Submitted:
14 June 2024
Posted:
14 June 2024
You are already at the latest version
Abstract
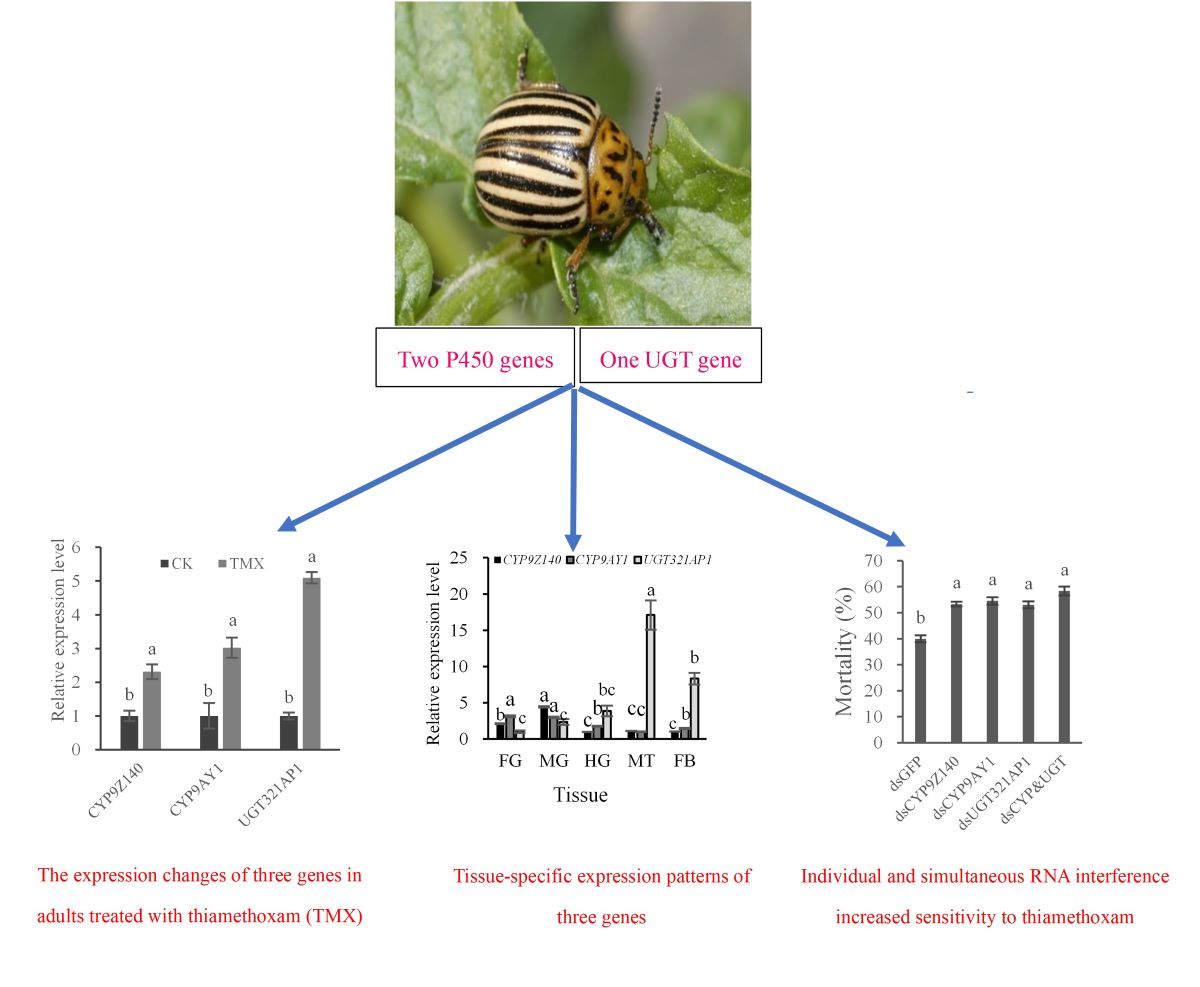
Keywords:
1. Introduction
2. Materials and Methods
2.1. Insects
2.2. Bioassay
2.3. RNA-Sequencing Data Analysis
2.4. Sequence and Phylogenetic Analysis
2.5. Preparation of Samples for Expression Analysis
