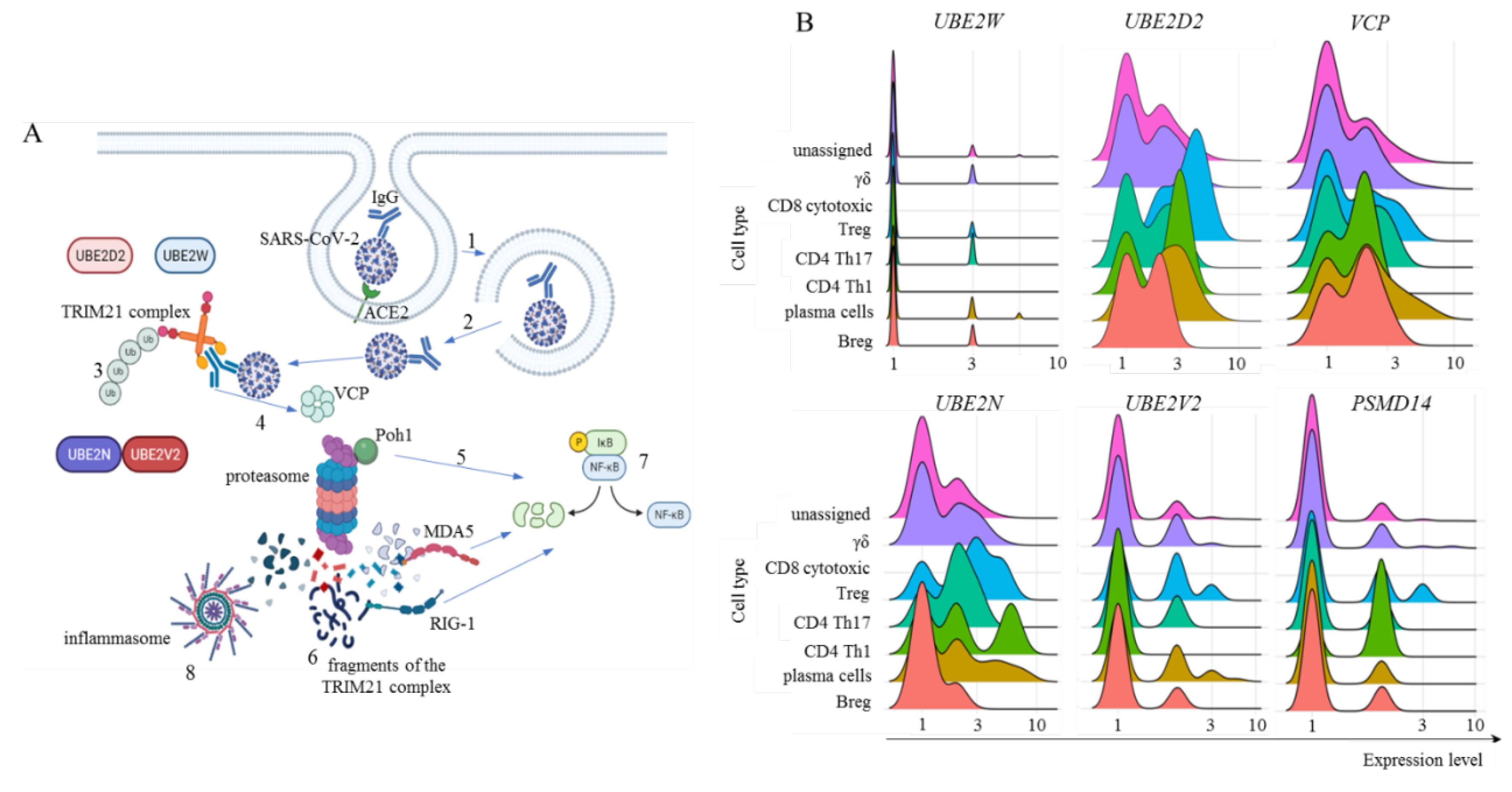
- S. Foss, M. Bottermann, A. Jonsson, I. Sandlie, L.C. James, and J.T. Andersen, (2019) TRIM21-From Intracellular Immunity to Therapy. Front Immunol 10 2049.
