Submitted:
08 July 2024
Posted:
10 July 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Experimental
Participants Profile
- I.
- The data quality rating was Q2 or Q3. This excluded 9 from the main dataset (see section on data quality).
- II.
- The Covid-19 PCR swab test was negative, but the patient was symptomatic (these were tagged as “suspected” Covid-19). This accounted for 1 case in the.
Data Collection
Machine Learning Algorithm
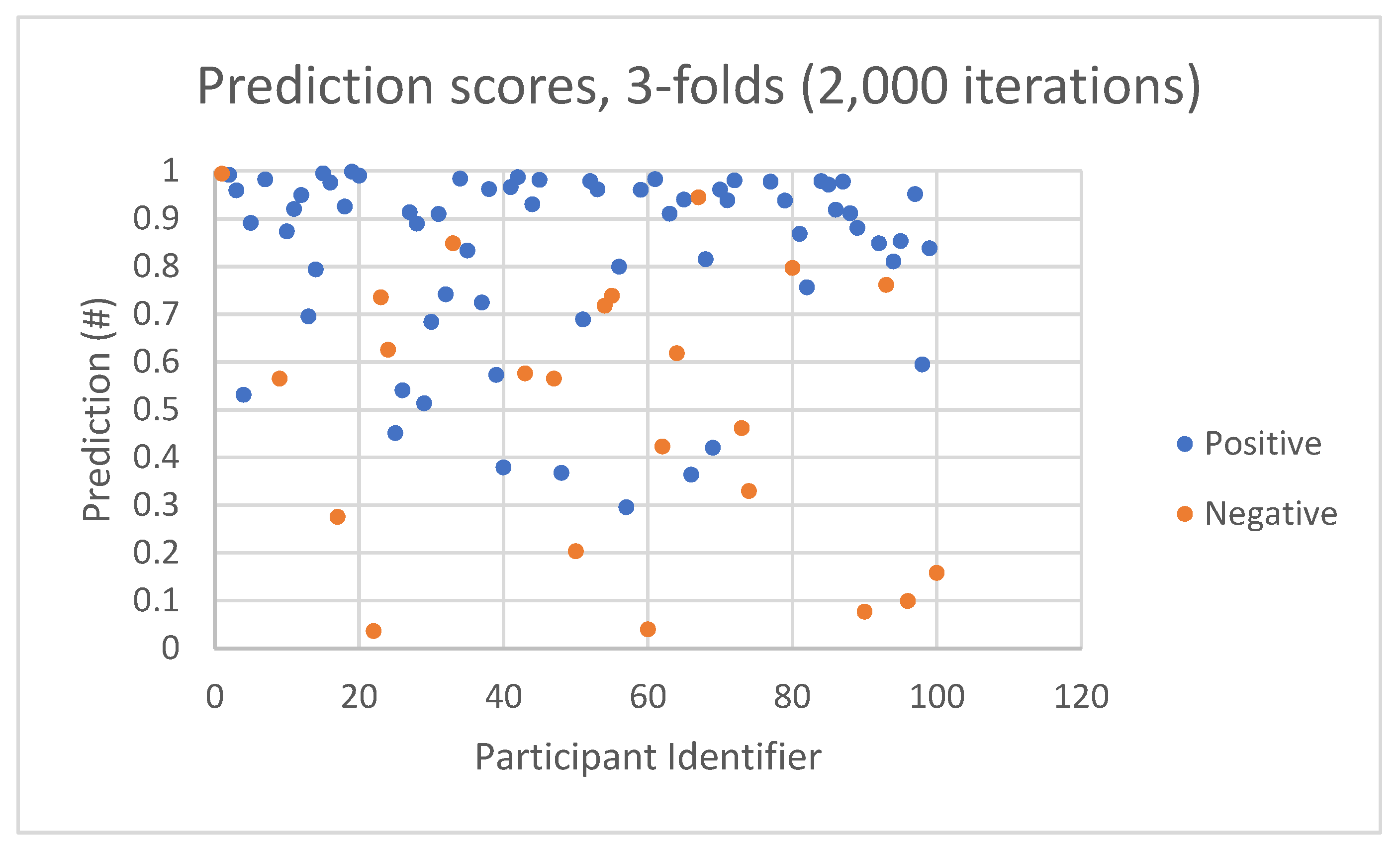
Results and Discussion
- The accuracy of the reference method’s diagnostics (RT-PCR) may not be 100%, see Supplementary Materials.
- The number of participants may be an issue. It’s well known that increasing the number of participants increases the accuracy of the prediction score.
Supplementary Materials
Acknowledgements
Conflicts of Interest
References
- Amman A, Smith D. Volatile biomarkers. Amsterdam: Elsevier BV; 2013.
- Arevalo-Rodriguez I, Buitrago-Garcia D, Simancas-Racines D, et al. (2020). False-negative results of initial RT-PCR assays for covid-19: a systematic review. medRxiv 20066787. 2020. [CrossRef]
- Barker, M., & Rayens, W. (2003). Partial least squares for discrimination. Journal of Chemometrics, 17(3), 166–173. [CrossRef]
- Beauchamp J, Davis C, Pleil J. (2020). Breathborne biomarkers and the human volatilome. Elsevier Science, 2nd Edition; 722p. ISBN: 9780128199671.
- Beliza’rio J. E., J Faintuch and M G Malpartida (2021). Breath Biopsy and Discovery of Exclusive Volatile Organic Compounds for Diagnosis of Infectious Diseases. Frontiers in Cellular and Infection Microbiology; 10. [CrossRef]
- Bourgon, R., Gentleman, R., & Huber, W. (2010). Independent filtering increases detection power for high-throughput experiments. Proceedings of the National Academy of Sciences of the United States of America, 107(21), 9546–9551. [CrossRef]
- Breiman, L. (2001). Random forests. Machine Learning, 45(1), 5–32.
