Submitted:
01 October 2024
Posted:
03 October 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Cultivation, Pathogen Material and Inoculation Procedure
2.2. Hyperspectral Imaging Measurement
2.3. Data Analysis
2.4. Supervised Machine Learning Methods
3. Results
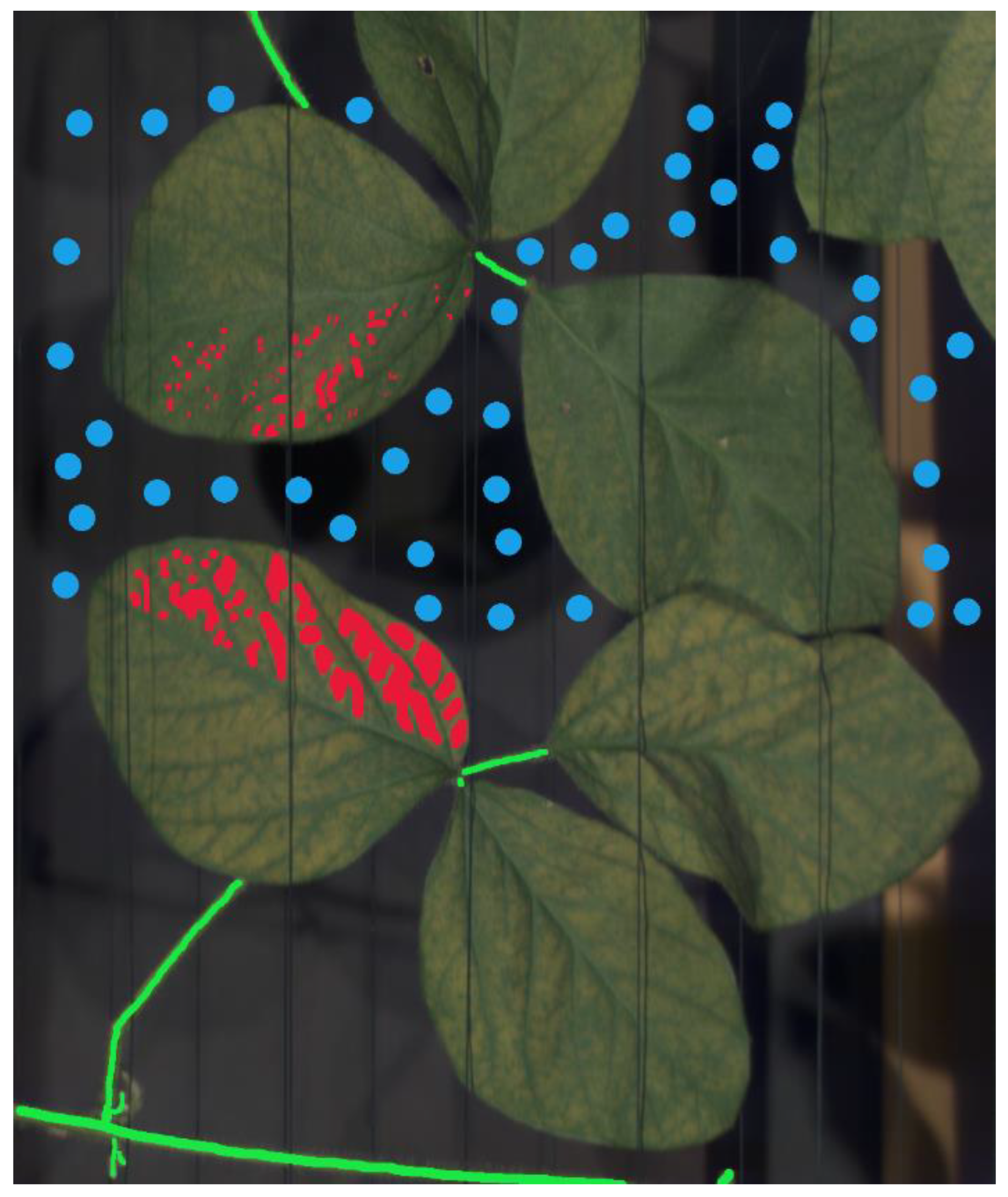
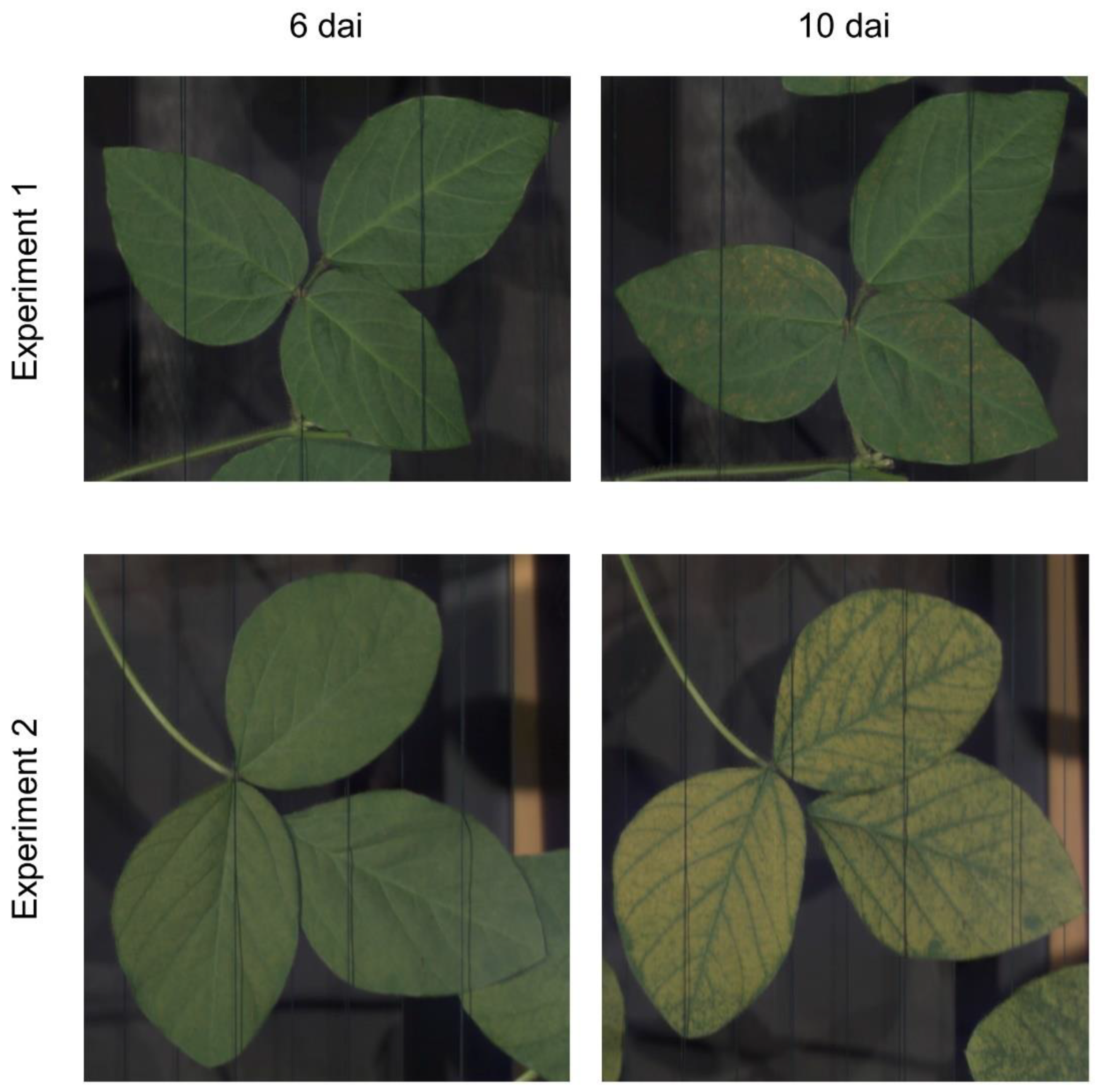
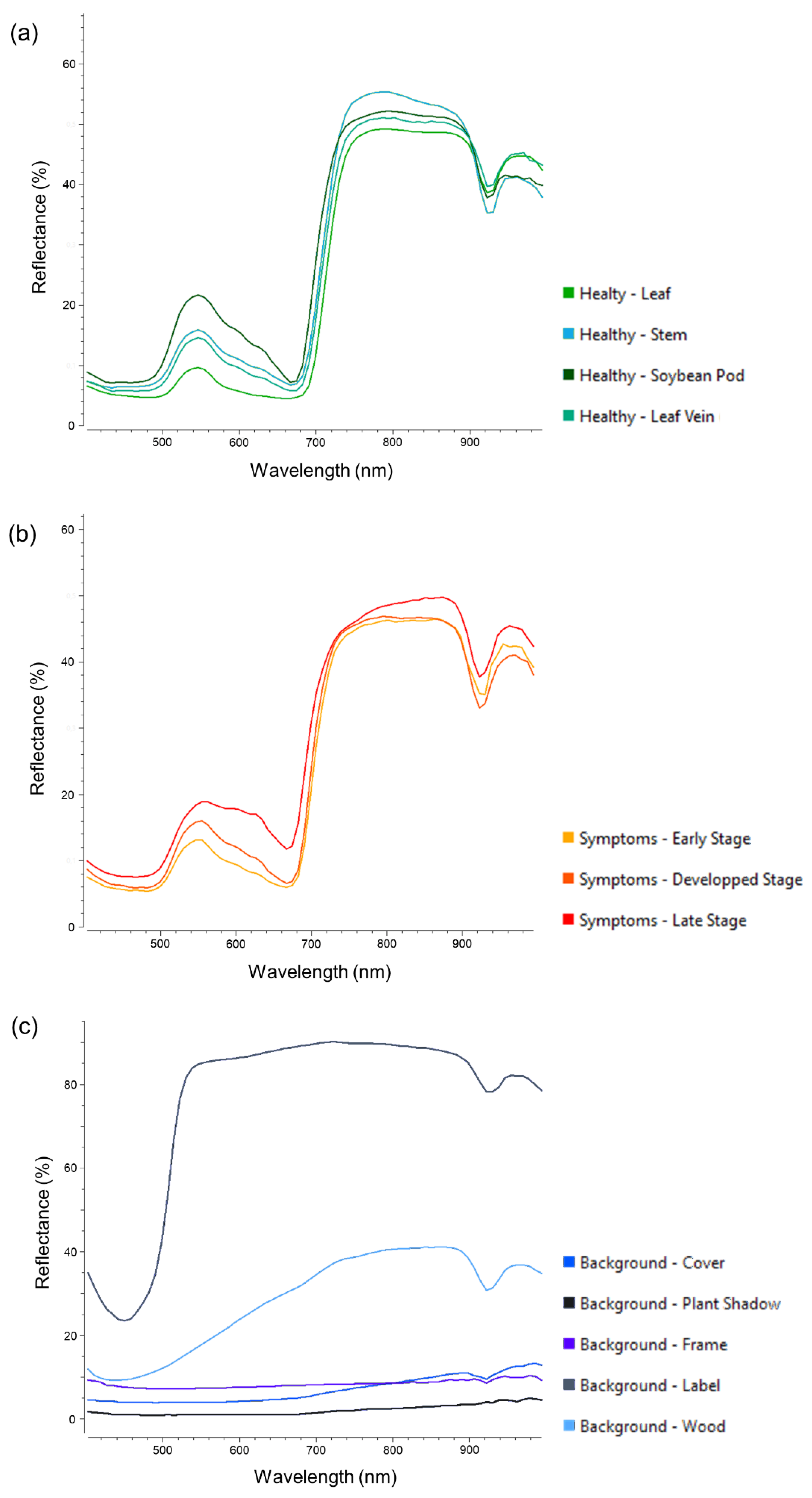
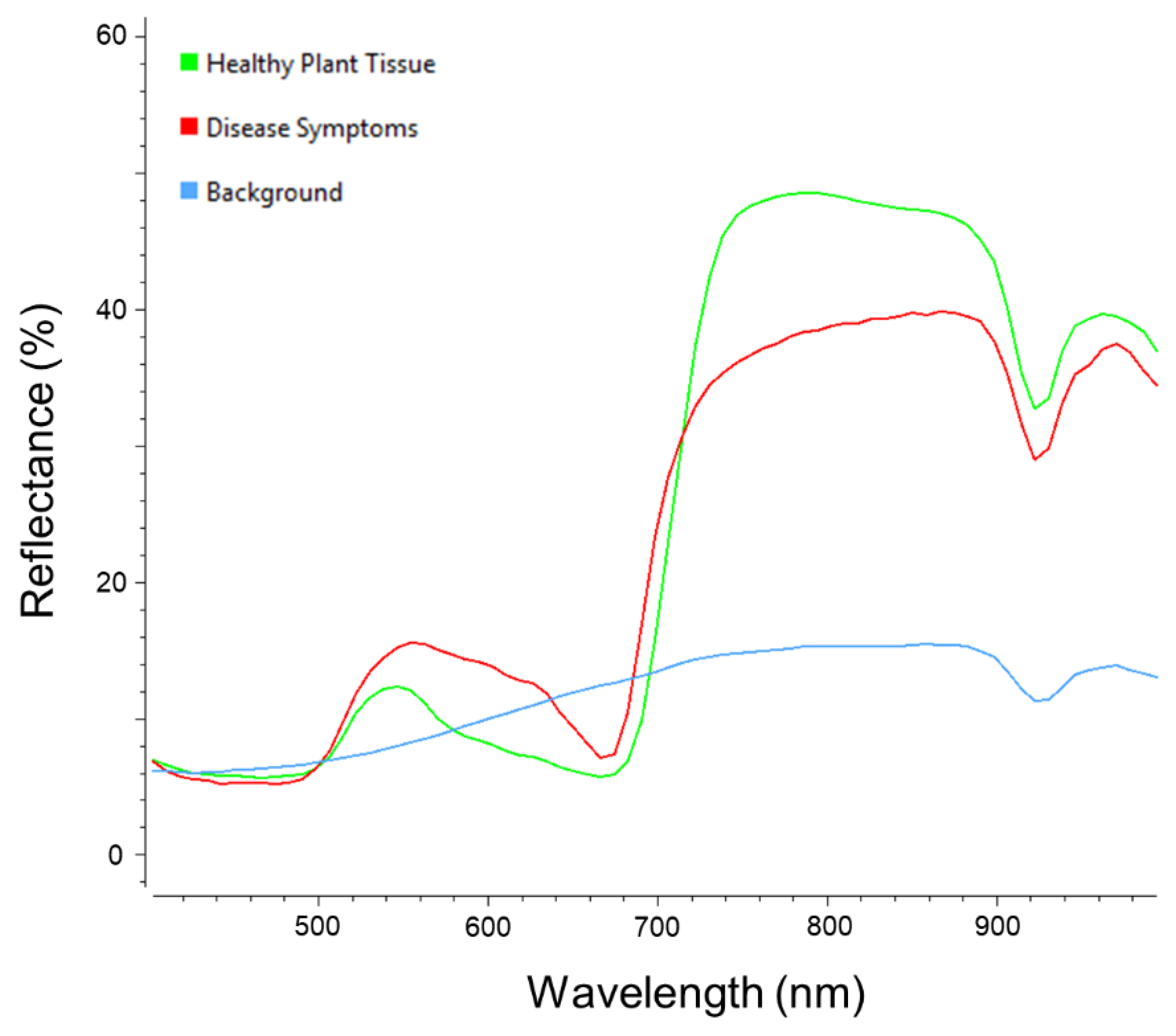
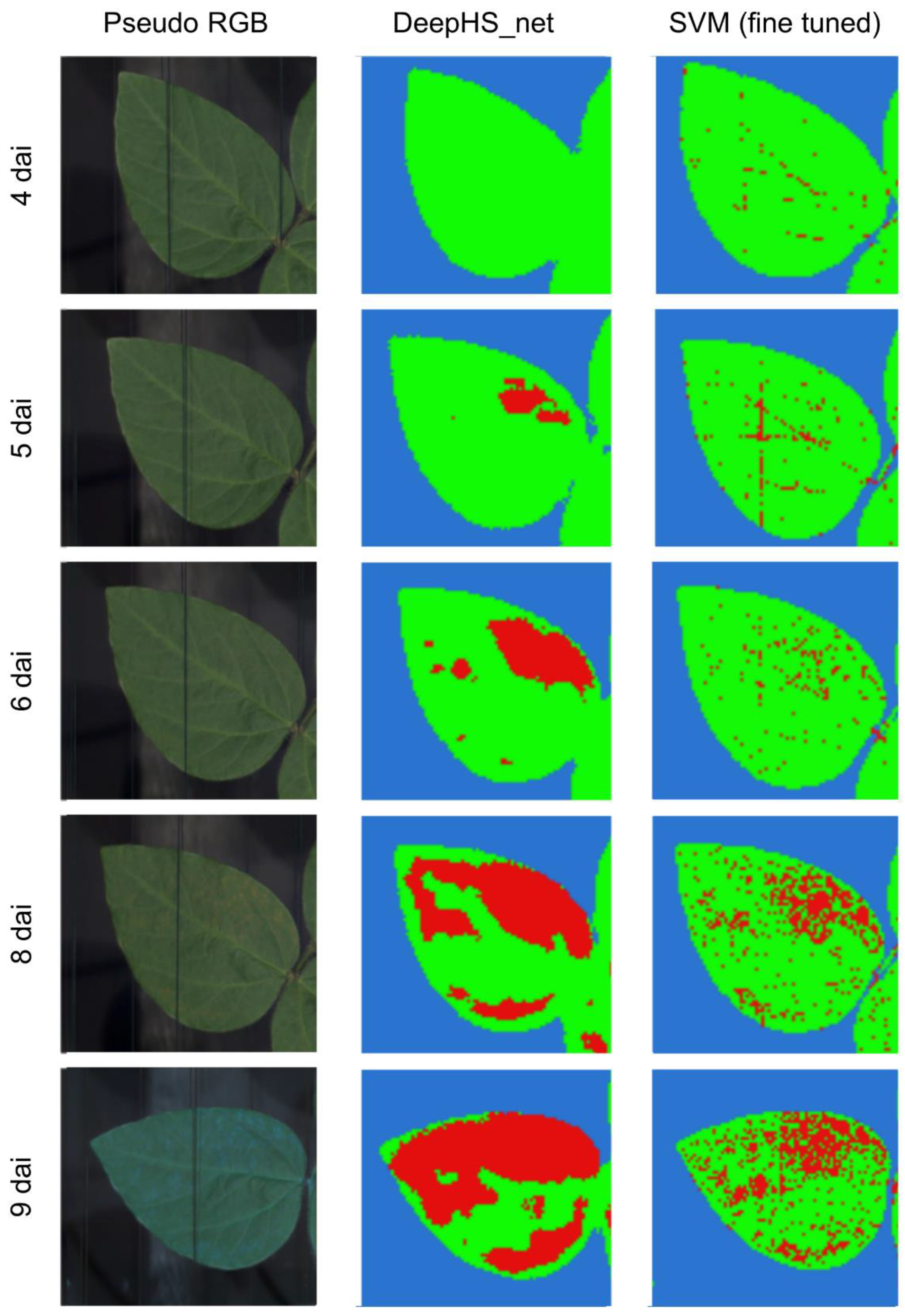
3.1. Visual Assessment of the Hyperspectral Datasets
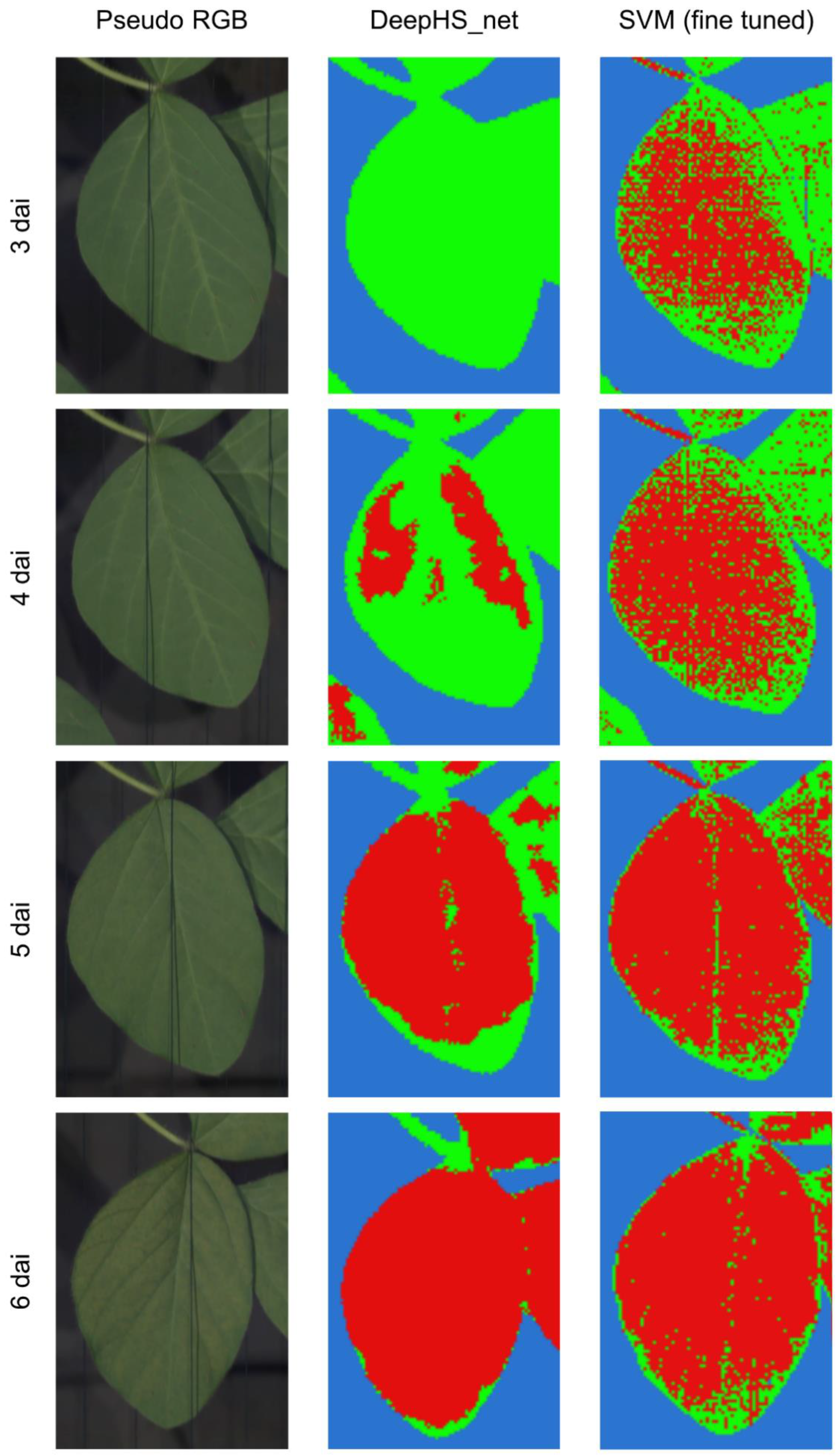
