1. Introduction
Our daily life is largely influenced by genetically engineered products, whether for industrial and medical purposes or for animal and human use. In fact, the food and agricultural industries are quickly integrating a variety of novel genetic technologies. In addition to traditional plant breeding methods, an increasing range of approaches, including genetic engineering and contemporary biotechniques, are becoming available for selecting and introducing desirable features in crop plants and consequently in the food products derived from them. The application of these different strategies in agriculture is a current new trend, and in particular, the possibility of precisely modifying the genome has had an extraordinary social impact since its origin in 2012 [
1]. Crop plants obtained by genome/gene editing (GE) technologies represent a notable and disruptive shift away from genetically modified or transgenic plants, outpacing public perception as well as regulatory frameworks [
2,
3]. However, one of the main goals in public and/or private contexts is to highlight possible conceptual and technical ambiguities, often due to the lack of simple definitional boundaries among different applicable approaches. During the last decade, the European Commission (EC)'s Group of Chief Scientific Advisors compared traditional breeding methods, well-established genetic modification methods, and new breeding methods, defined as new breeding techniques (NBT), and proposed the following conclusions: (i) realistically, safety assessments can only be made on a case-by-case basis and depend on the characteristics of the end product; (ii) genetically and phenotypically similar products resulting from the use of different techniques are not expected to present significant risks [
4]. In particular, regarding the application of NBTs, clear guidelines rather than border lines are increasingly necessary to highlight and define how the institutions intend to outline the concept of (non)canonical breeding strategies in view of the new emerging biotechnologies. In 2017, the GM-based agricultural industry celebrated its 22
nd year of commercialization, with 189.8 million hectares of GM crops farmed in 24 countries, including 19 developing and 5 industrialized nations. Compared to industrialized nations, which grow 47% of the world's biotech crop area, developing nations grow 53%, and 40 more nations officially import biotech crops for use in food, feed, and processing [
5]. In 2022, the global area planted to GM crops reached the new record of 202.2 million hectares, up 3.3% from the year before. Eleven distinct GM crops were grown in 27 different countries, with 98.9 million hectares planted with soybeans followed by maize at 66.2 million hectares. Since the initial release of genetically modified crops in 1996, there has been a fluctuation in the number of nations growing GM crops [
6].
For these reasons, which are combined with the advent of new editing technologies, a deeper delineation has become even more necessary to establish in a clearer and more defined way the boundary of what is or is not feasible and permitted from each specific governance, also in relation to the possibility of applying monitoring through molecular traceability. In fact, in addition to the administrative and legal aspects, increasingly frequent questions are becoming more common among the public; in particular, what are the real tangible advantages of these NBT-based technologies compared to conventional breeding approaches? Moreover, how can a technology based on genome editing be useful in a molecular traceability context? Is it possible to develop and use new technologies in this applied research area with direct repercussions for the safe food industry? To answer these questions, we reviewed and outlined the main aspects related to the general use of genome editing-based approaches, considering both their best-known roles in target editing of the genome and their new frontiers with specific reference to no editing implications, including the impacts (and influences) on the molecular traceability in the agri-food sector.
2. Genome-editing toolbox as a new frontier of precision breeding of food crops
The term genome editing (GE) reflects one of the most revolutionary and well-established strategies used in agricultural biotechnology and crop breeding in recent decades. Multiple aspects of GE have contributed to the advancement of modern agriculture and led to shifts in market structure because of the introduction of targeted changes within the existing genetic blueprint of an organism. Generally, the toolbox on which it is based is composed of various techniques that can be used to incorporate targeted, precise modifications into an organism’s genome [
7]. The three foundational GE technologies that have been developed and improved in recent decades include transcription activator-like effector nucleases (TALENs), zinc-finger nuclease (ZFN) technology, and clustered regularly interspaced short palindromic repeats (CRISPR)-associated protein (Cas) technology [
8]. Similarly, these gene editing tools are based primarily on the enzymatic activity of a protein complex being able to cleave DNA and on the subsequent exploitation of three biologically related mechanisms, such as the ability to find a specific DNA sequence in the genome, the ability to cleave DNA at that location, and the activity of the endogenous DNA repair machinery [
7]. Currently, among the three mechanisms mentioned above, the CRISPR system has become the dominant GE method, sweeping aside most other previously developed tools, such as ZFN technology and TALENs. It is indeed much more versatile than other systems since it can be very easily programmed to direct the nuclease to the desired location in the genome, becoming the system that is by far the most widely employed for GE applications [
9,
10]. Furthermore, since the CRISPR/Cas-based genome editing system was developed from a prokaryotic adaptive immune approach against certain viruses; it was introduced to the scientific community as only a modification of a natural mechanism. In addition, CRISPR/Cas-based technologies represent a considerable improvement over prior techniques, as they are simple and versatile in terms of vector design and construction for subsequent plant transformation, and they are regarded as more reliable and straightforward for targeting gene editing for several purposes [
11]. CRISPR/Cas has proven useful in a variety of sectors, including clinical diagnostics, food safety, and biological breeding. Because of their high specificity, sensitivity, and ease of use, class 2 systems, which include Cas9, Cas12a (Cpf1), Cas12b, Cas13a (C2c2), and Cas14a (Cas12f1), are currently the most studied and used single-protein effectors for numerous techniques [
12].
From a structural point of view, the CRISPR/Cas-based system consists of a short noncoding guide RNA (gRNA) made up of two basic elements represented by a target complementary CRISPR RNA (crRNA) and an auxiliary transactivating crRNA (tracrRNA) [
13]. The gRNA can guide the specific Cas endonuclease protein linked to it towards a specific genomic locus via base pairing between the crRNA sequence and the target sequence and cleaves the DNA to create a double-strand break [
14]. In bacteria, CRISPR loci are composed of a series of repeats separated by segments of ~30 bp in length of exogenous DNA called spacers. The repeat-spacer array is transcribed as a long precursor and processed within repeat sequences to generate small crRNAs that specify the target sequences, also known as protospacers, following cleavage by the Cas9 protein. CRISPR spacers are then used to recognize and silence exogenous genetic elements at the DNA level. Essential for cleavage is a three-nucleotide sequence motif (NGG) immediately downstream of the 3’ end of the target region, known as the protospacer-adjacent motif (PAM) [
13].
