Submitted:
01 July 2024
Posted:
02 July 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Sample Collection and Preservation
2.2. DNA sample preparation
2.3. Library Construction and high-throughput DNA sequencing of 16S rDNA
2.4. Bioinformatics analysis
2.5. Statistical Analysis
2.6. Microbial network Construction and Analysis
3. Results
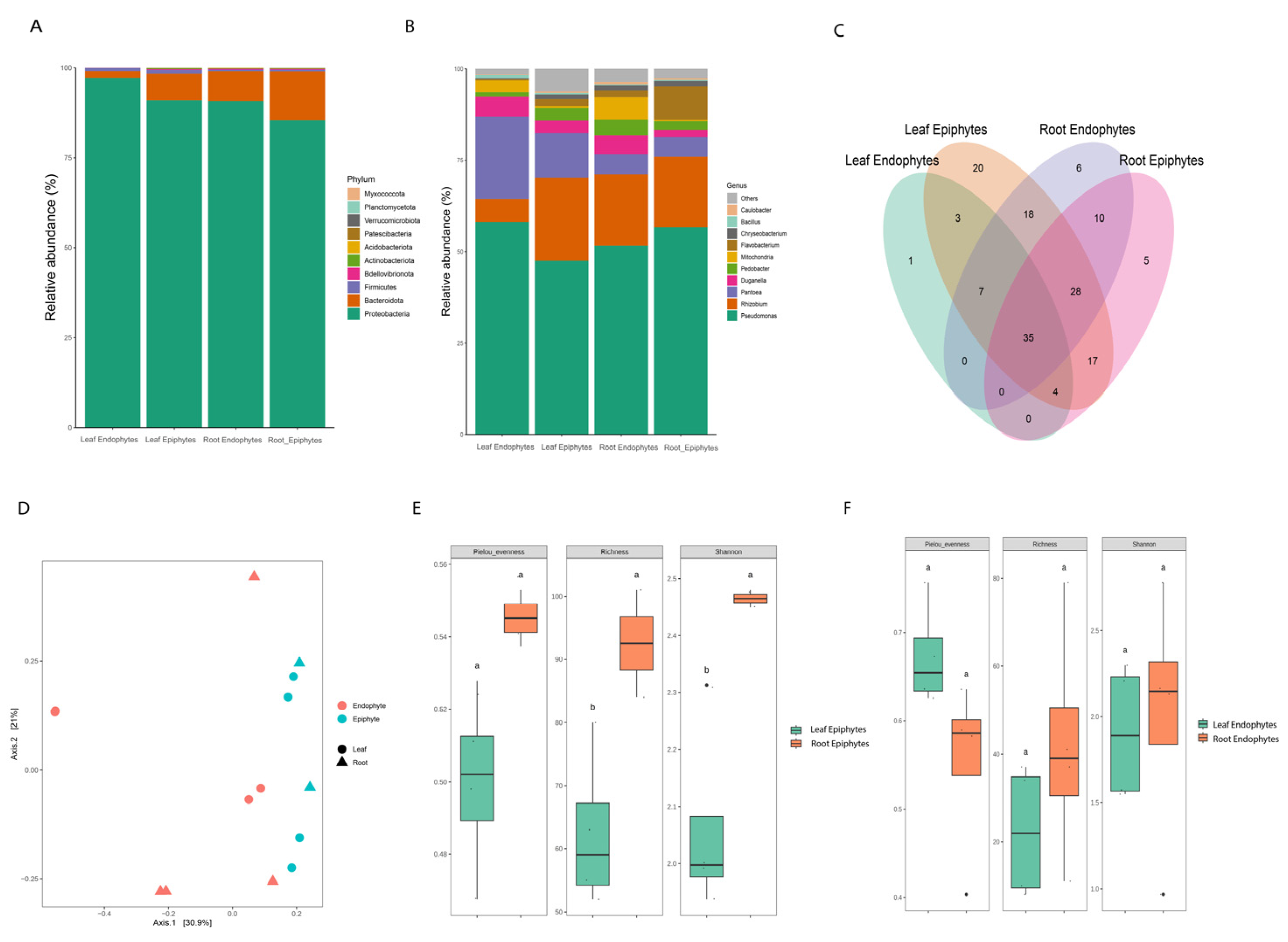
3.1. Microbial Community Composition and Diversity
3.2. Shared and Unique ASVs and Keystone Taxa
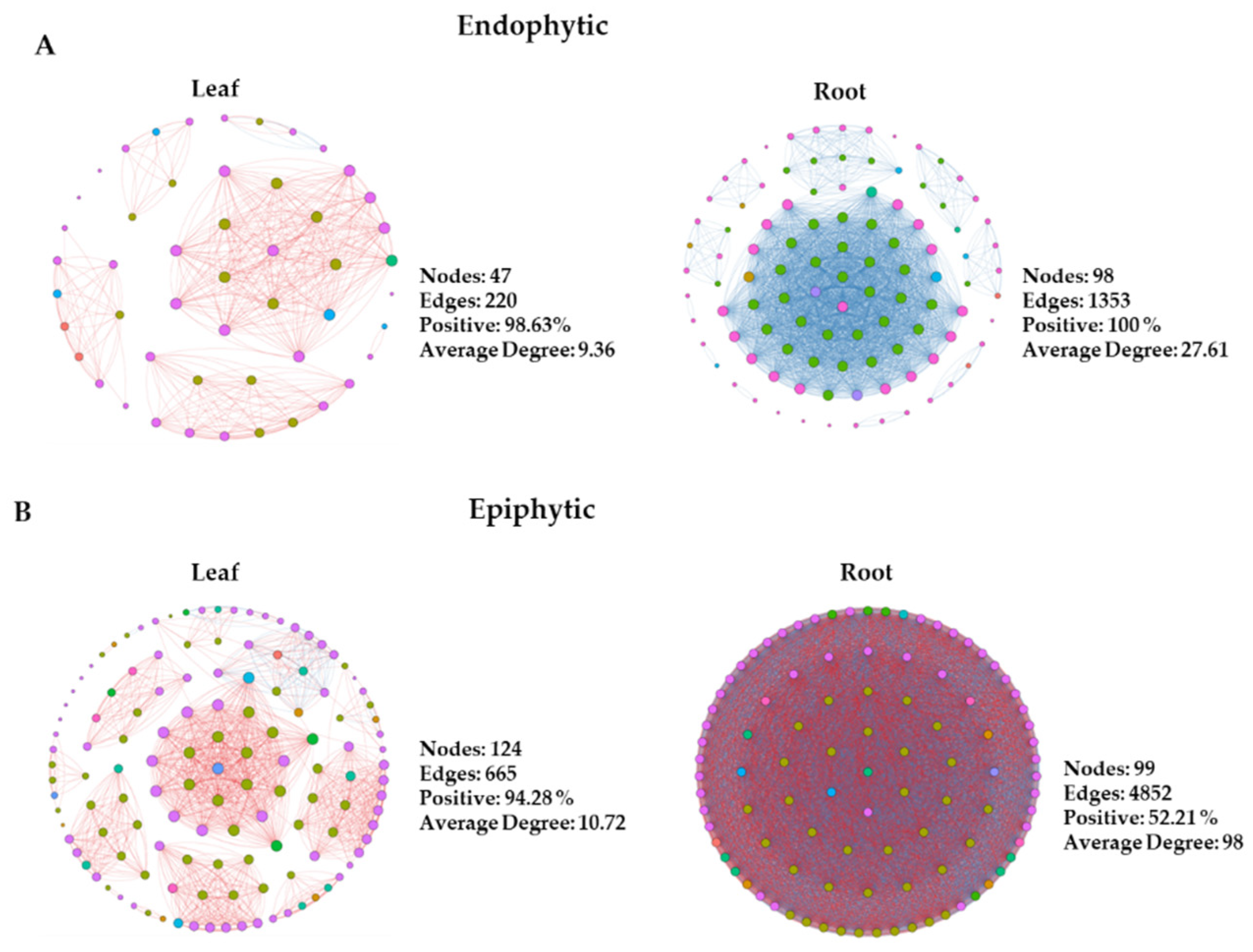
3.3. Microbial co-occurrence network
3.5. Bacterial Diversity Analyses Differentiate Communities among Habitats
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mcowen, C.J., et al., A global map of saltmarshes. Biodiversity data journal, 2017(5). [CrossRef]
- Rolando, J.L., et al., The core root microbiome of Spartina alterniflora is predominated by sulfur-oxidizing and sulfate-reducing bacteria in Georgia salt marshes, USA. Microbiome, 2022. 10(1): p. 37. [CrossRef]
- He, Q., B.R. Silliman, and B. Cui, Incorporating thresholds into understanding salinity tolerance: A study using salt-tolerant plants in salt marshes. Ecology and evolution, 2017. 7(16): p. 6326-6333. [CrossRef]
