Submitted:
24 September 2024
Posted:
25 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
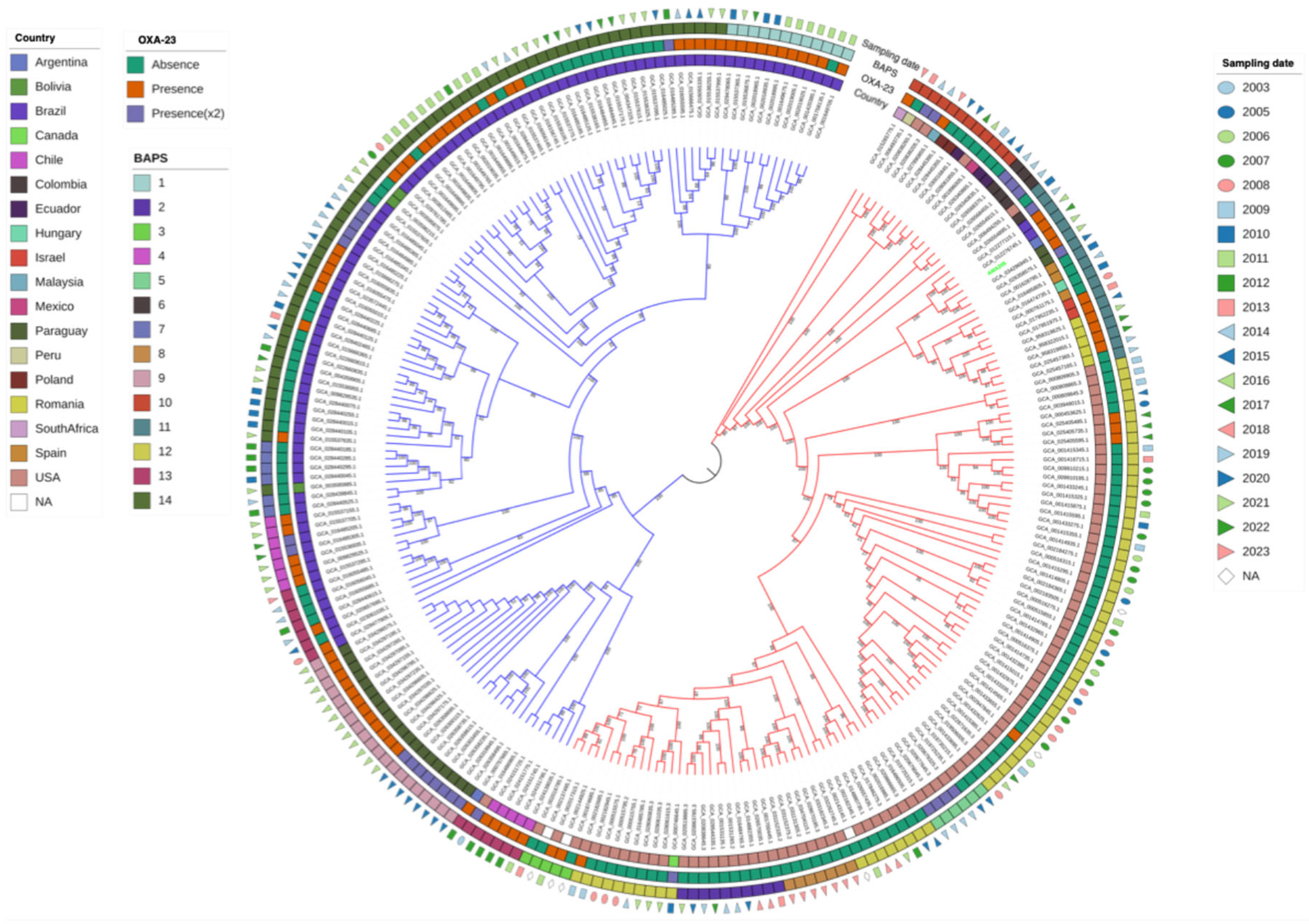
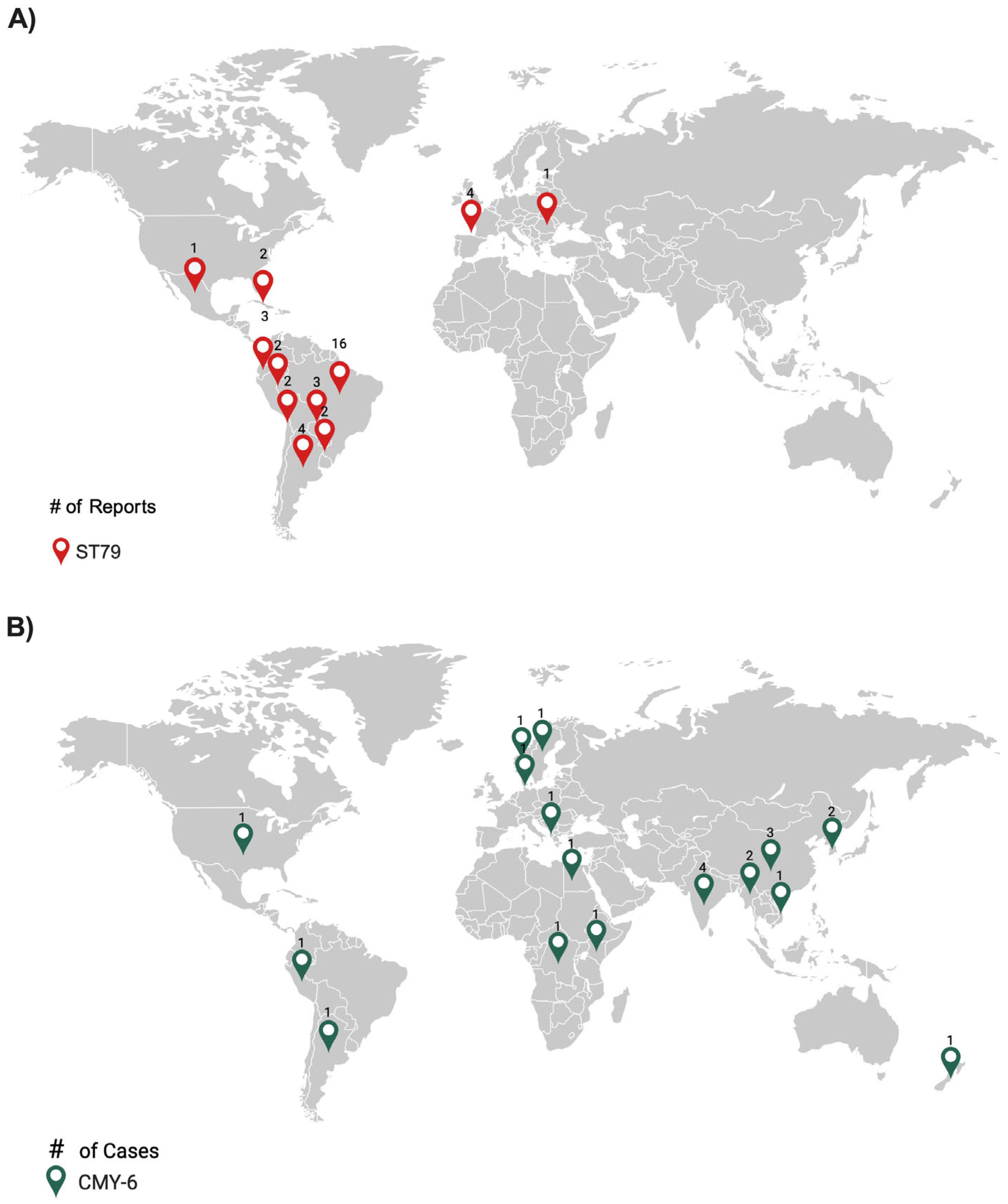
2.1. Genomic and Phylogenomic Comparative Analysis of A. baumannii AMA205 Reveals a Distinct Location of ST79
