Submitted:
22 October 2024
Posted:
22 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
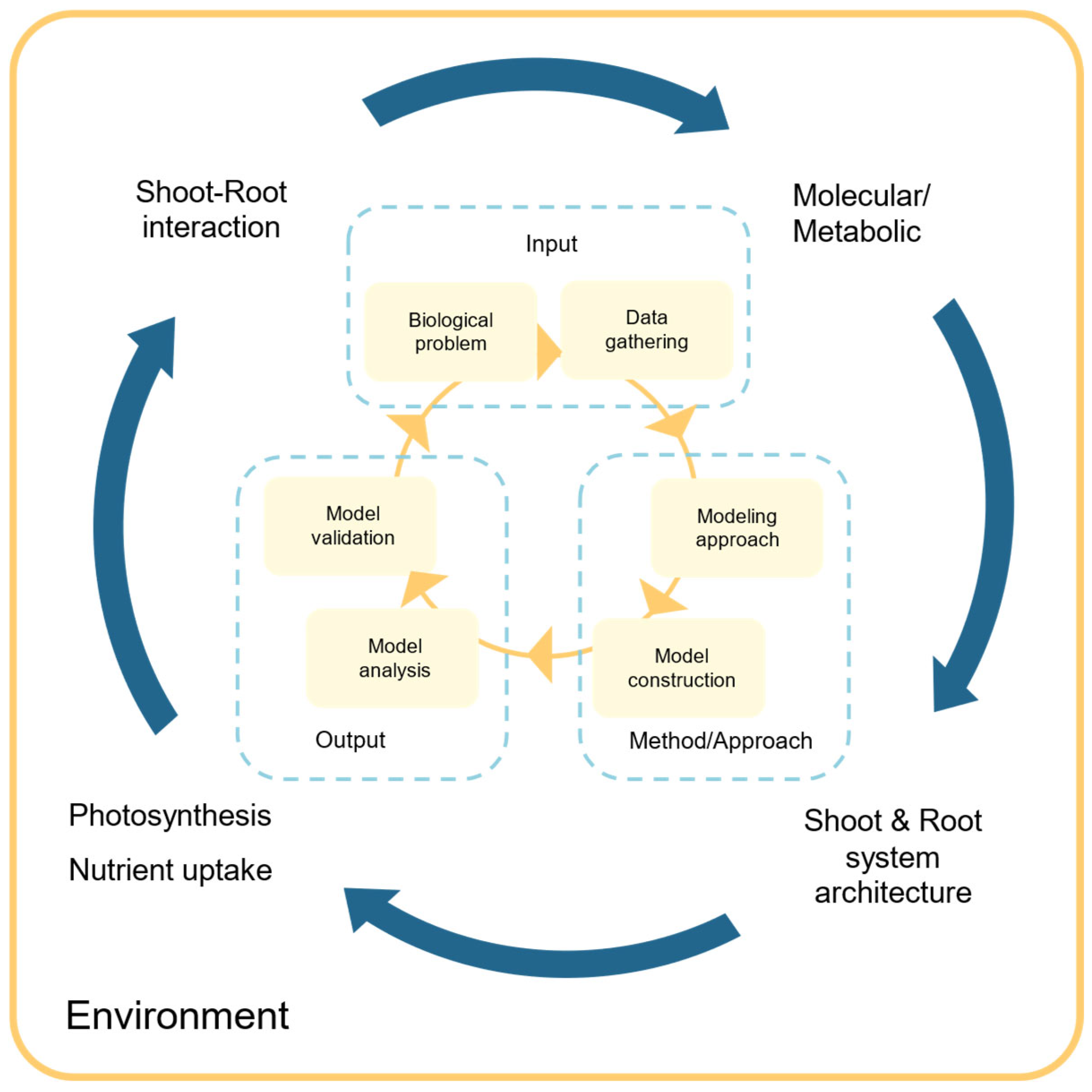
2. Model Building
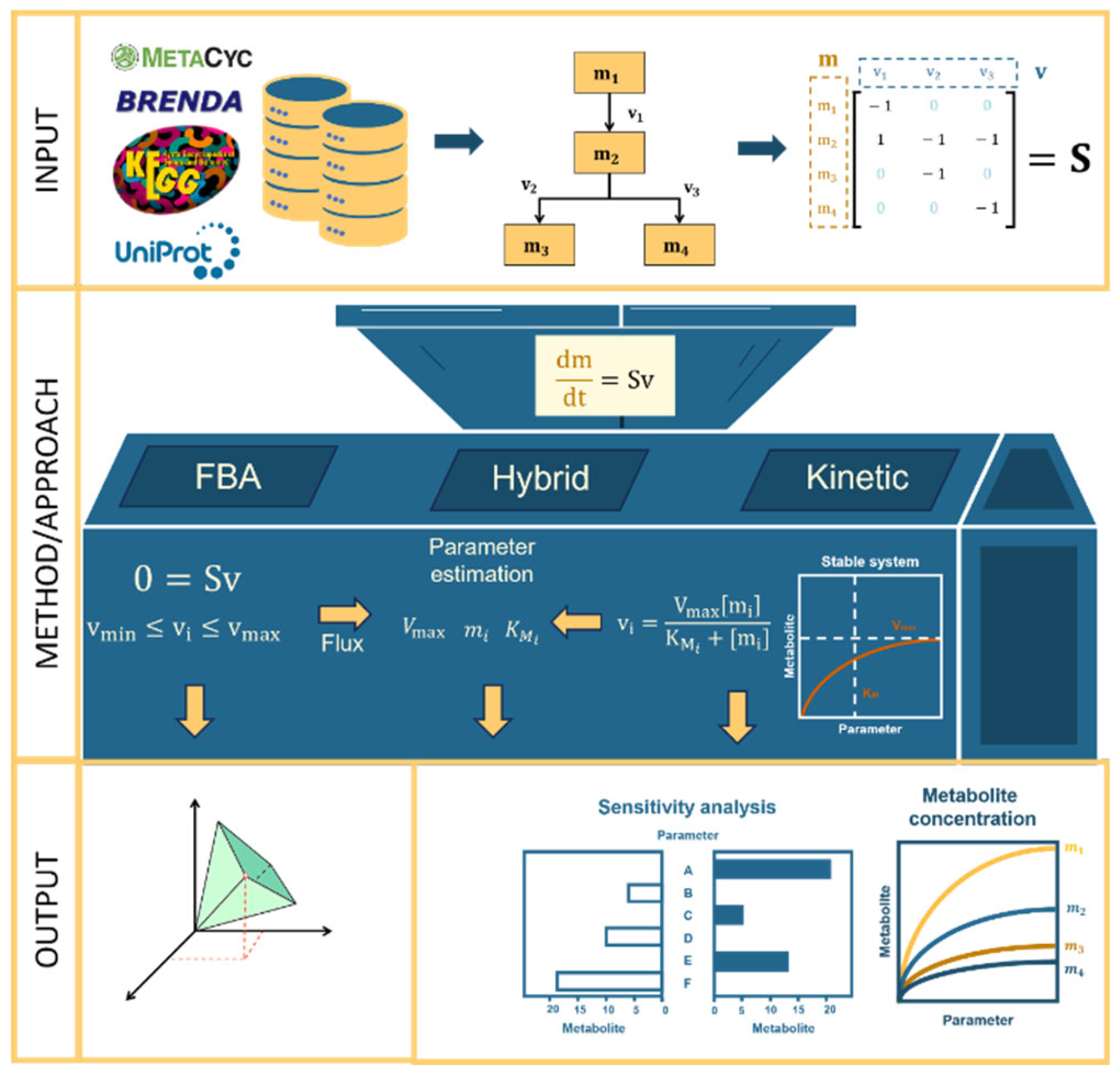
2.1. Metabolic Modeling with FBA, Kinetic, or Hybrid Approaches
2.1.1. Applications
2.1.2. Creating a Metabolic Model
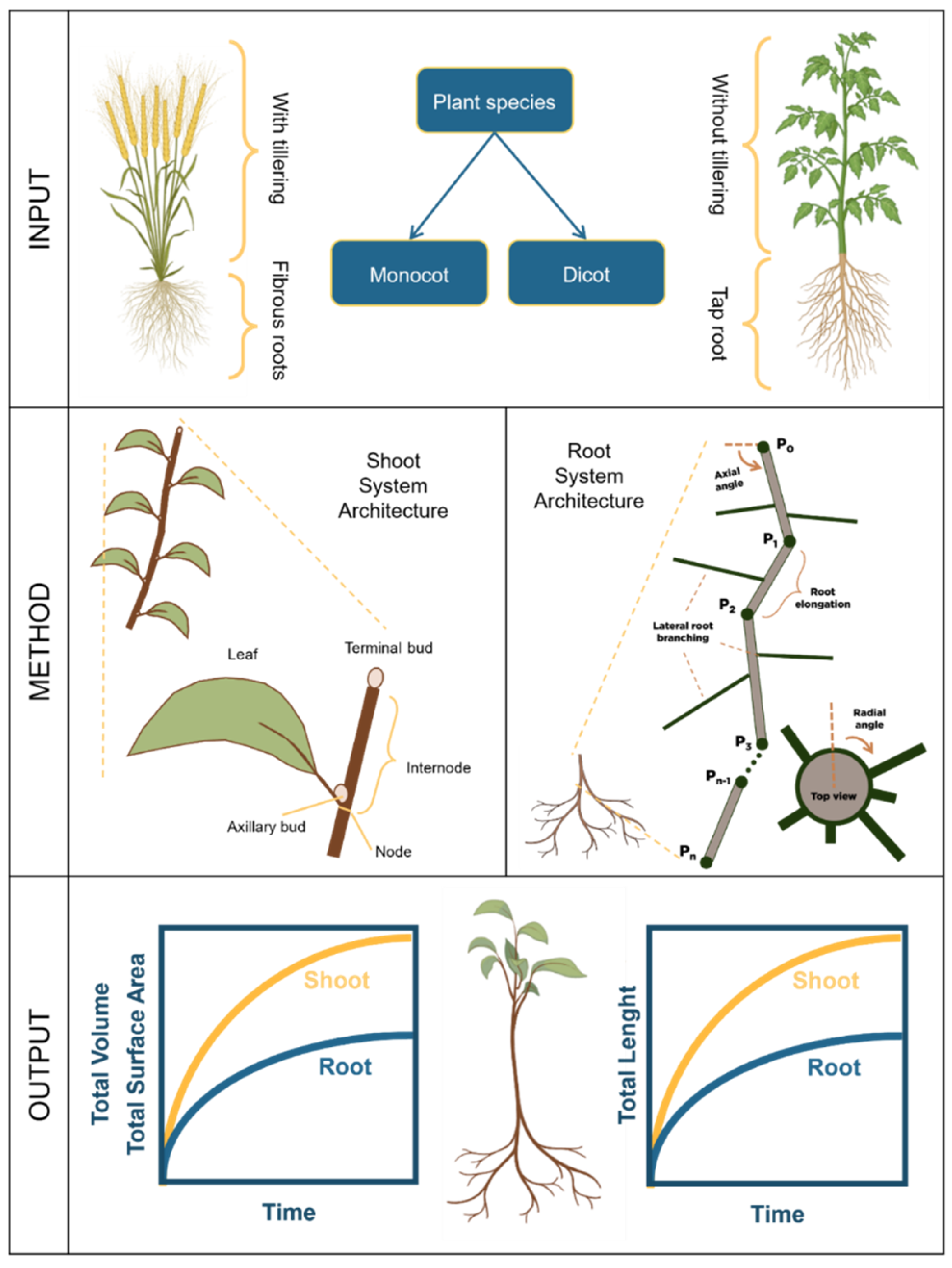
2.2. Shoot and Root System Architecture
2.2.1. Shoot System Architecture
2.2.1.1. Applications
2.2.1.2. Creating a Shoot System Architecture
2.2.2. Root System Architecture
2.2.2.1. Applications
2.2.2.2. Creating a Root System Architecture
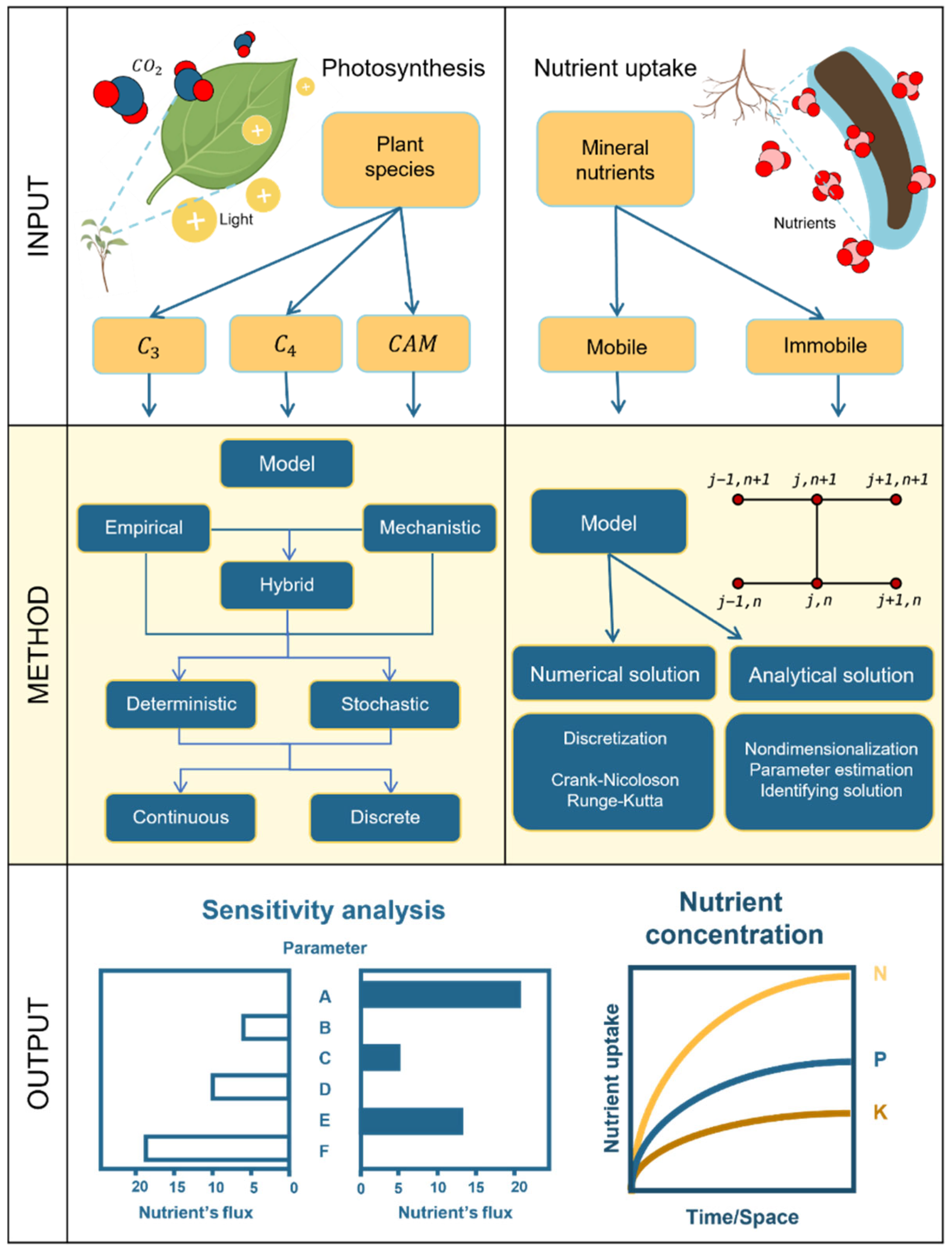
2.3. Resource Acquisition
2.3.1. Photosynthesis
2.3.1.1. Applications
2.3.1.2. Creating a Photosynthesis Model
2.3.2. Nutrient Uptake Model
2.3.2.1. Applications
2.3.2.2. Creating a Root Nutrient Uptake Model
2.4. Shoot-Root Interaction
2.4.1. Applications
2.4.2. Creating a Shoot-Interaction Model
3. Discussion
3.1. The Potential of Multiscale Mathematical Modeling
3.2. Limitations of Multiscale Mathematical Modeling
3.3. Concluding Remarks
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Food and Agriculture Organization of the United Nations Save and Grow in Practice: Maize, Rice, Wheat : A Guide to Sustainable Cereal Production; 2016; ISBN 978-92-5-108519-6.