2.6. Total RNA Isolation and cDNA Synthesis
2.7. Real-Time Quantitative PCR
2.8. RNA Interference
2.9. Statistical Analysis
3. Results
3.1. Resistance Levels of Leptinotarsa Decemlineata Populations to Thiamethoxam
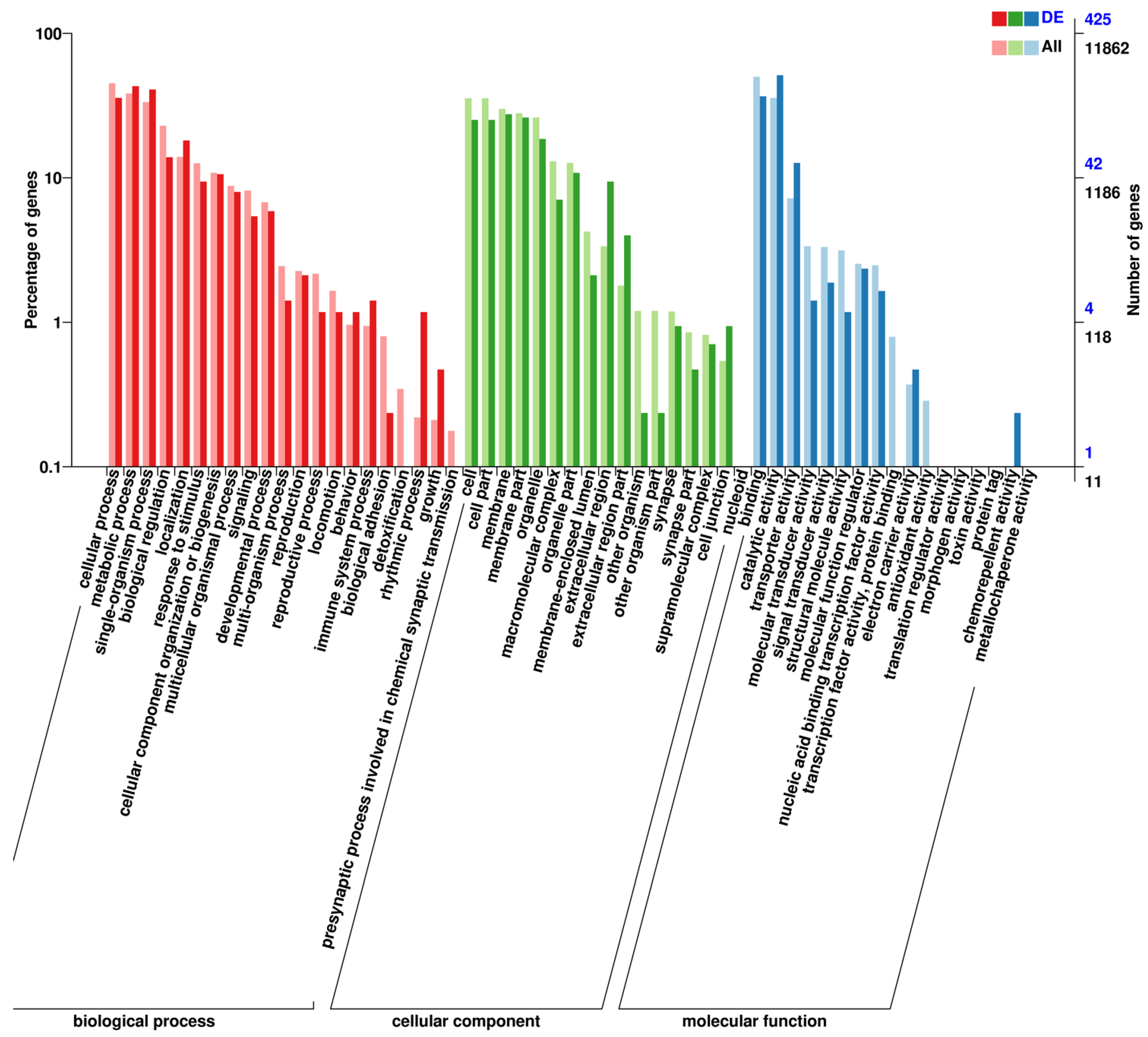
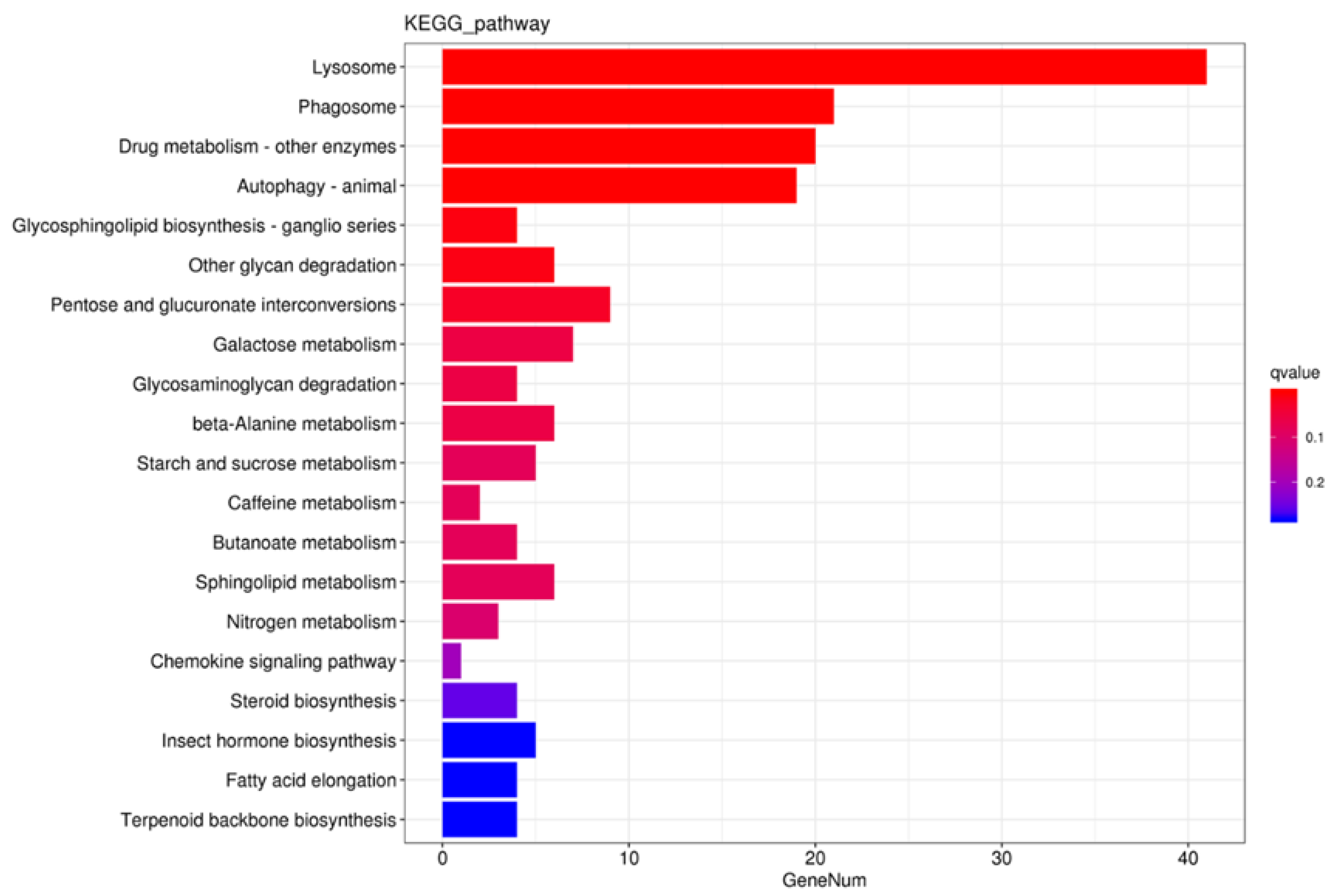