2.2. AMA205 Exhibits Resistance to Almost All Tested Antibiotics, Including Cefiderocol
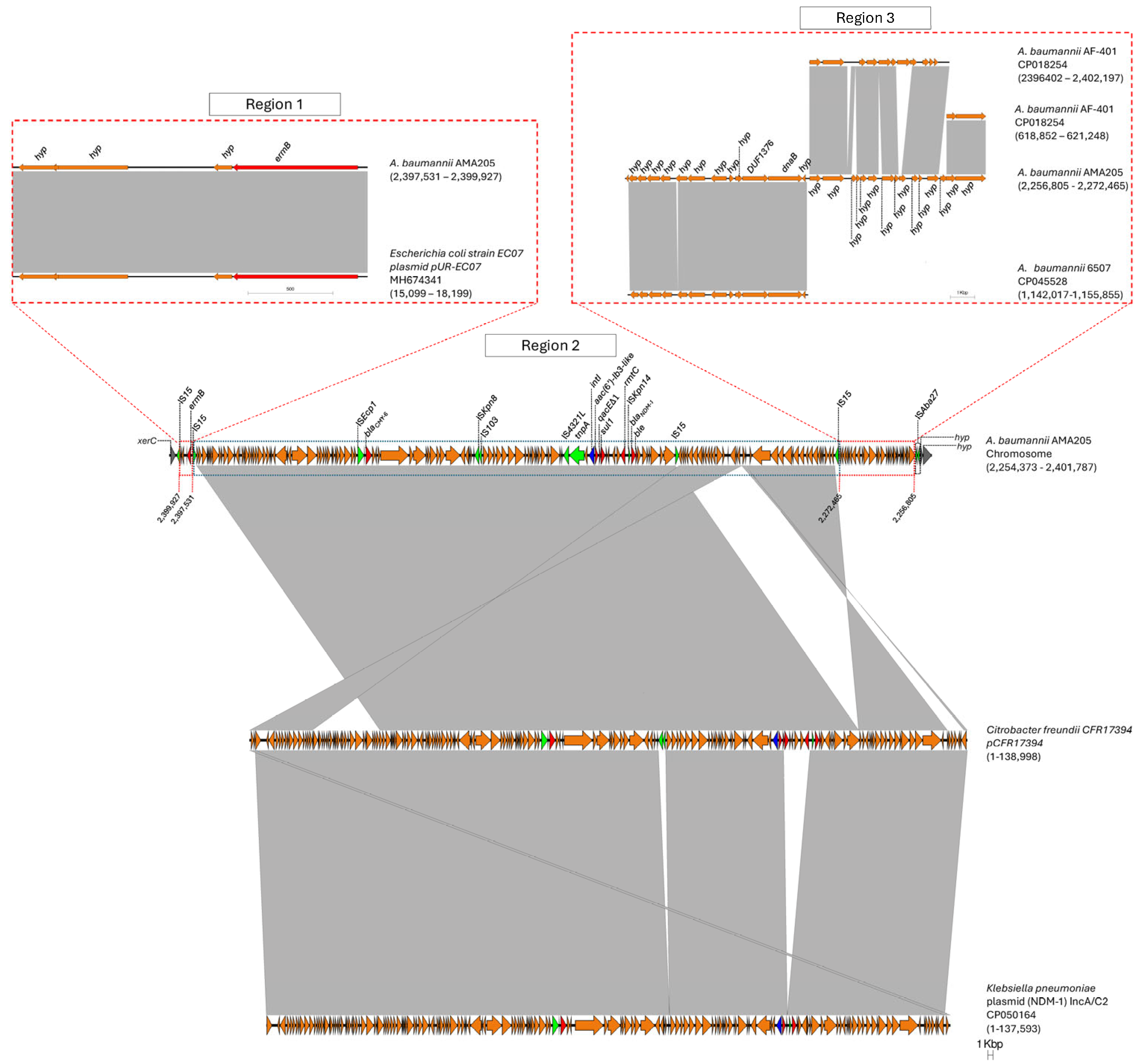
2.3. Genomic Studies Reveal the Presence of CMY-6 and Other Antimicrobial Resistance Genes in the AMA205 Genome
2.4. AMA205 Genomic Analysis Revealed the Presence of a Variety of Virulence Factors
3. Materials and Methods
3.1. Bacterial isolates
3.2. Whole Genomic Sequencing (WGS)
3.3. Comparative Genomic Analysis
3.4. Antibiotic Susceptibility Testing (AST)
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Henley, S.J.; King, J.B.; German, R.R.; Richardson, L.C.; Plescia, M. Centers for Disease Control & Prevention (CDC). 2010.