- Pingali, P.L. Green Revolution: Impacts, Limits, and the Path Ahead. Proceedings of the National Academy of Sciences 2012, 109, 12302–12308. [CrossRef]
- John, D.A.; Babu, G.R. Lessons From the Aftermaths of Green Revolution on Food System and Health. Frontiers in Sustainable Food Systems 2021, 5.
- Venkatappa, M.; Sasaki, N.; Han, P.; Abe, I. Impacts of Droughts and Floods on Croplands and Crop Production in Southeast Asia – An Application of Google Earth Engine. Science of The Total Environment 2021, 795, 148829. [CrossRef]
- Yacoubou, A.-M.; Zoumarou Wallis, N.; Menkir, A.; Zinsou, V.A.; Onzo, A.; Garcia-Oliveira, A.L.; Meseka, S.; Wende, M.; Gedil, M.; Agre, P. Breeding Maize (Zea Mays) for Striga Resistance: Past, Current and Prospects in Sub-Saharan Africa. Plant Breeding 2021, 140, 195–210. [CrossRef]
- Glauber, J.W.; Laborde Debucquet, D.; Swinnen, J. The Russia-Ukraine War’s Impact on Global Food Markets: A Historical Perspective; 0 ed.; International Food Policy Research Institute: Washington, DC, 2023;
- Martyshev, P.; Nivievskyi, O.; Bogonos, M. Regional War, Global Consequences: Mounting Damages to Ukraine’s Agriculture and Growing Challenges for Global Food Security Available online: https://ebrary.ifpri.org/digital/collection/p15738coll2/id/136766 (accessed on 4 October 2023).
- OECD; Food and Agriculture Organization of the United Nations OECD-FAO Agricultural Outlook 2021-2030; OECD-FAO Agricultural Outlook; OECD: Paris, France, 2021; ISBN 978-92-64-43607-7.
- Lynch, J.P. Roots of the Second Green Revolution. Aust. J. Bot. 2007, 55, 493–512. [CrossRef]
- Liu, W.; Stewart, C.N. Plant Synthetic Biology. Trends in Plant Science 2015, 20, 309–317. [CrossRef]
- Petzold, C.; Chan, L.J.; Nhan, M.; Adams, P. Analytics for Metabolic Engineering. Frontiers in Bioengineering and Biotechnology 2015, 3.
- Varadan, R.; SOLOMATIN, S.; HOLZ-SCHIETINGER, C.; COHN, E.; KLAPHOLZ-BROWN, A.; SHIU, J.W.-Y.; Kale, A.; KARR, J.; FRASER, R. Ground Meat Replicas 2015.
- Shankar, S.; Hoyt, M.A. Expression Constructs and Methods of Genetically Engineering Methylotrophic Yeast 2017.
- Mcnamara, J.; Harvey, J.D.; Graham, M.J.; Scherger, C. Optically Transparent Polyimides 2019.
- Temme, K.; TAMSIR, A.; BLOCH, S.; CLARK, R.; TUNG, E. Methods and Compositions for Improving Plant Traits 2017.
- Haun, W.; Coffman, A.; Clasen, B.M.; Demorest, Z.L.; Lowy, A.; Ray, E.; Retterath, A.; Stoddard, T.; Juillerat, A.; Cedrone, F.; et al. Improved Soybean Oil Quality by Targeted Mutagenesis of the Fatty Acid Desaturase 2 Gene Family. Plant Biotechnol J 2014, 12, 934–940. [CrossRef]
- Voigt, C.A. Synthetic Biology 2020–2030: Six Commercially-Available Products That Are Changing Our World. Nat Commun 2020, 11, 6379. [CrossRef]
- Nowacka, M.; Trusinska, M.; Chraniuk, P.; Piatkowska, J.; Pakulska, A.; Wisniewska, K.; Wierzbicka, A.; Rybak, K.; Pobiega, K. Plant-Based Fish Analogs—A Review. Applied Sciences 2023, 13, 4509. [CrossRef]
- Sun, T.; Song, J.; Wang, M.; Zhao, C.; Zhang, W. Challenges and Recent Progress in the Governance of Biosecurity Risks in the Era of Synthetic Biology. Journal of Biosafety and Biosecurity 2022, 4, 59–67. [CrossRef]
- Bohua, L.; Yuexin, W.; Yakun, O.; Kunlan, Z.; Huan, L.; Ruipeng, L. Ethical Framework on Risk Governance of Synthetic Biology. Journal of Biosafety and Biosecurity 2023, 5, 45–56. [CrossRef]
- Haseltine, E.L.; Arnold, F.H. Synthetic Gene Circuits: Design with Directed Evolution. Annual Review of Biophysics and Biomolecular Structure 2007, 36, 1–19. [CrossRef]
- Chandran, D.; Copeland, W.B.; Sleight, S.C.; Sauro, H.M. Mathematical Modeling and Synthetic Biology. Drug Discov Today Dis Models 2008, 5, 299–309. [CrossRef]
- Chew, Y.H.; Wenden, B.; Flis, A.; Mengin, V.; Taylor, J.; Davey, C.L.; Tindal, C.; Thomas, H.; Ougham, H.J.; Reffye, P. de; et al. Multiscale Digital Arabidopsis Predicts Individual Organ and Whole-Organism Growth. PNAS 2014, 111, E4127–E4136. [CrossRef]
- Karr, J.R.; Sanghvi, J.C.; Macklin, D.N.; Gutschow, M.V.; Jacobs, J.M.; Bolival, B.; Assad-Garcia, N.; Glass, J.I.; Covert, M.W. A Whole-Cell Computational Model Predicts Phenotype from Genotype. Cell 2012, 150, 389–401. [CrossRef]
- Carrera, J.; Estrela, R.; Luo, J.; Rai, N.; Tsoukalas, A.; Tagkopoulos, I. An Integrative, Multi-scale, Genome-wide Model Reveals the Phenotypic Landscape of Escherichia Coli. Molecular Systems Biology 2014, 10, 735. [CrossRef]
- Ye, C.; Xu, N.; Gao, C.; Liu, G.; Xu, J.; Zhang, W.; Chen, X.; Nielsen, J.; Liu, L. Comprehensive Understanding of Saccharomyces Cerevisiae Phenotypes with Whole-Cell Model WM_S288C. Biotechnology and Bioengineering 2020, 117, 1562–1574. [CrossRef]