- Soldan, R., et al., Bacterial endophytes of mangrove propagules elicit early establishment of the natural host and promote growth of cereal crops under salt stress. Microbiological research, 2019. 223: p. 33-43. [CrossRef]
- Berg, G., et al., The plant microbiome explored: implications for experimental botany. Journal of experimental botany, 2016. 67(4): p. 995-1002. [CrossRef]
- Zheng, Y., et al., Patterns in the microbial community of salt-tolerant plants and the functional genes associated with salt stress alleviation. Microbiology Spectrum, 2021. 9(2): p. e00767-21. [CrossRef]
- Bodenhausen, N., M.W. Horton, and J. Bergelson, Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PloS one, 2013. 8(2): p. e56329. [CrossRef]
- Yang, X., et al., Effects of salinity on assembly characteristics and function of microbial communities in the phyllosphere and rhizosphere of salt-tolerant Avicennia marina mangrove species. Microbiology Spectrum, 2023. 11(2): p. e03000-22. [CrossRef]
- Guo, J., et al., Roles of endophytic bacteria in Suaeda salsa grown in coastal wetlands: Plant growth characteristics and salt tolerance mechanisms. Environmental Pollution, 2021. 287: p. 117641. [CrossRef]
- Liu, H., et al., Microbiome-mediated stress resistance in plants. Trends in plant science, 2020. 25(8): p. 733-743. [CrossRef]
- Qu, X., et al., Response of Phyllosphere and Rhizosphere Microbial Communities to Salt Stress of Tamarix chinensis. Plants, 2024. 13(8): p. 1091. [CrossRef]
- Kim, K., et al., Alleviation of salt stress by Enterobacter sp. EJ01 in tomato and Arabidopsis is accompanied by up-regulation of conserved salinity responsive factors in plants. Molecules and cells, 2014. 37(2): p. 109-117. [CrossRef]
- Sultana, S., et al., Isolation and identification of salt-tolerant plant-growth-promoting rhizobacteria and their application for rice cultivation under salt stress. Canadian journal of microbiology, 2020. 66(2): p. 144-160. [CrossRef]
- Ullah, S. and A. Bano, Isolation of plant-growth-promoting rhizobacteria from rhizospheric soil of halophytes and their impact on maize (Zea mays L.) under induced soil salinity. Canadian journal of microbiology, 2015. 61(4): p. 307-313.