- Bzdok D, Altman N, Krzywinski M. (2018). Statistics versus Machine Learning. Nature Methods; 15:233-234.
- Chace, D. H. (2001). Mass spectrometry in the clinical laboratory. Chemical Reviews, 101(2), 445–477. [CrossRef]
- Cortes, C., & Vapnik, V. (1995). Support-vector networks. Machine Learning, 20(3), 273–297.
- Cover, T. M. and Thomas, J. A. Elements of Information Theory. John Wiley & Sons, 2nd edition, 2006.
- Davis C. E., M. Schivo, N. J. Kenyon (2021). A breath of fresh air; the potential for COVID-19 breath diagnostics. EBioMedicine, 63. [CrossRef]
- Eriksson, L., Johansson, E., Kettaneh-Wold, N., & Wold, S. (2001). Multi- and Megavariate Data Analysis Principles and Applications. Dublin: Umetrics Academy.
- Grassin-Delyle S., C. Roquencourt, P. Moine, G. Saffroy, Stanislas Carn, N. Heming, J. Fleuriet, H. Salvator, E. Naline, l-J Couderc, P. Devillier, E. A. Thevenot, D. Annane (2021). Metabolomics of exhaled breath in critically ill COVID-19 patients: A pilot study EBioMedicine, 63. [CrossRef]
- Jianguo Xia, David I. Broadhurst, Michael Wilson, David S. Wishart (2012). Translational biomarker discovery in clinical metabolomics: an introductory tutorial, Metabolomics. [CrossRef]
- Hackstadt, A. J., & Hess, A. M. (2009). Filtering for increased power for microarray data analysis. BMC Bioinformatics, 10, 11.
- Health Information and Quality Authority (2020). Evidence summary for SARS-CoV-2 viral load and infectivity over the course of an infection. 9 June 2020, Health Information and Quality Authority. Report. https://www.hiqa.ie/sites/default/files/2020-06/Evidence-Summary_SARS- CoV-2-duration-of-infectivity.pdf.
- Kohl, S. M., Klein, M. S., Hochrein, J., Oefner, P. J., Spang, R., & Gronwald, W. (2012). State-of-the art data normalization methods improve NMR-based metabolomic analysis. Metabolo- mics, 8(Suppl 1), 146–160.
- Kucirka L. M.; Lauer S. A.; Laeyendecker O.; Boon D.; and Lessler J. (2020). Variation in False-Negative Rate of Reverse Transcriptase Polymerase Chain Reaction–Based SARS-CoV-2 Tests by Time Since Exposure. Annals of Internal Medicine. [CrossRef]
- Ruszkiewicz DM, Sanders D, O’Brien R, Hempel F, Reed, MJ Riepe,AC, Bailie K, Brodrick E, Darnley, K Ellerkmann, R Mueller, O Skarysz A, Truss M, Wortelmann T, Yordanov S, Thomas CLP, Schaaf B & Eddleston M (2020). Diagnosis of COVID-19 by analysis of breath with gas chromatography-ion mobility spectrometry - a feasibility study. EClinicalMedicine, 29-30, 100609; (http://creativecommons.org/licenses/by-nc-nd/4.0/).
- Saeys, Y., Inza, I., & Larranaga, P. (2007). A review of feature selection techniques in bioinformatics. Bioinformatics, 23(19), 2507–2517.
- Sethuraman N, Sundararaj Stanleyraj J, Ryo A. (2020). Interpreting diagnostic tests for SARS-CoV-2. JAMA 2020. [CrossRef]
- Szymanska, E., Saccenti, E., Smilde, A. K., & Westerhuis, J. A. (2012). Double-check: Validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics, 8(Sup- pl 1), 3–16.
- Tang Y-W, J E. Schmitz, D H. Persing, C W. Strattonb (2020). Laboratory Diagnosis of COVID-19: Current Issues and Challenges; Journal of Clinical Microbiology, 58, p. 1-9.
- To, K.K.W., Tsang, O.T.Y., Leung, W.S., Tam, A.R., Wu, T.C., Lung, D.C., Yip, C.C.Y., Cai, J.P., Chan, J.M.C., Chik, T.S.H., Lau, D.P.L., Choi, C.Y.C., Chen, L.L., Chan, W.M., Chan, K.H., Ip, J.D., Ng, A.C.K., Poon, R.W.S., Luo, C.T.,... Yuen, K.Y. (2020). Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. Lancet Infect. Dis. 20, 565–574. [CrossRef]
- Trygg, J., Holmes, E., & Lundstedt, T. (2007). Chemometrics in metabolomics. Journal of Proteome Research, 6(2), 469–479.
- van den Berg, R. A., Hoefsloot, H. C., Westerhuis, J. A., Smilde, A. K., & van der Werf, M. J. (2006). Centering, scaling, and transformations: Improving the biological information content of metabolomics data. BMC Genomics, 7, 142.
- Wilcken, B., Wiley, V., Hammond, J., & Carpenter, K. (2003). Screening newborns for inborn errors of metabolism by tandem mass spectrometry. New England Journal of Medicine, 348(23), 2304–2312.
- Wölfel R, Corman VM, Guggemos W, et al. (2020). Virological assessment of hospitalized patients with COVID-2019. Nature 2020. [CrossRef]
| Trial location | ROCAUC (95% CI) | TPR % | FPR % | Precision % (±σ) | Accuracy % (±σ) |
|---|---|---|---|---|---|
| MUSC | 84.1 (1.3) | 76.0 | 14.8 | 93.6 (5.7) | 78.4 (4.8) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




