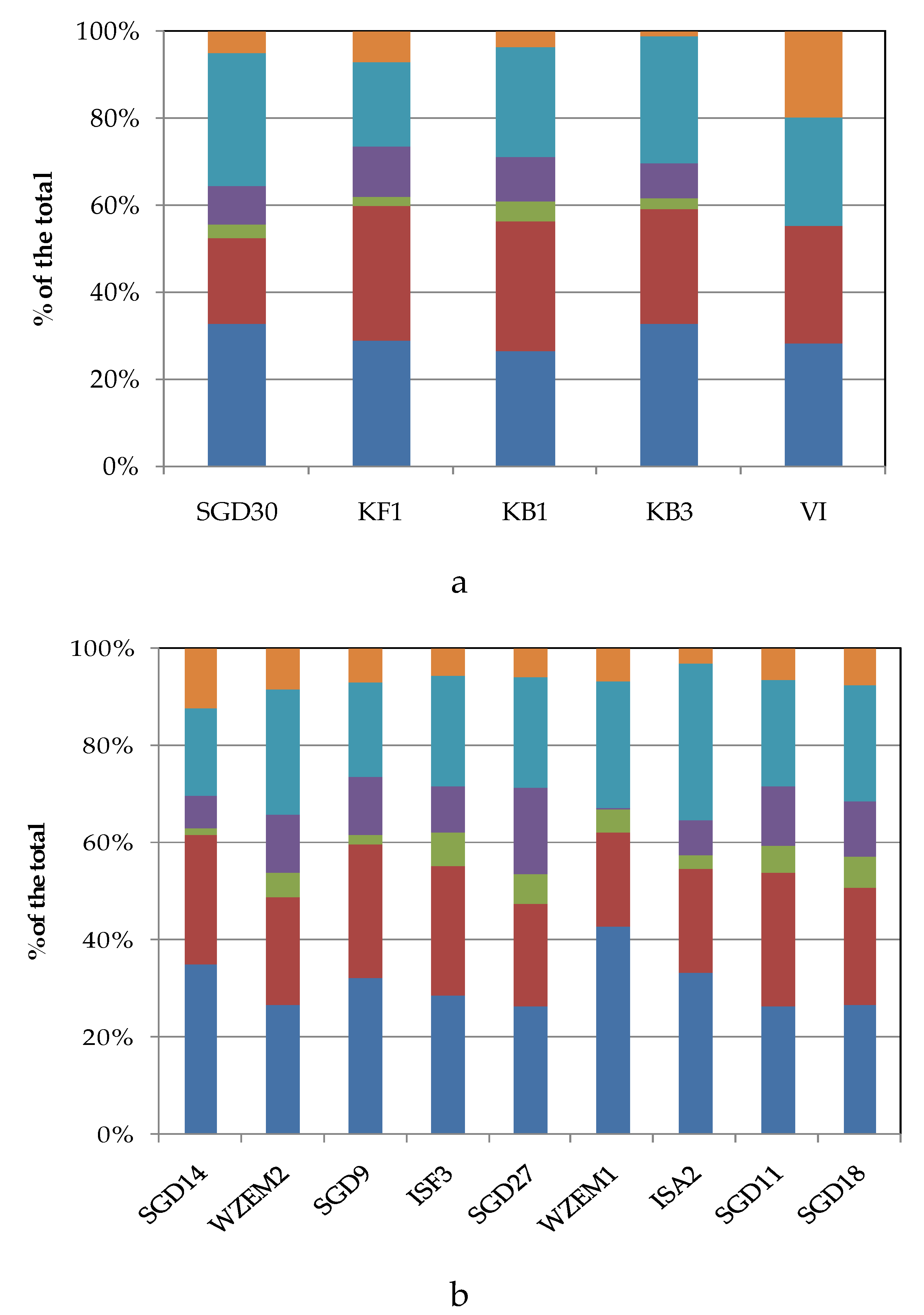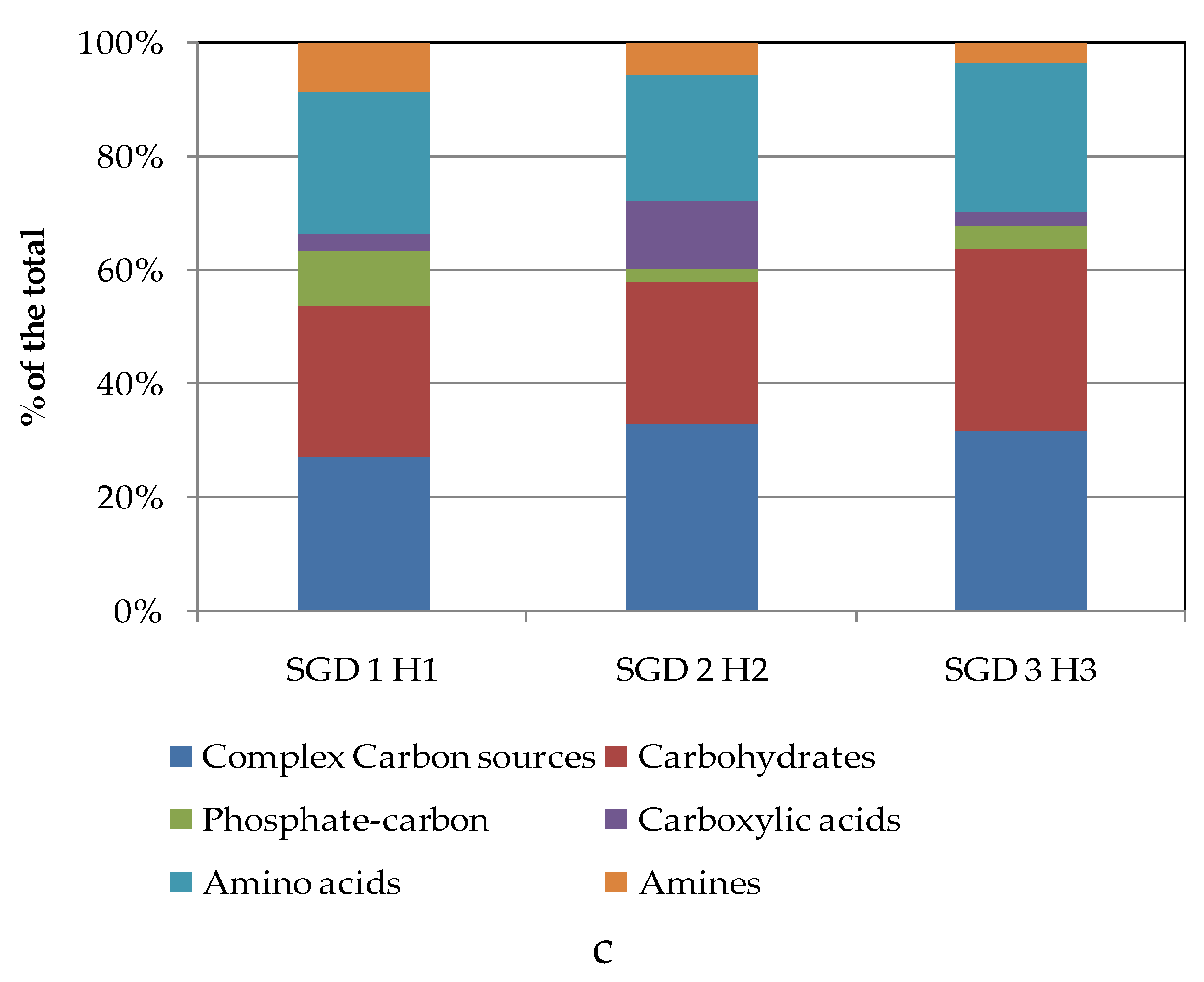1. Introduction
Svalbard Archipelago is one of the regions of the Earth most vulnerable to climate warming, but still unexplored in their microbiological features [
1]. Fjord ecosystems are considered to be sensitive sentinels of climate changes, affected by melting glaciers and warming marine water [
2]. Marine fjords influenced by active glacier freshwater inputs are key hot spots for organic matter burial in the sediments and subsequent microbial mineralization [
3]. Both freshwater supply and sedimentary processes drive the physical characteristics of fjords, making them natural laboratories where the effects of environmental changes can be detected and monitored. Particularly, fjord systems where temperature, salinity, nutrient and light gradients occur can serve as suitable models to study the response of microbial communities to the environmental variability [
4]. In this scenario, a clear knowledge of the patterns of microbial taxonomical and functional diversity and their environmental drivers is still lacking, while assessing how microbial structure and function are modulated by freshwater, terrestrial and marine organic carbon inputs would be critical for understanding microbial role in the whole fjord ecosystems’ functioning. Marine sediments act as the largest reservoirs of organic carbon on the planet [
5]; many natural compounds produced by the living biota in the water column, as well as contaminants, are buried into the benthic compartment, stressing the role of sediments as environmental archives.
Cultivable heterotrophic microorganisms represent the metabolically active fraction of the microbial community, playing a pivotal role in the biogeochemical and energy fluxes [
6]. The rapid climate warming experienced by the Arctic region is expected to result in the mobilization/transport of organic matter through glacier melting waters; currently, however, only fragmentary data are available on the local contamination and the possible effects caused by contaminants derived from the increased human pressure and industrial activity. In the continental Svalbard area, mining, sewage wastes and dumpsites are the main sources of terrestrial contamination, that following runoff may be discharged into the surrounding water bodies (including the sea), making the whole region particularly vulnerable to potential impacts of anthropic activities [