- Roca, I.; Espinal, P.; Vila-Farrés, X.; Vila, J. The Acinetobacter baumannii Oxymoron: Commensal Hospital Dweller Turned Pan-Drug-Resistant Menace. Front. Microbiol. 2012, 3. [CrossRef]
- Perez, F.; Stiefel, U. The Impact of Natural Transformation on the Acquisition of Antibiotic Resistance Determinants. MBio 2022, 13. [CrossRef]
- Antunes, L.C.S.; Imperi, F.; Carattoli, A.; Visca, P. Deciphering the Multifactorial Nature of Acinetobacter baumannii Pathogenicity. PLoS One 2011, 6, e22674. [CrossRef]
- Nasr, P. Genetics, Epidemiology, and Clinical Manifestations of Multidrug-Resistant Acinetobacter baumannii. J. Hosp. Infect. 2020, 104, 4–11. [CrossRef]
- Godeux, A.-S.; Svedholm, E.; Barreto, S.; Potron, A.; Venner, S.; Charpentier, X.; Laaberki, M.-H. Interbacterial Transfer of Carbapenem Resistance and Large Antibiotic Resistance Islands by Natural Transformation in Pathogenic Acinetobacter. MBio 2022, 13. [CrossRef]
- Traglia, G.M.; Place, K.; Dotto, C.; Fernandez, J.S.; Montaña, S.; Bahiense, C. dos S.; Soler-Bistue, A.; Iriarte, A.; Perez, F.; Tolmasky, M.E.; et al. Interspecies DNA Acquisition by a Naturally Competent Acinetobacter baumannii Strain. Int. J. Antimicrob. Agents 2019, 53, 483–490. [CrossRef]
- Traglia, G.M.; Pasteran, F.; Escalante, J.; Nishimura, B.; Tuttobene, M.R.; Subils, T.; Nuñez, M.R.; Rivollier, M.G.; Corso, A.; Tolmasky, M.E.; et al. Genomic Comparative Analysis of Two Multi-Drug Resistance (MDR) Acinetobacter Baumannii Clinical Strains Assigned to International Clonal Lineage II Recovered Pre- and Post-COVID-19 Pandemic. Biology (Basel). 2023, 12, 358. [CrossRef]
- López, C.; Ayala, J.A.; Bonomo, R.A.; González, L.J.; Vila, A.J. Protein Determinants of Dissemination and Host Specificity of Metallo-β-Lactamases. Nat. Commun. 2019, 10, 3617. [CrossRef]
- Bauernfeind, A.; Stemplinger, I.; Jungwirth, R.; Wilhelm, R.; Chong, Y. Comparative Characterization of the Cephamycinase BlaCMY-1 Gene and Its Relationship with Other Beta-Lactamase Genes. Antimicrob. Agents Chemother. 1996, 40, 1926–1930. [CrossRef]
- Lee, K.; Lee, M.; Shin, J.H.; Lee, M.H.; Kang, S.H.; Park, A.J.; Yong, D.; Chong, Y. Prevalence of Plasmid-Mediated AmpC β -Lactamases in Escherichia coli and Klebsiella Pneumoniae in Korea. Microb. Drug Resist. 2006, 12, 44–49. [CrossRef]
- Sekar, R.; Mahalakshmi, D.; Srivani, R.; Shankar, E.M.; Vignesh, R. High Rate of Detection of High-Level Aminoglycoside-Resistant Enterococci from Urinary Tract Specimens in South India. Int. J. Antimicrob. Agents 2008, 31, 383–385. [CrossRef]