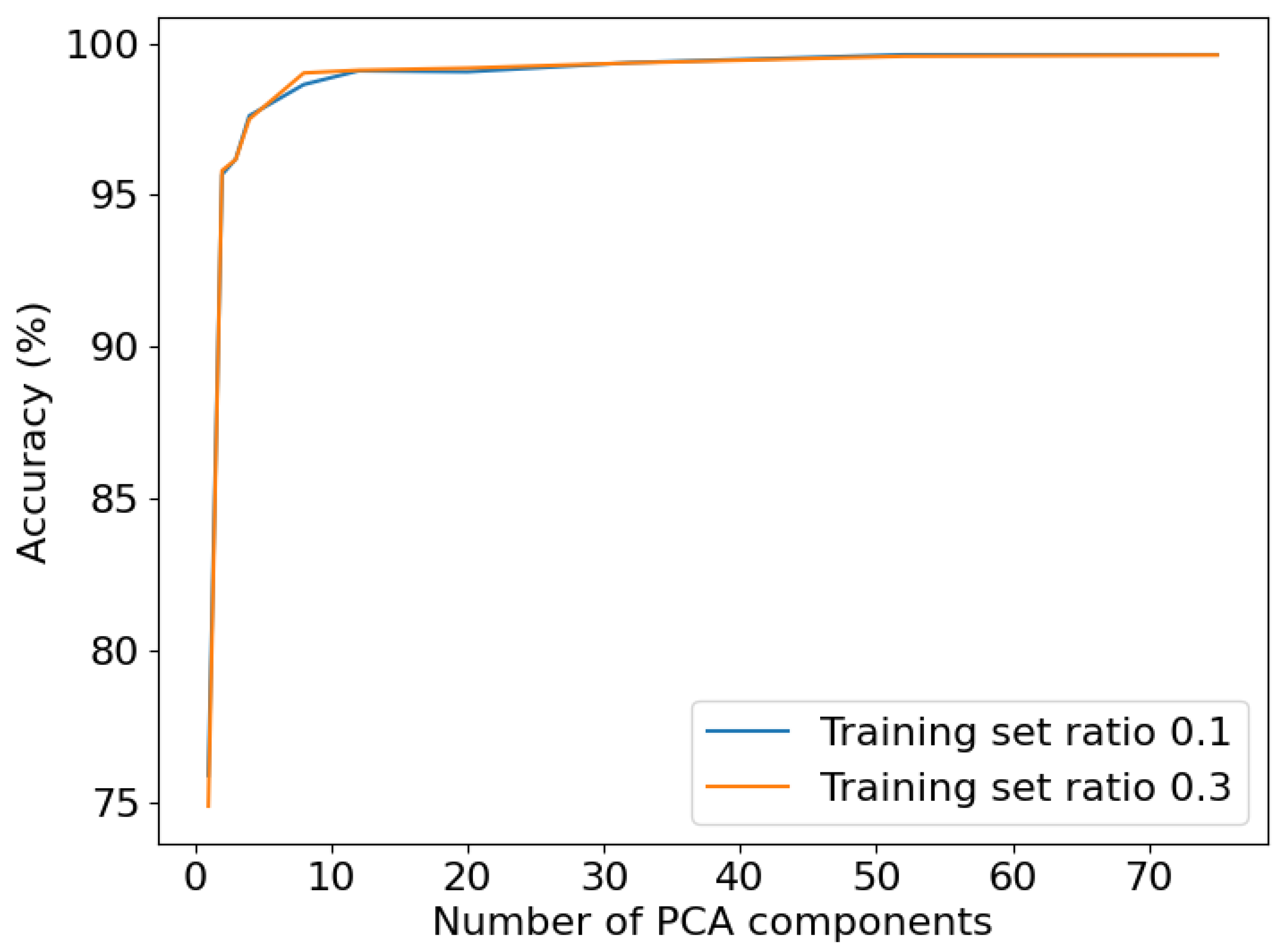
3.2. Analysis of the Hyperspectral Datasets through Supervised Machine Learning and Neural Networks
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- United States Department of Agriculture, Foreign Agricultural Service. Oilseeds: World Market and Trade. Available online: https://apps.fas.usda.gov/psdonline/circulars/oilseeds.pdf (accessed on 14 October 2022).
- Barro, J. P.; Alves, K. S.; Godoy, C. V.; Dias, A. R.; Forcelini, C. A.; Utiamada, C. M.; de Andrade Junior, E. R.; Juliatti, F. C.; Grigolli, F. J.; Feksa, H. R.; Campos, H. D.; Chaves, I. C.; Júnior, I. P. A.; Roy, J. M. T.; Nunes, J.; de R. Belufi, L. M.; Carneiro, L. C.; da Silva, L. H. C. P.; Canteri, M. G.; Júnior, M. M. G.; Senger, M.; Meyer, M. C.; Dias, M. D.; Müller, M. A.; Martins, M. C.; Debortoli, M. P.; Tormen, N. R.; Furlan, S. H.; Konageski, T. F.; Carlin, V. J.; Venâncio, W. S.; Del Ponte, E. M. Temporal and regional performance of dual and triple premixes for soybean rust management in Brazil: A meta-analysis update. Available online: https://osf.io/g94yn (accessed on 14 October 2022).