7,
8]. In consideration of the role of sediments as a conservative matrix, characterizing and quantifying the metabolic potential of microbial communities in the benthic environments is of great interest to get insights on the physiological role of prokaryotic assemblage in the carbon cycling: To this end, the Biolog Ecoplate method provides a simple and rapid tool to determine the carbon substrates utilization patterns, and to get a picture of physiological profiles at a community level, as documented by previous studies performed in temperate and polar marine ecosystems [
9,
10,
11]. This information on biogeochemical role of microbial communities might have a crucial role for understanding how future changes in benthic communities dynamics may influence the efficiency of the biological carbon pump and, in ultimate, the overall ecosystem functioning.
During the AREX survey carried out in summer 2021 in Svalbard by the Institute of Oceanology PAS (Sopot, Poland), a multidisciplinary set of physical-chemical and biological variables were measured in order to get an updated picture of the environmental state. In this framework, a quantitative and qualitative microbiological study was carried out in fjords differently affected by natural and anthropogenic factors, with the objectives of:
- (i)
drawing a first characterization of the abundance and phenotypic characteristics of the benthic heterotrophic microbial community;
- (ii)
determining microbial metabolic patterns through extracellular enzymatic activity rates and community-level physiological profiles;
- (iii)
to assess the susceptibility of bacterial isolated strains to commonly used antibiotic molecules, and the occurrence of (multiple) antibiotic resistant bacteria.
Our working hypothesis was that microbial functional patterns and antibiotic susceptibility profiles varied differently in the various northern, central and southern areas of the West Spitsbergen and therefore these microbial characteristics could be suitable descriptors of the health status of the fjord sediments.
