Submitted:
04 September 2024
Posted:
05 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
- Type of study
- Samples
- RNA extraction
- qRT-PCR
- RT-LAMP
- RT-LAMP to detect DENV in samples without prior extraction
- Data analysis
3. Results
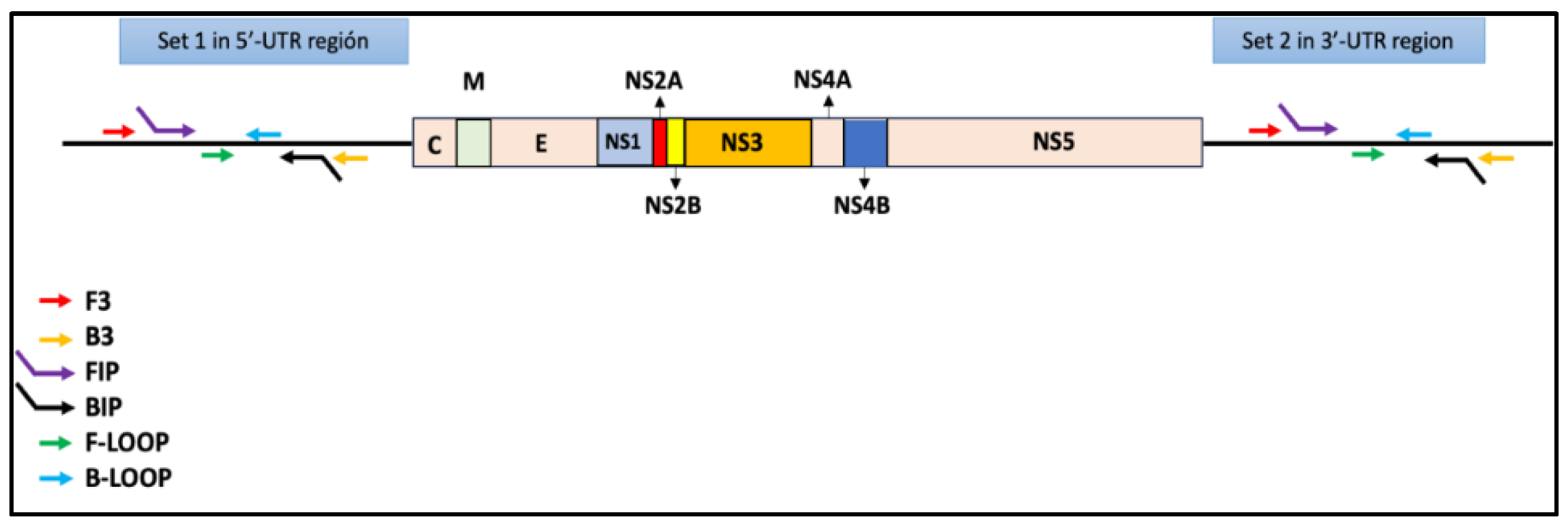
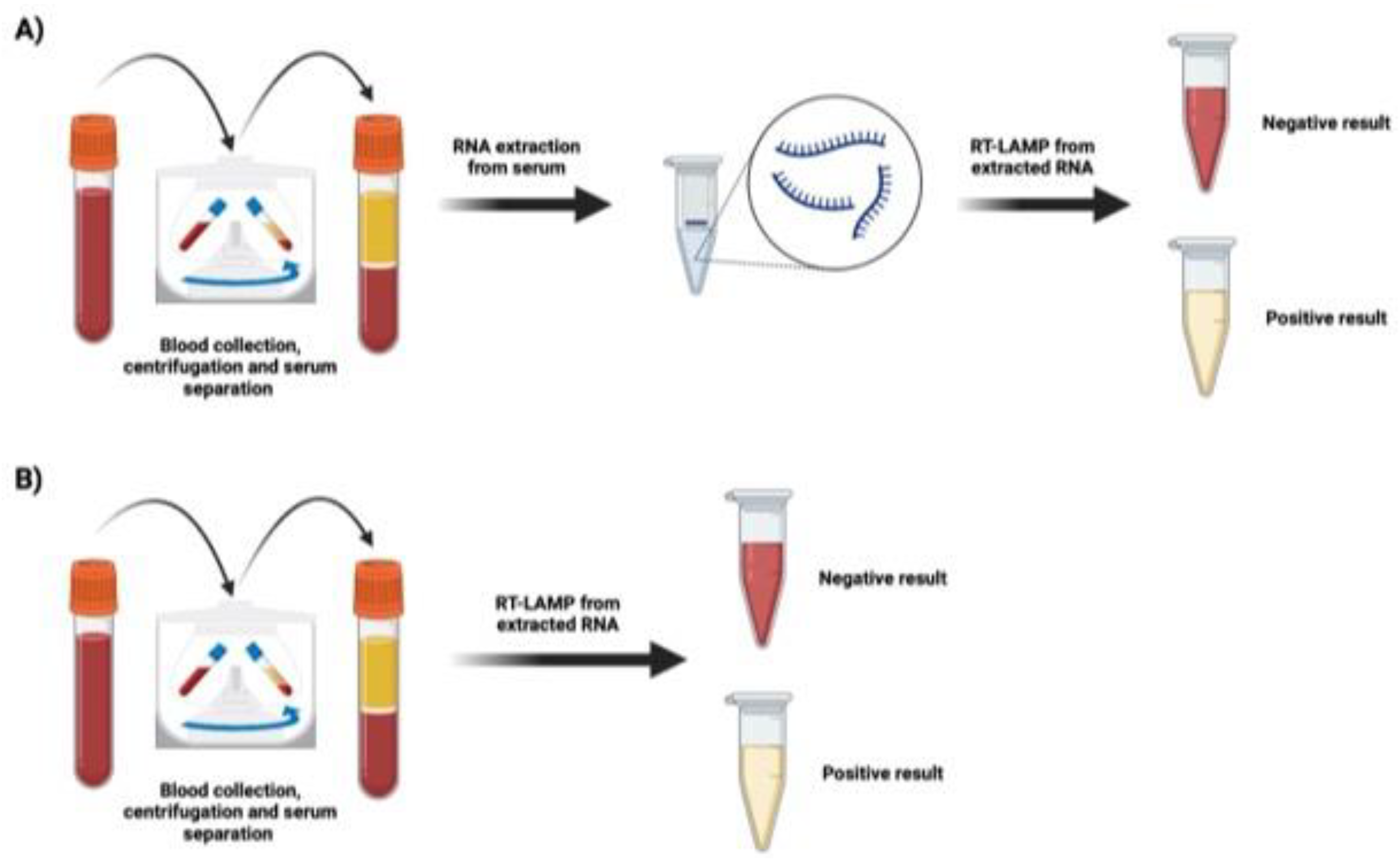
- VPP= A/(A+B)
- VPP= 45 / (45+4) =0.91.
- NPV= D/(C+D)
- NPV= 107/ (2+107) =0.98.
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- CDC, «Clinical Testing Guidance for Dengue», Dengue. Accedido: 18 de agosto de 2024. [En línea]. Disponible en: https://www.cdc.gov/dengue/hcp/diagnosis-testing/index.html.
- I. Gjenero-Margan et al., «Autochthonous dengue fever in Croatia, August-September 2010», Euro Surveill. Bull. Eur. Sur Mal. Transm. Eur. Commun. Dis. Bull., vol. 16, n.o 9, p. 19805, mar. 2011.