- J. Zeng, A.F. Santos, A.S. Mukadam, M. Osswald, D.A. Jacques, C.F. Dickson, S.H. McLaughlin, C.M. Johnson, L. Kiss, J. Luptak, N. Renner, M. Vaysburd, W.A. McEwan, E. Morais-de-Sa, D. Clift, and L.C. James, (2021) Target-induced clustering activates Trim-Away of pathogens and proteins. Nat Struct Mol Biol 28 278-289. [CrossRef]
- W.A. McEwan, F. Hauler, C.R. Williams, S.R. Bidgood, D.L. Mallery, R.A. Crowther, and L.C. James, (2012) Regulation of virus neutralization and the persistent fraction by TRIM21. J Virol 86 8482-91. [CrossRef]
- S.R. Bidgood, J.C. Tam, W.A. McEwan, D.L. Mallery, and L.C. James, (2014) Translocalized IgA mediates neutralization and stimulates innate immunity inside infected cells. Proc Natl Acad Sci U S A 111 13463-8. [CrossRef]
- D.L. Mallery, W.A. McEwan, S.R. Bidgood, G.J. Towers, C.M. Johnson, and L.C. James, (2010) Antibodies mediate intracellular immunity through tripartite motif-containing 21 (TRIM21). Proc Natl Acad Sci U S A 107 19985-90. [CrossRef]
- A.J. Fletcher, D.L. Mallery, R.E. Watkinson, C.F. Dickson, and L.C. James, (2015) Sequential ubiquitination and deubiquitination enzymes synchronize the dual sensor and effector functions of TRIM21. Proc Natl Acad Sci U S A 112 10014-9. [CrossRef]
- T.E.T. Mevissen, A.V. Prasad, and J.C. Walter, (2023) TRIM21-dependent target protein ubiquitination mediates cell-free Trim-Away. Cell Rep 42 112125. [CrossRef]
- D.A. Rhodes, and D.A. Isenberg, (2017) TRIM21 and the Function of Antibodies inside Cells. Trends Immunol 38 916-926. [CrossRef]
- F. Hauler, D.L. Mallery, W.A. McEwan, S.R. Bidgood, and L.C. James, (2012) AAA ATPase p97/VCP is essential for TRIM21-mediated virus neutralization. Proc Natl Acad Sci U S A 109 19733-8.
- T. Yao, and R.E. Cohen, (2002) A cryptic protease couples deubiquitination and degradation by the proteasome. Nature 419 403-7. [CrossRef]
- F. Full, and M.U. Gack, (2015) Delicate coordination of TRIM21’s dual activity in virus neutralization and signaling. Proc Natl Acad Sci U S A 112 9797-8. [CrossRef]
- S. Mao, X. Cai, S. Niu, J. Wei, N. Jiang, H. Deng, W. Wang, J. Zhang, S. Shen, Y. Ma, X. Wu, Q. Peng, A. Huang, and D. Wang, (2023) TRIM21 promotes ubiquitination of SARS-CoV-2 nucleocapsid protein to regulate innate immunity. J Med Virol 95 e28719.