Several Cas enzymes have been put into use because of their peculiar nuclease activities. Because they originate from various CRISPR systems, they may be beneficial for editing techniques due to their slightly varying abilities to recognize and change PAM sites [
15,
16]. In general, the merits of these systems include swift, broad acceptance for editing applications in a wide variety of plant species, as confirmed by an increasing number of studies in which an expanded use of Cas9 nuclease for editing beyond double strand breaks has been applied [
17,
18,
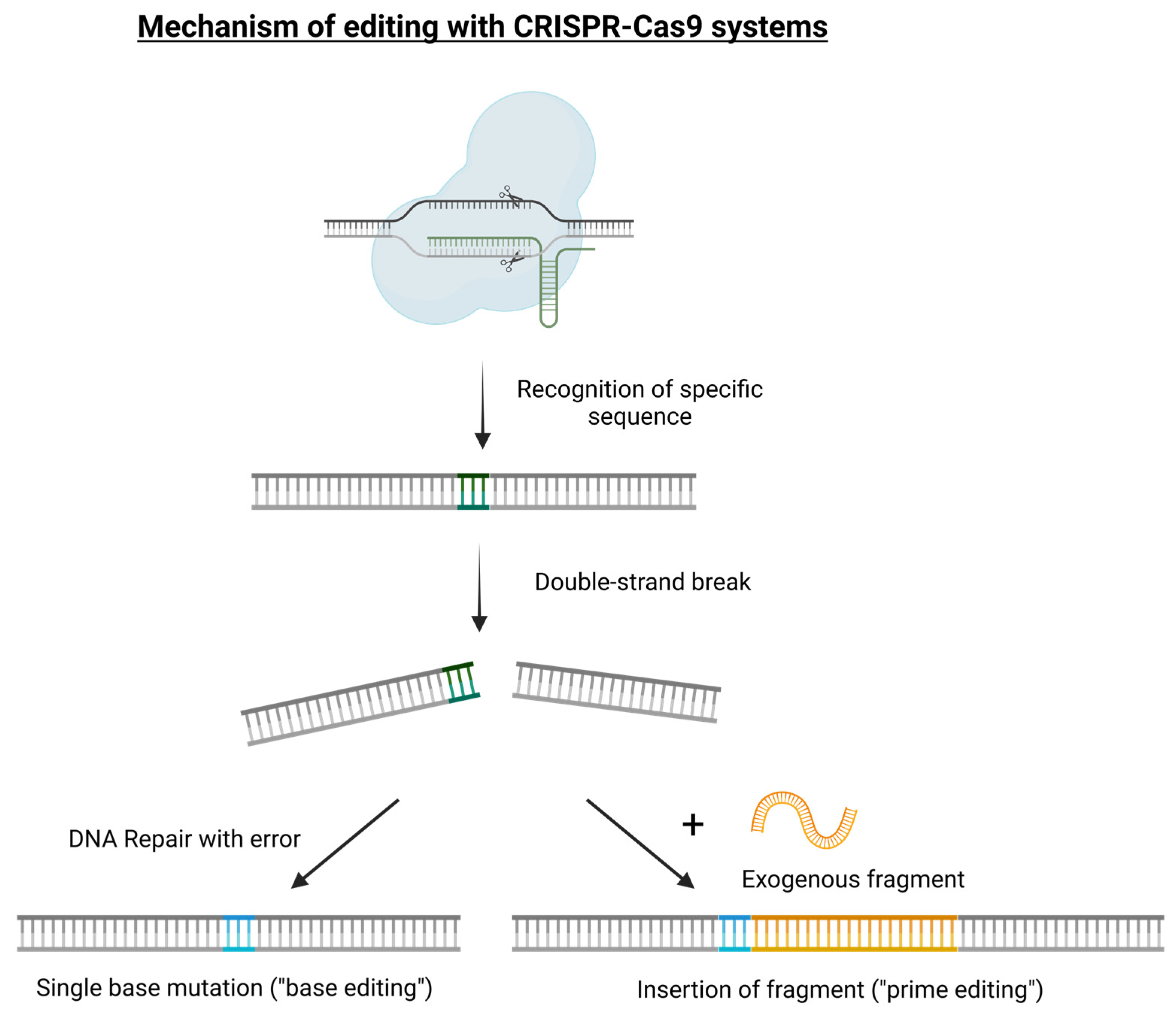
19]. Errors occur during the subsequent repair step, making the DNA not identical to the original DNA. This allows for the insertion of a permanent minor change to the DNA, known as a mutation. Alternatively, additional variants of CRISPR/Cas-based technology have been developed, supporting the precise editing of a single nucleotide (“base editing”) or rewriting a given sequence of DNA (“prime editing”), as in the case in which a DNA template fragment can be added and introduced in association with the gRNA and Cas protein (
Figure 1). With these strategies, a DNA template can also be used to introduce foreign genes and sequences into the genomes of plant species, leading to the formation of a transgenic crop but avoiding any positional effect [
20]. In plant research, this technology is currently exploited for gene knock-out (loss-of-function) or gene replacement (gain-of-function) and it is referred to as precision-type plant breeding technology and it actually represents a technology-assisted plant evolution strategy. The scientific community and specific legislation refer to the commonly known new genomic techniques (NGTs) or new breeding techniques (NBTs) [
21,
22,
23], which are increasingly utilized to design new genotypes and to adjust plant traits, including a significant number of food crops [
24]. Many reviews and investigations have addressed the fundamentals and different aspects of using CRISPR/Cas in functional genomics and crop improvement, particularly in regard to producing target mutants of the genes responsible for several traits related to agronomical characteristics and/or physio-biological responses: applications have mostly concentrated on characteristics linked to disease resistance, environmental stress tolerance, food quality enhancement, and increased crop yields with less inputs [
11,
17,
18,
25,
26,
27]. All this progress has also been made possible by the development of high-throughput sequencing (HTS) platforms that promote and support the study of gene regulators of important traits or specific responses to the environment, as attested by many studies in which the enormous potential of editing approach-based technology has been reported. Some examples range from the examination of stress response control in roots at the cellular level to allele replacement for quantitative trait locus (QTL) validation, all traits related to roots crucial for crop improvement utilizing CRISPR/Cas-based genome editing [
28]. Additionally, developing cereal crops with root systems that can absorb irregularly distributed water and nutrient resources in times of resource scarcity and climate instability is another main application example of a primary breeding goal for CRISPR/Cas genome editing. To date, studies employing CRISPR/Cas-mediated strategies to increase crop abiotic stress tolerance have been the main research focus for food crop improvement using GE technology [
29,
30,
31,
32,
33,
34,
35,
36,
37,
38]; the development of crop resistance to pathogens (e.g., bacteria, viruses, and fungi); the modulation of targeting susceptibility systems, as demonstrated for the tomato yellow leaf curl virus in
Nicotiana benthamiana [
39,
40], bean yellow dwarf virus in
Nicotiana benthamiana [
41], and beet severe curly top virus in Arabidopsis and
Nicotiana benthamiana [
19,
40,
42,
43,
44,
45,
46,
47,
48,
49,
50,
51]. Furthermore, CRISPR/Cas technology has produced a successful substitute for the production of plant material that is resistant to a variety of herbicides; for example, CRISPR/Cas has been used to target the ALS gene in crops, including rice, wheat, and other cereals, and a number of herbicide-tolerant polymers have been created [
52,
53,
54,
55].