2. Materials and Methods
2.1. Study Area and Sample Collection
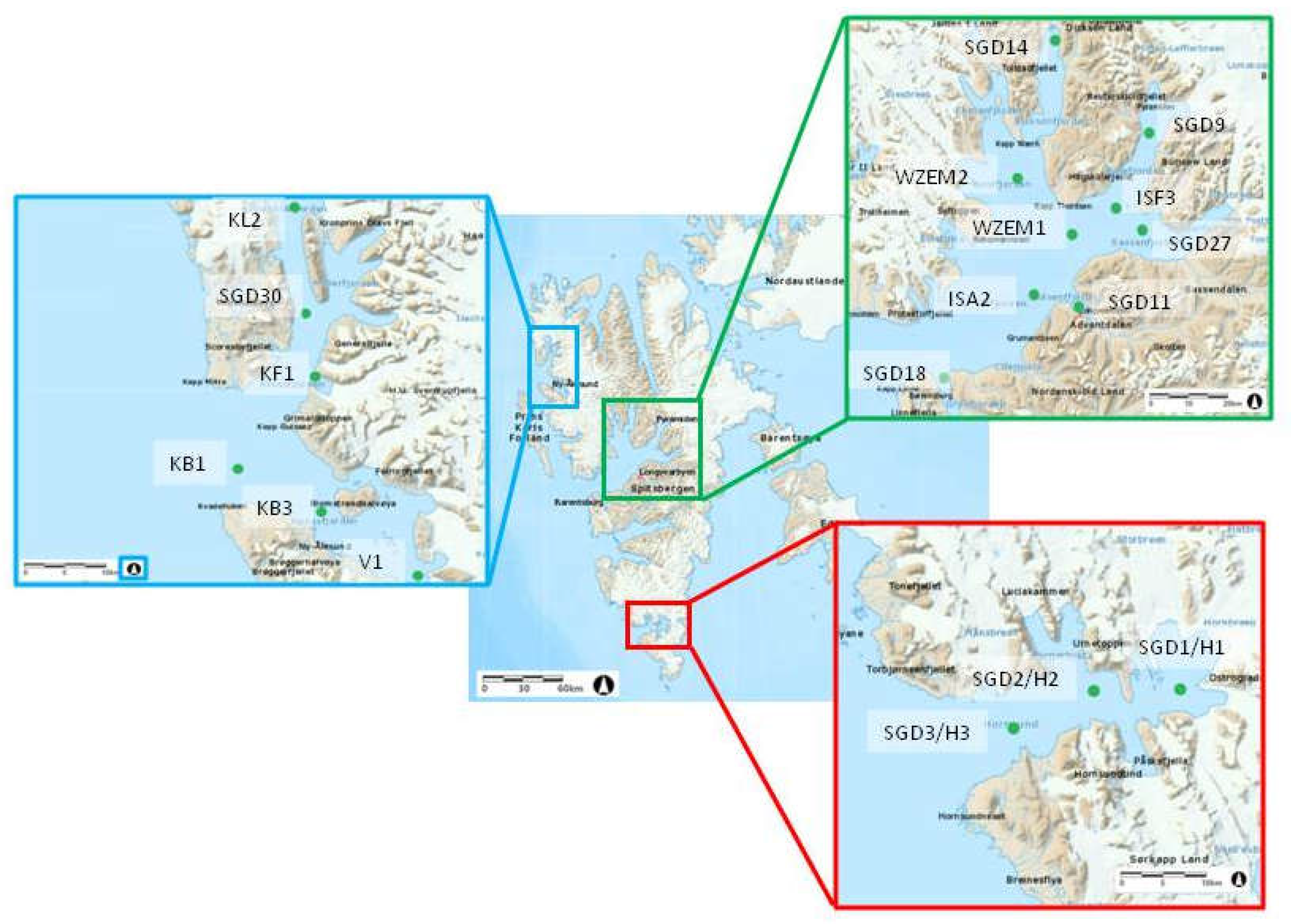
Several fjords of the Svalbard Archipelago on the western side of Spitsbergen covering from northernmost to the southernmost areas were investigated in this study: In the northern area, Krossfjorden (with stations KL2, SGD30 and KF1, from the inner to the outer ones) and Kongsfjorden (stations VI, KB3 and KB1) were studied; in the central area, sampling was performed in Dicksonfjorden (station SGD14) and Isfjorden, with its branches of Nordfjorden (station WZEM2), Sassenfjorden (stations SGD9, ISF3, SGD 27), central Isfjorden (stations WZEM1 and ISA2), Adventfjorden (station SGD 11); Gronfjorden (station SGD18). In the southern area, Hornsund (stations SGD3 H3, SGD2 H2 and SGD1 H1) was studied.
Krossfjorden and Kongsfjorden are two fjords located in the northern area on the western side of the Spitsbergen; during summer both receive sediment inputs from glacial meltwater leading to decreased surface salinity and increased turbidity, all these variables were found to affect the distribution of the bacterial community in the water column [
1,
12].
Isfjorden, with a length of 100 km, a mean width of 24 km and length of 107 km, is the largest fjord in the western Spitsbergen; it receives freshwater from tidewater glaciers as well as from rivers fed by alpine glaciers and precipitation. Isfjorden seawater is of both Atlantic and Arctic origin. In this fjord, local processes produce the Local Water and the Winter- Cooled Water; mixed water masses such as the Intermediate Water and the Transformed Atlantic Water are also found [
13].
Hornsund is a large 200 m-deep fjord located in the southernmost area of Spitsbergen, opening to the Greeland Sea; it receives seasonal sediment inputs through melting water discharge from eight major tidewater glaciers. Different sedimentation rates are observed inside this fjord and it is affected by the Atlantic Core Water, Transformed Atlantic Water and Arctic Water[
14].
The geographical coordinates of the sampled stations and the main characteristics of the sediments are shown in
Table 1 and
Figure 1. Surface (1-2 cm) sediment samples were collected using a van-Veen grab by the researchers of the Institute of Oceanology PAS (Sopot, Poland); for the microbiological study, aliquots of the samples were stored in 50 mL sterile Falcon tubes at -20°C for further processing, carried out at the arrival at the CNR-ISP laboratories in Messina (Italy).