3.2. Transcriptome Analysis
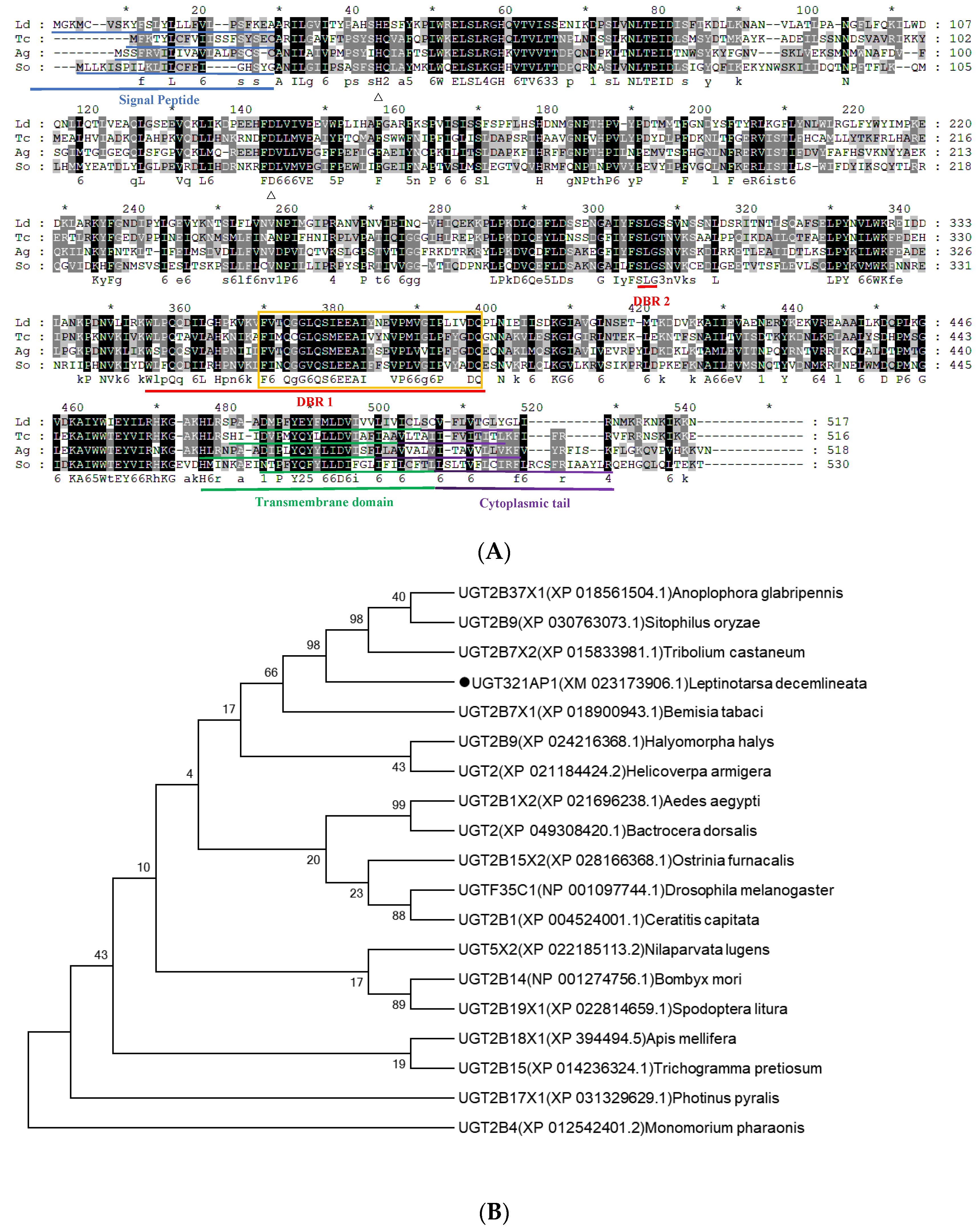
3.3. Gene Structure and Phylogenetic Analysis
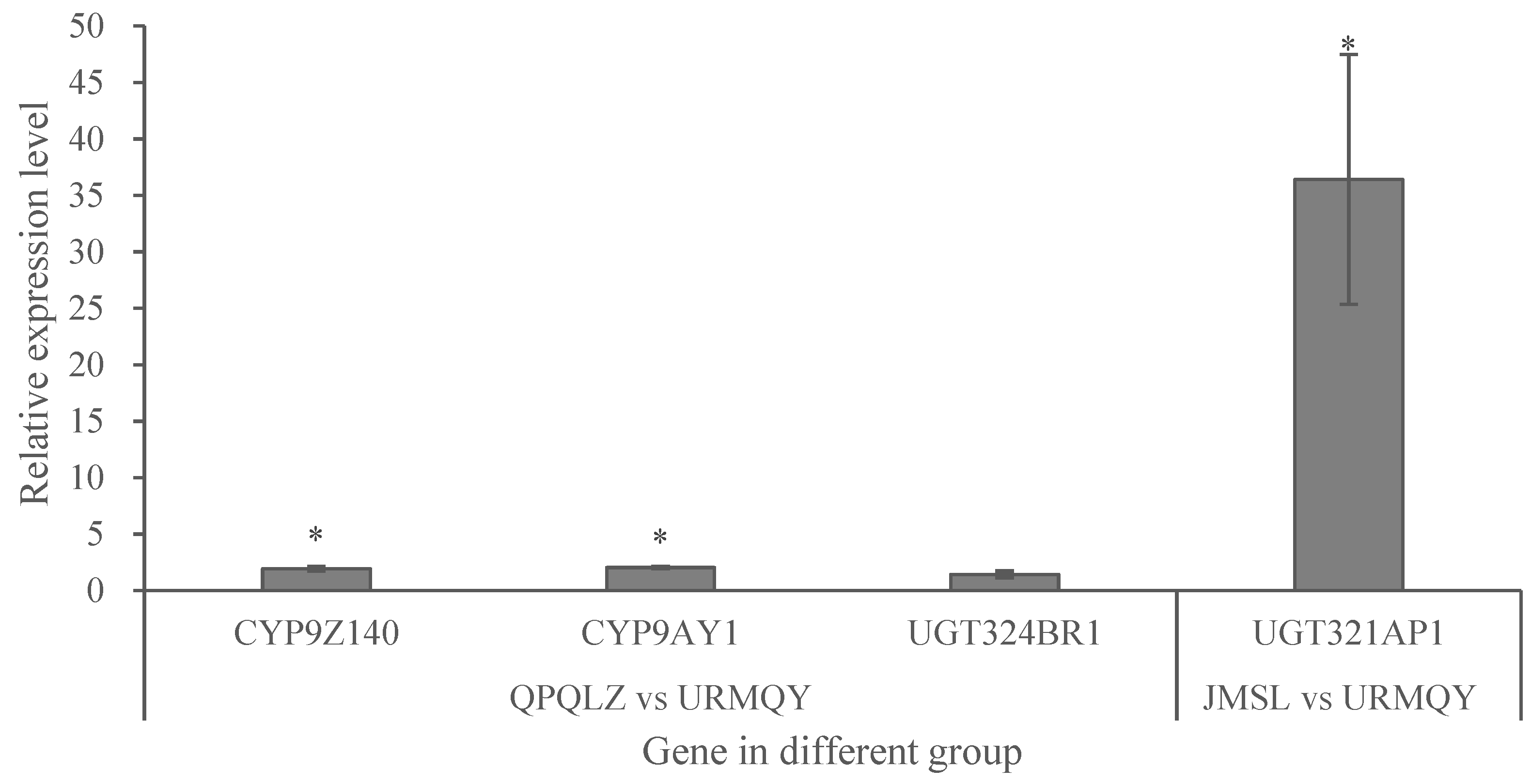
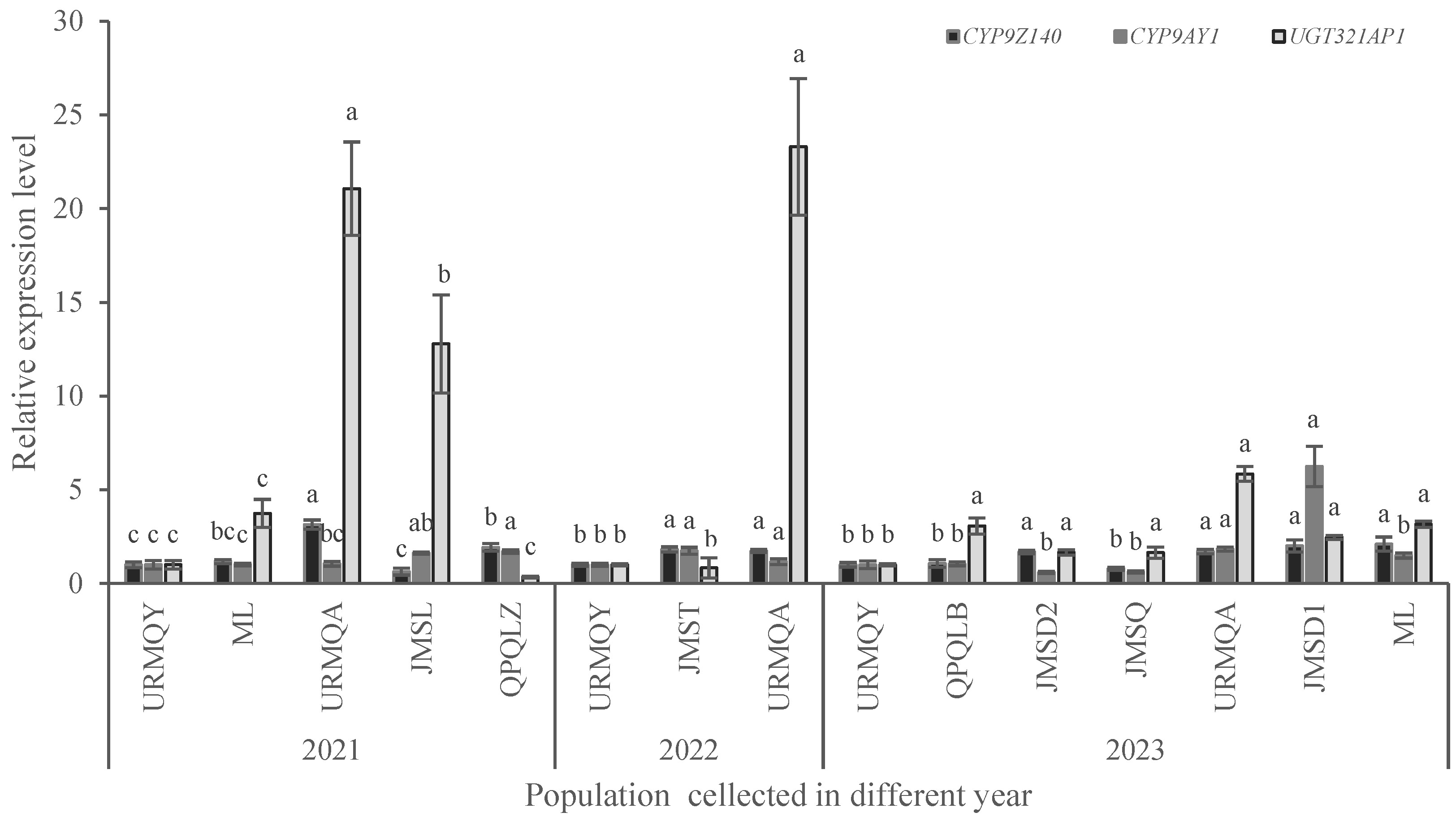
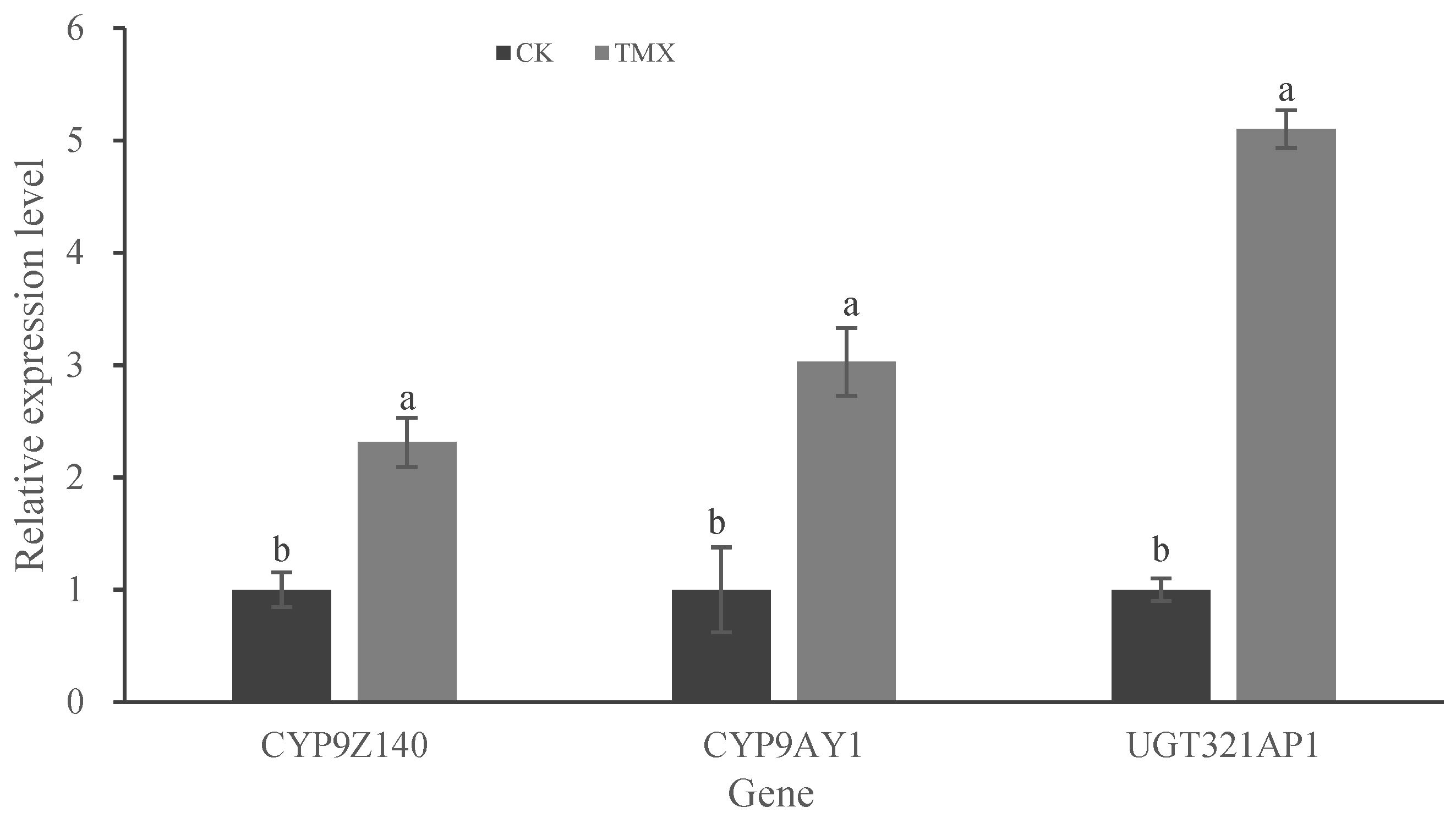
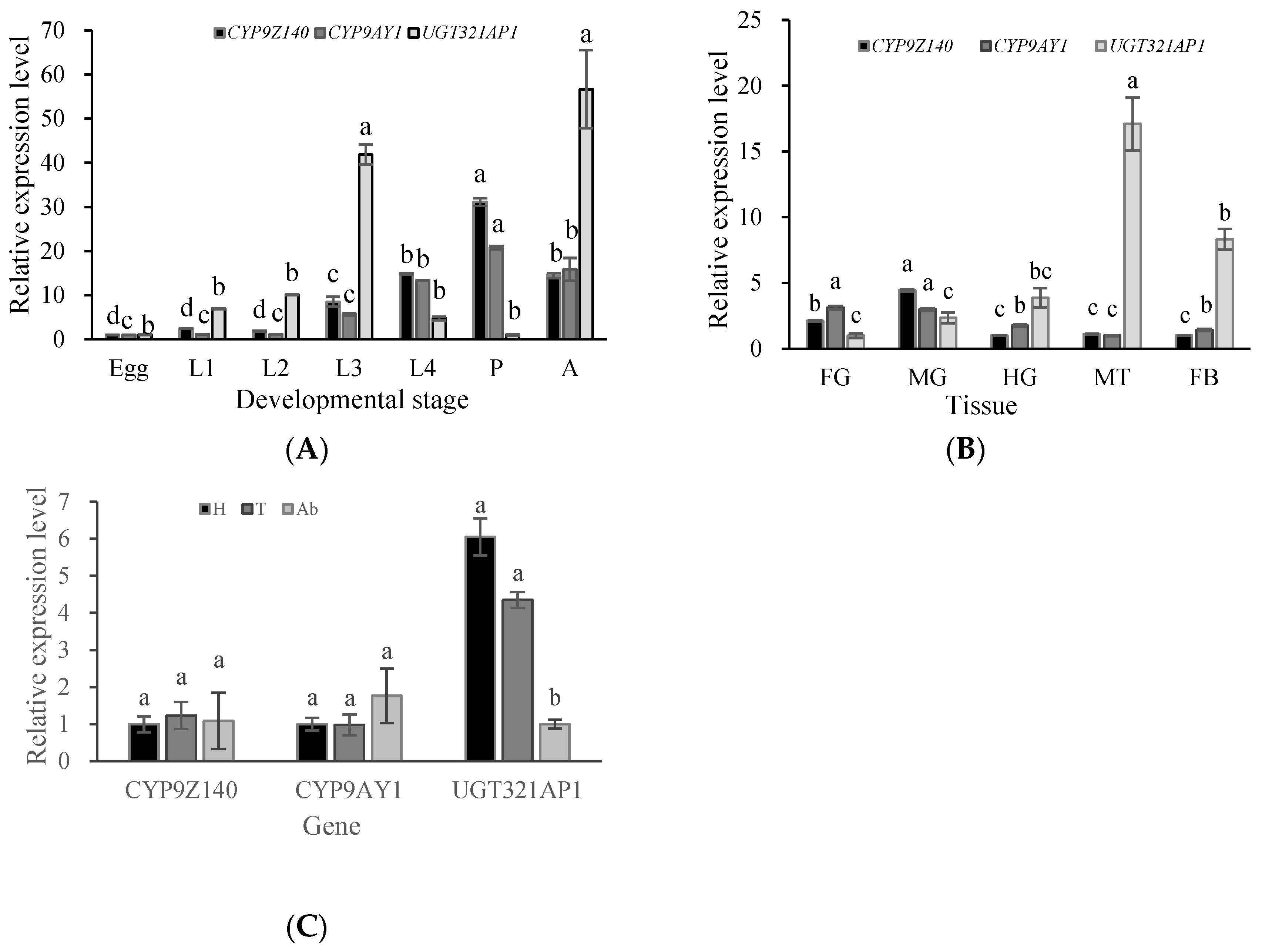
3.4. Expression Analysis of P450 and UGT Genes