- S. D. Sekaran y H. Artsob, «Molecular diagnostics for the detection of human flavivirus infections», Expert Opin. Med. Diagn., vol. 1, n.o 4, pp. 521-530, dic. 2007. [CrossRef]
- B. L. Ligon, «Dengue fever and dengue hemorrhagic fever: A review of the history, transmission, treatment, and prevention», Semin. Pediatr. Infect. Dis., vol. 16, n.o 1, pp. 60-65, ene. 2005. [CrossRef]
- M. G. Guzman et al., «Dengue: a continuing global threat», Nat. Rev. Microbiol., vol. 8, n.o S12, pp. S7-S16, dic. 2010. [CrossRef]
- C. Paupy, H. Delatte, L. Bagny, V. Corbel, y D. Fontenille, «Aedes albopictus, an arbovirus vector: From the darkness to the light», Microbes Infect., vol. 11, n.o 14-15, pp. 1177-1185, dic. 2009. [CrossRef]
- R. E. Whitmire, D. S. Burke, A. Nisalak, B. A. Harrison, y D. M. Watts, «Effect of Temperature on the Vector Efficiency of Aedes aegypti for Dengue 2 Virus», Am. J. Trop. Med. Hyg., vol. 36, n.o 1, pp. 143-152, ene. 1987. [CrossRef]
- T. Notomi, «Loop-mediated isothermal amplification of DNA», Nucleic Acids Res., vol. 28, n.o 12, pp. 63e-663, jun. 2000. [CrossRef]
- P. C. G. Nunes, M. R. Q. Lima, y F. B. Dos Santos, «Molecular Diagnosis of Dengue», en Dengue Virus, vol. 2409, R. Mohana-Borges, Ed., en Methods in Molecular Biology, vol. 2409. , New York, NY: Springer US, 2022, pp. 157-171. [CrossRef]
- A. L. Dauner et al., «Development of a pan-serotype reverse transcription loop-mediated isothermal amplification assay for the detection of dengue virus», Diagn. Microbiol. Infect. Dis., vol. 83, n.o 1, pp. 30-36, sep. 2015. [CrossRef]