- Domingues, S.; Harms, K.; Fricke, W.F.; Johnsen, P.J.; da Silva, G.J.; Nielsen, K.M. Natural Transformation Facilitates Transfer of Transposons, Integrons and Gene Cassettes between Bacterial Species. PLoS Pathog. 2012, 8, e1002837. [CrossRef]
- Da Silva, G.; Domingues, S. Insights on the Horizontal Gene Transfer of Carbapenemase Determinants in the Opportunistic Pathogen Acinetobacter baumannii. Microorganisms 2016, 4, 29. [CrossRef]
- Héritier, C.; Poirel, L.; Fournier, P.-E.; Claverie, J.-M.; Raoult, D.; Nordmann, P. Characterization of the Naturally Occurring Oxacillinase of Acinetobacter baumannii. Antimicrob. Agents Chemother. 2005, 49, 4174–4179. [CrossRef]
- Figueiredo, S.; Poirel, L.; Croize, J.; Recule, C.; Nordmann, P. In Vivo Selection of Reduced Susceptibility to Carbapenems in Acinetobacter baumannii Related to IS Aba1 -Mediated Overexpression of the Natural Bla OXA-66 Oxacillinase Gene. Antimicrob. Agents Chemother. 2009, 53, 2657–2659. [CrossRef]
- Nigro, S.; Hall, R.M. Distribution of the Bla OXA-23-Containing Transposons Tn 2006 and Tn 2008 in Australian Carbapenem-Resistant Acinetobacter baumannii Isolates. J. Antimicrob. Chemother. 2015, 70, 2409–2411. [CrossRef]
- Yoon, E.-J.; Kim, J.O.; Yang, J.W.; Kim, H.S.; Lee, K.J.; Jeong, S.H.; Lee, H.; Lee, K. The BlaOXA-23-Associated Transposons in the Genome of Acinetobacter Spp. Represent an Epidemiological Situation of the Species Encountering Carbapenems. J. Antimicrob. Chemother. 2017, 72, 2708–2714. [CrossRef]
- Traglia, G.; Chiem, K.; Quinn, B.; Fernandez, J.S.; Montaña, S.; Almuzara, M.; Mussi, M.A.; Tolmasky, M.E.; Iriarte, A.; Centrón, D.; et al. Genome Sequence Analysis of an Extensively Drug-Resistant Acinetobacter baumannii Indigo-Pigmented Strain Depicts Evidence of Increase Genome Plasticity. Sci. Rep. 2018, 8, 16961. [CrossRef]
- Martino, F.; Tijet, N.; Melano, R.; Petroni, A.; Heinz, E.; De Belder, D.; Faccone, D.; Rapoport, M.; Biondi, E.; Rodrigo, V.; et al. Isolation of Five Enterobacteriaceae Species Harbouring BlaNDM-1 and mcr-1 Plasmids from a Single Paediatric Patient. PLoS One 2019, 14, e0221960. [CrossRef]
- Malik, S.; Kaminski, M.; Landman, D.; Quale, J. Cefiderocol Resistance in Acinetobacter baumannii: Roles of β-Lactamases, Siderophore Receptors, and Penicillin Binding Protein 3. Antimicrob. Agents Chemother. 2020, 64. [CrossRef]
- Yamano, Y.; Ishibashi, N.; Kuroiwa, M.; Takemura, M.; Sheng, W.-H.; Hsueh, P.-R. Characterisation of Cefiderocol-Non-Susceptible Acinetobacter baumannii Isolates from Taiwan. J. Glob. Antimicrob. Resist. 2022, 28, 120–124. [CrossRef]