3. CRISPR/Cas technology as a precision tool for food crop traceability
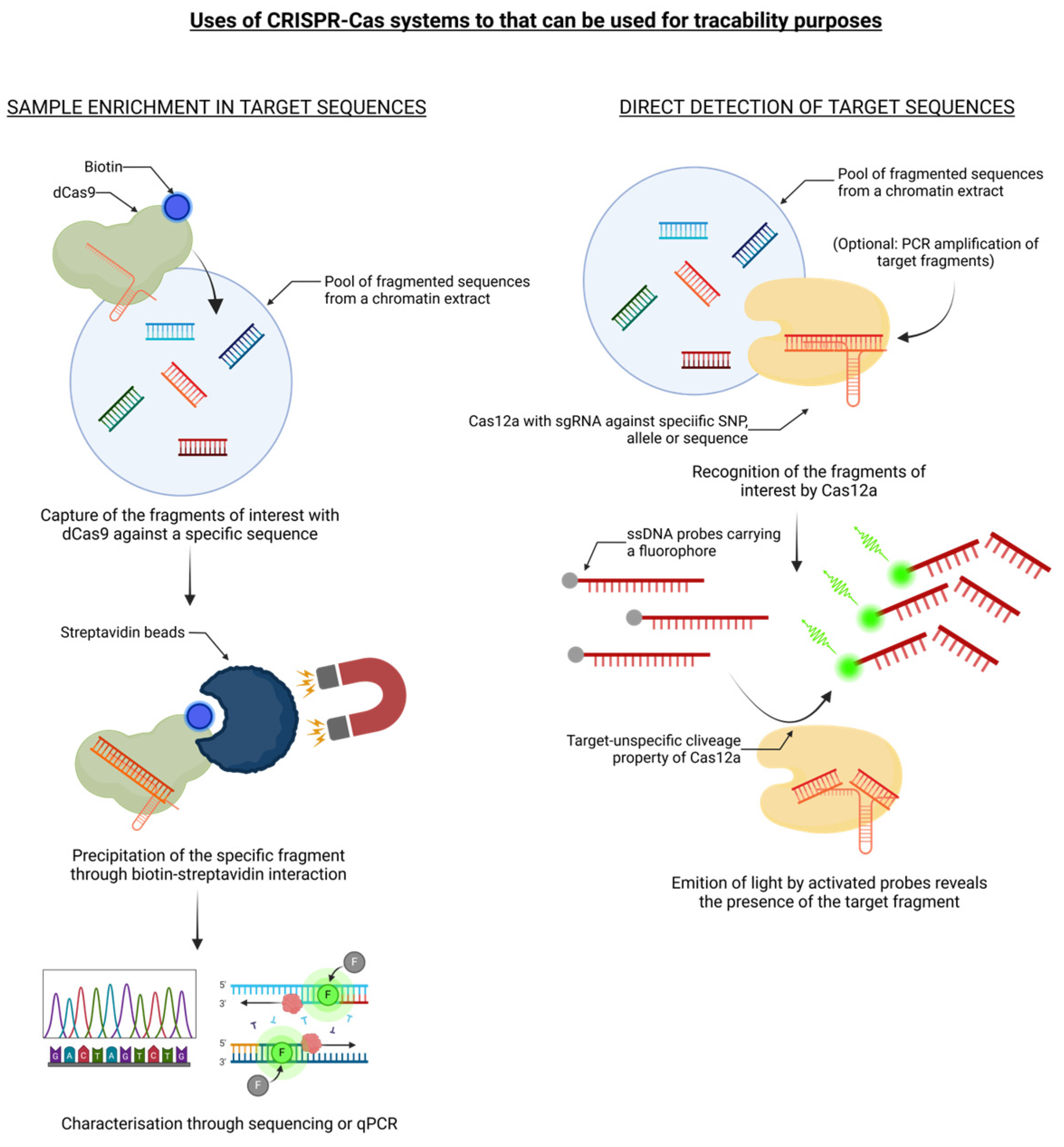
CRISPR/Cas technology, due to its enormous potential related to its precision and reliability, ease of use and low cost, is now considered a valuable alternative to conventional plant breeding methods for intraspecific gene mutation (e.g. mutagenesis) or interspecific gene introgression (e.g. backcross strategy). As documented above, CRISPR/Cas has been used for an increasing number of applications, which has led to the development of a range of editing tools in crop plant science. However, since a genetic engineering approach may be affected by a highly contested and controversial societal issue, as was the case for conventional genetic transformation techniques (i.e. GMOs produced via transgenesis and intragenesis), the use of CRISPR/Cas as a new breeding technique poses new questions regarding the preferences of consumers and producers, food ethics and political standpoints of governance leaders. Since its discovery in 2012 as a novel tool for genome editing, the CRISPR/Cas system has spawned a plethora of literature, but in-depth evaluations of its potentials and/or limitations in relation to food production and authentication by means of molecular or genetic traceability applications are just beginning to take shape. Indeed, these emerging genomic engineering approaches can offer a valid key for the effective prevention and control of food safety risks, in which the development of specific, sensitive, and rapid nucleic acid detection methods, such as foodborne pathogenic bacteria, genetically modified crops, and product adulteration, is imperative (
Figure 2).
The sequencing of nucleic acids is the main approach for identifying one or more specific species of any kingdom since DNA acts as a natural biomarker for validation in both (un)processed food products and derivatives. The DNA barcoding technique, which refers to the identification of plant species using a short domain of the plastidial genome from a specific genic or intergenic target sequence, is currently available and still appears to be among the most useful food authentication applications based on DNA sequencing [
56]. For similar reasons, recent isothermal amplification strategies and advances in DNA and omics-based technology (i.e., genomics, metabolomics, and proteomics) for the detection of specific nucleic acid regions have been developed, allowing for potential use in food testing, identification and authentication [
57,
58,
59]. Nevertheless, ascertaining food species and even varieties or genotypes by sequencing DNA barcodes or genotyping DNA markers is mainly performed through targeted polymerase chain reaction (PCR)-based and real-time quantitative PCR (qPCR)-based techniques coupled with methods to improve the specificity of PCR methods and increase the applicability and reliability/reproducibility of PCR for food authenticity applications [
59,
60,
61,
62].
Despite DNA barcoding is to date widely used as routine molecular tool, at least when it comes to species-level identification strategies, there are still drawbacks to its use, including i) the difficulty in creating universal primers for amplifying genomic regions that can specifically discriminant plant varieties or genotypes, and ii) the low resolution of the technology when it comes to the identification of closely related species/botanical varieties/cultivars [
63]. To surpasses these limitations, the CRISPR-Cas system can now be used to detect food adulteration, mislabelling, the presence of allergens, and allergen sources due to recent technological breakthroughs, but also variety identification for food-destinated crops [
64]. The growing number of studies highlighting the potential of the CRISPR-Cas system in the investigation of food authenticity and its ability to improve PCR method detection performance is a testimony of the ongoing development of various technical solutions [
65]. The combination of CRISPR-Cas with nucleic acid amplification techniques greatly improves the specificity and sensitivity of nucleic acid identification not only for food crops but also for detecting infections and nucleic acid biomarkers [
66,
67,
68,
69]. Several targeted approaches exploiting both collaboration and competition between CRISPR- and omics-based technologies have been developed, leading to the development of universal nucleic acid testing approaches based on the CRISPR‒Cas system to identify DNA barcodes in a very sensitive and reliable way regarding food safety and authenticity. The resulting CRISPR‒Cas-based DNA barcoding method showed great potential for food authenticity and broadened the application of the CRISPR-Cas system to these sectors [
70,
71]. Furthermore, most applications have demonstrated the potential of CRISPR/Cas systems to target nucleic acid portions due to the specific recognition of their sequence, which is also useful for non-editing applications. In the past decade, several new methods have emerged in the field of genome studies that involve the use of an interesting version of the Cas9 protein, referred to as dead-Cas9 (dCas9), which was developed to target DNA sequences through the use of a specific guide without cleaving them [
72]. Indeed, this peculiar Cas9 version is deprived of its catalytic activity because of the two point mutations (D10A and H841A) that its amino acid chain harbours and that induce the loss of function of the RuvC1 and HNH nuclease domains [
73].
In addition to the Cas9 protein, an alternative endonuclease, which is largely employed for these purposes, Cas12a, also known as Cpf1, is a typical V-A CRISPR system. Instead of producing blunt ends when it cleaves DNA, Cas12a can produce 5' overhangs, which is more useful for some genome editing applications, such as inserting a DNA sequence at a particular point [
74]. Furthermore, when activated by the highly specific binding of the DNA sequence indicated by the gRNA, Cas12a exhibits both sequence-independent single-stranded DNA degradation and sequence-specific double-stranded DNA cleavage [
75,
76].
3.1. CRISPR/Cas as an enrichment tool for next-generation sequencing
Selective enrichment of nucleotide regions of interest using CRISPR/Cas technology for sequencing purposes has predominantly emerged in the traceability field due to its importance. The need to target specific genetic loci is crucial and at the base of traceability purposes themselves, often conditioned/influenced by genome-wide approaches. In general, for the application of techniques based on genome-wide analyses, it is imperative to screen for signals from high-abundance undesirable species before sequencing [
77]. The identification or enrichment of certain regions of interest has progressively increased as a crucial tool for traceable sequencing, prompting the creation of multiple procedures suitable for various experimental purposes. It is therefore presumable that in the coming years, additional tools that take advantage of the CRISPR/Cas system and its characteristics will be created in the scope of increasing the possibility of studying nucleic acid characteristics, such as the detection of structural variations at the haplotype level using long-read sequencing. These tools are mainly based on the high specificity of targeting through sgRNA and the existence of a plethora of available Cas proteins with the ability to perform a series of molecular actions on the nucleotide sequence of interest. For example, CRISPR-assisted targeted enrichment sequencing (CATE-seq) consists of the fragmentation and subsequent specific ligation of adaptors to sample DNA, after which the targets are bound by dCas9 and purified for allele-specific PCR or library preparation, resulting in a highly sensitive alternative approach able to reach over 3000-fold enrichment of the target sequences [
78]. In addition, ultrasensitive CRISPR–Cas9-triggered nicking endonuclease-mediated strand displacement amplification (CRISDA) [
79], which can achieve attomolar sensitivity, combines Cas9 cleavage activity with highly specific target site amplification and annealing of biotin and Cy5-labelled PNA (a peptide nucleic acid) probes to the amplicon. In contrast, CRISPR-mediated isolation of specific megabase-sized regions of the genome (CISMR) enables the isolation of regions of interest through Cas9-driven cleavage at flanking sites by combining a pulse-gel electrophoresis step for sequence isolation and an amplification step, followed by long-read sequencing [
80].