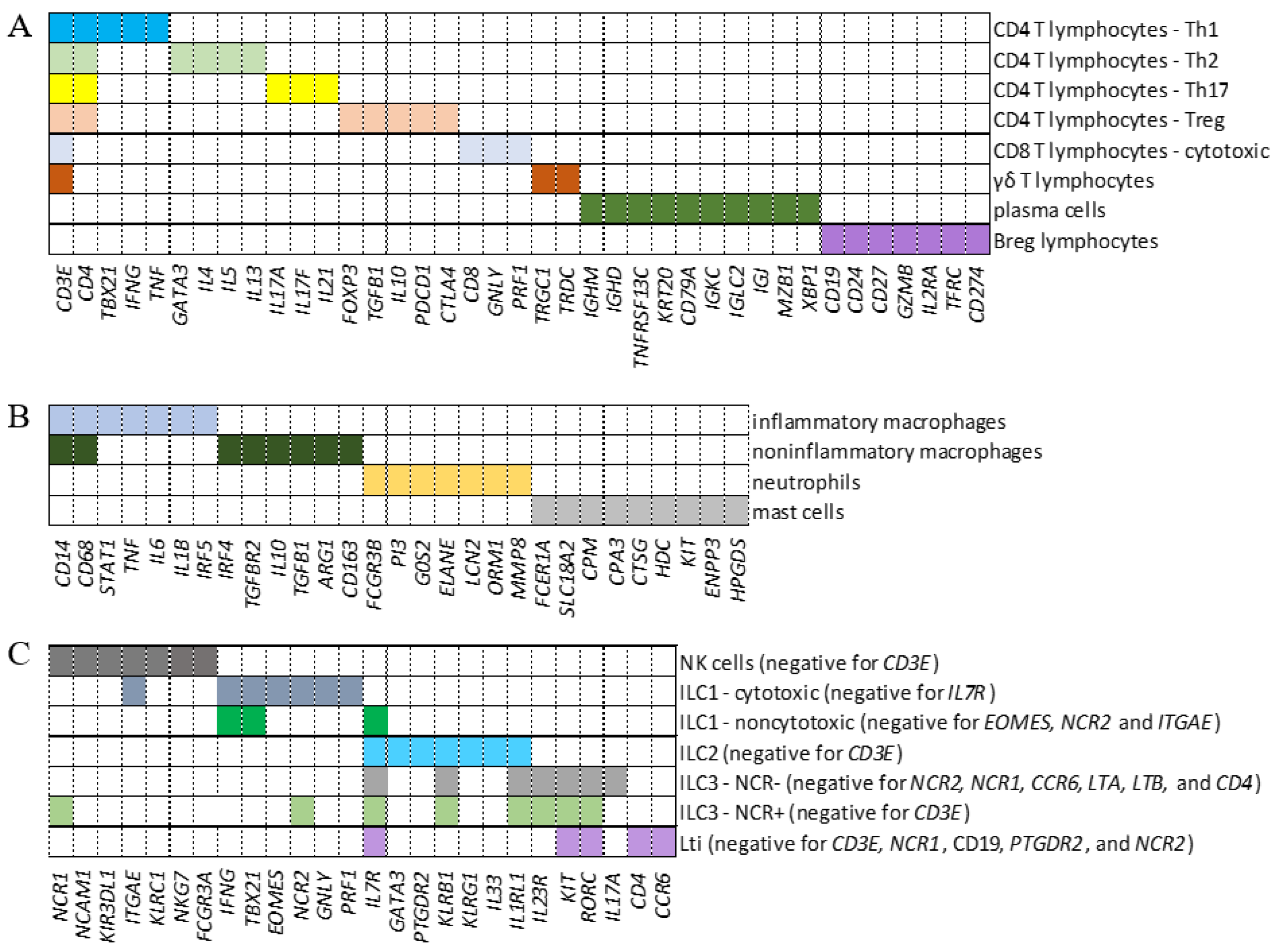
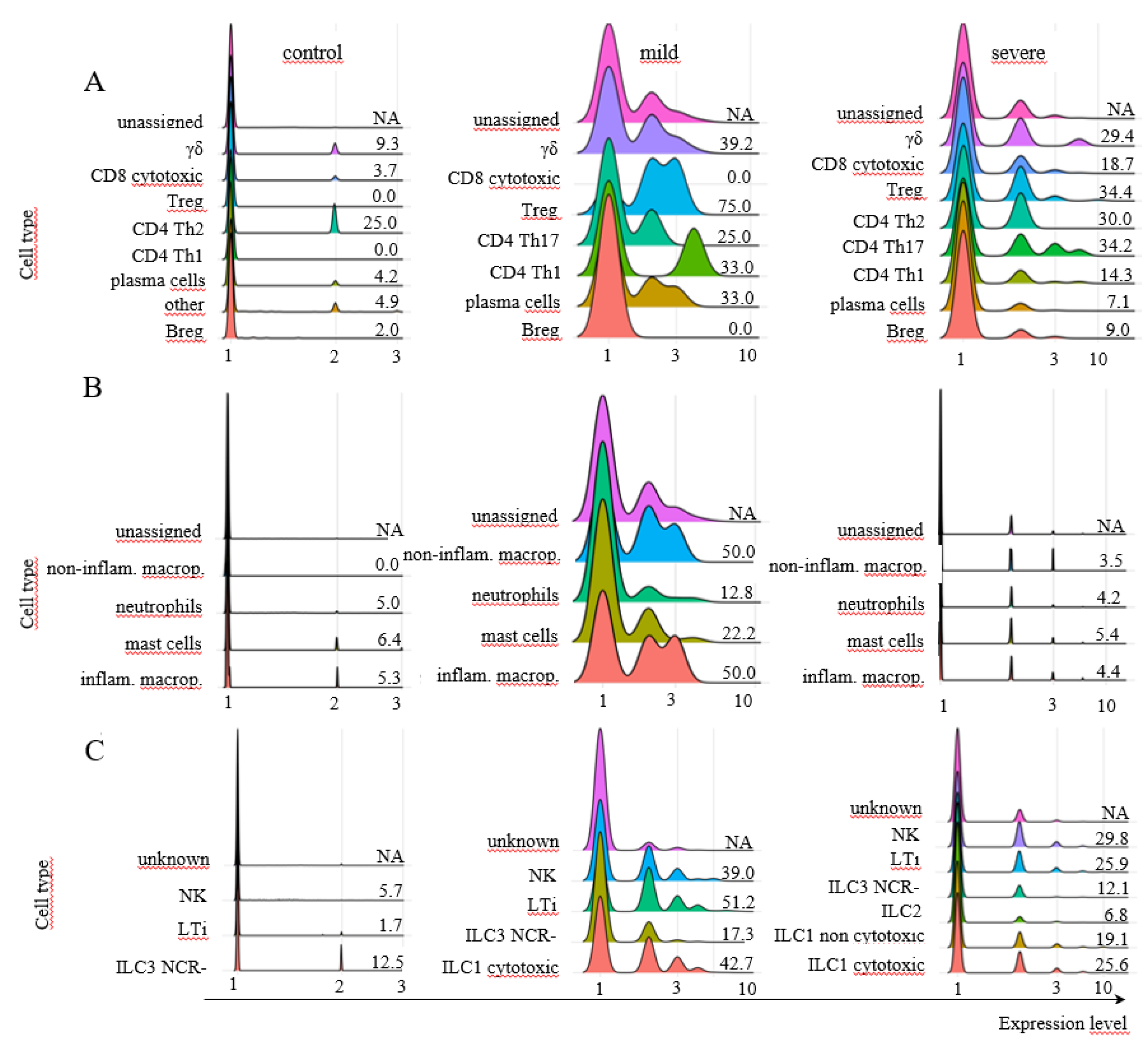
- M. Liao, Y. Liu, J. Yuan, Y. Wen, G. Xu, J. Zhao, L. Cheng, J. Li, X. Wang, F. Wang, L. Liu, I. Amit, S. Zhang, and Z. Zhang, (2020) Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat Med 26 842-844. [CrossRef]
- Bost, P., De Sanctis, F., Canè, S., Stefano Ugel, Donadello, K., Castellucci, M., Eyal, D., Fiore, A., Anselmi, C., Barouni, R. M., Trovato, R., Caligola, S., Lamolinara, A., Iezzi, M., Facciotti, F., Mazzariol, A., Gibellini, D., De Nardo, P., Tacconelli, E., Gottin, L., Polati, E., Schwikowski, B., Amit, I., Bronte, V., (2021) Deciphering the state of immune silence in fatal COVID-19 patients. Nat Commun 12, 1428. [CrossRef]
- V.S. Silva, M.O.C. Costa, M.C.S. Castro, H.S. Silva, M.E.M.T. Walter, A.C.M.A. Melo, K.A.C. Ocana, M.T. dos Santos, M.F. Nicolas, A.C.C. Carvalho, A. Henriques-Pons, and F.A.B. Silva, (2021) CellHeap: A Workflow for Optimizing COVID-19 Single-Cell RNA-Seq Data Processing in the Santos Dumont Supercomputer. Lect Notes Comput Sc 13063 41-52.