- Lucidi, M.; Visaggio, D.; Migliaccio, A.; Capecchi, G.; Visca, P.; Imperi, F.; Zarrilli, R. Pathogenicity and Virulence of Acinetobacter baumannii : Factors Contributing to the Fitness in Healthcare Settings and the Infected Host. Virulence 2024, 15. [CrossRef]
- Ramirez, M.S.; Penwell, W.F.; Traglia, G.M.; Zimbler, D.L.; Gaddy, J.A.; Nikolaidis, N.; Arivett, B.A.; Adams, M.D.; Bonomo, R.A.; Actis, L.A.; et al. Identification of Potential Virulence Factors in the Model Strain Acinetobacter baumannii A118. Front. Microbiol. 2019, 10. [CrossRef]
- Tomaras, A.P.; Dorsey, C.W.; Edelmann, R.E.; Actis, L.A. Attachment to and Biofilm Formation on Abiotic Surfaces by Acinetobacter baumannii: Involvement of a Novel Chaperone-Usher Pili Assembly System. Microbiology 2003, 149, 3473–3484. [CrossRef]
- Gaddy, J.A.; Actis, L.A. Regulation of Acinetobacter baumannii Biofilm Formation. Future Microbiol. 2009, 4, 273–278. [CrossRef]
- Kishii, K.; Hamada, M.; Aoki, K.; Ito, K.; Onodera, J.; Ishii, Y.; Tateda, K. Differences in Biofilm Formation and Transcription of Biofilm-Associated Genes among Acinetobacter baumannii Clinical Strains Belonging to the International Clone II Lineage. J. Infect. Chemother. 2020, 26, 693–698. [CrossRef]
- Hamidian, M.; Wick, R.R.; Hartstein, R.M.; Judd, L.M.; Holt, K.E.; Hall, R.M. Insights from the Revised Complete Genome Sequences of Acinetobacter baumannii Strains AB307-0294 and ACICU Belonging to Global Clones 1 and 2. Microbiol. Hamidian, RR Wick, RM Harts. LM Judd, KE Holt, RM HallMicrobial genomics, 2019•microbiologyresearch.org 2019, 5. [CrossRef]
- Fattahian, Y.; Rasooli, I.; Gargari, S.; … M.R.-M.; 2011, undefined Protection against Acinetobacter baumannii Infection via Its Functional Deprivation of Biofilm Associated Protein (Bap). Elsevier.
- Crosa, J.H. Genetics and Molecular Biology of Siderophore-Mediated Iron Transport in Bacteria. Microbiol. Rev. 1989, 53, 517–530. [CrossRef]
- Zimbler, D.L.; Penwell, W.F.; Gaddy, J.A.; Menke, S.M.; Tomaras, A.P.; Connerly, P.L.; Actis, L.A. Iron Acquisition Functions Expressed by the Human Pathogen Acinetobacter baumannii. BioMetals 2009, 22, 23–32. [CrossRef]
- Klebba, P.E.; Newton, S.M.C.; Six, D.A.; Kumar, A.; Yang, T.; Nairn, B.L.; Munger, C.; Chakravorty, S. Iron Acquisition Systems of Gram-Negative Bacterial Pathogens Define TonB-Dependent Pathways to Novel Antibiotics. Chem. Rev. 2021, 121, 5193–5239. [CrossRef]