- Matthews, M.L.; Marshall-Colón, A. Multiscale Plant Modeling: From Genome to Phenome and Beyond. Emerging Topics in Life Sciences 2021, 5, 231–237. [CrossRef]
- Benes, B.; Guan, K.; Lang, M.; Long, S.P.; Lynch, J.P.; Marshall-Colón, A.; Peng, B.; Schnable, J.; Sweetlove, L.J.; Turk, M.J. Multiscale Computational Models Can Guide Experimentation and Targeted Measurements for Crop Improvement. The Plant Journal 2020, 103, 21–31. [CrossRef]
- Messina, C.D.; Technow, F.; Tang, T.; Totir, R.; Gho, C.; Cooper, M. Leveraging Biological Insight and Environmental Variation to Improve Phenotypic Prediction: Integrating Crop Growth Models (CGM) with Whole Genome Prediction (WGP). European Journal of Agronomy 2018, 100, 151–162. [CrossRef]
- Falco, B. de; Giannino, F.; Carteni, F.; Mazzoleni, S.; Kim, D.-H. Metabolic Flux Analysis: A Comprehensive Review on Sample Preparation, Analytical Techniques, Data Analysis, Computational Modelling, and Main Application Areas. RSC Adv. 2022, 12, 25528–25548. [CrossRef]
- Clark, T.J.; Guo, L.; Morgan, J.; Schwender, J. Modeling Plant Metabolism: From Network Reconstruction to Mechanistic Models. Annual Review of Plant Biology 2020, 71, 303–326. [CrossRef]
- Foster, C.J.; Wang, L.; Dinh, H.V.; Suthers, P.F.; Maranas, C.D. Building Kinetic Models for Metabolic Engineering. Current Opinion in Biotechnology 2021, 67, 35–41. [CrossRef]
- Gudmundsson, S.; Nogales, J. Recent Advances in Model-Assisted Metabolic Engineering. Current Opinion in Systems Biology 2021, 28, 100392. [CrossRef]
- Orth, J.D.; Thiele, I.; Palsson, B.Ø. What Is Flux Balance Analysis? Nat Biotechnol 2010, 28, 245–248. [CrossRef]
- Poolman, M.G.; Miguet, L.; Sweetlove, L.J.; Fell, D.A. A Genome-Scale Metabolic Model of Arabidopsis and Some of Its Properties. Plant Physiology 2009, 151, 1570–1581. [CrossRef]
- Poolman, M.G.; Kundu, S.; Shaw, R.; Fell, D.A. Responses to Light Intensity in a Genome-Scale Model of Rice Metabolism. Plant Physiology 2013, 162, 1060–1072. [CrossRef]
- Dal’Molin, C.G. de O.; Quek, L.-E.; Palfreyman, R.W.; Brumbley, S.M.; Nielsen, L.K. AraGEM, a Genome-Scale Reconstruction of the Primary Metabolic Network in Arabidopsis. Plant Physiology 2010, 152, 579–589. [CrossRef]
- Williams, T.C.R.; Poolman, M.G.; Howden, A.J.M.; Schwarzlander, M.; Fell, D.A.; Ratcliffe, R.G.; Sweetlove, L.J. A Genome-Scale Metabolic Model Accurately Predicts Fluxes in Central Carbon Metabolism under Stress Conditions. Plant Physiology 2010, 154, 311–323. [CrossRef]
- Saha, R.; Suthers, P.F.; Maranas, C.D. Zea Mays iRS1563: A Comprehensive Genome-Scale Metabolic Reconstruction of Maize Metabolism. PLOS ONE 2011, 6, e21784. [CrossRef]
- Mintz-Oron, S.; Meir, S.; Malitsky, S.; Ruppin, E.; Aharoni, A.; Shlomi, T. Reconstruction of Arabidopsis Metabolic Network Models Accounting for Subcellular Compartmentalization and Tissue-Specificity. PNAS 2012, 109, 339–344. [CrossRef]
- Cheung, C.Y.M.; Williams, T.C.R.; Poolman, M.G.; Fell, David.A.; Ratcliffe, R.G.; Sweetlove, L.J. A Method for Accounting for Maintenance Costs in Flux Balance Analysis Improves the Prediction of Plant Cell Metabolic Phenotypes under Stress Conditions. The Plant Journal 2013, 75, 1050–1061. [CrossRef]
- Yen, J.Y.; Nazem-Bokaee, H.; Freedman, B.G.; Athamneh, A.I.M.; Senger, R.S. Deriving Metabolic Engineering Strategies from Genome-Scale Modeling with Flux Ratio Constraints. Biotechnology Journal 2013, 8, 581–594. [CrossRef]
- Simons, M.; Saha, R.; Amiour, N.; Kumar, A.; Guillard, L.; Clément, G.; Miquel, M.; Li, Z.; Mouille, G.; Lea, P.J.; et al. Assessing the Metabolic Impact of Nitrogen Availability Using a Compartmentalized Maize Leaf Genome-Scale Model. Plant Physiology 2014, 166, 1659–1674. [CrossRef]
- Yuan, H.; Cheung, C.Y.M.; Poolman, M.G.; Hilbers, P.A.J.; Riel, N.A.W. van A Genome-Scale Metabolic Network Reconstruction of Tomato (Solanum Lycopersicum L.) and Its Application to Photorespiratory Metabolism. The Plant Journal 2016, 85, 289–304. [CrossRef]
- Wanichthanarak, K.; Boonchai, C.; Kojonna, T.; Chadchawan, S.; Sangwongchai, W.; Thitisaksakul, M. Deciphering Rice Metabolic Flux Reprograming under Salinity Stress via in Silico Metabolic Modeling. Computational and Structural Biotechnology Journal 2020, 18, 3555–3566. [CrossRef]
- Sugimoto, K.; Zager, J.J.; Aubin, B.S.; Lange, B.M.; Howe, G.A. Flavonoid Deficiency Disrupts Redox Homeostasis and Terpenoid Biosynthesis in Glandular Trichomes of Tomato. Plant Physiology 2022, 188, 1450–1468. [CrossRef]
- Savageau, M.A. Biochemical Systems Analysis. A Study of Function and Design in Molecular Biology. In ADDISON WESLEY PUBL.; 1976.