- X. Zhang, T. Li, F. Liu, Y. Chen, J. Yao, Z. Li, Y. Huang, and J. Wang, (2019) Comparative Analysis of Droplet-Based Ultra-High-Throughput Single-Cell RNA-Seq Systems. Mol Cell 73 130-142 e5. [CrossRef]
- M.D. Young, and S. Behjati, (2020) SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9. [CrossRef]
- Y. Hao, S. Hao, E. Andersen-Nissen, W.M. Mauck, 3rd, S. Zheng, A. Butler, M.J. Lee, A.J. Wilk, C. Darby, M. Zager, P. Hoffman, M. Stoeckius, E. Papalexi, E.P. Mimitou, J. Jain, A. Srivastava, T. Stuart, L.M. Fleming, B. Yeung, A.J. Rogers, J.M. McElrath, C.A. Blish, R. Gottardo, P. Smibert, and R. Satija, (2021) Integrated analysis of multimodal single-cell data. Cell 184 3573-3587 e29.
- M.D. Luecken, and F.J. Theis, (2019) Current best practices in single-cell RNA-seq analysis: a tutorial. Mol Syst Biol 15 e8746.
- A.W. Zhang, C. O’Flanagan, E.A. Chavez, J.L.P. Lim, N. Ceglia, A. McPherson, M. Wiens, P. Walters, T. Chan, B. Hewitson, D. Lai, A. Mottok, C. Sarkozy, L. Chong, T. Aoki, X. Wang, A.P. Weng, J.N. McAlpine, S. Aparicio, C. Steidl, K.R. Campbell, and S.P. Shah, (2019) Probabilistic cell-type assignment of single-cell RNA-seq for tumor microenvironment profiling. Nat Methods 16 1007-1015. [CrossRef]
- A. Ianevski, A.K. Giri, and T. Aittokallio, (2022) Fully automated and ultrafast cell-type identification using specific marker combinations from single-cell transcriptomic data. Nat Commun 13, 1246. [CrossRef]
- Wauters, E., Van Mol, P., Garg, A.D., Jansen, S., Herck, Y.V., Vanderbeke, L., Bassez, A., Boeckx, B., Malengier-Devlies, B., Timmerman, A., Van Brussel, T., Buyten, T. V., Schepers, R., Heylen, E., Dauwe, D., Dooms, C., Gunst, J., Hermans, G., Meersseman, P., Testelmans, D., Yserbyt, J., Tejpar, S., De Wever, W., Matthys, P., (2021). Discriminating mild from critical COVID-19 by innate and adaptive immune single-cell profiling of bronchoalveolar lavages. Cell Res 31, 272–290. [CrossRef]
- Mkadeem, S. B., Benhamou, M., and Monteiro R. C. (2019) Understanding Fc Receptor Involvement in Inflammatory Diseases: From Mechanisms to New Therapeutic Tools. Frontiers in Immunology 10 1-12. [CrossRef]
- Borger, J. G., Lau, M., and Hibbs, M. L., (2019) The Influence of Innate Lymphoid Cells and Unconventional T Cells in Chronic Inflammatory Lung Disease. Frontiers in Immunology 10 1-16. [CrossRef]
- Kumar, A., Cao, W., Endrias, K., Kuchipudi, S. V., Mittal, S. K., and Sambhara, S., (2021) Innate lymphoid cells (ILC) in SARS-CoV-2 infection. Molecular Aspects of Medicine 80 1-8.
- A.M.C. Gomes, G.B. Farias, M. Dias-Silva, J. Laia, A.C. Trombetta, A. Godinho-Santos, P. Rosmaninho, D.F. Santos, C.M. Conceicao, R. Costa-Reis, M. Adao-Serrano, C. Mota, A.R.M. Almeida, A.E. Sousa, and S.M. Fernandes, (2021) SARS-CoV-2 pneumonia recovery is linked to expansion of innate lymphoid cells type 2 expressing CCR10. Eur J Immunol 51 3194-3201.