- B.-T. Teoh et al., «Detection of dengue viruses using reverse transcription-loop-mediated isothermal amplification», BMC Infect. Dis., vol. 13, n.o 1, p. 387, dic. 2013. [CrossRef]
- L. Hurtado et al., «Validación clínica de la prueba RT-LAMP para el diagnóstico rápido del SARS-CoV-2», Biomédica, vol. 42, pp. 59-72, oct. 2022. [CrossRef]
- K. J. Land, D. I. Boeras, X.-S. Chen, A. R. Ramsay, y R. W. Peeling, «REASSURED diagnostics to inform disease control strategies, strengthen health systems and improve patient outcomes», Nat. Microbiol., vol. 4, n.o 1, pp. 46-54, dic. 2018. [CrossRef]
- S. Hu et al., «Development of reverse-transcription loop-mediated isothermal amplification assay for rapid detection and differentiation of dengue virus serotypes 1–4», BMC Microbiol., vol. 15, p. 265, nov. 2015. [CrossRef]
- Y.-L. Lau et al., «Colorimetric Detection of Dengue by Single Tube Reverse-Transcription-Loop-Mediated Isothermal Amplification», PLOS ONE, vol. 10, n.o 9, p. e0138694, sep. 2015. [CrossRef]
- S. Li et al., «Simultaneous detection and differentiation of dengue virus serotypes 1-4, Japanese encephalitis virus, and West Nile virus by a combined reverse-transcription loop-mediated isothermal amplification assay», Virol. J., vol. 8, n.o 1, p. 360, dic. 2011. [CrossRef]
- B. Lopez-Jimena et al., «Development and validation of four one-step real-time RT-LAMP assays for specific detection of each dengue virus serotype», PLoS Negl. Trop. Dis., vol. 12, n.o 5, p. e0006381, may 2018. [CrossRef]
