Submitted:
24 September 2024
Posted:
24 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Homology Search and Window Analyses
2.2. Phylogenetic Tree Analysis
3. Discussion
4. Materials and Methods
4.1. Homology Search and Window Analyses
4.2. Phylogenetic Tree Reconstruction
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- N. Dalbeth, T. R. Merriman, L. K. Stamp, Gout. Lancet 388, 2039-2052 (2016).
- T. R. Merriman, An update on the genetic architecture of hyperuricemia and gout. Arthritis Res Ther 17, 98 (2015). [CrossRef]
- T. J. Major, N. Dalbeth, E. A. Stahl, T. R. Merriman, An update on the genetics of hyperuricaemia and gout. Nat Rev Rheumatol 14, 341-353 (2018). [CrossRef]
- N. Otani, M. Ouchi, K. Misawa, I. Hisatome, N. Anzai, Hypouricemia and Urate Transporters. Biomedicines 10, (2022). [CrossRef]
- I. Mihaljevic, M. Popovic, R. Zaja, T. Smital, Phylogenetic, syntenic, and tissue expression analysis of slc22 genes in zebrafish (Danio rerio). BMC Genomics 17, 626 (2016). [CrossRef]
- A. Enomoto et al., Molecular identification of a renal urate anion exchanger that regulates blood urate levels. Nature 417, 447-452 (2002). [CrossRef]
- K. Ichida et al., Clinical and molecular analysis of patients with renal hypouricemia in Japan-influence of URAT1 gene on urinary urate excretion. J Am Soc Nephrol 15, 164-173 (2004). [CrossRef]
- N. Iwai et al., A high prevalence of renal hypouricemia caused by inactive SLC22A12 in Japanese. Kidney Int 66, 935-944 (2004). [CrossRef]
- A. Mancikova et al., Functional analysis of novel allelic variants in URAT1 and GLUT9 causing renal hypouricemia type 1 and 2. Clin Exp Nephrol 20, 578-584 (2016). [CrossRef]
- K. Misawa et al., Contribution of rare variants of the SLC22A12 gene to the missing heritability of serum urate levels. Genetics 214, 1079-1090 (2020). [CrossRef]
- D. Oliveira et al., Identification of a novel nucleobase-ascorbate transporter family member in fish and amphibians. Fishes 4, 1 (2019). [CrossRef]
- J. Morales et al., A joint NCBI and EMBL-EBI transcript set for clinical genomics and research. Nature 604, 310-315 (2022). [CrossRef]
- G. Stecher, K. Tamura, S. Kumar, Molecular Evolutionary Genetics Analysis (MEGA) for macOS. Mol Biol Evol 37, 1237-1239 (2020). [CrossRef]
- K. Tamura, G. Stecher, S. Kumar, MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 38, 3022-3027 (2021). [CrossRef]
- J. Felsenstein, Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783-791 (1985). [CrossRef]
- K. Tamura, M. Nei, Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10, 512-526 (1993). [CrossRef]
- N. Saitou, M. Nei, The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4, 406-425 (1987). [CrossRef]
- O. Gascuel, BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol 14, 685-695 (1997). [CrossRef]
- M. Nei, S. Kumar, Molecular Evolution and Phylogenetics. (Oxford University Press, New York, 2000).
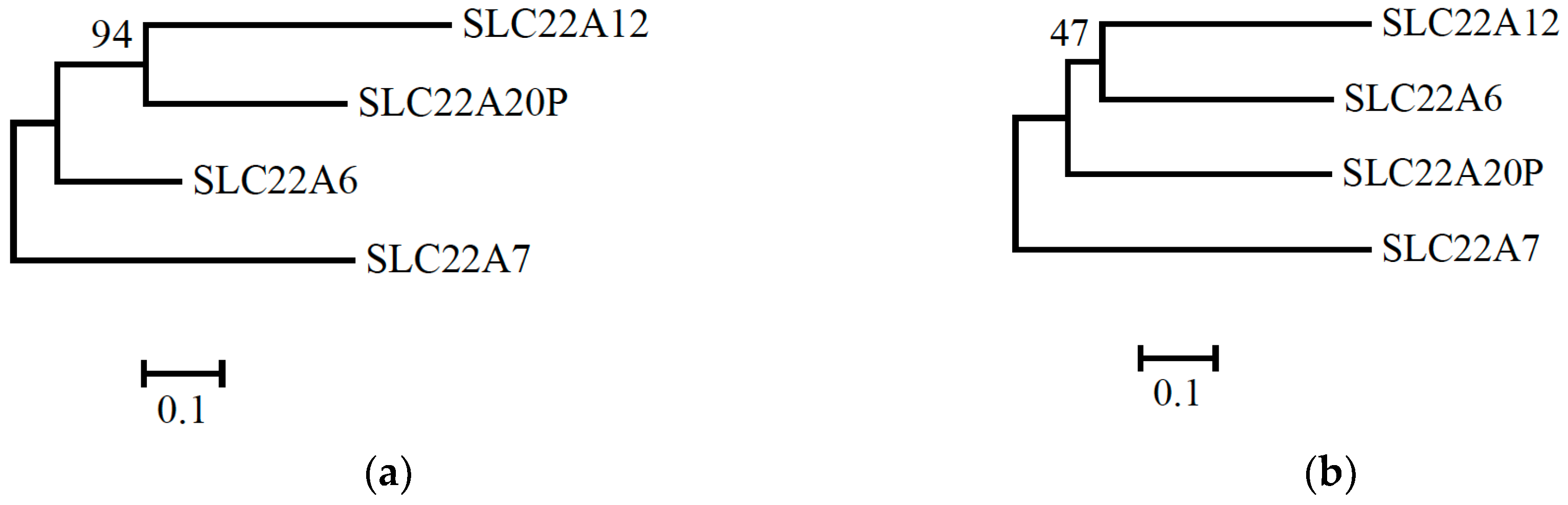
| SLC22A12 | SLC22A20P | SLC22A6 | SLC22A7 | |
| SLC22A12 | 0.387 | 0.398 | 0.497 | |
| SLC22A20P | 0.438 | 0.345 | 0.441 | |
| SLC22A6 | 0.413 | 0.424 | 0.393 | |
| SLC22A7 | 0.484 | 0.488 | 0.484 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