- Comas, J.; Benfeitas, R.; Vilaprinyo, E.; Sorribas, A.; Solsona, F.; Farré, G.; Berman, J.; Zorrilla, U.; Capell, T.; Sandmann, G.; et al. Identification of Line-Specific Strategies for Improving Carotenoid Production in Synthetic Maize through Data-Driven Mathematical Modeling. Plant J 2016, 87, 455–471. [CrossRef]
- Basallo, O.; Perez, L.; Lucido, A.; Sorribas, A.; Marin-Saguino, A.; Vilaprinyo, E.; Perez-Fons, L.; Albacete, A.; Martínez-Andújar, C.; Fraser, P.D.; et al. Changing Biosynthesis of Terpenoid Percursors in Rice through Synthetic Biology. Front Plant Sci 2023, 14, 1133299. [CrossRef]
- Yang, Z.; Midmore, D.J. A Model for the Circadian Oscillations in Expression and Activity of Nitrate Reductase in Higher Plants. Annals of Botany 2005, 96, 1019–1026. [CrossRef]
- Curien, G.; Bastien, O.; Robert-Genthon, M.; Cornish-Bowden, A.; Cárdenas, M.L.; Dumas, R. Understanding the Regulation of Aspartate Metabolism Using a Model Based on Measured Kinetic Parameters. Molecular Systems Biology 2009, 5, 271. [CrossRef]
- Colón, A.M.; Sengupta, N.; Rhodes, D.; Dudareva, N.; Morgan, J. A Kinetic Model Describes Metabolic Response to Perturbations and Distribution of Flux Control in the Benzenoid Network of Petunia Hybrida. The Plant Journal 2010, 62, 64–76. [CrossRef]
- Nägele, T.; Henkel, S.; Hörmiller, I.; Sauter, T.; Sawodny, O.; Ederer, M.; Heyer, A.G. Mathematical Modeling of the Central Carbohydrate Metabolism in Arabidopsis Reveals a Substantial Regulatory Influence of Vacuolar Invertase on Whole Plant Carbon Metabolism. Plant Physiology 2010, 153, 260–272. [CrossRef]
- Groenenboom, M.; Gomez-Roldan, V.; Stigter, H.; Astola, L.; Daelen, R. van; Beekwilder, J.; Bovy, A.; Hall, R.; Molenaar, J. The Flavonoid Pathway in Tomato Seedlings: Transcript Abundance and the Modeling of Metabolite Dynamics. PLOS ONE 2013, 8, e68960. [CrossRef]
- Valancin, A.; Srinivasan, B.; Rivoal, J.; Jolicoeur, M. Analyzing the Effect of Decreasing Cytosolic Triosephosphate Isomerase on Solanum Tuberosum Hairy Root Cells Using a Kinetic–Metabolic Model. Biotechnology and Bioengineering 2013, 110, 924–935. [CrossRef]
- Pokhilko, A.; Bou-Torrent, J.; Pulido, P.; Rodríguez-Concepción, M.; Ebenhöh, O. Mathematical Modelling of the Diurnal Regulation of the MEP Pathway in Arabidopsis. New Phytologist 2015, 206, 1075–1085. [CrossRef]
- Lucido, A.; Basallo, O.; Sorribas, A.; Marin-Sanguino, A.; Vilaprinyo, E.; Alves, R. A Mathematical Model for Strigolactone Biosynthesis in Plants. Frontiers in Plant Science 2022, 13.
- Olsen, K.M.; Slimestad, R.; Lea, U.S.; Brede, C.; Løvdal, T.; Ruoff, P.; Verheul, M.; Lillo, C. Temperature and Nitrogen Effects on Regulators and Products of the Flavonoid Pathway: Experimental and Kinetic Model Studies. Plant, Cell & Environment 2009, 32, 286–299. [CrossRef]
- Sorribas, A.; Hernández-Bermejo, B.; Vilaprinyo, E.; Alves, R. Cooperativity and Saturation in Biochemical Networks: A Saturable Formalism Using Taylor Series Approximations. Biotechnology and Bioengineering 2007, 97, 1259–1277. [CrossRef]
- Alves, R.; Vilaprinyo, E.; Hernádez-Bermejo, B.; Sorribas, A. Mathematical Formalisms Based on Approximated Kinetic Representations for Modeling Genetic and Metabolic Pathways. Biotechnol Genet Eng Rev 2008, 25, 1–40. [CrossRef]
- Faraji, M.; Fonseca, L.L.; Escamilla-Treviño, L.; Barros-Rios, J.; Engle, N.L.; Yang, Z.K.; Tschaplinski, T.J.; Dixon, R.A.; Voit, E.O. A Dynamic Model of Lignin Biosynthesis in Brachypodium Distachyon. Biotechnology for Biofuels 2018, 11, 253. [CrossRef]
- Lee, Y.; Voit, E.O. Mathematical Modeling of Monolignol Biosynthesis in Populus Xylem. Mathematical Biosciences 2010, 228, 78–89. [CrossRef]
- Tran, L.M.; Rizk, M.L.; Liao, J.C. Ensemble Modeling of Metabolic Networks. Biophysical Journal 2008, 95, 5606–5617. [CrossRef]
- Reder, C. Metabolic Control Theory: A Structural Approach. Journal of Theoretical Biology 1988, 135, 175–201. [CrossRef]
- Savageau, M.A. Biochemical Systems Analysis: A Study of Function and Design in Molecular Biology; CreateSpace Independent Publishing Platform, 2010; ISBN 978-1-4495-9076-5.
- Voit, E. A First Course in Systems Biology; New York, NY, 2012; ISBN 978-0-8153-4467-4.
- Resendis-Antonio, O. Stoichiometric Matrix. In Encyclopedia of Systems Biology; Dubitzky, W., Wolkenhauer, O., Cho, K.-H., Yokota, H., Eds.; Springer: New York, NY, 2013; pp. 2014–2014 ISBN 978-1-4419-9863-7.
- Paez-Garcia, A.; Motes, C.M.; Scheible, W.-R.; Chen, R.; Blancaflor, E.B.; Monteros, M.J. Root Traits and Phenotyping Strategies for Plant Improvement. Plants 2015, 4, 334–355. [CrossRef]
- Tajima, R. Importance of Individual Root Traits to Understand Crop Root System in Agronomic and Environmental Contexts. Breed Sci 2021, 71, 13–19. [CrossRef]
- Vergauwen, D.; De Smet, I. From Early Farmers to Norman Borlaug — the Making of Modern Wheat. Current Biology 2017, 27, R858–R862. [CrossRef]
- Reynolds, M.P.; Braun, H.-J. Wheat Improvement. In Wheat Improvement: Food Security in a Changing Climate; Reynolds, M.P., Braun, H.-J., Eds.; Springer International Publishing: Cham, 2022; pp. 3–15 ISBN 978-3-030-90673-3.
- Ramstein, G.P.; Jensen, S.E.; Buckler, E.S. Breaking the Curse of Dimensionality to Identify Causal Variants in Breeding 4. Theor Appl Genet 2019, 132, 559–567. [CrossRef]
- Yan, H.; KANG, M.Z.; DE REFFYE, P.; DINGKUHN, M. A Dynamic, Architectural Plant Model Simulating Resource-dependent Growth. Ann Bot 2004, 93, 591–602. [CrossRef]
- McSteen, P.; Leyser, O. Shoot Branching. Annu Rev Plant Biol 2005, 56, 353–374. [CrossRef]
- Evers, J.B.; Vos, J. Modeling Branching in Cereals. Front. Plant Sci. 2013, 4. [CrossRef]
- Wen, W.; Wang, Y.; Wu, S.; Liu, K.; Gu, S.; Guo, X. 3D Phytomer-Based Geometric Modelling Method for Plants—the Case of Maize. AoB PLANTS 2021, 13, plab055. [CrossRef]
- Buck-Sorlin, G.H.; Kniemeyer, O.; Kurth, W. Barley Morphology, Genetics and Hormonal Regulation of Internode Elongation Modelled by a Relational Growth Grammar. New Phytologist 2005, 166, 859–867. [CrossRef]
- Buck-Sorlin, G.; de Visser, P.H.B.; Henke, M.; Sarlikioti, V.; van der Heijden, G.W.A.M.; Marcelis, L.F.M.; Vos, J. Towards a Functional–Structural Plant Model of Cut-Rose: Simulation of Light Environment, Light Absorption, Photosynthesis and Interference with the Plant Structure. Annals of Botany 2011, 108, 1121–1134. [CrossRef]
- Groer, C.; Kniemeyer, O.; Hemmerling, R.; Kurth, W.; Becker, H.; Buck-Sorlin, G. A Dynamic 3D Model of Rape (Brassica Napus L.) Computing Yield Components under Variable Nitrogen Fertilization Regimes. 2007.