3.2. CRISPR/Cas as a detection system
As described in the previous paragraphs, PCR-based methods, including qualitative PCR [
81], quantitative PCR (qPCR) [
82], droplet digital PCR (ddPCR) [
83], and LAMP [
11], have long been the primary and undisputed DNA detection strategies and have been utilized for traceability in all research sectors, despite being costly, labour-intensive, and time-consuming. Therefore, the improvement of rapid, simple, and sensitive new detection methods for nucleic acids without special technical expertise and ancillary equipment is increasingly necessary.
The introduction of the CRISPR/Cas system, as a novel gene editing tool, has allowed biotechnology to enter a new era. In addition to the widely documented purposes, ranging from cell imaging to expression regulation and DNA assembly [
84,
85,
86,
87,
88], a plethora of additional uses, defined as CRISPR/Cas-based nucleic acid detection system assays, have also been described. They mainly exploit the capacities of the Cas12a protein [
67,
76] to perform nonspecific ssDNA cleavage upon recognition of a target that matches its crRNA [
14,
74,
85] and to produce a 5′ sticky end on the target itself [
76,
89,
90]. Recently, recombinase polymerase amplification (RPA) and the nonspecific ssDNA cleavage property of Cas12a have been combined to create a novel detection technology platform known as RPA-Cas12a-FS: the combination of RPA with guide crRNA targeting and detection of FAMBHQ1-labelled reporter ssDNA allowed the development of an on-site detection method for the molecular identification of various biological contaminants, genetically modified crops, and adulteration [
71,
91,
92]. This innovative method offers an alternative tool for DNA-level detection and assists as a front-line nucleic acid quick detection system for food safety. It may be applied at the end of the food inspection and traceability/authentication industry chain. In relation to Cas12 protein use, a new method combining CRISPR‒Cas-based PCR DNA barcoding (CAPCOD) was developed, in which the specificity of the CRISPR/Cas12 system and the high amplification efficiency of the PCR method for identifying DNA barcodes to improve the accuracy of sample authenticity are combined [
93]. The nonspecific endonuclease activity of dCas9 allows the random cleavage of adjacent single-strand nucleic acids by trans-cleavage upon recognition of the target sequence [
84,
94]; based on this understanding of its activity, CRISPR-Cas12 has been applied to nucleic acid testing because it can precisely identifying amplicons via isothermal amplification or PCR [
95,
96,
97].
Since CRISPR/Cas can be considered an effective method for molecular detection based on trans-cleavage activity, this ultrasensitive detection method can usually be exploited for the detection of GMOs, gene-edited products and SNPs [
71,
91,
98]. Regarding GMO detection or edited products, with and without exogenous sequences, the strategies may differ based on the employed approach. In contrast, the methods for detecting mutations of a single or a few bases can differ from those used to detect SNPs. A combination of LAMP and CRISPR/Cas12a was used for visual detection of GM soybean powders by UV light [
99]. A portable biosensor for visual dual detection of the
CaMV35S promoter and
lectin gene in soybean powders, named Cas12a-PB, has also been developed [
99]. Furthermore, in 2022, Cao and colleagues [
100] established the MPT-Cas12a/13a technology, which combines multiplex PCR and transcription for simultaneous but distinct detection of
CaMV35S and T-
nos elements. Many different additional approaches have been established with unique analyses (particularly fluorescence detection and gold nanoparticle-based colorimetry assays) combined with CRISPR/Cas systems, and they have been extensively reviewed by Wang et al. [
16].
For edited sequences in which universal components cannot be identified in the same manner as for GMOs, it is necessary to pick a specific motif with an appropriate PAM site for subsequent detection. An interesting method, called PCR/ribonucleoprotein (RNP), to detect mutations in gene-edited diploid and polyploid plants is to assemble CRISPR/Cas9 and CRISPR/Cpf1. This method outcomes able to differentiate between homozygous and biallelic mutations as well as mutagenesis induced by TALEN protein and mutant screening that is unaffected by background noise SNPs, resulting particularly advantageous for screening polyploid plants [
101]. In rice, a Cas12aFVD platform for biosensing and for the visible detection of gene-edited mutants was also developed [
102]. For gene-edited products with known editing sites and sequences, methodologies like ACT-PCR, ddPCR, AS-PCR, CRISPR/Cas, etc., can be used for preliminary screening, whereas in the presence of known editing sites and sequences, they can be detected according to the current GMO detection strategy. In contrast, for gene-edited products with unknown editing sites and sequences, one or more methods can be used for preliminary screening.. In general, Sanger sequencing, NGS, T7EI, and RFLP are currently the most extensively used methods; the use of other molecular-based techniques is less prevalent because the choice of detection techniques is strongly correlated with the kind of mutation, plant ploidy, and efficiency of gene editing [
16,
103].
The application of the CRISPR/Cas system for SNP detection has been employed with the purpose of providing different strategies that are useful for the discrimination of single nucleotide mismatches (SNMs). For example, when combined with asymmetric PCR, Cas12b successfully distinguished the SNP locus without the PAM sequence. This means that it can cleave ssDNA without a PAM sequence [
85]. Several other reports have confirmed the success of different combination strategies for distinguishing SNPs and achieving single-base resolution detection in all kingdoms [
69,
104,
105].
CRISPR/Cas detection systems can also benefit from the application of microfluidic technology. In 2021, Chen et al. [
98] included a nucleotide mismatch to increase the universality of SNP detection. To automate the procedure, CRISPR/Cas12a reagents were preloaded onto the biochip to differentiate between the homozygous wild type, homozygous mutant type, and heterozygous mutant type strains.
4. Future opportunities and trends
This communication highlights the wide application potential of the CRISPR-Cas technology as a support for molecular traceability purposes in several research fields related to crop production (i.e. plant varieties) and food commercialization (i.e. food products and their derivatives). Using the CRISPR/Cas-based method, numerous genes have been edited and plant traits have been improved, and the resulting commercial items have been introduced into the market.
CRISPR/Cas detection systems display numerous outstanding features, including low cost, easy operation, quick application. and high accuracy and reliability. However, most of the currently available detection methods are not suitable for prompt on-site detection because they require labour-intensive specialized methodologies, dedicated instruments, and sophisticated sample processing and analysis. In addition to gene editing, CRISPR/Cas systems have recently been employed for target detection and can also be used to efficiently identify proteins, metal ions, nucleic acids, and other chemicals via a variety of technologies. The high sensitivity and specificity of these methods are two of the highest strengths of the CRISPR/Cas system, and laborious experimental procedures and time-consuming analyses are not needed. To accomplish flexible detection of nucleic acid and non-nucleic acid targets in the domains of clinical diagnostics, environmental testing, biological breeding, food safety, and others, the CRISPR/Cas system can be coupled with a range of amplification techniques, readout techniques, and devices. Although CRISPR testing has been applied extensively, CRISPR nucleic acid testing remains in its infancy and has much room for development. CRISPR/Cas systems have many potential applications in the future due to research into their use with nanomaterials, 3D printing, the internet, big data, automation, and artificial intelligence. The industrial revolution and the biotechnological revolution are currently accelerating. Therefore, rapid, efficient and accurate detection methods will become a major challenge for standard detection as more diverse traits and products continue to emerge, molecular characterization information and related databases are extremely limited and imperfect, and the optimization of mutation detection technology remains an ongoing attempt. This should be accomplished to guarantee researchers’ intellectual property rights, to expedite the development of testing standards and procedures for biotechnology goods, and to offer robust technical assistance for the oversight and monitoring of national security.