- M. Parida et al., «Rapid Detection and Differentiation of Dengue Virus Serotypes by a Real-Time Reverse Transcription-Loop-Mediated Isothermal Amplification Assay», J. Clin. Microbiol., vol. 43, n.o 6, pp. 2895-2903, jun. 2005. [CrossRef]
- D. W. Vaughn et al., «Dengue Viremia Titer, Antibody Response Pattern, and Virus Serotype Correlate with Disease Severity», J. Infect. Dis., vol. 181, n.o 1, pp. 2-9, ene. 2000. [CrossRef]
- J. Yamagishi et al., «Serotyping dengue virus with isothermal amplification and a portable sequencer», Sci. Rep., vol. 7, n.o 1, p. 3510, jun. 2017. [CrossRef]
- P. Matangkasombut et al., «Dengue viremia kinetics in asymptomatic and symptomatic infection», Int. J. Infect. Dis., vol. 101, pp. 90-97, dic. 2020. [CrossRef]
- Y. Zhou et al., «A Mismatch-Tolerant Reverse Transcription Loop-Mediated Isothermal Amplification Method and Its Application on Simultaneous Detection of All Four Serotype of Dengue Viruses», Front. Microbiol., vol. 10, p. 1056, may 2019. [CrossRef]
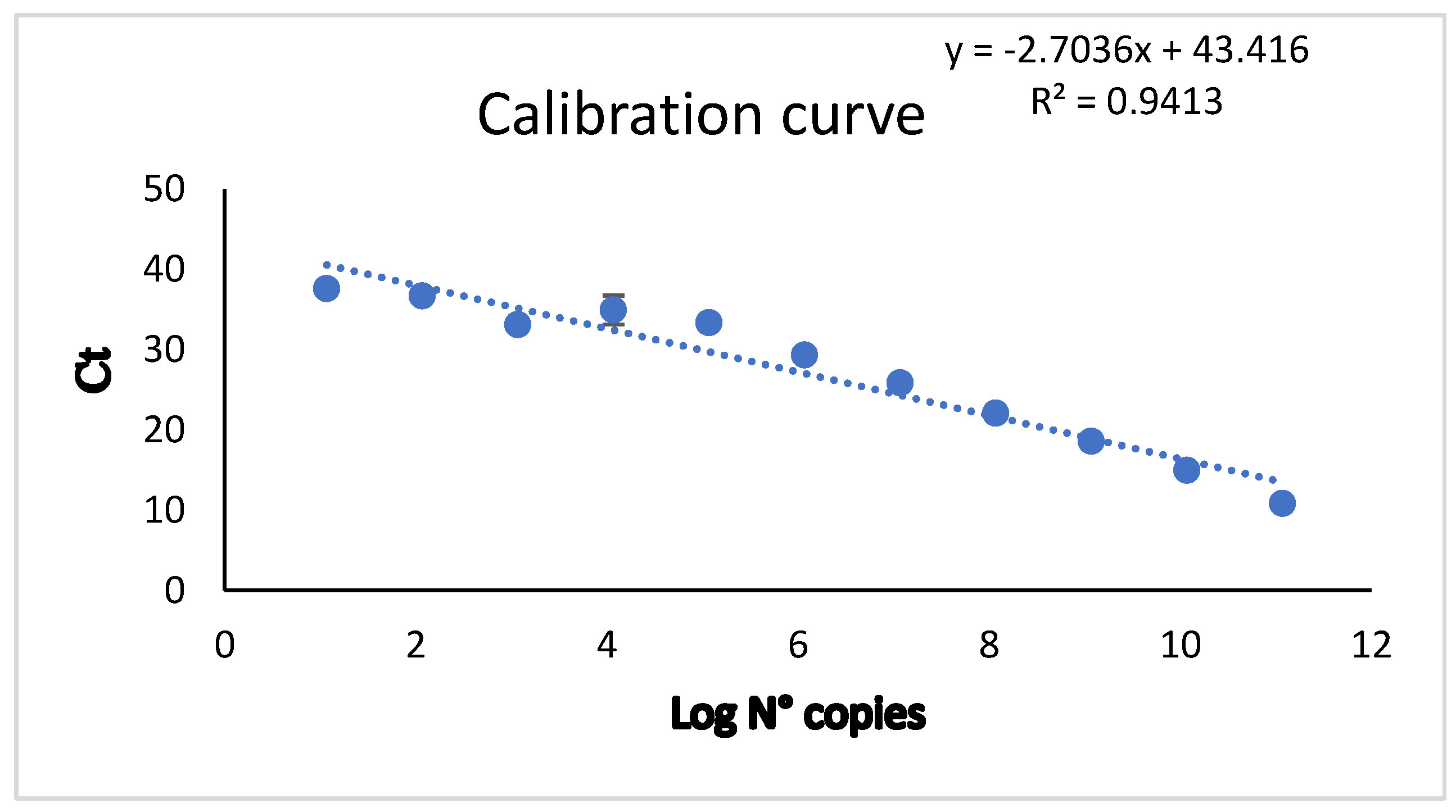
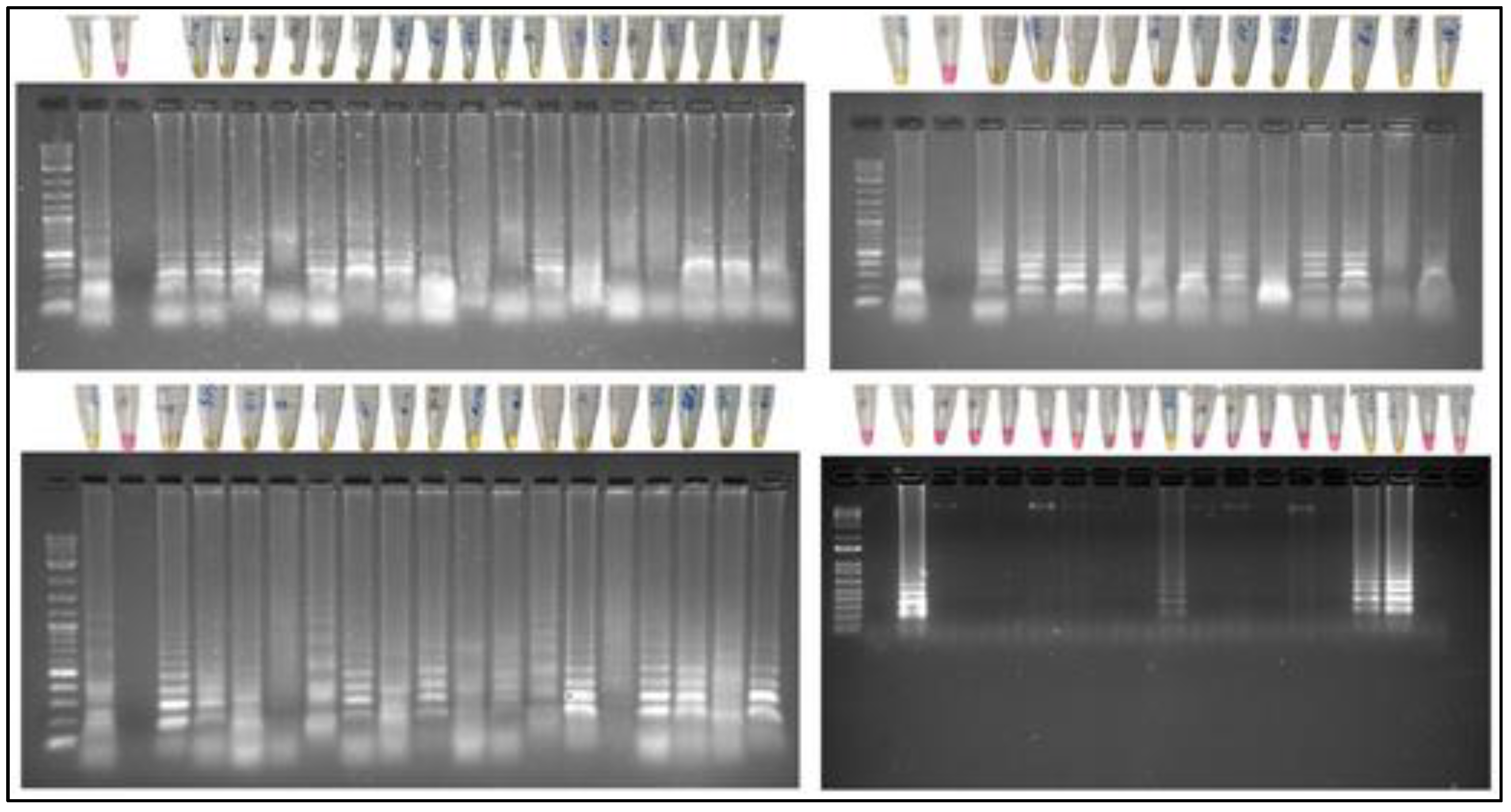
| Primers Set | Oligos name | Primers Sequence |
|---|---|---|
|
SET 1 [11] |
FIP-DENV1/3 | 5'-GGTTATTCATCAGAGATCTGCTCTCTTTTTTTCGAATCGGAAGCTTCGT -3' |
| FIP-DENV2 | 5'-GGTTATTCATCAGAGATCTGCTCTCTTTTATTCTTTGAGGGAGCTAAGC-3' | |
| FIP-DENV4 | 5'-TTCATTTTTCCAGAGATCTGCTCTCTTTTTTTCGAATCGGAAGCTTCGT-3 ' | |
| B1P-DENV1/3 | 5'-AACGGAAAAAGACGGGTCAACCGTTTTTCTTTGTCAGCTGTTGCACAGT-3 ' | |
| B1P-DENV2 | 5'-AACGGAAAAAGGCGAGAAA TACGCTTTTCTTTGTCAGCTGTTGCACAGT -3' | |