- Thornby, D.; Spencer, D.; Hanan, J.; Sher, A. L-DONAX, a Growth Model of the Invasive Weed Species, Arundo Donax L. Aquatic Botany 2007, 87, 275–284. [CrossRef]
- Barczi, J.-F.; Rey, H.; Caraglio, Y.; de Reffye, P.; Barthélémy, D.; Dong, Q.X.; Fourcaud, T. AmapSim: A Structural Whole-Plant Simulator Based on Botanical Knowledge and Designed to Host External Functional Models. Annals of Botany 2008, 101, 1125–1138. [CrossRef]
- Kahlen, K.; Wiechers, D.; Stützel, H. Modelling Leaf Phototropism in a Cucumber Canopy. Functional Plant Biol. 2008, 35, 876–884. [CrossRef]
- Jallas, E.; Sequeira, R.; Martin, P.; Turner, S.; Papajorgji, P. Mechanistic Virtual Modeling: Coupling a Plant Simulation Model with a Three-Dimensional Plant Architecture Component. Environ Model Assess 2009, 14, 29–45. [CrossRef]
- Chen, T.-W.; Nguyen, T.M.N.; Kahlen, K.; Stützel, H. Quantification of the Effects of Architectural Traits on Dry Mass Production and Light Interception of Tomato Canopy under Different Temperature Regimes Using a Dynamic Functional–Structural Plant Model. Journal of Experimental Botany 2014, 65, 6399–6410. [CrossRef]
- Gu, S.; Evers, J.B.; Zhang, L.; Mao, L.; Zhang, S.; Zhao, X.; Liu, S.; van der Werf, W.; Li, Z. Modelling the Structural Response of Cotton Plants to Mepiquat Chloride and Population Density. Annals of Botany 2014, 114, 877–887. [CrossRef]
- Henke, M.; Kurth, W.; Buck-Sorlin, G.H. FSPM-P: Towards a General Functional-Structural Plant Model for Robust and Comprehensive Model Development. Front. Comput. Sci. 2016, 10, 1103–1117. [CrossRef]
- Boudon, F.; Persello, S.; Jestin, A.; Briand, A.-S.; Grechi, I.; Fernique, P.; Guédon, Y.; Léchaudel, M.; Lauri, P.-É.; Normand, F. V-Mango: A Functional–Structural Model of Mango Tree Growth, Development and Fruit Production. Annals of Botany 2020, 126, 745–763. [CrossRef]
- Prusinkiewicz, P. A Look at the Visual Modeling of Plants Using L-Systems. In Proceedings of the Bioinformatics; Hofestädt, R., Lengauer, T., Löffler, M., Schomburg, D., Eds.; Springer: Berlin, Heidelberg, 1997; pp. 11–29.
- Lobet, G.; Draye, X.; Périlleux, C. An Online Database for Plant Image Analysis Software Tools. Plant Methods 2013, 9, 38. [CrossRef]
- Teichmann, T.; Muhr, M. Shaping Plant Architecture. Front. Plant Sci. 2015, 6. [CrossRef]
- Zhang, W.; Wang, G.; Han, J.; Li, F.; Zhang, Q.; Doonan, J. A Functional-Structural Model for Alfalfa That Accurately Integrates Shoot and Root Growth and Development. In Proceedings of the 2018 6th International Symposium on Plant Growth Modeling, Simulation, Visualization and Applications (PMA); November 2018; pp. 134–140.
- Trachsel, S.; Kaeppler, S.M.; Brown, K.M.; Lynch, J.P. Shovelomics: High Throughput Phenotyping of Maize (Zea Mays L.) Root Architecture in the Field. Plant Soil 2011, 341, 75–87. [CrossRef]
- Teramoto, S.; Kitomi, Y.; Nishijima, R.; Takayasu, S.; Maruyama, N.; Uga, Y. Backhoe-Assisted Monolith Method for Plant Root Phenotyping under Upland Conditions. Breed Sci 2019, 69, 508–513. [CrossRef]
- Teramoto, S.; Takayasu, S.; Kitomi, Y.; Arai-Sanoh, Y.; Tanabata, T.; Uga, Y. High-Throughput Three-Dimensional Visualization of Root System Architecture of Rice Using X-Ray Computed Tomography. Plant Methods 2020, 16, 66. [CrossRef]
- Leitner, D.; Klepsch, S.; Bodner, G.; Schnepf, A. A Dynamic Root System Growth Model Based on L-Systems. Plant Soil 2010, 332, 177–192. [CrossRef]
- Stützel, H.; Kahlen, K. Editorial: Virtual Plants: Modeling Plant Architecture in Changing Environments. Front. Plant Sci. 2016, 7. [CrossRef]
- Lungley, D.R. The Growth of Root Systems — A Numerical Computer Simulation Model. Plant Soil 1973, 38, 145–159. [CrossRef]
- Hackett, C.; Rose, D.A. A MODEL OF THE EXTENSION AND BRANCHING OF A SEMINAL ROOT OF BARLEY, AND ITS USE IN STUDYING RELATIONS BETWEEN ROOT DIMENSIONS. 1972.
- Diggle, A.J. ROOTMAP—a Model in Three-Dimensional Coordinates of the Growth and Structure of Fibrous Root Systems. Plant Soil 1988, 105, 169–178. [CrossRef]
- Pagès, L.; Jordan, M.O.; Picard, D. A Simulation Model of the Three-Dimensional Architecture of the Maize Root System. Plant Soil 1989, 119, 147–154. [CrossRef]
- Pagès, L.; Vercambre, G.; Drouet, J.-L.; Lecompte, F.; Collet, C.; Le Bot, J. Root Typ: A Generic Model to Depict and Analyse the Root System Architecture. Plant and Soil 2004, 258, 103–119. [CrossRef]
- Barczi, J.-F.; Rey, H.; Griffon, S.; Jourdan, C. DigR: A Generic Model and Its Open Source Simulation Software to Mimic Three-Dimensional Root-System Architecture Diversity. Annals of Botany 2018, 121, 1089–1104. [CrossRef]
- Wu, L.; McGechan, M.B.; McRoberts, N.; Baddeley, J.A.; Watson, C.A. SPACSYS: Integration of a 3D Root Architecture Component to Carbon, Nitrogen and Water Cycling—Model Description. Ecological Modelling 2007, 200, 343–359. [CrossRef]
- Lynch, J.P.; Nielsen, K.L.; Davis, R.D.; Jablokow, A.G. SimRoot: Modelling and Visualization of Root Systems. Plant and Soil 1997, 188, 139–151. [CrossRef]
- Postma, J.A.; Kuppe, C.; Owen, M.R.; Mellor, N.; Griffiths, M.; Bennett, M.J.; Lynch, J.P.; Watt, M. OpenSimRoot: Widening the Scope and Application of Root Architectural Models. New Phytologist 2017, 215, 1274–1286. [CrossRef]
- Schnepf, A.; Leitner, D.; Landl, M.; Lobet, G.; Mai, T.H.; Morandage, S.; Sheng, C.; Zörner, M.; Vanderborght, J.; Vereecken, H. CRootBox: A Structural–Functional Modelling Framework for Root Systems. Annals of Botany 2018, 121, 1033–1053. [CrossRef]
- Lucido, A.; Andrade, F.; Basallo, O.; Eleiwa, A.; Marin-Sanguino, A.; Vilaprinyo, E.; Sorribas, A.; Alves, R. Modeling the Effects of Strigolactone Levels on Maize Root System Architecture. Frontiers in Plant Science 2024, 14.