Author Contributions
Conceptualization, S.F. and G.B.; investigation, S.F. and A.D.; resources, G.B.; data curation, S.F. and A.D.; writing—original draft preparation, S.F.; writing—review and editing, S.F., A.D. and F.S; visualization, S.F., F.S., F.P. and G.B.; supervision, A.V. and G.B.; project administration, G.B.; funding acquisition, G.B. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the research contract signed by the Gruppo Padana S.S. company (Paese, TV, Italy), and Department of Agronomy, Food, Natural resources, Animals and Environment (DAFNAE), University of Padua (Italy), within action IV.5_GREEN, PON 2014–2021, and within the Agritech National Research Centre, for which it was received funding from the European Union Next-Generation EU (PIANO NAZIONALE DI RIPRESA E RESILIENZA (PNRR)—MISSIONE 4 COMPONENTE 2, INVESTIMENTO 1.4—D.D. 1032 17/06/2022, CN00000022, Spoke 4—Multifunctional and resilient agricultural and forest systems for the mitigation of climate change risks, PI: Gianni Barcaccia). In particular, the study represents a baseline for the fulfilment of the milestones within Task 1.3.3 titled “Development and implementation of novel biotechnological approaches, including cisgenesis and genome editing, for accelerated precision breeding”, and Task 1.3.5 titled “Genome-wide strategies for fast-forward molecular breeding aimed at assessment of genetic distinctiveness, uniformity and stability (DUS), and identity of pre-commercial varieties in the main horticultural crop species for the fresh-cut market” related to the Spoke 1 “Plant and animal genetic resources and adaptation to climate changes”.
Acknowledgments
In this section, you can acknowledge any support given which is not covered by the author contribution or funding sections. This may include administrative and technical support, or donations in kind (e.g., materials used for experiments).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Shukla-Jones, A.; Friedrichs, S.; Winickoff, D.E. Gene editing in an international context. 2018. [Google Scholar] [CrossRef]
- Schiemann, J.; Dietz-Pfeilstetter, A.; Hartung, F.; Kohl, C.; Romeis, J.; Sprink, T. Risk Assessment and Regulation of Plants Modified by Modern Biotechniques: Current Status and Future Challenges. Annu Rev Plant Biol 2019, 70, 699–726. [Google Scholar] [CrossRef] [PubMed]
- Bujnicki, J.; Dykstra, P.; Wegener, H. New techniques in agricultural biotechnology. Publications Office of the European Union: Luxembourg 2017.
- Briefs. Global status of commercialized biotech/GM crops in 2017: Biotech crop adoption surges as economic benefits accumulate in 22 years. ISAAA 2017, 53, 25–26. [Google Scholar]
- AgbioInvestor. Global GM Global area review. 2023.
- Songstad, D.D.; Petolino, J.F.; Voytas, D.F.; Reichert, N.A. Genome Editing of Plants. Critical Reviews in Plant Sciences 2017, 36, 1–23. [Google Scholar] [CrossRef]
- Gay, G.; Braun, L.; Brenier-Pinchart, M.P.; Vollaire, J.; Josserand, V.; Bertini, R.L.; Varesano, A.; Touquet, B.; De Bock, P.J.; Coute, Y. , et al. Toxoplasma gondii TgIST co-opts host chromatin repressors dampening STAT1-dependent gene regulation and IFN-gamma-mediated host defenses. J Exp Med 2016, 213, 1779–1798. [Google Scholar] [CrossRef]
- Menz, J.; Modrzejewski, D.; Hartung, F.; Wilhelm, R.; Sprink, T. Genome Edited Crops Touch the Market: A View on the Global Development and Regulatory Environment. Front Plant Sci 2020, 11, 586027. [Google Scholar] [CrossRef] [PubMed]
- Parisi, C.; Rodríguez-Cerezo, E. Current and future market applications of new genomic techniques. Publications Office of the European Union: Luxembourg 2021, JRC123830.
- Panozzo, S.; Farinati, S.; Sattin, M.; Scarabel, L. Can allele-specific loop-mediated isothermal amplification be used for rapid detection of target-site herbicide resistance in Lolium spp.? Plant Methods 2023, 19, 14. [Google Scholar] [CrossRef] [PubMed]
- Sashital, D.G. Pathogen detection in the CRISPR-Cas era. Genome Med 2018, 10, 32. [Google Scholar] [CrossRef]
- Hassan, M.M.; Zhang, Y.; Yuan, G.; De, K.; Chen, J.G.; Muchero, W.; Tuskan, G.A.; Qi, Y.; Yang, X. Construct design for CRISPR/Cas-based genome editing in plants. Trends Plant Sci 2021, 26, 1133–1152. [Google Scholar] [CrossRef]
- Ma, E.; Harrington, L.B.; O'Connell, M.R.; Zhou, K.; Doudna, J.A. Single-Stranded DNA Cleavage by Divergent CRISPR-Cas9 Enzymes. Mol Cell 2015, 60, 398–407. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Dong, H.; Cui, Y.; Cong, L.; Zhang, D. Application of different types of CRISPR/Cas-based systems in bacteria. Microb Cell Fact 2020, 19, 172. [Google Scholar] [CrossRef]
- Wang, M.; Wang, H.; Li, K.; Li, X.; Wang, X.; Wang, Z. Review of CRISPR/Cas Systems on Detection of Nucleotide Sequences. In Foods, 2023; Vol. 12.
- Bortesi, L.; Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol Adv 2015, 33, 41–52. [Google Scholar] [CrossRef]
- Quetier, F. The CRISPR-Cas9 technology: Closer to the ultimate toolkit for targeted genome editing. Plant Sci 2016, 242, 65–76. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Simmonds, J.; Pan, Q.; Davidson, D.; He, F.; Battal, A.; Akhunova, A.; Trick, H.N.; Uauy, C.; Akhunov, E. Gene editing and mutagenesis reveal inter-cultivar differences and additivity in the contribution of TaGW2 homoeologues to grain size and weight in wheat. Theor Appl Genet 2018, 131, 2463–2475. [Google Scholar] [CrossRef]
- Gong, X.; Zhang, T.; Xing, J.; Wang, R.; Zhao, Y. Positional effects on efficiency of CRISPR/Cas9-based transcriptional activation in rice plants. aBIOTECH 2020, 1, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Whelan, A.I.; Lema, M.A. Regulatory framework for gene editing and other new breeding techniques (NBTs) in Argentina. GM Crops Food 2015, 6, 253–265. [Google Scholar] [CrossRef]
- He, Y.; Mudgett, M.; Zhao, Y. Advances in gene editing without residual transgenes in plants. Plant Physiol 2022, 188, 1757–1768. [Google Scholar] [CrossRef] [PubMed]
- Romeo Lironcurti, S.; Demaria, F.; Quarto, A.; Solazzo, R. The ongoing debate on NBTs and possible roads for the future. Frontiers in Political Science 2024, 5. [Google Scholar] [CrossRef]
- Gaillochet, C.; Develtere, W.; Jacobs, T.B. CRISPR screens in plants: approaches, guidelines, and future prospects. Plant Cell 2021, 33, 794–813. [Google Scholar] [CrossRef] [PubMed]
- Belhaj, K.; Chaparro-Garcia, A.; Kamoun, S.; Patron, N.J.; Nekrasov, V. Editing plant genomes with CRISPR/Cas9. Curr Opin Biotechnol 2015, 32, 76–84. [Google Scholar] [CrossRef]
- Shan, Q.; Zhang, Y.; Chen, K.; Zhang, K.; Gao, C. Creation of fragrant rice by targeted knockout of the OsBADH2 gene using TALEN technology. Plant Biotechnol J 2015, 13, 791–800. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Chen, W.; Li, Y.; Sun, L.; Chai, Y.; Chen, H.; Nie, H.; Huang, C. CRISPR/Cas9 Technology and Its Utility for Crop Improvement. In International Journal of Molecular Sciences, 2022; Vol. 23.