- Jia, B.; Raphenya, A.R.; Alcock, B.; Waglechner, N.; Guo, P.; Tsang, K.K.; Lago, B.A.; Dave, B.M.; Pereira, S.; Sharma, A.N.; et al. CARD 2017: Expansion and Model-Centric Curation of the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2017, 45, D566–D573. [CrossRef]
- Penwell, W.F.; DeGrace, N.; Tentarelli, S.; Gauthier, L.; Gilbert, C.M.; Arivett, B.A.; Miller, A.A.; Durand-Reville, T.F.; Joubran, C.; Actis, L.A. Discovery and Characterization of New Hydroxamate Siderophores, Baumannoferrin A and B, Produced by Acinetobacter baumannii. ChemBioChem 2015, 16, 1896–1904. [CrossRef]
- Sheldon, J.R.; Skaar, E.P. Acinetobacter Baumannii Can Use Multiple Siderophores for Iron Acquisition, but Only Acinetobactin Is Required for Virulence. PLOS Pathog. 2020, 16, e1008995. [CrossRef]
- Kenyon, J.J.; Hall, R.M. Variation in the Complex Carbohydrate Biosynthesis Loci of Acinetobacter baumannii Genomes. PLoS One 2013, 8, e62160. [CrossRef]
- Tickner, J.; Hawas, S.; Totsika, M.; Kenyon, J.J. The Wzi Outer Membrane Protein Mediates Assembly of a Tight Capsular Polysaccharide Layer on the Acinetobacter baumannii Cell Surface. Sci. Rep. 2021, 11, 21741. [CrossRef]
- Seemann, T. Prokka: Rapid Prokaryotic Genome Annotation. Bioinformatics 2014, 30, 2068–2069. [CrossRef]
- Richter, M.; Rosselló-Móra, R.; Oliver Glöckner, F.; Peplies, J. JSpeciesWS: A Web Server for Prokaryotic Species Circumscription Based on Pairwise Genome Comparison. Bioinformatics 2016, 32, 929–931. [CrossRef]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [CrossRef]
- Tonkin-Hill, G.; Lees, J.A.; Bentley, S.D.; Frost, S.D.W.; Corander, J. Fast Hierarchical Bayesian Analysis of Population Structure. Nucleic Acids Res. 2019, 47, 5539–5549. [CrossRef]
- Lowe, T.M.; Eddy, S.R. TRNAscan-SE: A Program for Improved Detection of Transfer RNA Genes in Genomic Sequence. Nucleic Acids Res. 1997, 25, 955–964. [CrossRef]
- Liu, B.; Zheng, D.; Jin, Q.; Chen, L.; Yang, J. VFDB 2019: A Comparative Pathogenomic Platform with an Interactive Web Interface. Nucleic Acids Res. 2019, 47, D687–D692. [CrossRef]
- Gupta, S.K.; Padmanabhan, B.R.; Diene, S.M.; Lopez-Rojas, R.; Kempf, M.; Landraud, L.; Rolain, J.-M. ARG-ANNOT, a New Bioinformatic Tool To Discover Antibiotic Resistance Genes in Bacterial Genomes. Antimicrob. Agents Chemother. 2014, 58, 212–220. [CrossRef]
- Wayne, A. Clinical and Laboratory Standards Institute; CLSI. 2011. Perform. Stand. Antimicrob. susceptibility testing. 20th Informational Suppl. CLSI Doc. 2017.