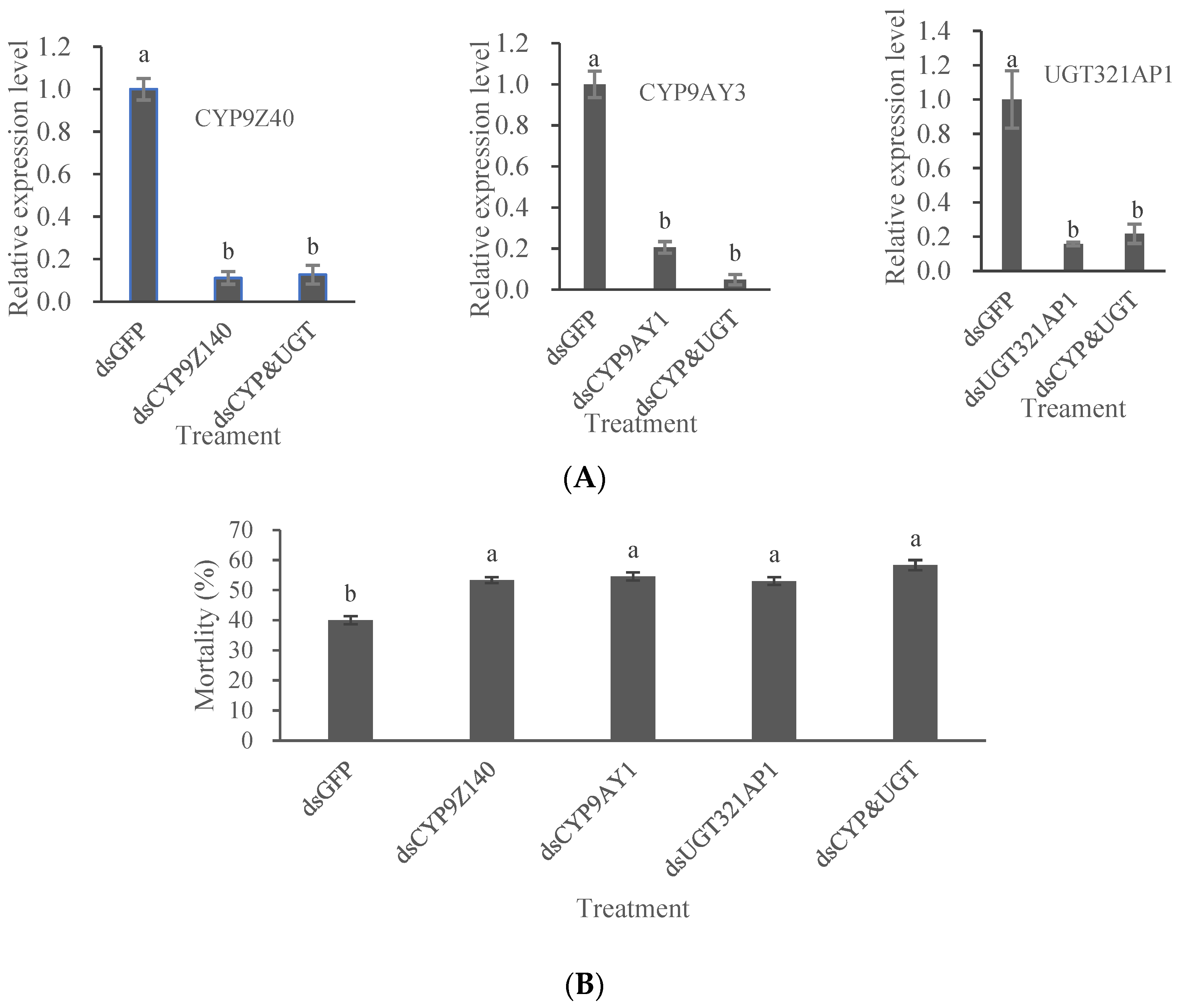
3.5. RNAi Effects of CYP9Z140, CYP9AY1, and UGT321AP on Leptinotarsa Decemlineata
4. Discussion
5. Conclusions
Funding
References
- Yan, J.J., Guo, W.C., Li, G.Q., Pan, H.P., Chen, B., Hou, W.W., Li, S.H., Gao, Y.L. Current status and prospects of the management of important insect pests on potato in China. Plant Protec. 2023, 49(5):190-195. [CrossRef]
- Liu, P., Jiang, W.H., Lu, W.P., Li, G.Q. Susceptibility of Colorado potato beetle Leptinotarsa decemlineata adults from northern Xinjiang Uygur autonomous region to four neonicotinoids. Chinese J. Pestic. Sci. 2011, (13), 271-275.