- N.J. Silverstein, Y. Wang, Z. Manickas-Hill, C. Carbone, A. Dauphin, B.P. Boribong, M. Loiselle, J. Davis, M.M. Leonard, L. Kuri-Cervantes, M.C.-. Collection, T. Processing, N.J. Meyer, M.R. Betts, J.Z. Li, B.D. Walker, X.G. Yu, L.M. Yonker, and J. Luban, (2022) Innate lymphoid cells and COVID-19 severity in SARS-CoV-2 infection. Elife 11.
- M. Garcia, E. Kokkinou, A. Carrasco Garcia, T. Parrot, L.M. Palma Medina, K.T. Maleki, W. Christ, R. Varnaite, I. Filipovic, H.G. Ljunggren, N.K. Bjorkstrom, E. Folkesson, O. Rooyackers, L.I. Eriksson, A. Sonnerborg, S. Aleman, K. Stralin, S. Gredmark-Russ, J. Klingstrom, J. Mjosberg, and K.I.K.C.-S.G. Karolinska, (2020) Innate lymphoid cell composition associates with COVID-19 disease severity. Clin Transl Immunology 9 e1224. [CrossRef]
- Nagasawa, M., Spits, H., and Ros, X. R., (2018) Innate Lymphoid Cells (ILCs): Cytokine Hubs Regulating Immunity and Tissue Homeostasis. Cold Spring Harb Perspect Biol 2018;10:a030304. [CrossRef]
- Walker, J., Barlow, J., and McKenzie, A., (2013) Innate lymphoid cells - how did we miss them?. Nat Rev Immunol 13, 75–87.
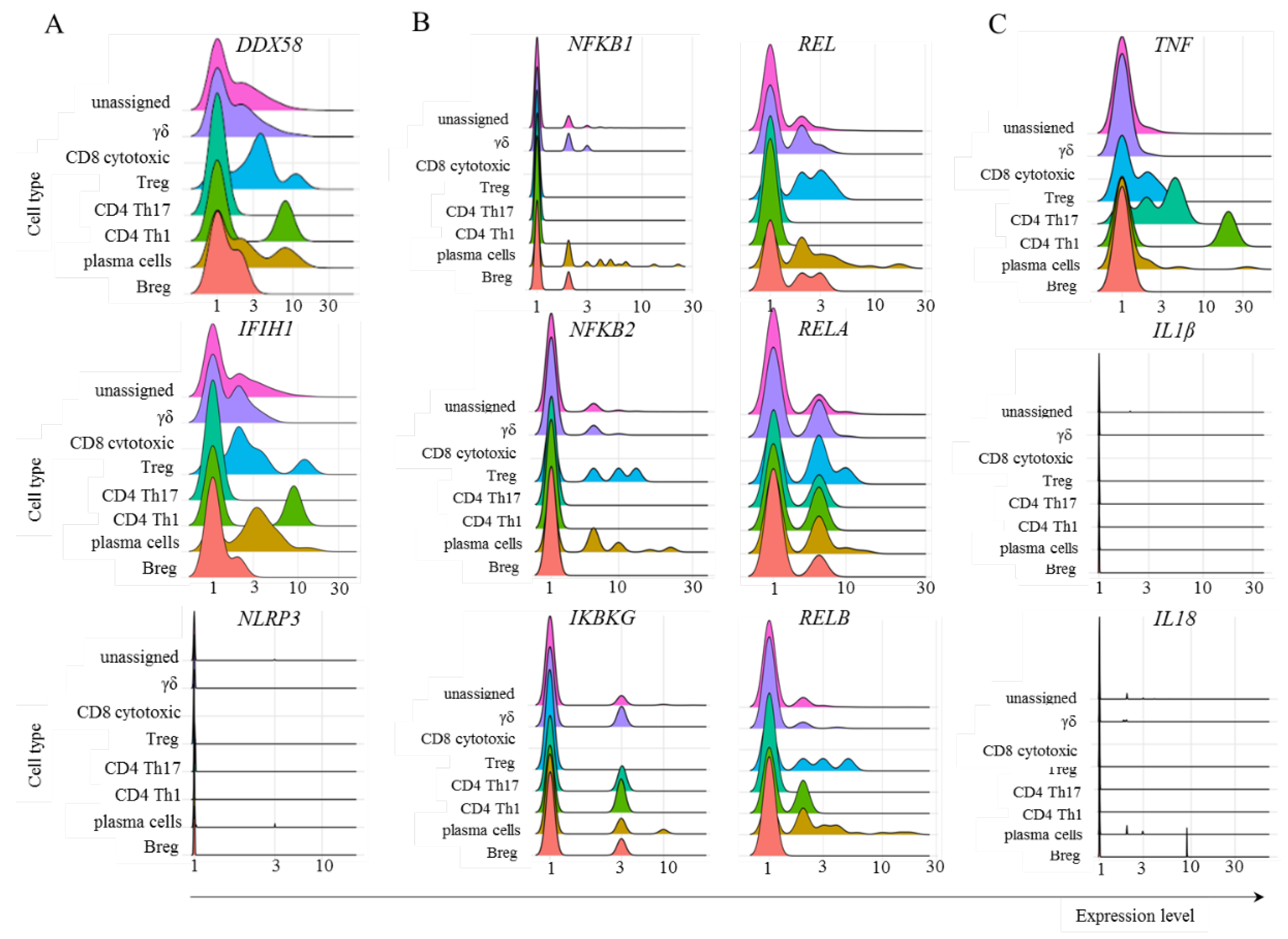
- K. Schroder, and J. Tschopp, (2010) The inflammasomes. Cell 140 821-32.