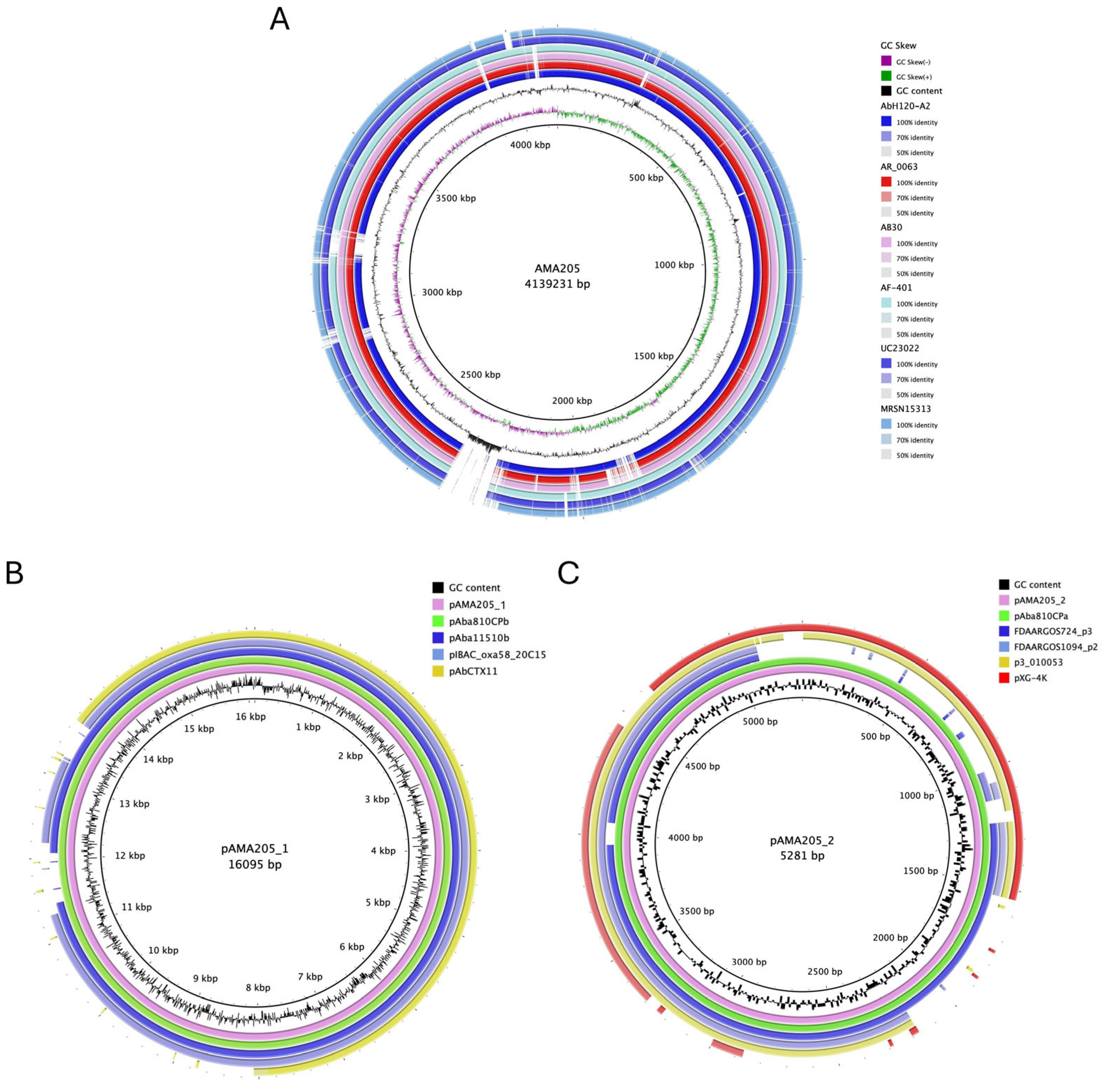
| AMA205 Chromosome | pAMA205_1Plasmid | pAMA205_2Plasmid | |
|---|---|---|---|
| Size (bp) | 4,139,231 | 16,095 | 5281 |
| G + C content (%) | 50.9 | 35.32 | 36.37 |
| Protein-coding sequences (CDSs) | 4007 | 19 | 7 |
| Insertion sequences | 71 | 0 | 0 |
| rRNA operons | 18 | 0 | 0 |
| tRNA | 73 | 0 | 0 |
| Antimicrobial agent | MIC (mg/L) |
|---|---|
| CAZ | > 256 |
| FDC | 4* (IHC) |
| IMP | 96 |
| MEM | 128 |
| AMS | > 256 |
| AK | > 256 |
| CN | > 1024 |
| TIG | 0.50 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