- Shi, X., Li, S., Wang, Z.M., Fu, K.Y., Fu, W.J., Jiang, W.H. Resistance monitoring to thiamethoxam and expression analysis of cytochrome P450 genes in Leptinotarsa decemlineata from Xinjiang. Scientia Agricultura Sinica. 2021, 54(14), 3004-3016. [CrossRef]
- Liu, Z.W., Williamson, M.S., Lansdell, S.J., Denholm, I., Han, Z.J., Millar, N.S. A nicotinic acetylcholine receptor mutation conferring target-site resistance to imidacloprid in Nilaparvata lugens (brown planthopper). Proc. Natl. Acad. Sci. USA 2005, 102(24), 8420-8425. [CrossRef]
- Hirata, K., Jouraku, A., Kuwazaki, S., Kanazawa, J., Iwasa, T. The R81T mutation in the nicotinic acetylcholine receptor of Aphis gossypii is associated with neonicotinoid insecticide resistance with differential effects for cyano- and nitro-substituted neonicotinoids. Pestic. Biochem. Physiol. 2017,143, 57-65. [CrossRef]
- Qu, Y., Chen, J.H., Li, C.G., Wang, Q., Guo, W.C., Han, Z.J., Jiang, W.H. The subunit gene Ldα1 of nicotinic acetylcholine receptors plays important roles in the toxicity of imidacloprid and thiamethoxam against Leptinotarsa decemlineata. Pestic. Biochem. Physiol. 2016, 127(2), 51-58. [CrossRef]
- Wang, Z.M., Li, S., Shi, C.C., Xie, L.J., Fu, K.Y., Jiang, W.H. The actions of neonicotinoid insecticides on nicotinic acetylcholine subunits Ldα1 and Ldα8 of Leptinotarsa decemlineata and assembled receptors. Insect Sci. 2022, 29,1387-1400. [CrossRef]
- Shi, C.C., Tian, Y.T., Wang, Y.Q., Guo, W.C., Jiang, W.H. The interaction of nicotinic acetylcholine receptor subunits Ldα3, Ldα8 and Ldβ1 with neonicotinoids in Colorado potato beetle, Leptinotarsa decemlineata. Pestic. Biochem. Physiol. 2023, 195,105558. [CrossRef]
- Li, X.C., Schuler, M.A., Berenbaum, M.R. Molecular mechanisms of metabolic resistance to synthetic and natural xenobiotics. Annu. Rev. Entomol. 2007, 52, 231–253. . [CrossRef]
- Feyereisen, R., Dermauw, W., Leeuwen, T.V. Genotype to phenotype, the molecular and physiological dimensions of resistance in arthropods. Pestic. Biochem. Physiol. 2015, 121, 61–77. . [CrossRef]
- Karunker, I., Morou, E., Nikou, D., Nauen, R., Sertchook, R., Stevenson, B.J., Paine, M.J.I., Morin, S., Vontas, J. Structural model and functional characterization of the Bemisia tabaci CYP6CM1vQ, a cytochrome P450 associated with high levels of imidacloprid resistance. Insect. Biochem. Mol. Biol. 2009, 39, 697–706. [CrossRef]
- Zhang, Y.X., Yang, Y.X., Sun, H.H., Liu, Z.W. Metabolic imidacloprid resistance in the brown planthopper, Nilaparvata lugens, relies on multiple P450 enzymes. Insect Biochem. Mol. Biol. 2016, 79, 50–56. [CrossRef]
- Wu, Y.Q., Xu, H.F., Pan, Y., Gao, X.W., Xi, J.H., Zhang, J.H., Shang, Q.L. Expression profile changes of cytochrome P450 genes between thiamethoxam susceptible and resistant strains of Aphis gossypii Glover. Pestic. Biochem. Physiol. 2018,149, 1-7. [CrossRef]
- Chen, C.Y., Shan, T.S., Liu, Y., Wang, C.C., Shi, X.Y., Gao, X.W. Identification and functional analysis of a cytochrome P450 gene involved in imidacloprid resistance in Bradysia odoriphaga Yang et Zhang. Pestic. Biochem. Physiol. 2019,153,129-135. [CrossRef]
- Zhang, H.H., Yang, H.L., Dong, W.Y., Gu, Z.X., Wang, C.C., Chen, A.Q., Shi, X.Y., Gao, X.W. Mutations in the nAChR β1 subunit and overexpression of P450 genes are associated with high resistance to thiamethoxam in melon aphid, Aphis gossypii Glover. Comp. Biochem. Physiol. Part B. 2022, 258, 110682. [CrossRef]
- Wei, X.G., Hu, J.Y., Yang, J., Yin, C., Du, T.H., Huang, M.J., Fu, B.L., Gong, P.P., Liang, J.J., Liu, S.N., Xue, H., He, C., Ji, Y., Du, H., Zhang, R., Wang, C., Li, J.K., Yang, X., Zhang YJ. Cytochrome P450 CYP6DB3 was involved in thiamethoxam and imidacloprid resistance in Bemisia tabaci Q (Hemiptera: Aleyrodidae). Pestic. Biochem. Physiol. 2023, 194, 105468. [CrossRef]
- Zhu, F., Moural, T.W., Nelson, D.R., Palli, S.R. A specialist herbivore pest adaptation to xenobiotics through up-regulation of multiple cytochrome P450s. Sci. Rep. 2016, 6 (1), 20421. [CrossRef]
- Clements, J., Schoville, S., Peterson, N., Lan, Q., Groves, R.L. Characterizing molecular mechanisms of imidacloprid resistance in melect populations of Leptinotarsa decemlineata in the central sands region of Wisconsin. PLOS ONE 2016,11 (1), e0147844. [CrossRef]
- Clements, J., Sanchez-Sedillo, B., Bradfield, A.C., Groves, R.L. Transcriptomic analysis reveals similarities in genetic activation of detoxification mechanisms resulting from imidacloprid and chlorothalonil exposure. PLOS ONE 2018,13(10), e0205881. [CrossRef]