- Niinemets, Ü.; Tenhunen, J.D. A Model Separating Leaf Structural and Physiological Effects on Carbon Gain along Light Gradients for the Shade-Tolerant Species Acer Saccharum. Plant, Cell & Environment 1997, 20, 845–866. [CrossRef]
- Hinojo-Hinojo, C.; Castellanos, A.E.; Llano-Sotelo, J.; Peñuelas, J.; Vargas, R.; Romo-Leon, J.R. High Vcmax, Jmax and Photosynthetic Rates of Sonoran Desert Species: Using Nitrogen and Specific Leaf Area Traits as Predictors in Biochemical Models. Journal of Arid Environments 2018, 156, 1–8. [CrossRef]
- Liu, Q.; Xie, L.; Li, F. Dynamic Simulation of the Crown Net Photosynthetic Rate for Young Larix Olgensis Henry Trees. Forests 2019, 10, 321. [CrossRef]
- Barley, K.P. The Configuration of the Root System in Relation to Nutrient Uptake. In Advances in Agronomy; Brady, N.C., Ed.; Academic Press, 1970; Vol. 22, pp. 159–201.
- Johnson, M.P. Photosynthesis. Essays Biochem 2016, 60, 255–273. [CrossRef]
- García-Rodríguez, L. del C.; Prado-Olivarez, J.; Guzmán-Cruz, R.; Rodríguez-Licea, M.A.; Barranco-Gutiérrez, A.I.; Perez-Pinal, F.J.; Espinosa-Calderon, A. Mathematical Modeling to Estimate Photosynthesis: A State of the Art. Applied Sciences 2022, 12, 5537. [CrossRef]
- Farquhar, G.D.; von Caemmerer, S.; Berry, J.A. A Biochemical Model of Photosynthetic CO2 Assimilation in Leaves of C 3 Species. Planta 1980, 149, 78–90. [CrossRef]
- von Caemmerer, S. Updating the Steady-State Model of C4 Photosynthesis. Journal of Experimental Botany 2021, 72, 6003–6017. [CrossRef]
- Yin, X.; Struik, P.C. C3 and C4 Photosynthesis Models: An Overview from the Perspective of Crop Modelling. NJAS: Wageningen Journal of Life Sciences 2009, 57, 27–38. [CrossRef]
- Sage, R.F. The Evolution of C4 Photosynthesis. New Phytologist 2004, 161, 341–370. [CrossRef]
- Burgos, A.; Miranda, E.; Vilaprinyo, E.; Meza-Canales, I.D.; Alves, R. CAM Models: Lessons and Implications for CAM Evolution. Front Plant Sci 2022, 13, 893095. [CrossRef]
- Nungesser, D.; Kluge, M.; Tolle, H.; Oppelt, W. A Dynamic Computer Model of the Metabolic and Regulatory Processes in Crassulacean Acid Metabolism. Planta 1984, 162, 204–214.
- Lüttge, U.; Beck, F. Endogenous Rhythms and Chaos in Crassulacean Acid Metabolism. Planta 1992, 188, 28–38.
- Blasius, B.; Neif, R.; Beck, F.; Lüttge, U. Oscillatory Model of Crassulacean Acid Metabolism with a Dynamic Hysteresis Switch. Proceedings of the Royal Society of London. Series B: Biological Sciences 1999, 266, 93–101. [CrossRef]
- Cheung, C.Y.M.; Poolman, M.G.; Fell, David.A.; Ratcliffe, R.G.; Sweetlove, L.J. A Diel Flux Balance Model Captures Interactions between Light and Dark Metabolism during Day-Night Cycles in C3 and Crassulacean Acid Metabolism Leaves. Plant Physiology 2014, 165, 917–929. [CrossRef]
- Javaux, M.; Schröder, T.; Vanderborght, J.; Vereecken, H. Use of a Three-Dimensional Detailed Modeling Approach for Predicting Root Water Uptake. Vadose Zone Journal 2008, 7, 1079–1088. [CrossRef]
- Lacointe, A.; Minchin, P.E.H. A Mechanistic Model to Predict Distribution of Carbon Among Multiple Sinks. In Phloem: Methods and Protocols; Liesche, J., Ed.; Springer: New York, NY, 2019; pp. 371–386 ISBN 978-1-4939-9562-2.
- Rengel, Z. Mechanistic Simulation Models of Nutrient Uptake: A Review. Plant Soil 1993, 152, 161–173. [CrossRef]
- Willigen, P. de; Noordwijk, M. van Modelling Nutrient Uptake : From Single Roots to Complete Root Systems. In Simulation and systems analysis for rice production (SARP) : selected papers presented at workshops on crop simulation of a network of National and International Research Centres of several Asian countries and The Netherlands, 1990 - 1991; Pudoc, 1991; pp. 277–295.
- Nye, P.H. The Effect of the Nutrient Intensity and Buffering Power of a Soil, and the Absorbing Power, Size and Root Hairs of a Root, on Nutrient Absorption by Diffusion. Plant Soil 1966, 25, 81–105. [CrossRef]
- Nye, P.H.; Marriott, F.H.C. A Theoretical Study of the Distribution of Substances around Roots Resulting from Simultaneous Diffusion and Mass Flow. Plant Soil 1969, 30, 459–472. [CrossRef]
- Baldwin, J.P.; Nye, P.H.; Tinker, P.B. Uptake of Solutes by Multiple Root Systems from Soil: Iii — a Model for Calculating the Solute Uptake by a Randomly Dispersed Root System Developing in a Finite Volume of Soil. Plant and Soil 1973, 38, 621–635.
- Baldwin, J.P.; Nye, P.H. A Model to Calculate the Uptake by a Developing Root System or Root Hair System of Solutes with Concentration Variable Diffusion Coefficients. Plant and Soil 1974, 40, 703–706.
- Bhat, K.K.S.; Nye, P.H.; Baldwin, J.P. Diffusion of Phosphate to Plant Roots in Soil. Plant Soil 1976, 44, 63–72. [CrossRef]
- Itoh, S.; Barber, S.A. A Numerical Solution of Whole Plant Nutrient Uptake for Soil-Root Systems with Root Hairs. Plant Soil 1983, 70, 403–413. [CrossRef]
- Barber, S.A. Soil Nutrient Bioavailability: A Mechanistic Approach; John Wiley & Sons, 1995; ISBN 978-0-471-58747-7.
- Tinker, P.B.; Nye, P.H. Solute Movement in the Rhizosphere; 2nd edition.; Oxford University Press: New York, 2000; ISBN 978-0-19-512492-7.