- Y.S. Choi, M.K. Jung, J. Lee, S.J. Choi, S.H. Choi, H.W. Lee, J.J. Lee, H.J. Kim, S.H. Ahn, D.H. Lee, W. Kim, S.H. Park, J.R. Huh, H.P. Kim, J.Y. Park, and E.C. Shin, (2018) Tumor Necrosis Factor-producing T-regulatory Cells Are Associated With Severe Liver Injury in Patients With Acute Hepatitis A. Gastroenterology 154 1047-1060. [CrossRef]
- L.M. Weiner, J.C. Murray, and C.W. Shuptrine, (2012) Antibody-based immunotherapy of cancer. Cell 148 1081-4. [CrossRef]
- S. Jiang, X. Zhang, Y. Yang, P.J. Hotez, and L. Du,(2020) Neutralizing antibodies for the treatment of COVID-19. Nat Biomed Eng 4 1134-1139. [CrossRef]
- L. Landeck, R. Sabat, K. Ghoreschi, X.Y. Man, K. Fuhrmeister, E. Gonzalez-Martinez, and K. Asadullah, (2021) Immunotherapy in psoriasis. Immunotherapy 13 605-619.
- M. Ballow, and C.L. Haga, (2021) Why Do Some People Develop Serious COVID-19 Disease After Infection, While Others Only Exhibit Mild Symptoms? J Allergy Clin Immunol Pract 9 1442-1448.
- S. Bournazos, A. Gupta, and J.V. Ravetch, (2020) The role of IgG Fc receptors in antibody-dependent enhancement. Nat Rev Immunol 20 633-643. [CrossRef]
- L. Liu, Q. Wei, Q. Lin, J. Fang, H. Wang, H. Kwok, H. Tang, K. Nishiura, J. Peng, Z. Tan, T. Wu, K.W. Cheung, K.H. Chan, X. Alvarez, C. Qin, A. Lackner, S. Perlman, K.Y. Yuen, and Z. Chen, (2019) Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight 4.