| B1P DENV4 | 5' AACGAAAAAAGGTGGTTAGACCACTTTTCTTCACCAACCCTTGAGGGGT-3 ' | |
| F3 | 5'-GTGGACCGACAAAGACAG -3' | |
| B3 DENV1/3 | 5'-GTGAGCAATCCTTTTGAG-3' | |
| B3 DENV2 | 5'-TGCAGCA TTCCAAGTGAG-3' | |
| B3 DENV4 | 5'-GAAAAAAGTCCGGTTGAG-3' | |
| LoopB | 5' GCGAGAGAAACCGCGTGTC·3' | |
|
SET 2 [10] |
F3/134 | 5'-CAAACCGTGCTGCCTGT -3' |
| F3/2 | 5'-TGAGTAAACTATGCAGCCTGT-3 ' | |
| B3/123 | 5'-ACCTGTTGA TTCAACAGCACC-3 ' | |
| B3/4 | 5' ACCTGTTGGATCAACAACACC· 3' | |
| FIP/123 | 5'-AGGGGTCTCCTCT AACCRCTAGTCTTTCAAACCRTGGAAGCTGTACGC-3' | |
| FIP/4 | 5' AGGGGTCTCCTCTAACCRCTAGTCTTTTTTGCCACGGAAGCTGTACGC· 3' | |
| B1P/123 | 5'-ACAGCATATTGACGCTGGGARAGACGTTCTGTGCCTGGAA TGATGCTG-3' | |
| B1P/4 | 5'-ACAGCATATTGACGCTGGGARAGACGCTCTGTGCCTGGATTGATGTTG-3 ' | |
| BLP/1234 | 5' CAGAGATCCTGCTGTCTC-3' |
| Set primers by Authors | Oligos name | 10X working solution | 1X final concentration in the LAMP reaction |
|---|---|---|---|
|
SET 1 [11] |
FIP-DENV1/3 | 16 μM | 1,6 μM |
| FIP-DENV2 | |||
| FIP-DENV4 | |||
| B1P-DENV1/3 | 8 μM | 0,8 μM | |
| B1P-DENV2 | |||
| B1P DENV4 | |||
| F3 | 2 μM | 0,2 μM | |
| B3 DENV1/3 | |||
| B3 DENV2 | |||
| B3 DENV4 | |||
| LoopB | 8 μM | 0,8 μM | |
|
SET 2 [10] |
F3/134 | 2 μM | 0,2 μM |
| F3/2 | |||
| B3/123 | |||
| B3/4 | |||
| FIP/123 | 13 μM | 1,3 μM | |
| FIP/4 | |||
| B1P/123 | |||
| B1P/4 | |||
| BLP/1234 | 8 μM | 0,8 μM |
| DENV2 | 79pb | Copy number | cq | desv cq | Log number copy |
|---|---|---|---|---|---|
| Dilution factor | ng/μl | ||||