- Childs, S.P.; Buck, J. W.; Li, Z. Breeding soybeans with resistance to soybean rust (Phakopsora pachyrhizi). Plant Breed. 2018, 137, 250–261. [Google Scholar] [CrossRef]
- Goellner, K.; Loehrer, M.; Langenbach, C.; Conrath, U.; Koch, E.; Schaffrath, U. Phakopsora pachyrhizi, the causal agent of Asian soybean rust. Mol. Plant Path. 2010, 11, 169–177. [Google Scholar] [CrossRef]
- Ralf T., V. Uromyces fabae: development, metabolism, and interactions with its host Vicia faba. FEMS Microbiology Letters 2006, 259, 165–173. [Google Scholar] [CrossRef]
- Koch, E.; Ebrahim-Nesbat, F.; Hoppe, H. H. Light and Electron Microscopic Studies on the Development of Soybean Rust (Phakopsora pachyrhizi Syd.) in Susceptible Soybean Leaves. Journal of Phytopathology 1983, 106, 302–320. [Google Scholar] [CrossRef]
- Mahlein, A.-K. Plant Disease Detection by Imaging Sensors – Parallels and Specific Demands for Precision Agriculture and Plant Phenotyping. Plant Disease 2016, 100, 241–251. [Google Scholar] [CrossRef]
- Roitsch, T.; Cabrera-Bosquet, L.; Fournier, A.; Ghamkhar, K.; Jiménez-Berni, J.; Pinto, F.; Ober, E. S. Review: New sensors and data-driven approaches—A path to next generation phenomics. Plant Science 2019, 282, 2–10. [Google Scholar] [CrossRef]
- Alisaac, E.; Behmann, J.; Kuska, M. T.; Dehne, H.-W.; Mahlein, A.-K. Hyperspectral quantification of wheat resistance to Fusarium head blight: comparison of two Fusarium species. Eur J Plant Pathol 2018, 152, 869–884. [Google Scholar] [CrossRef]
- Thomas, S.; Kuska, M. T.; Bohnenkamp, D.; Brugger, A.; Alisaac, E.; Wahabzada, M.; Behmann, J.; Mahlein, A.-K. Benefits of hyperspectral imaging for plant disease detection and plant protection: a technical perspective. J Plant Dis Prot 2018, 125, 5–20. [Google Scholar] [CrossRef]
- Mahlein, A.-K.; Kuska, M. T.; Thomas, S.; Wahabzada, M.; Behmann, J.; Rascher, U.; Kersting, K. Quantitative and qualitative phenotyping of disease resistance of crops by hyperspectral sensors: seamless interlocking of phytopathology, sensors, and machine learning is needed! Current Opinion in Plant Biology 2019, 50, 156–162. [Google Scholar] [CrossRef]
- Cristianini, N.; Shawe-Taylor, J. An Introduction to Support Vector Machines and Other Kernel-Based Learning Methods. Cambridge: Cambridge University Press, England, 2000, 93-124. [CrossRef]
- Fix, E.; Hodges, J. L. Discriminatory Analysis. Nonparametric Discrimination: Consistency Properties. International Statistical Review / Revue Internationale de Statistique 1989, 57, 238–247. [Google Scholar] [CrossRef]
- Kuo, B.-C.; Ho, H.-H.; Li, C.-H.; Hung, C.-C.; Taur, J.-S. A Kernel-Based Feature Selection Method for SVM With RBF Kernel for Hyperspectral Image Classification. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing 2014, 7, 317–326. [Google Scholar] [CrossRef]
- Guo, Y.; Han, S.; Li, Y.; Zhang, C.; Bai, Y. K-Nearest Neighbor combined with guided filter for hyperspectral image classification. Procedia Computer Science 2018, 129, 159–165. [Google Scholar] [CrossRef]
- Li, S.; Song, W.; Fang, L.; Chen, Y.; Ghamisi, P.; Benediktsson, J. A. Deep Learning for Hyperspectral Image Classification: An Overview. IEEE Transactions on Geoscience and Remote Sensing 2019, 57, 6690–6709. [Google Scholar] [CrossRef]
- Matsugu, M.; Mori, K.; Mitari, Y.; Kaneda, Y. Subject independent facial expression recognition with robust face detection using a convolutional neural network. Neural Networks 2003, 16, 555–559. [Google Scholar] [CrossRef]
- Pearson, K. On lines and planes of closest fit to systems of points in space. The London, Edinburgh, and Dublin Philosophical Magazine and Journal of Science 1901, 2, 559–572. [Google Scholar] [CrossRef]
- Nair, V.; Hinton, G. E. Rectified Linear Units Improve Restricted Boltzmann Machines. In Proceedings of the 27th International Conference on Machine Learning (ICML-10), Haifa, Israel, 21-24 June 2010; Available online: https://icml.cc/Conferences/2010/papers/432.pdf.
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. In Proceedings of the 32nd International Conference on Machine Learning {ICML}, Lille, France, 6-11 July 2015; Available online: http://proceedings.mlr.press/v37/ioffe15.html.