- Ahmadi, N.; Audebert, A.; Bennett, M.J.; Bishopp, A.; de Oliveira, A.C.; Courtois, B.; Diedhiou, A.; Dievart, A.; Gantet, P.; Ghesquiere, A. , et al. The roots of future rice harvests. Rice (N Y) 2014, 7, 29. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Wang, Y.; Zhang, R.; Zhang, H.; Gao, C. CRISPR/Cas Genome Editing and Precision Plant Breeding in Agriculture. Annu Rev Plant Biol 2019, 70, 667–697. [Google Scholar] [CrossRef]
- Voss-Fels, K.P.; Stahl, A.; Hickey, L.T. Q&A: modern crop breeding for future food security. BMC Biol 2019, 17, 18. [Google Scholar] [CrossRef]
- Liu, Q.; Yang, F.; Zhang, J.; Liu, H.; Rahman, S.; Islam, S.; Ma, W.; She, M. Application of CRISPR/Cas9 in Crop Quality Improvement. In International Journal of Molecular Sciences, 2021; Vol. 22.
- Zhu, H.; Li, C.; Gao, C. Applications of CRISPR-Cas in agriculture and plant biotechnology. Nat Rev Mol Cell Biol 2020, 21, 661–677. [Google Scholar] [CrossRef]
- Lu, K.; Wu, B.; Wang, J.; Zhu, W.; Nie, H.; Qian, J.; Huang, W.; Fang, Z. Blocking amino acid transporter OsAAP3 improves grain yield by promoting outgrowth buds and increasing tiller number in rice. Plant Biotechnol J 2018, 16, 1710–1722. [Google Scholar] [CrossRef]
- Ma, X.; Feng, F.; Zhang, Y.; Elesawi, I.E.; Xu, K.; Li, T.; Mei, H.; Liu, H.; Gao, N.; Chen, C. , et al. A novel rice grain size gene OsSNB was identified by genome-wide association study in natural population. PLoS Genet 2019, 15, e1008191. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liang, Z.; Zong, Y.; Wang, Y.; Liu, J.; Chen, K.; Qiu, J.L.; Gao, C. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat Commun 2016, 7, 12617. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Tu, M.; Wang, D.; Liu, J.; Li, Y.; Li, Z.; Wang, Y.; Wang, X. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol J 2018, 16, 844–855. [Google Scholar] [CrossRef] [PubMed]
- Dahan-Meir, T.; Filler-Hayut, S.; Melamed-Bessudo, C.; Bocobza, S.; Czosnek, H.; Aharoni, A.; Levy, A.A. Efficient in planta gene targeting in tomato using geminiviral replicons and the CRISPR/Cas9 system. Plant J 2018, 95, 5–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, G.; Gao, Y.; Lu, G.; Habben, J.E.; Mao, G.; Chen, G.; Wang, J.; Yang, F.; Zhao, X. , et al. A cytokinin-activation enzyme-like gene improves grain yield under various field conditions in rice. Plant Mol Biol 2020, 102, 373–388. [Google Scholar] [CrossRef]
- Ali, M.S.; Kim, K.W.; Dhakal, R.; Choi, D.; Baek, K.H. Accumulation of high contents of free amino acids in the leaves of Nicotiana benthamiana by the co-suppression of NbClpC1 and NbClpC2 genes. Plant Cell Rep 2015, 34, 355–365. [Google Scholar] [CrossRef]
- Ali, M.S.; Yu, Y.; Oh, W.; Cho, J.Y.; Choi, J.; Dhakal, R.; Park, Y.-I.; Baek, K.-H. Co-suppression of “NbClpC1” and “NbClpC2” in “Nicotiana benthamiana” lowers photosynthetic capacity via altered leaf structure. Plant Omics 2015, 8, 508. [Google Scholar] [CrossRef]
- Baltes, N.J.; Hummel, A.W.; Konecna, E.; Cegan, R.; Bruns, A.N.; Bisaro, D.M.; Voytas, D.F. Conferring resistance to geminiviruses with the CRISPR-Cas prokaryotic immune system. Nat Plants 2015, 1, 15145. [Google Scholar] [CrossRef] [PubMed]
- Borrelli, V.M.G.; Brambilla, V.; Rogowsky, P.; Marocco, A.; Lanubile, A. The Enhancement of Plant Disease Resistance Using CRISPR/Cas9 Technology. Front Plant Sci 2018, 9, 1245. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cheng, X.; Shan, Q.; Zhang, Y.; Liu, J.; Gao, C.; Qiu, J.L. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 2014, 32, 947–951. [Google Scholar] [CrossRef]
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci Rep 2017, 7, 482. [Google Scholar] [CrossRef] [PubMed]
- Dong, O.X.; Ronald, P.C. Genetic Engineering for Disease Resistance in Plants: Recent Progress and Future Perspectives. Plant Physiol 2019, 180, 26–38. [Google Scholar] [CrossRef]
- Pu Yan, L.I.U.C.L.I.J.-Y.A.G.T.H.U.Y.L.I.U.X. Different<em> SlU6</em> Promoters Cloning and Establishment of CRISPR/Cas9 Mediated Gene Editing System in Tomato. Scientia Agricultura Sinica 2018, 51, 315–326. [Google Scholar] [CrossRef]
- Wang, F.; Wang, C.; Liu, P.; Lei, C.; Hao, W.; Gao, Y.; Liu, Y.G.; Zhao, K. Enhanced Rice Blast Resistance by CRISPR/Cas9-Targeted Mutagenesis of the ERF Transcription Factor Gene OsERF922. PLoS One 2016, 11, e0154027. [Google Scholar] [CrossRef]
- Ma, J.; Chen, J.; Wang, M.; Ren, Y.; Wang, S.; Lei, C.; Cheng, Z.; Sodmergen. Disruption of OsSEC3A increases the content of salicylic acid and induces plant defense responses in rice. J Exp Bot 2018, 69, 1051–1064. [Google Scholar] [CrossRef]
- Oliva, R.; Ji, C.; Atienza-Grande, G.; Huguet-Tapia, J.C.; Perez-Quintero, A.; Li, T.; Eom, J.S.; Li, C.; Nguyen, H.; Liu, B. , et al. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotechnol 2019, 37, 1344–1350. [Google Scholar] [CrossRef]
- Malnoy, M.; Viola, R.; Jung, M.H.; Koo, O.J.; Kim, S.; Kim, J.S.; Velasco, R.; Nagamangala Kanchiswamy, C. DNA-Free Genetically Edited Grapevine and Apple Protoplast Using CRISPR/Cas9 Ribonucleoproteins. Front Plant Sci 2016, 7, 1904. [Google Scholar] [CrossRef]
- Mishra, R.; Mohanty, J.N.; Mahanty, B.; Joshi, R.K. A single transcript CRISPR/Cas9 mediated mutagenesis of CaERF28 confers anthracnose resistance in chilli pepper (Capsicum annuum L.). Planta 2021, 254, 5. [Google Scholar] [CrossRef]
- Zong, Y.; Song, Q.; Li, C.; Jin, S.; Zhang, D.; Wang, Y.; Qiu, J.L.; Gao, C. Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat Biotechnol 2018, 36, 950–953. [Google Scholar] [CrossRef]
- Zhang, R.; Liu, J.; Chai, Z.; Chen, S.; Bai, Y.; Zong, Y.; Chen, K.; Li, J.; Jiang, L.; Gao, C. Generation of herbicide tolerance traits and a new selectable marker in wheat using base editing. Nat Plants 2019, 5, 480–485. [Google Scholar] [CrossRef]
- Kuang, Y.; Li, S.; Ren, B.; Yan, F.; Spetz, C.; Li, X.; Zhou, X.; Zhou, H. Base-Editing-Mediated Artificial Evolution of OsALS1 In Planta to Develop Novel Herbicide-Tolerant Rice Germplasms. Mol Plant 2020, 13, 565–572. [Google Scholar] [CrossRef]