2.2. Heterotrophic (Marine and Not-Marine) Bacterial Abundance
Volumes (100 μL) of sediment suspension (obtained by sediment dilution in sterile physiological saline in a ratio 1:10 w/v) were spread onto Marine Agar 2216 and Triptic Soy Agar (Difco) plates in duplicate, which were incubated at + 5 °C for at least 14 days in order to allow the growth of marine and non-marine bacteria, respectively The colonies grown on both culture media were then counted. Bacterial strains were isolated, by streaking on plates of the same culture media, until axenic cultures were obtained. During the isolation, colonies showing different phenotypic characteristics (i.e. morphology, size and pigmentation), and at least one colony per each typology were streaked, in order to get a collection of bacterial isolates as much as possible representative of the whole culturable bacterial community present in each sample.
2.3. Extracellular Enzymatic Activities (Leucine Aminopeptidase, LAP, Beta-Glucosidase, Beta-GLU and Alkaline Phosphatase, AP)
Microbial ectoenzymatic activity measurements were performed to estimate the potential activity rates of leucine aminopeptidase (LAP), b-glucosidase (GLU) and alkaline phosphatase (AP), enzymes involved in the decomposition mediated by the microbial community of proteins, polysaccharides and organic phosphates respectively [
1]. The enzymatic assay relies on the hydrolysis of specific fluorogenic substrates, L-leucine-7 amido-4-methylcoumarin hydrochloride (Leu-MCA), 4-methylumbelliferyl-b-d-glucoside and methylumbelliferyl phosphate, respectively, that are derivatives of methylcoumarin (MCA) and of methylumbelliferone (MUF), following the method reported by Hoppe [
15]. Increasing amounts (from 20 to 400 µmol) of substrates were added to 10 ml sub-volumes of sediment supernatant and fluorometric measurements were performed at the initial time and after incubation at ‘in situ’ temperature for 2 h. Through calibration with the standard curves obtained with known amounts of MCA for LAP and MUF for GLU and AP, the enzymatic values were expressed in term of maximum velocity of hydrolysis (Vmax, in nmol of substrate hydrolysed per gram and per hour, nmol/g/h).
2.4. Community-Level Physiological Profiles (CLPP) by Biolog Ecoplates
Microbial community metabolism was studied through the analysis of the carbon substrate utilization patterns [
16,
17,
18]. Biolog Ecoplates (Rigel Life Sciences, Rome, Italy) were used to determine the differences of the metabolic potentials of microbial assemblages hosted in the samples. Each plate is a 96-well microtiter plate, containing 31 carbon substrates and a control in triplicate together with the redox dye tetrazolium violet. Biolog plates were inoculated with 150 μl of sample and incubated at 22 °C in the dark under aerobic conditions. The oxidation of the carbon substrates into formazan was quantified by absorbance measurements at 590 nm using a compact plate reader Byonoy Absorbance 9 (Hamburg, Germany). The optical density (OD) of the reaction product was measured at time zero (T0, namely immediately after inoculation), and at fixed times intervals until 35 days of incubation. The colour development as violet pigment in each plate well was expressed in terms of the averaged well colour development (AWCD), which was calculated using the formula
where R is the averaged absorbance of the three wells with substrate and C is the averaged absorbance of the control wells (without substrate) (according to Sala et al. [
17]).
The absorbance percentages of each substrate were determined in agreement with Sala et al. [
18], setting a value of 2% absorbance of the total absorbance measured per plate as a threshold for substrate utilisation.
The carbon substrates were grouped into the following six guilds: complex carbon sources (polymers), carbohydrates, phosphate carbon sources, carboxylic and acetic acids, amino acids and amines.
2.5. Phenotypic and Biochemical Characterization of Bacterial Isolates
Bacterial isolates were phenotypically characterized for their presumptive identification, and classified according to cell morphology, Gram’s staining, pigment production, oxidase production, colony morphology, catalase test [
19]. Biochemical characterization of isolates such as sugar fermentation tests on Triple Sugar Iron agar for glucose, lactose and sucrose utilization was also performed.
2.6. Antibiotic Susceptibility Profiles of Bacterial Isolates
The screening of bacterial isolates for their susceptibility to antibiotics was performed according to the procedure reported by Laganà et al. [
20]. Briefly, the bacterial isolates were screened for antibiotic susceptibility by the Bauer’s test [
21], performed with three replicates per each isolate. The isolates were grown for 48 h on plates of Tryptic Soy Agar (TSA, Oxoid), harvested and then suspended in sterile water adjusted to a 0.5 McFarland turbidity standard (bioMérieux), corresponding to 1.5 × 10
8 CFU mL
−1. The inoculum was streaked onto plates of Mueller-Hinton agar using a cotton swab; the produced diameters of inhibition were measured after 48 h of incubation at 5 °C and averaged. Commercially available antibacterial disks (Oxoid) were used to determine the resistance patterns of the isolates against 34 different antibacterials (dose/disk as found in the common reagents), grouped into the following categories according to their mechanisms of action:
- -
Cell wall antibiotics such as beta-lactams, including penicillins [ampicillin (10 μg, code CT0003B)],
- -
Nucleic acid inhibitors: fluoroquinolones [enrofloxacin (5 μg, CT0639B)],
- -
Protein synthesis inhibitors: 1) Aminoglycoside antibiotics [gentamycin (10 μg, CT0024B)]; 2) lincosamides [clindamycin (2 μg, CT0064B)] 3) Tetracyclines [tetracycline (30 μg, CT0054B)],
The diameter of the zone of inhibition around each antibiotic disk was measured with a precision calliper (Mitutoyo, Andover, UK). Each bacterial isolate was classified as resistant (R) or sensitive (S) according to the breakpoints established by EUCAST [
22].