- L. Zhang, F. Zhang, W. Yu, T. He, J. Yu, C.E. Yi, L. Ba, W. Li, M. Farzan, Z. Chen, K.Y. Yuen, and D. Ho,(2006) Antibody responses against SARS coronavirus are correlated with disease outcome of infected individuals. J Med Virol 78 1-8. [CrossRef]
- Sender R, Weiss Y, Navon Y, Milo I, Azulay N, Keren L, Fuchs S, Ben-Zvi D, Noor E, Milo R. The total mass, number, and distribution of immune cells in the human body. Proc Natl Acad Sci USA. 2023 Oct 31;120(44):e2308511120. [CrossRef]
- C.K. Li, H. Wu, H. Yan, S. Ma, L. Wang, M. Zhang, X. Tang, N.J. Temperton, R.A. Weiss, J.M. Brenchley, D.C. Douek, J. Mongkolsapaya, B.H. Tran, C.L. Lin, G.R. Screaton, J.L. Hou, A.J. McMichael, and X.N. Xu, (2008) T cell responses to whole SARS coronavirus in humans. J Immunol 181 5490-500. [CrossRef]
- P.J. Hotez, M.E. Bottazzi, and D.B. Corry, (2020) The potential role of Th17 immune responses in coronavirus immunopathology and vaccine-induced immune enhancement. Microbes Infect 22 165-167. [CrossRef]
- W.C. Hsieh, E.Y. Lai, Y.T. Liu, Y.F. Wang, Y.S. Tzeng, L. Cui, Y.J. Lai, H.C. Huang, J.H. Huang, H.C. Ni, D.Y. Tsai, J.J. Liang, C.C. Liao, Y.T. Lu, L. Jiang, M.T. Liu, J.T. Wang, S.Y. Chang, C.Y. Chen, H.C. Tsai, Y.M. Chang, G. Wernig, C.W. Li, K.I. Lin, Y.L. Lin, H.K. Tsai, Y.T. Huang, and S.Y. Chen, (2021) NK cell receptor and ligand composition influences the clearance of SARS-CoV-2. J Clin Invest 131.
- G. Qin, S. Liu, L. Yang, W. Yu, and Y. Zhang, (2021) Myeloid cells in COVID-19 microenvironment. Signal Transduct Target Ther 6 372. [CrossRef]
- S. Bhaskar, A. Sinha, M. Banach, S. Mittoo, R. Weissert, J.S. Kass, S. Rajagopal, A.R. Pai, and S. Kutty, (2020) Cytokine Storm in COVID-19-Immunopathological Mechanisms, Clinical Considerations, and Therapeutic Approaches: The REPROGRAM Consortium Position Paper. Front Immunol 11 1648.
- S. Montazersaheb, S.M. Hosseiniyan Khatibi, M.S. Hejazi, V. Tarhriz, A. Farjami, F. Ghasemian Sorbeni, R. Farahzadi, and T. Ghasemnejad, (2022) COVID-19 infection: an overview on cytokine storm and related interventions. Virol J 19 92. [CrossRef]
- G. Scaramuzzo, F. Nucera, A. Asmundo, R. Messina, M. Mari, F. Montanaro, M.D. Johansen, F. Monaco, G. Fadda, G. Tuccari, N.G. Hansbro, P.M. Hansbro, T.T. Hansel, I.M. Adcock, A. David, P. Kirkham, G. Caramori, C.A. Volta, and S. Spadaro, (2023) Cellular and molecular features of COVID-19 associated ARDS: therapeutic relevance. J Inflamm (Lond) 20 11. [CrossRef]
- M.H. Shakiba, I. Gemund, M. Beyer, and L. Bonaguro, (2023) Lung T cell response in COVID-19. Front Immunol 14 1108716. [CrossRef]
- M. Menon, T. Hussell, and H. Ali Shuwa, (2021) Regulatory B cells in respiratory health and diseases. Immunol Rev 299 61-73. [CrossRef]
- M. Dhawan, A.A. Rabaan, S. Alwarthan, M. Alhajri, M.A. Halwani, A. Alshengeti, M.A. Najim, A.S.S. Alwashmi, A.A. Alshehri, S.A. Alshamrani, B.M. AlShehail, M. Garout, S. Al-Abdulhadi, S.H. Al-Ahmed, N. Thakur, and G. Verma, (2023) Regulatory T Cells (Tregs) and COVID-19: Unveiling the Mechanisms, and Therapeutic Potentialities with a Special Focus on Long COVID. Vaccines (Basel) 11.
- E. Vivier, D. Artis, M. Colonna, A. Diefenbach, J.P. Di Santo, G. Eberl, S. Koyasu, R.M. Locksley, A.N.J. McKenzie, R.E. Mebius, F. Powrie, and H. Spits, (2018) Innate Lymphoid Cells: 10 Years On. Cell 174 1054-1066. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