| 1 | 10 | 117234664070 | 11 | 0,24 | 11,06 |
| 2 | 1 | 11723466407 | 15 | 0,15 | 10,06 |
| 3 | 0,1 | 1172346641 | 19 | 0,01 | 9,06 |
| 4 | 0,01 | 117234664 | 22 | 0,18 | 8,06 |
| 5 | 0,001 | 11723466 | 26 | 0,34 | 7,06 |
| 6 | 0,0001 | 1172347 | 29 | 0,27 | 6,06 |
| 7 | 0,00001 | 117235 | 33 | 0,21 | 5,06 |
| 8 | 0,000001 | 11723 | 35 | 1,80 | 4,06 |
| 9 | 0,0000001 | 1172 | 33 | 0,11 | 3,06 |
| 10 | 0,00000001 | 117 | 37 | 0,23 | 2,06 |
| 11 | 0,000000001 | 12 | 38 | 0,22 | 1,06 |
| 12 | 1E-10 | 1 | 35 | 0,17 | 0,06 |
| Table 2*2 | Positive RT-qPCR |
Negative RT-qPCR |
total |
|---|---|---|---|
| Positive LAMP | 45 | 4 | 49 |
| Negative LAMP | 2 | 107 | 109 |
| 47 | 111 | 158 |
| Kappa Values | Interpretation |
|---|---|
| <0 | No agreement |
| 0-0.19 | Poor agreement |
| 0.20-0.39 | Fair agreement |
| 0.40-0.59 | Moderate agreement |
| 0.60-0.79 | Substantial agreement |
| 0.80-1.00 | Almost perfect agreement |
| Sensitivity | Specificity | Positive Predictive Value | Negative Predictive Value | Kappa index |
Po | Pe |
|---|---|---|---|---|---|---|
| 0.96 (0.85, 0.99) |
0.96 (0.91, 0.99) |
0.92 (0.80, 0.98) |
0.98 (0.94, 1.00) |
0.91 | 0.962 | 0.576 |
| RT-LAMP | ||||||
|---|---|---|---|---|---|---|
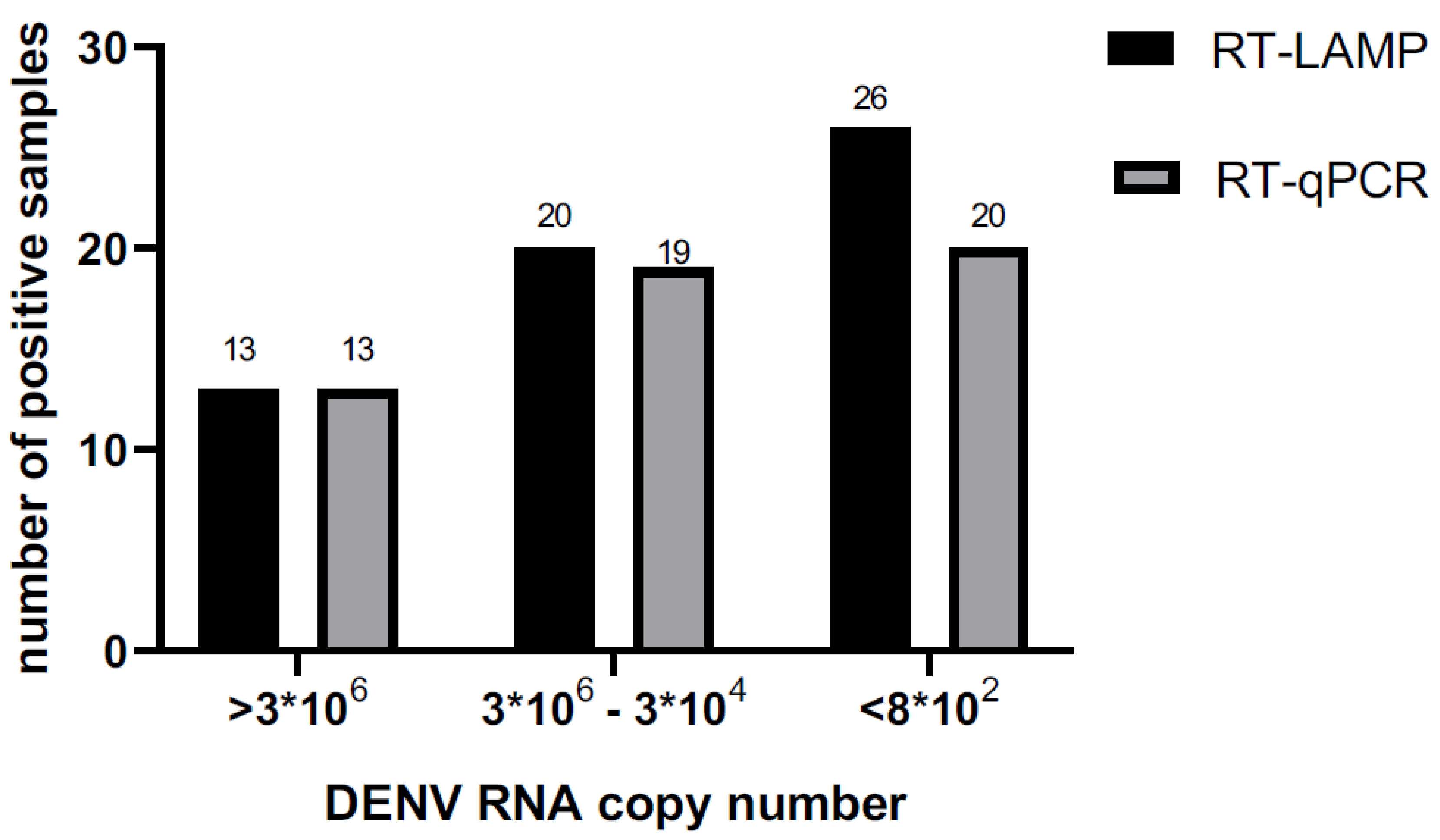
| RT-qPCR | Copies number/µl | CT | Positives | Negatives | Total | Sensitivity |
| >3*106 | 0-25 | 13 | 0 | 13 | 1 | |
| 3*106-3*104 | 25-30 | 19 | 1 | 20 | 0.95 | |
| < 8*102 | > 35 | 19 | 5 | 24 | 0.79 | |
| Total | 51 | 6 | 57 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