3. Results
According to a spatial scale, the results of the microbiological characterization of fjord sediments are reported referring to northern, central and southern areas separately.
3.1. Heterotrophic Bacterial (Marine and Non-Marine) Abundance
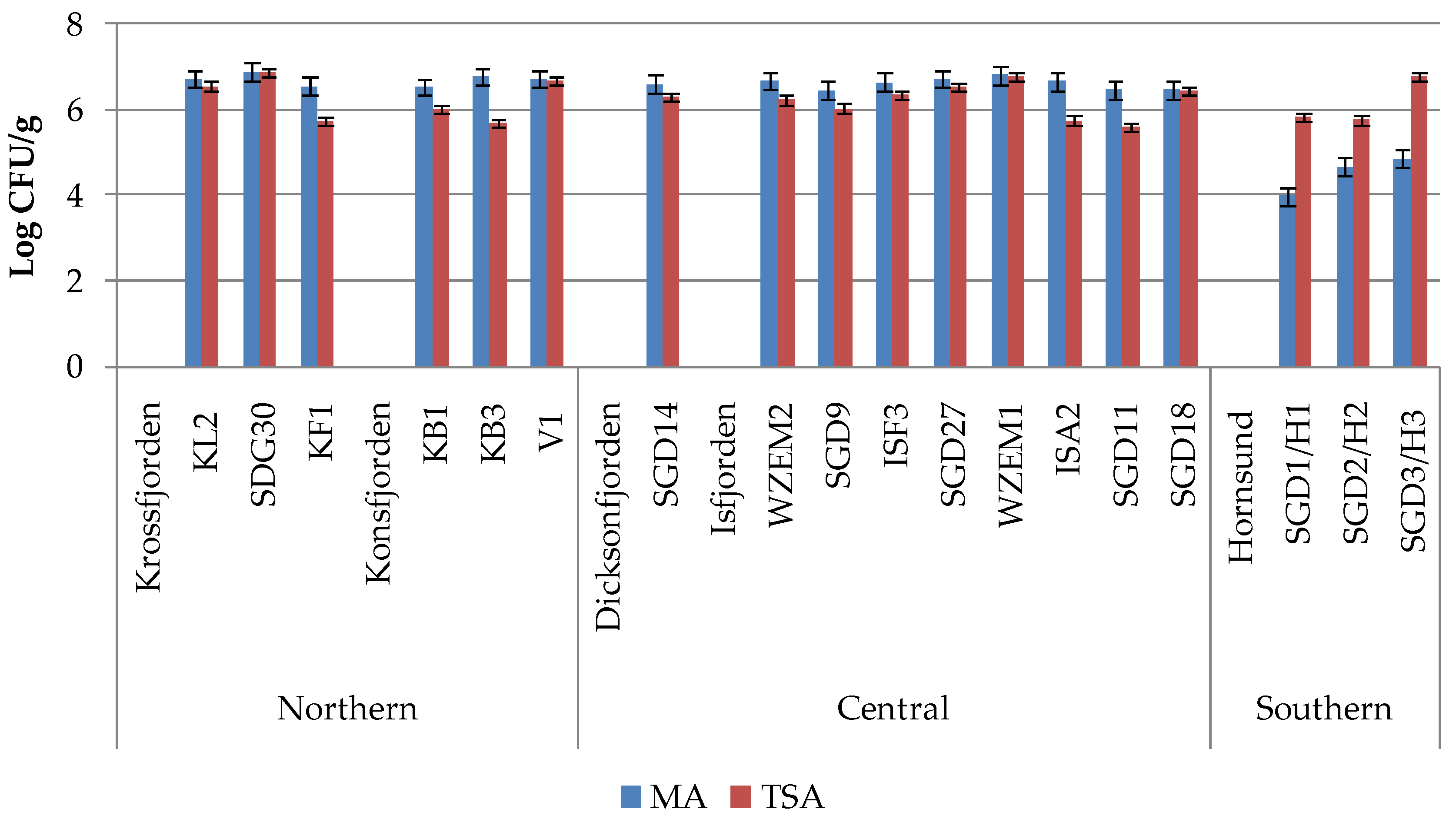
Heterotrophic bacterial abundance varied between 9.6 x 10
3 and 7.64 x 10
4 CFU/g of sediment for marine bacteria (MA) and between 4.05 x 10
3 and 7.44 x 10
4 CFU/g of sediment for non-marine bacteria (TSA), with mean ± standard deviation values of 4.56 x 10
4 ± 1.71 x 10
4 CFU/g and 2.43 x 10
4 ± 2.28 x 10
4 CFU/g, respectively (
Figure 2).
The spatial distribution of marine and non-marine bacteria pointed out high abundances of both microorganisms in the northern and central areas (increasing from SDG9 to WZEM1), while lower abundances detected in the Hornsund region. Within Krossfjorden and Konsfjorden, non marine bacteria increased from the outer to the inner zone of the fjords (stations KL2 and V1, respectively). While marine and non-marine bacteria occurred in similar quantitative ratios in the sediments of the northern and central areas (1.07 ± 0.08 and 1.06 ± 0.05, respectively), an abrupt decrease in the abundance of marine bacteria was observed in the southern Hornsund fjord (with a ratio of 0.74 ± 0.07), especially at its western station SGD1/H1.