- Li, Z.; Liu, Z.B.; Xing, A.; Moon, B.P.; Koellhoffer, J.P.; Huang, L.; Ward, R.T.; Clifton, E.; Falco, S.C.; Cigan, A.M. Cas9-Guide RNA Directed Genome Editing in Soybean. Plant Physiol 2015, 169, 960–970. [Google Scholar] [CrossRef]
- Galimberti, A.; De Mattia, F.; Losa, A.; Bruni, I.; Federici, S.; Casiraghi, M.; Martellos, S.; Labra, M. DNA barcoding as a new tool for food traceability. Food Research International 2013, 50, 55–63. [Google Scholar] [CrossRef]
- Hu, Y.; Lu, X. Rapid Pomegranate Juice Authentication Using a Simple Sample-to-Answer Hybrid Paper/Polymer-Based Lab-on-a-Chip Device. ACS Sens 2020, 5, 2168–2176. [Google Scholar] [CrossRef]
- Hu, Y.; Huang, S.Y.; Hanner, R.; Levin, J.; Lu, X. Study of fish products in Metro Vancouver using DNA barcoding methods reveals fraudulent labeling. Food Control 2018, 94, 38–47. [Google Scholar] [CrossRef]
- Skouridou, V.; Tomaso, H.; Rau, J.; Bashammakh, A.S.; El-Shahawi, M.S.; Alyoubi, A.O.; O'Sullivan, C.K. Duplex PCR-ELONA for the detection of pork adulteration in meat products. Food Chem 2019, 287, 354–362. [Google Scholar] [CrossRef] [PubMed]
- Ng, C.C.; Chang, C.C.; Wu, I.C.; Kotwal, S.; Shyu, Y.T. Rapid molecular identification of freshly squeezed and reconstituted orange juice. International Journal of Food Science & Technology 2006, 41, 646–651. [Google Scholar] [CrossRef]
- Li, T.; Wang, J.; Wang, Z.; Qiao, L.; Liu, R.; Li, S.; Chen, A. Quantitative determination of mutton adulteration with single-copy nuclear genes by real-time PCR. Food Chem 2021, 344, 128622. [Google Scholar] [CrossRef] [PubMed]
- Valentini, P.; Galimberti, A.; Mezzasalma, V.; De Mattia, F.; Casiraghi, M.; Labra, M.; Pompa, P.P. DNA Barcoding Meets Nanotechnology: Development of a Universal Colorimetric Test for Food Authentication. Angew Chem Int Ed Engl 2017, 56, 8094–8098. [Google Scholar] [CrossRef] [PubMed]
- Dawan, J.; Ahn, J. Application of DNA barcoding for ensuring food safety and quality. Food Sci Biotechnol 2022, 31, 1355–1364. [Google Scholar] [CrossRef]
- Kumar, P.; Rani, A.; Singh, S.; Kumar, A. Recent advances on DNA and omics-based technology in Food testing and authentication: A review. Journal of Food Safety 2022, 42, e12986. [Google Scholar] [CrossRef]
- Lanubile, A.; Stagnati, L.; Marocco, A.; Busconi, M. DNA-based techniques to check quality and authenticity of food, feed and medicinal products of plant origin: A review. Trends in Food Science & Technology 2024, 149, 104568. [Google Scholar] [CrossRef]
- Gootenberg, J.S.; Abudayyeh, O.O.; Lee, J.W.; Essletzbichler, P.; Dy, A.J.; Joung, J.; Verdine, V.; Donghia, N.; Daringer, N.M.; Freije, C.A. , et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 2017, 356, 438–442. [Google Scholar] [CrossRef] [PubMed]
- Gootenberg, J.S.; Abudayyeh, O.O.; Kellner, M.J.; Joung, J.; Collins, J.J.; Zhang, F. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 2018, 360, 439–444. [Google Scholar] [CrossRef]
- Shen, J.; Zhou, X.; Shan, Y.; Yue, H.; Huang, R.; Hu, J.; Xing, D. Sensitive detection of a bacterial pathogen using allosteric probe-initiated catalysis and CRISPR-Cas13a amplification reaction. Nat Commun 2020, 11, 267. [Google Scholar] [CrossRef]
- Wang, D.; Chen, G.; Lyu, Y.; Feng, E.; Zhu, L.; Pan, C.; Zhang, W.; Liu, X.; Wang, H. A CRISPR/Cas12a-based DNAzyme visualization system for rapid, non-electrically dependent detection of Bacillus anthracis. Emerg Microbes Infect 2022, 11, 428–437. [Google Scholar] [CrossRef]
- Sun, Y.; Li, J.; Zhu, L.; Jiang, L. Cooperation and competition between CRISPR- and omics-based technologies in foodborne pathogens detection: a state of the art review. Current Opinion in Food Science 2022, 44, 100813. [Google Scholar] [CrossRef]
- Wu, Y.; Liu, J.; Li, H.-t.; Zhang, T.; Dong, Y.; Deng, S.; Lv, Y.; He, Q.; Deng, R. CRISPR-Cas system meets DNA barcoding: Development of a universal nucleic acid test for food authentication. Sensors and Actuators B: Chemical 2022, 353, 131138. [Google Scholar] [CrossRef]
- Devillars, A.; Magon, G.; Pirrello, C.; Palumbo, F.; Farinati, S.; Barcaccia, G.; Lucchin, M.; Vannozzi, A. Not Only Editing: A Cas-Cade of CRISPR/Cas-Based Tools for Functional Genomics in Plants and Animals. In International Journal of Molecular Sciences, 2024; Vol. 25.
- Qi, L.S.; Larson, M.H.; Gilbert, L.A.; Doudna, J.A.; Weissman, J.S.; Arkin, A.P.; Lim, W.A. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 2013, 152, 1173–1183. [Google Scholar] [CrossRef]
- Park, H.M.; Liu, H.; Wu, J.; Chong, A.; Mackley, V.; Fellmann, C.; Rao, A.; Jiang, F.; Chu, H.; Murthy, N. , et al. Extension of the crRNA enhances Cpf1 gene editing in vitro and in vivo. Nat Commun 2018, 9, 3313. [Google Scholar] [CrossRef] [PubMed]
- Pickar-Oliver, A.; Gersbach, C.A. The next generation of CRISPR-Cas technologies and applications. Nat Rev Mol Cell Biol 2019, 20, 490–507. [Google Scholar] [CrossRef]
- Chen, J.S.; Ma, E.; Harrington, L.B.; Da Costa, M.; Tian, X.; Palefsky, J.M.; Doudna, J.A. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 2018, 360, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Schultzhaus, Z.; Wang, Z.; Stenger, D. CRISPR-based enrichment strategies for targeted sequencing. Biotechnol Adv 2021, 46, 107672. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Xia, Q.; Zhang, S.; Gao, J.; Dai, W.; Wu, J.; Wang, J. CRISPR-assisted targeted enrichment-sequencing (CATE-seq). bioRxiv 2019, 672816. [Google Scholar] [CrossRef]
- Zhou, W.; Hu, L.; Ying, L.; Zhao, Z.; Chu, P.K.; Yu, X.F. A CRISPR-Cas9-triggered strand displacement amplification method for ultrasensitive DNA detection. Nat Commun 2018, 9, 5012. [Google Scholar] [CrossRef] [PubMed]
- Bennett-Baker, P.E.; Mueller, J.L. CRISPR-mediated isolation of specific megabase segments of genomic DNA. Nucleic Acids Res 2017, 45, e165. [Google Scholar] [CrossRef] [PubMed]
- Delong, R.K.; Zhou, Q. Introductory Experiments on Biomolecules and their Interactions; Academic Press: 2015.