- Kalsi, M., Palli, SR. Transcription factor cap n collar C regulates multiple cytochrome P450 genes conferring adaptation to potato plant allelochemicals and resistance to imidacloprid in Leptinotarsa decemlineata (Say). Insect Biochem. Mol. Biol. 2017, 83, 1–12. [CrossRef]
- Kaplanoglu, E., Chapman, P., Scott, I.M., Donly, C. Overexpression of a cytochrome P450 and a UDP-glycosyltransferase is associated with imidacloprid resistance in the Colorado potato beetle, Leptinotarsa decemlineata. Sci. Rep. 2017, 7, 1762. [CrossRef]
- Zeng, X.C., Pan, Y., Tian, F.Y., Li, J.Y., Xu, H.F., Liu, X.M., Chen, X.W., Gao, X.W., Peng, T.F., Bi, R., Shang, Q.L. Functional validation of key cytochrome P450 monooxygenase and UDP-glycosyltransferase genes conferring cyantraniliprole resistance in Aphis gossypii Glover. Pestic. Biochem. Physiol. 2021, 176, 104879. [CrossRef]
- Wang, J., Wan, Y.R., Zhang, Y., Yuan, J.J., Zheng, X.B., Cao, H.Y., Qian, K.H., Feng, J.M., Tang, Y.X., Chen, S.R., Zhang, Y.J., Zhou, X.G., Liang, P., Wu, Q.J. Uridine diphosphate glucosyltransferases are involved in spinosad resistance in western flower thrips Frankliniella occidentalis (Pergande). J. Hazard Mater. 2024, 466, 133575. [CrossRef]
- Du, T.H., Fu, B.L., Wei, X.G., Yin, C., Yang, J., Huang, M.J., Liang, J.J., Gong, P.P., Liu, S.N., Xue, H., Hu, J.Y., Diao, Y.C., Gui, L.Y., Yang, X., Zhang, Y.J. Knockdown of UGT352A5 decreases the thiamethoxam resistance in Bemisia tabaci (Hemiptera: Gennadius). Int. J. Biol. Macromol. 2021,186, 100-108. [CrossRef]
- Livak, K.J., Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods (San Diego, Calif.) 2011, 25, 402–408. [CrossRef]
- Naqqash, M.N., Gӧkce, A., Aksoy, E., Bakhsh, A. Downregulation of imidacloprid resistant genes alters the biological parameters in Colorado potato beetle, Leptinotarsa decemlineata (Say) (chrysomelidae: Coleoptera). Chemosphere 2020, 124857. [CrossRef]
- Clements, J., Olson, J.M., Sedillo, B.S., Bradford, B., Groves, R.L. Changes in emergence phenology, fatty acid composition, and xenobiotic-metabolizing enzyme expression is associated with increased insecticide resistance in the Colorado potato beetle. Arch. Insect Biochem. Physiol. 2020,103, e21630. [CrossRef]
- Yang, Z.F., Yang, H.Y., He, G.C. Cloning and characterization of two cytochrome P450 CYP6AX1 and CYP6AY1 cDNAs from Nilaparvata lugens Stl (Homoptera: Delphacidae). Arch. Insect Biochem. Physiol. 2010, 64(2), 88-99. [CrossRef]
- Wang, R., Zhu, Y., Deng, L., Zhang, H., Wang, Q., Yin, M., Song, P., Elzaki, M.E.A., Han, Z., Wu, M. Imidacloprid is hydroxylated by Laodelphax striatellus CYP6AY3v2. Insect Mol. Biol. 2017, 26(5), 543-551. [CrossRef]
- Bouafoura, R., Bastarache, P., Ouédraogo, B.C., Dumas, P., Moffat, C. E., Vickruck, J.L., Morin, P.J. Characterization of insecticide response-associated transcripts in the Colo rado potato beetle: relevance of selected cytochrome P450s and clothianidin. Insects 2022, 13(5), 505. [CrossRef]
- Oppert, B., Guedes, R.N.C., Aikins, M.J., Perkin, L., Chen, Z., Phillips, T.W., Zhu, K.Y., Opit, G.P., Hoon, K., Sun, Y.M., Meredith, G, Bramlett, K., Hernandez, N.S., Sanderson, B., Taylor, M.W., Dhingra, D., Blakey, B., Lorenzen, M., Adedipe, F., Arthur, F. Genes related to mitochondrial functions are differentially expressed in phosphine-resistant and -susceptible Tribolium castaneum. BMC Genomics 2015, 16, 968. [CrossRef]
- Jiang, W.J., He, X.J., Wang, Z.L., Yan, W.Y., Zeng, Z.J., Wu, X.B. Cloning and expression analysis of cytochrome CYP9E2 gene in the Chinese honeybee, Apis cerana cerana. Acta Entomologica Sinica 2016, 59 (10), 1050 -1057. [CrossRef]
- Gao, Y., Kim, K., Kwon, D.H., Jeong, I.H, Clark, J.M., Lee, S.H. Transcriptome-based identification and characterization of genes commonly responding to five different insecticides in the diamondback moth, Plutella xylostella. Pestic. Biochem. Physiol. 2018, 144, 1–9. [CrossRef]
- Wu, L.X., Jia, Q.Q., Zhang, X.B., Zhang, X.Y., Liu, S.N., Park, Y., Feyereisen, R., Zhu, K., Ma, E., Zhang, J.Z., Li, S. CYP303A1 has a conserved function in adult eclosion in Locusta migratoria and Drosophila melanogaster. Insect Biochem. Mol. Biol. 2019, 113, 103210. [CrossRef]
- Du, T.H., Yin, C., Gui, L.Y., Liang, J.J., Liu, S.N., Fu, B.L., He, C., Yang, J., Wei, X.G., Gong, P.P., Huang, M.J., Xue, H., Hu, J.Y., Du, H., Ji, Y., Zhang, R., Wang, C., Zhang, C.J., Yang, X., Zhang, Y.J. Over-expression of UDP-glycosyltransferase UGT353G2 confers resistance to neonicotinoids in whitefly (Bemisia tabaci). Pestic. Biochem. Physiol. 2023, 196,105635. [CrossRef]