3.3. Enzymatic Activities
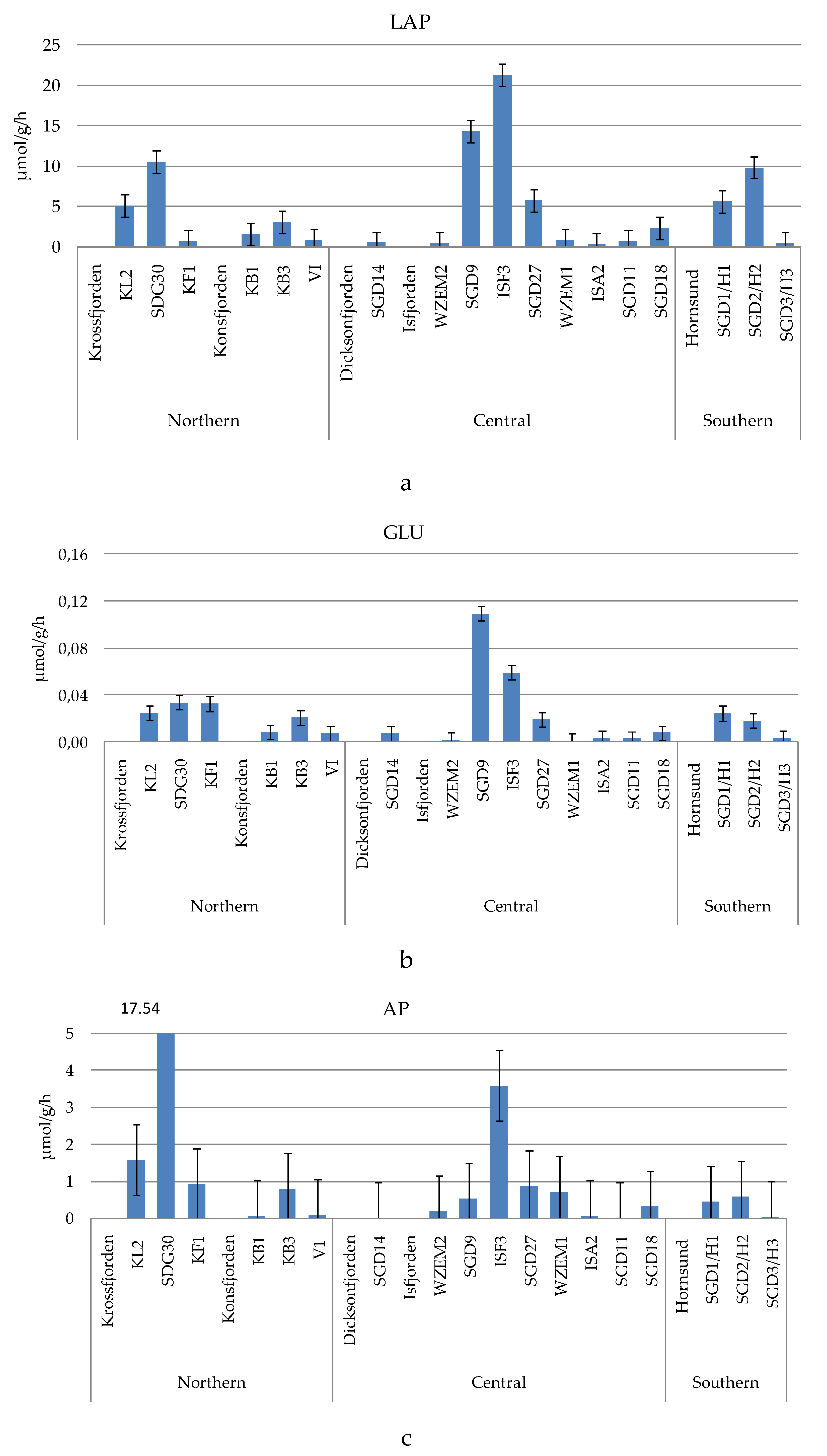
The measured enzyme activity rates (
Figure 3a–c) were generally in the order of magnitude LAP>AP>GLU, with average values ± sd of 4.618 ± 6.058 µmol/g/h, 1.574 ± 4.198 µmol/g/h and 0.021 ± 0.027 µmol/g/h respectively.
LAP ranged from 0.296 to 21.27 µmol/g/h, measured at stations ISA2 and ISF3, respectively. Enhanced proteolytic activity was observed in the central area at the inner Isfjorden stations SGD9 and ISF3, while similar levels of proteolytic activity were found in the Krossfjorden-Kongsfjorden and Hornsund fjords, where LAP peaked at station SGD30 and at the inner station (SGD2/H2), respectively. An abrupt decrease of microbial metabolism was recorded in the central area at stations ISA2, SGD11, WZEM1, WZEM2 and SGD14.
GLU was comprised between 0.0011 and 0.109 µmol/g/h, recorded at stations WZEM1 and SGD9, respectively. Like LAP, the spatial patterns of this glycolytic enzyme depicted high metabolic rates in the central area at the inner Isfjorden stations SGD9 and ISF3, and a decreasing trend moving towards stations WZEM1 and WZEM2. Low activity rates were measured in the southern Hornsund fjord, where GLU decreased across a western-eastern gradient, as well as in the northern area at Kongsfjorden. At the entrance and in the central part of Krossfjorden GLU was still more active, while this enzyme reached minima at the stations of the central area (ISA2, SGD11, SGD14).
AP varied from 0.008 to 17.54 µmol/g/h, measured respectively at stations SGD11 and SGD30. Excepting for the high activity rates observed in the northern and central areas at stations SGD30 and ISF3 respectively, AP values were generally <1 µmol/g/h, with minimal enzyme activity at stations ISA2, SGD11 and SGD14. In the southern Horsund area, the eastern station was characterised by the lowest AP activity, like what observed for LAP and GLU.