- Roose, T.; Fowler, A.C.; Darrah, P.R. A Mathematical Model of Plant Nutrient Uptake. J Math Biol 2001, 42, 347–360. [CrossRef]
- Roose, T.; Fowler, A.C. A Mathematical Model for Water and Nutrient Uptake by Plant Root Systems. Journal of Theoretical Biology 2004, 228, 173–184. [CrossRef]
- Roose, T.; Fowler, A.C. A Model for Water Uptake by Plant Roots. Journal of Theoretical Biology 2004, 228, 155–171. [CrossRef]
- Roose, T.; Kirk, G.J.D. The Solution of Convection–Diffusion Equations for Solute Transport to Plant Roots. Plant Soil 2009, 316, 257–264. [CrossRef]
- Ou, Z. Approximate Nutrient Flux and Concentration Solutions of the Nye–Tinker–Barber Model by the Perturbation Expansion Method. Journal of Theoretical Biology 2019, 476, 19–29. [CrossRef]
- Wang, Y.; Lin, W.; Ou, Z. Analytical Solution of Nye–Tinker–Barber Model by Laplace Transform. Biosystems 2023, 225, 104845. [CrossRef]
- Jones Jr., J.B. Plant Nutrition and Soil Fertility Manual; 2nd ed.; CRC Press: Boca Raton, 2012; ISBN 978-0-429-13081-6.
- Edelstein-Keshet, L. Mathematical Models in Biology; Classics in Applied Mathematics; Society for Industrial and Applied Mathematics, 2005; ISBN 978-0-89871-554-5.
- Kuppe, C.W.; Schnepf, A.; von Lieres, E.; Watt, M.; Postma, J.A. Rhizosphere Models: Their Concepts and Application to Plant-Soil Ecosystems. Plant Soil 2022, 474, 17–55. [CrossRef]
- Conejo, A.N. Non-Dimensionalization of Differential Equations. In Fundamentals of Dimensional Analysis; Springer Singapore: Singapore, 2021; pp. 347–378 ISBN 9789811616013.
- Kuppe, C.W.; Huber, G.; Postma, J.A. Comparison of Numerical Methods for Radial Solute Transport to Simulate Uptake by Plant Roots. Rhizosphere 2021, 18, 100352. [CrossRef]
- Sultan, S.E. Phenotypic Plasticity for Plant Development, Function and Life History. Trends in Plant Science 2000, 5, 537–542. [CrossRef]
- Thornley, J.H.M. A Balanced Quantitative Model for Root: Shoot Ratios in Vegetative Plants. Annals of Botany 1972, 36, 431–441. [CrossRef]
- Thornley, J.H.M. Shoot: Root Allocation with Respect to C, N and P: An Investigation and Comparison of Resistance and Teleonomic Models. Annals of Botany 1995, 75, 391–405.
- Thornley, J.H.M. Modelling Shoot[Ratio]Root Relations: The Only Way Forward? Annals of Botany 1998, 81, 165–171. [CrossRef]
- Dewar, R.C. A Root-Shoot Partitioning Model Based on Carbon-Nitrogen-Water Interactions and Munch Phloem Flow. Functional Ecology 1993, 7, 356–368. [CrossRef]
- Feller, C.; Favre, P.; Janka, A.; Zeeman, S.C.; Gabriel, J.-P.; Reinhardt, D. Mathematical Modeling of the Dynamics of Shoot-Root Interactions and Resource Partitioning in Plant Growth. PLOS ONE 2015, 10, e0127905. [CrossRef]
- Zhou, X.-R.; Schnepf, A.; Vanderborght, J.; Leitner, D.; Lacointe, A.; Vereecken, H.; Lobet, G. CPlantBox, a Whole-Plant Modelling Framework for the Simulation of Water- and Carbon-Related Processes. in silico Plants 2020, 2, diaa001. [CrossRef]
- Lacointe, A.; Minchin, P.E.H. Modelling Phloem and Xylem Transport within a Complex Architecture. Functional Plant Biol. 2008, 35, 772–780. [CrossRef]
- Wilson, J.B. A Review of Evidence on the Control of Shoot: Root Ratio, in Relation to Models. Annals of Botany 1988, 61, 433–449.
- Reynolds, J.F.; Thornley, J.H.M. A Shoot: Root Partitioning Model. Annals of Botany 1982, 49, 585–597.
- Li, X.; Feng, Y.; Boersma, L. Partition of Photosynthates between Shoot and Root in Spring Wheat (Triticum Aestivum L.) as a Function of Soil Water Potential and Root Temperature. Plant Soil 1994, 164, 43–50. [CrossRef]
- Bertheloot, J.; Cournède, P.-H.; Andrieu, B. NEMA, a Functional–Structural Model of Nitrogen Economy within Wheat Culms after Flowering. I. Model Description. Annals of Botany 2011, 108, 1085–1096. [CrossRef]
- Gupta, D.; Sharma, G.; Saraswat, P.; Ranjan, R. Synthetic Biology in Plants, a Boon for Coming Decades. Mol Biotechnol 2021, 63, 1138–1154. [CrossRef]
- Deneer, A.; Fleck, C. Mathematical Modelling in Plant Synthetic Biology. In Methods in molecular biology (Clifton, N.J.); 2022; Vol. 2379, pp. 209–251 ISBN 978-1-07-161790-8.
- Roell, M.-S.; Zurbriggen, M.D. The Impact of Synthetic Biology for Future Agriculture and Nutrition. Current Opinion in Biotechnology 2020, 61, 102–109. [CrossRef]
- Andres, J.; Blomeier, T.; Zurbriggen, M.D. Synthetic Switches and Regulatory Circuits in Plants. Plant Physiology 2019, 179, 862–884. [CrossRef]
- Scheible, W.-R.; Lauerer, M.; Schulze, E.-D.; Caboche, M.; Stitt, M. Accumulation of Nitrate in the Shoot Acts as a Signal to Regulate Shoot-Root Allocation in Tobacco†. The Plant Journal 1997, 11, 671–691. [CrossRef]
- Lemoine, R.; La Camera, S.; Atanassova, R.; Dédaldéchamp, F.; Allario, T.; Pourtau, N.; Bonnemain, J.-L.; Laloi, M.; Coutos-Thévenot, P.; Maurousset, L.; et al. Source-to-Sink Transport of Sugar and Regulation by Environmental Factors. Front. Plant Sci. 2013, 4. [CrossRef]
- Wang, L.; Ruan, Y.-L. Shoot–Root Carbon Allocation, Sugar Signalling and Their Coupling with Nitrogen Uptake and Assimilation. Functional Plant Biol. 2015, 43, 105–113. [CrossRef]
- Alves, R.; Savageau, M.A. Effect of Overall Feedback Inhibition in Unbranched Biosynthetic Pathways. Biophys J 2000, 79, 2290–2304. [CrossRef]
- Vazquez-Vilar, M.; Selma, S.; Orzaez, D. The Design of Synthetic Gene Circuits in Plants: New Components, Old Challenges. Journal of Experimental Botany 2023, 74, 3791–3805. [CrossRef]
- Marshall-Colon, A.; Long, S.P.; Allen, D.K.; Allen, G.; Beard, D.A.; Benes, B.; von Caemmerer, S.; Christensen, A.J.; Cox, D.J.; Hart, J.C.; et al. Crops In Silico: Generating Virtual Crops Using an Integrative and Multi-Scale Modeling Platform. Front. Plant Sci. 2017, 8. [CrossRef]
- Baldazzi, V.; Bertin, N.; Jong, H. de; Génard, M. Towards Multiscale Plant Models: Integrating Cellular Networks. Trends in Plant Science 2012, 17, 728–736. [CrossRef]
- Herrero-Huerta, M.; Meline, V.; Iyer-Pascuzzi, A.S.; Souza, A.M.; Tuinstra, M.R.; Yang, Y. 4D Structural Root Architecture Modeling from Digital Twins by X-Ray Computed Tomography. Plant Methods 2021, 17, 123. [CrossRef]
- York, L.M. Functional Phenomics: An Emerging Field Integrating High-Throughput Phenotyping, Physiology, and Bioinformatics. Journal of Experimental Botany 2019, 70, 379–386. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