- Cao, X.; Zhao, L.; Zhang, J.; Chen, X.; Shi, L.; Fang, X.; Xie, H.; Chang, Y.; Wang, L. Detection of viable but nonculturable Vibrio parahaemolyticus in shrimp samples using improved real-time PCR and real-time LAMP methods. Food Control 2019, 103, 145–152. [Google Scholar] [CrossRef]
- Bogozalec Kosir, A.; Demsar, T.; Stebih, D.; Zel, J.; Milavec, M. Digital PCR as an effective tool for GMO quantification in complex matrices. Food Chem 2019, 294, 73–78. [Google Scholar] [CrossRef]
- Lei, C.; Li, S.Y.; Liu, J.K.; Zheng, X.; Zhao, G.P.; Wang, J. The CCTL (Cpf1-assisted Cutting and Taq DNA ligase-assisted Ligation) method for efficient editing of large DNA constructs in vitro. Nucleic Acids Res 2017, 45, e74. [Google Scholar] [CrossRef]
- Li, S.Y.; Cheng, Q.X.; Liu, J.K.; Nie, X.Q.; Zhao, G.P.; Wang, J. CRISPR-Cas12a has both cis- and trans-cleavage activities on single-stranded DNA. Cell Res 2018, 28, 491–493. [Google Scholar] [CrossRef]
- Rath, D.; Amlinger, L.; Rath, A.; Lundgren, M. The CRISPR-Cas immune system: biology, mechanisms and applications. Biochimie 2015, 117, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Swiat, M.A.; Dashko, S.; den Ridder, M.; Wijsman, M.; van der Oost, J.; Daran, J.M.; Daran-Lapujade, P. FnCpf1: a novel and efficient genome editing tool for Saccharomyces cerevisiae. Nucleic Acids Res 2017, 45, 12585–12598. [Google Scholar] [CrossRef]
- Zalatan, J.G.; Lee, M.E.; Almeida, R.; Gilbert, L.A.; Whitehead, E.H.; La Russa, M.; Tsai, J.C.; Weissman, J.S.; Dueber, J.E.; Qi, L.S. , et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 2015, 160, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Fonfara, I.; Richter, H.; Bratovic, M.; Le Rhun, A.; Charpentier, E. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA. Nature 2016, 532, 517–521. [Google Scholar] [CrossRef] [PubMed]
- Creutzburg, S.C.A.; Swartjes, T.; van der Oost, J. Medium-throughput in vitro detection of DNA cleavage by CRISPR-Cas12a. Methods 2020, 172, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, J.; Zeng, H.; Liu, X.; Jiang, W.; Wang, Y.; Ouyang, W.; Tang, X. RPA-Cas12a-FS: A frontline nucleic acid rapid detection system for food safety based on CRISPR-Cas12a combined with recombinase polymerase amplification. Food Chem 2021, 334, 127608. [Google Scholar] [CrossRef] [PubMed]
- Deng, R.; Xu, L.; Zhang, Y.; Zhang, X.; Yuan, Z.; Chen, J.; Xia, X. CRISPR-based nucleic acid assays for food authentication. Trends in Food Science & Technology 2024, 145, 104351. [Google Scholar] [CrossRef]
- Zetsche, B.; Gootenberg, J.S.; Abudayyeh, O.O.; Slaymaker, I.M.; Makarova, K.S.; Essletzbichler, P.; Volz, S.E.; Joung, J.; van der Oost, J.; Regev, A. , et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015, 163, 759–771. [Google Scholar] [CrossRef] [PubMed]
- Li, S.Y.; Zhao, G.P.; Wang, J. C-Brick: A New Standard for Assembly of Biological Parts Using Cpf1. ACS Synth Biol 2016, 5, 1383–1388. [Google Scholar] [CrossRef] [PubMed]
- Myhrvold, C.; Freije, C.A.; Gootenberg, J.S.; Abudayyeh, O.O.; Metsky, H.C.; Durbin, A.F.; Kellner, M.J.; Tan, A.L.; Paul, L.M.; Parham, L.A. , et al. Field-deployable viral diagnostics using CRISPR-Cas13. Science 2018, 360, 444–448. [Google Scholar] [CrossRef]
- Broughton, J.P.; Deng, X.; Yu, G.; Fasching, C.L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J.A.; Granados, A.; Sotomayor-Gonzalez, A. , et al. CRISPR-Cas12-based detection of SARS-CoV-2. Nat Biotechnol 2020, 38, 870–874. [Google Scholar] [CrossRef]
- Chen, Y.; Wu, S.; Wu, H.; Cheng, P.; Wang, X.; Qian, S.; Zhang, M.; Xu, J.; Ji, F.; Wu, J. CRISPR/Cas12a-Based Versatile Method for Checking Quantitative Polymerase Chain Reaction Samples with Cycles of Threshold Values in the Gray Zone. ACS Sens 2021, 6, 1963–1970. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Mei, Y.; Jiang, X. Universal and high-fidelity DNA single nucleotide polymorphism detection based on a CRISPR/Cas12a biochip. Chem Sci 2021, 12, 4455–4462. [Google Scholar] [CrossRef]
- Wu, H.; He, J.S.; Zhang, F.; Ping, J.; Wu, J. Contamination-free visual detection of CaMV35S promoter amplicon using CRISPR/Cas12a coupled with a designed reaction vessel: Rapid, specific and sensitive. Anal Chim Acta 2020, 1096, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Cao, G.; Dong, J.; Chen, X.; Lu, P.; Xiong, Y.; Peng, L.; Li, J.; Huo, D.; Hou, C. Simultaneous detection of CaMV35S and T-nos utilizing CRISPR/Cas12a and Cas13a with multiplex-PCR (MPT-Cas12a/13a). Chem Commun (Camb) 2022, 58, 6328–6331. [Google Scholar] [CrossRef]
- Liang, Z.; Chen, K.; Yan, Y.; Zhang, Y.; Gao, C. Genotyping genome-edited mutations in plants using CRISPR ribonucleoprotein complexes. Plant Biotechnol J 2018, 16, 2053–2062. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Liu, X.; Yang, J.; Wang, Z.; Wang, H.; Wang, X. CRISPR/Cas12a-based biosensing platform for the on-site detection of single-base mutants in gene-edited rice. Front Plant Sci 2022, 13, 944295. [Google Scholar] [CrossRef] [PubMed]
- Schaart, J.G.; van de Wiel, C.C.M.; Smulders, M.J.M. Genome editing of polyploid crops: prospects, achievements and bottlenecks. Transgenic Res 2021, 30, 337–351. [Google Scholar] [CrossRef] [PubMed]
- Teng, F.; Guo, L.; Cui, T.; Wang, X.G.; Xu, K.; Gao, Q.; Zhou, Q.; Li, W. CDetection: CRISPR-Cas12b-based DNA detection with sub-attomolar sensitivity and single-base specificity. Genome Biol 2019, 20, 132. [Google Scholar] [CrossRef]
- Harrington, L.B.; Burstein, D.; Chen, J.S.; Paez-Espino, D.; Ma, E.; Witte, I.P.; Cofsky, J.C.; Kyrpides, N.C.; Banfield, J.F.; Doudna, J.A. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 2018, 362, 839–842. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).