- Sun, X.Q., Gong, Y.H., Ali, S., Hou, M.L. Mechanisms of resistance to thiamethoxam and dinotefuran compared to imidacloprid in the brown planthopper: Roles of cytochrome P450 monooxygenase and a P450 gene CYP6ER1. Pestic. Biochem. Physiol. 2018, 150, 17–26. [CrossRef]
- Clements, J., Schoville, S., Peterson, N., Huseth, A.S., Lan, Q., Groves. R.L. RNA interference of three up-regulated transcripts associated with insecticide resistance in an imidacloprid resistant population of Leptinotarsa decemlineata. Pestic. Biochem. Physiol. 2017, 135: 35-40. [CrossRef]
- Kaplanoglu, E., Scott, I. M., Vickruck, J., Donly, C. Role of CYP9E2 and a long non-coding RNA gene in resistance to a spinosad insecticide in the Colorado potato beetle, Leptinotarsa decemlineata. PLOS ONE, 2024, 19(5), e0304037. [CrossRef]


| Sampling date | Population | Sampling location |
| 2021.6 | QPQLZ | Development zone of Zakuqiniulu Town, Qapqal County, Yili Prefecture |
| 2021.6 | ML | Dongcheng Town, Mulei County, Changji Prefecture |
| 2021.6 | JMSL | Louzhuangzi Village, Jimsar County, Changji Prefecture |
| 2021.6 | URMQA | Anningqu Town, new urban area of Urumqi City |
| 2021,7 | URMQY | Yongfeng Town, Urumqi County |
| 2022, 6 | JMST | Taiping Village, Jimsar County, Changji Prefecture |
| 20022,7 | URMQA | Anningqu town, new urban area of Urumqi City |
| 2023.6 | URMQA | Anningqu Town, new urban area of Urumqi City |
| 2023.6 | JMSQ | Quanzijie Town, Jimsar County, Changji Prefecture |
| 2023.6 | ML | Dongcheng Town, Mulei County, Changji Prefecture |
| 2023.7 | QPQLB | Development zone of Ba Town, Qapqal County, Yili Prefecture |
| 2023.7 | JMSD1 | Dayou Town, Jimsar County, Changji Prefecture |
| 2023.8 | JMSD2 | Dayou Town, Jimsar County, Changji Prefecture |
| Gene | GenBank accession | Primer sequence (5'–3') | Product size (bp) | Application |
| CYP9Z140 | XP_023030479.1 | F: TAACGAGTTTAGCGTCAG | 1881 | Cloning |
| R: CAATTGTTAATATGGAAGAC | ||||
| CYP9AY1 | XP_023020330.1 | F: TCGGTGGAATACCCATAT | 1916 | |
| R: CAAACCAAATCCAAAACA | ||||
| UGT321AP1 | XM_023173906.1 | F:TCGAAACAGTGTTGGATATT | 1663 | |
| R:AGTTTGACATGGCAACTTAG | ||||
| CYP9Z140 | F: ACATGGCCCGAGGAATTGTA | 157 | qPCR | |
| R: TTTTCAACGGCAAGGACCAC | ||||
| CYP9AY1 | F: CATTCGGCATTGGTCCAAGA | 163 | ||
| R: CCTTCTGGGCGCATATTGAA | ||||
| UGT321AP1 | F: CATCAGGAAATGGCTACCGC R: AGACCCACAGCTATGCCTTT |
189 | ||
| RPL4 | EB761170 | F: AAAGAAACGAGCATTGCCCTTCC | 119 | |
| R: TTGTCGCTGACACTGTAGGGTTGA | ||||
| Ef1α | EB754313 | F: AAGGTTCCTTCAAGTATGCGTGGG | 184 | |
| R: GCACAATCAGCTTGCGATGTACCA | ||||
| CYP9Z140 | F: AGATCAGCAAACAGCCAGTAGTCAC | 394 | RNAi | |
| R: TATTAGCCCACAATGGATCAACATC | ||||
| CYP9AY1 | F: TCGCAAATGATGTTATAGCTTCTTG | 231 | ||
| R: ATGGTACTATGGATGAGGTCGTGAA | ||||
| UGT321AP1 | F: CGTCGCTGGTTAATCTCA R: GGGTGCGTAGGGTTGC |
337 |
| Year | Population | Slope±SE | LD50 (µg/beetle) / (95% FL) | Resistance ratio |
| 2021 | URMQY | 2.1167±0.0528 | 0.0311 (0.0238-0.0408) | 1.00 |
| QPQLZ | 1.7757±0.1442 | 0.2592 (0.1797-0.3741) | 8.33 | |
| JMSL | 3.0319±0.2467 | 0.2234 (0.1870-0.2669) | 7.18 | |
| URMQA | 2.0484±0.0546 | 0.0944 (0.0717-0.1243) | 3.04 | |
| ML | 2.5969±0.1200 | 0.0679 (0.0549-0.0842) | 2.18 | |
| 2022 | URMQA | 1.5184±0.0336 | 0.2963 (0.1505-0.5834) | 9.52 |
| JMST | 2.6984±0.1913 | 0.1006 (0.0823-0.1228) | 3.23 | |
| 2023 | ML | 1.0797±0.0213 | 0.2309 (0.1057-0.5044) | 7.42 |
| JMSD1 | 1.9694±0.1116 | 0.2072 (0.1445-0.2970) | 6.66 | |
| URMQA | 1.5521±0.0887 | 0.1440 (0.0838-0.2083) | 4.63 | |
| JMSQ | 3.5502±0.1599 | 0.1310 (0.1068-0.1485) | 4.21 | |
| JMSD2 | 1.9202±0.0577 | 0.0946 (0.0630-0.1422) | 3.04 | |
| QPQLB | 1.3473±0.0956 | 0.0690 (0.0355-0.1319) | 2.22 |
| Samples | Clean reads | Clean bases | GC content (%) | Q30 (%) |
| URMQY21 | 26,114,887 | 7,748,786,787 | 41.23 | 94.17 |
| JMSL | 24,582,542 | 7,263,561,882 | 40.52 | 93.39 |
| QPQLZ | 21,000,522 | 6,225,521,885 | 40.94 | 94.16 |
| Gene function | Gene ID | URMQY21 vs JMSL | URMQY21 vs QPQLZ | ||
| log2FC | FDR value | log2FC | FDR value | ||
| CYP4C1-like | 111503441 | 1.47 | 0.0018 | ||
| CYP12a5 | 111514589 | 1.64 | 0.0167 | ||
| CYP9e2-like | 111508872 | 1.30 | 0.0035 | ||
| CYP9e2-like | 111518298 | 2.28 | 1.44E-17 | ||
| CYP9e2-like | 111508919 | 1.56 | 5.11E-07 | ||
| CYP4V2-like | 111510743 | 1.46 | 0.0453 | ||
| CYP6a13 | 111506689 | 1.33 | 0.0004 | ||
| CYP4c3-like | 111504218 | 1.47 | 0.0132 | ||
| UTG2B4-like | 111517685 | 2.43 | 0.004 | ||
| UTG2B7-like | 111518183 | 2.50 | 0.008 | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