- Varga, L. A.; Makowski, J.; Zell, A. Measuring the Ripeness of Fruit with Hyperspectral Imaging and Deep Learning. 2021 International Joint Conference on Neural Networks (IJCNN) 2021, 1–8. [Google Scholar] [CrossRef]
- Varga, L. A.; Messmer, M.; Benbarka, N.; Zell, A. Wavelength-aware 2D Convolutions for Hyperspectral Imaging. arXiv preprint 2022, accepted. arXiv:2209.03136. [Google Scholar]
- Kingma, D. P.; Ba, J. Adam: {A} Method for Stochastic Optimization. In Proceedings of the 3rd International Conference on Learning Representations {ICLR}, San Diego, CA, USA, 7-9 May 2015. [Google Scholar]
- Huang, L.; Ding, W.; Liu, W.; Zhao, J.; Huang, W.; Xu, C.; Zhang, D.; Liang, D. Identification of wheat powdery mildew using in-situ hyperspectral data and linear regression and support vector machines. J Plant Pathol 2019, 101, 1035–1045. [Google Scholar] [CrossRef]
- Nagasubramanian, K.; Jones, S.; Sarkar, S.; Singh, A. K.; Singh, A.; Ganapathysubramanian, B. Hyperspectral band selection using genetic algorithm and support vector machines for early identification of charcoal rot disease in soybean stems. Plant Methods 2018, 14, 86. [Google Scholar] [CrossRef]
- Rumpf, T.; Mahlein, A.-K.; Steiner, U.; Oerke, E.-C.; Dehne, H.-W.; Plümer, L. Early detection and classification of plant diseases with Support Vector Machines based on hyperspectral reflectance. Computers and Electronics in Agriculture 2010, 74, 91–99. [Google Scholar] [CrossRef]
- Thomas, S.; Behmann, J.; Rascher, U.; Mahlein, A.-K. Evaluation of the benefits of combined reflection and transmission hyperspectral imaging data through disease detection and quantification in plant–pathogen interactions. J Plant Dis Prot 2022, 129, 505–520. [Google Scholar] [CrossRef]
- Thomas, S.; Behmann, J.; Steier, A.; Kraksa, T.; Muller, O.; Rascher, U.; Mahlein, A.-K. Quantitative assessment of disease severity and rating of barley cultivars based on hyperspectral imaging in a non-invasive, automated phenotyping platform. Plant Methods 2018, 14, 45. [Google Scholar] [CrossRef]
- Lowe, A.; Harrison, N.; French, A. P. Hyperspectral image analysis techniques for the detection and classification of the early onset of plant disease and stress. Plant Methods 2017, 13, 80. [Google Scholar] [CrossRef]
- Bauriegel, E.; Herppich, W. B. Hyperspectral and Chlorophyll Fluorescence Imaging for Early Detection of Plant Diseases, with Special Reference to Fusarium spec. Infections on Wheat. Agriculture 2014, 4, 32–57. [Google Scholar] [CrossRef]
- Khan, I. H.; Liu, H.; Li, W.; Cao, A.; Wang, X.; Liu, H.; Cheng, T.; Tian, Y.; Zhu, Y.; Cao, W.; Yao, X. Early Detection of Powdery Mildew Disease and Accurate Quantification of Its Severity Using Hyperspectral Images in Wheat. Remote Sens. 2021, 13, 3612. [Google Scholar] [CrossRef]
- Wang, D.; Vinson, R.; Holmes, M.; Seibel, G.; Bechar, A.; Nof, S.; Tao, Y. Early Detection of Tomato Spotted Wilt Virus by Hyperspectral Imaging and Outlier Removal Auxiliary Classifier Generative Adversarial Nets (OR-AC-GAN). Sci Rep 2019, 9, 4377. [Google Scholar] [CrossRef]
- Bohnenkamp, D.; Kuska, M.T.; Mahlein, A.-K.; Behmann, J. Hyperspectral signal decomposition and symptom detection of wheat rust disease at the leaf scale using pure fungal spore spectra as reference. Plant Pathol 2019, 68, 1188–1195. [Google Scholar] [CrossRef]



| Method | % of annotated data used as training data | Accuracy (%) |
|---|---|---|
| k-Nearest Neighbor | 10 | 86.80 |
| k-Nearest Neighbor | 30 | 87.21 |
| k-Nearest Neighbor (fine-tuned) | 10 | 99.50 |
| k-Nearest Neighbor (fine-tuned) | 30 | 99.58 |
| Support Vector Machine | 10 | 61.94 |
| Support Vector Machine | 30 | 78.73 |
| Support Vector Machine (fine-tuned) | 10 | 99.69 |
| Support Vector Machine (fine-tuned) | 30 | 99.69 |
| Fully connected network | 10 | 99.94 |
| DeepHS_net | 10 | 99.99 |
| DeepHS_net + HyveConv++ | 10 | 99.99 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