- Yasmin, H., et al., Halotolerant rhizobacteria Pseudomonas pseudoalcaligenes and Bacillus subtilis mediate systemic tolerance in hydroponically grown soybean (Glycine max L.) against salinity stress. PLoS One, 2020. 15(4): p. e0231348. [CrossRef]
- Travis, S.E., M.R. Simon, and G.P. Zogg, Environmental Sensitivity of Soil Microbial Communities is Altered in Association with Plant Roots in Saltmarsh Ecosystems. Northeastern Naturalist, 2024. 31(2): p. 146-162. [CrossRef]
- Jiang, H., et al., High-throughput 16S rRNA gene-based amplicon sequencing reveals the functional divergence of halophilic bacterial communities in the Suaeda salsa root compartments on the Eastern Coast of China. Science of The Total Environment, 2024: p. 173775. [CrossRef]
- Rush, G.I., B. Clark, and C.Y. Lumibao, Biotic versus abiotic factors shaping culturable root endosymbionts of the saltmarsh halophyte, Batis maritima and implications for plant stress tolerance. Wetlands Ecology and Management, 2024: p. 1-9. [CrossRef]
- Chen, S., et al., fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics, 2018. 34(17): p. i884-i890. [CrossRef]
- Bolyen, E., et al., Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature biotechnology, 2019. 37(8): p. 852-857. [CrossRef]
- McDonald, D., et al., An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. The ISME journal, 2012. 6(3): p. 610-618. [CrossRef]
- Oksanen, J., et al., Package ‘vegan’. Community ecology package, version, 2018. 2(3).
- Liu, C., et al., microeco: an R package for data mining in microbial community ecology. FEMS microbiology ecology, 2021. 97(2): p. fiaa255. [CrossRef]
- Wen, T., et al., ggClusterNet: An R package for microbiome network analysis and modularity-based multiple network layouts. Imeta, 2022. 1(3): p. e32. [CrossRef]
- Tian, R., et al., Small and mighty: adaptation of superphylum Patescibacteria to groundwater environment drives their genome simplicity. Microbiome, 2020. 8: p. 1-15. [CrossRef]
- Mateus, M., et al., Conflictive uses of coastal areas: A case study in a southern European coastal lagoon (Ria de Alvor, Portugal). Ocean & coastal management, 2016. 132: p. 90-100. [CrossRef]
- Zheng, W., et al., The responses and adaptations of microbial communities to salinity in farmland soils: a molecular ecological network analysis. Applied Soil Ecology, 2017. 120: p. 239-246. [CrossRef]
- Szymańska, S., et al., Bacterial microbiome of root-associated endophytes of Salicornia europaea in correspondence to different levels of salinity. Environmental Science and Pollution Research, 2018. 25: p. 25420-25431. [CrossRef]
- Chen, X., et al., Salt-tolerant plant moderates the effect of salinity on soil organic carbon mineralization in a subtropical tidal wetland. Science of The Total Environment, 2022. 837: p. 155855. [CrossRef]
- Lenk, S., et al., Novel groups of Gammaproteobacteria catalyse sulfur oxidation and carbon fixation in a coastal, intertidal sediment. Environmental Microbiology, 2011. 13(3): p. 758-774.
- Wang, K., et al., Potential ecological impacts of physical control on Spartina alterniflora in coastal wetland: Migration and transformation of nutrients and the response of bacterial community structure. Journal of Cleaner Production, 2023. 398: p. 136556. [CrossRef]
- Kim, C., et al., Microbial community succession along a chronosequence in constructed salt marsh soils. Microbial ecology, 2023. 85(3): p. 931-950. [CrossRef]
- An, X., et al., Rhizosphere bacterial diversity and environmental function prediction of wild salt-tolerant plants in coastal silt soil. Ecological Indicators, 2022. 134: p. 108503. [CrossRef]
- Petrosyan, K., et al., Diversity and potential plant growth promoting capacity of seed endophytic bacteria of the holoparasite Cistanche phelypaea (Orobanchaceae). Scientific Reports, 2023. 13(1): p. 11835. [CrossRef]
- Doni, F., et al., Associations of Pantoea with rice plants: as friends or foes? Agriculture, 2021. 11(12): p. 1278. [CrossRef]
- Guo, B., et al., Microbial co-occurrence network topological properties link with reactor parameters and reveal importance of low-abundance genera. npj Biofilms and Microbiomes, 2022. 8(1): p. 3.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