3.4. Community-Level Physiological Profiles (CLPP) by Biolog Ecoplates
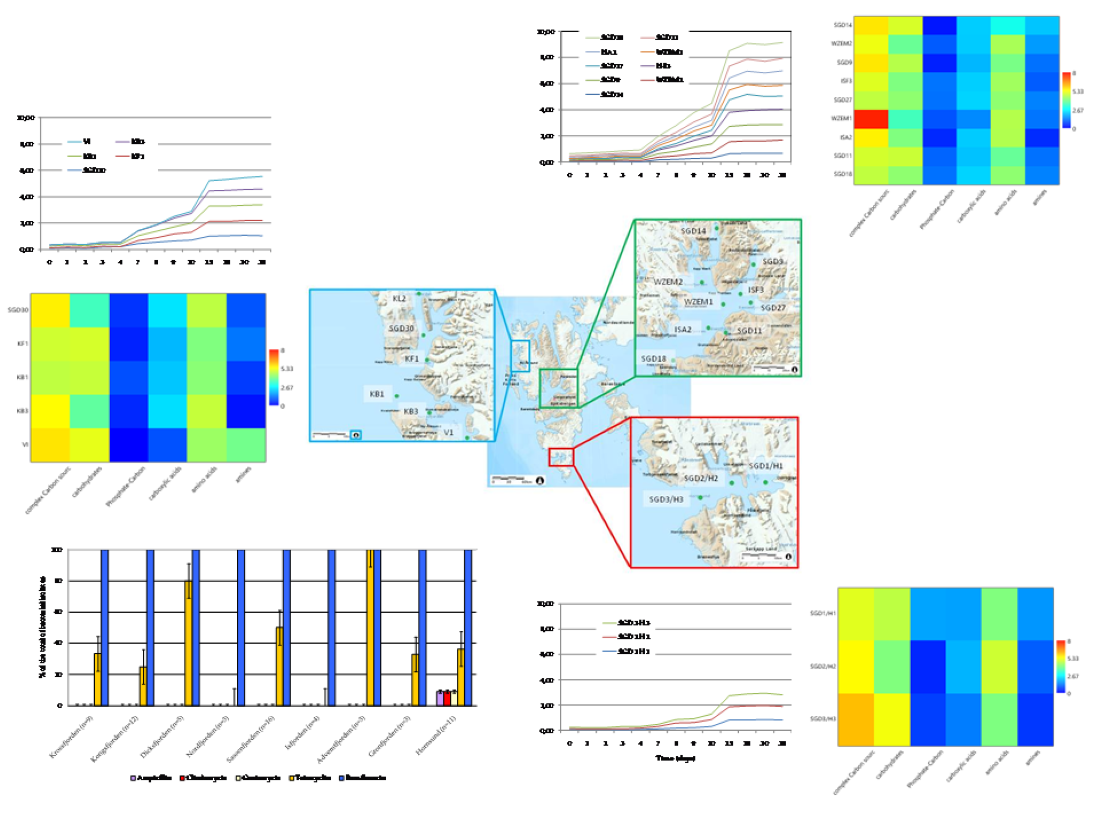
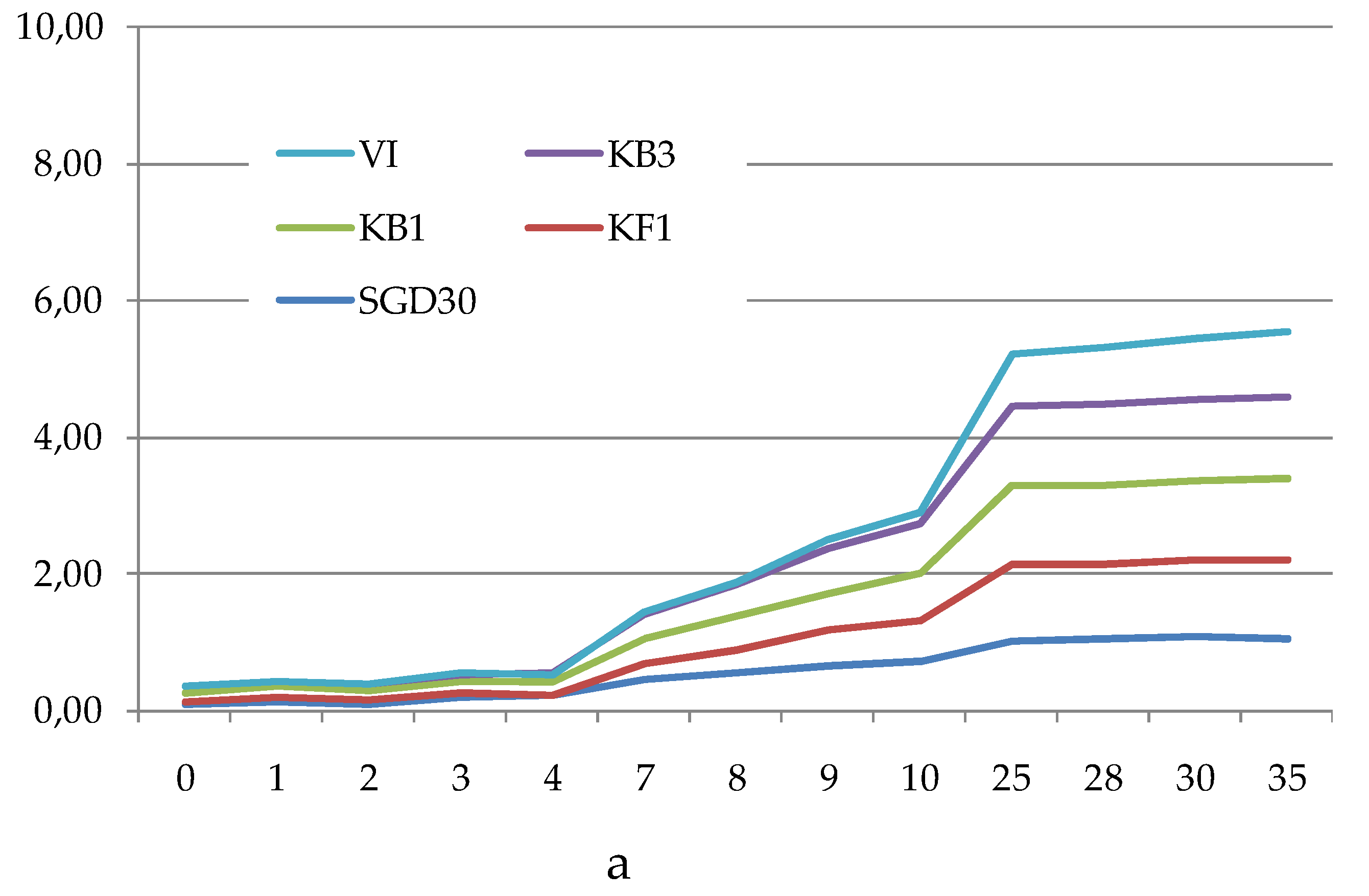
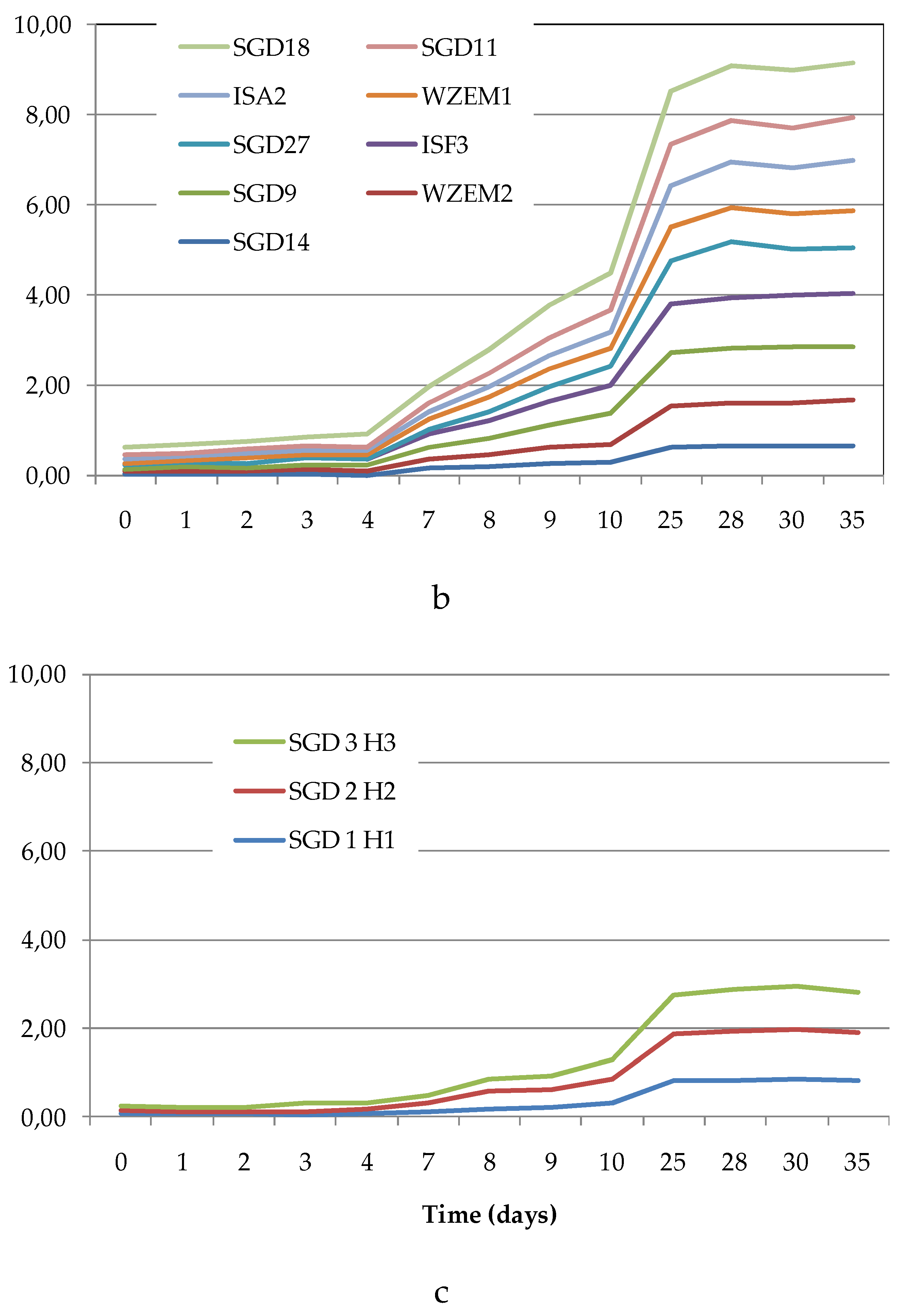
The Average Well Colour Development (AWCD), expressed as absorbance values (
Figure 4a–c), showed that during the incubation, the absorbance values gradually increased, reaching after 25 days a peak value sometimes followed by a plateau until the end of the readings (35 days).
To compare the levels of microbial metabolism recorded at the different stations, only the peak values of absorbance recorded during the incubation were considered. CLPP pointed out a generalized spatial variability (
Figure 5a–c), with the lowest activity levels recorded at station SGD14 and the highest ones at stations SGD18, SGD11 and ISA2. In the northern area (Kongsfjorden), microbial metabolism decreased from inner to outer stations. In Hornsund, the inner station displayed the highest microbial metabolism. Considering the main guilds of carbon substrates, complex Carbon sources, carbohydrates and amino acids were the substrates sources preferentially metabolized, while amines, carboxylic acids and phosphate-Carbon compounds were the less utilized ones (
Figure 5a–c).
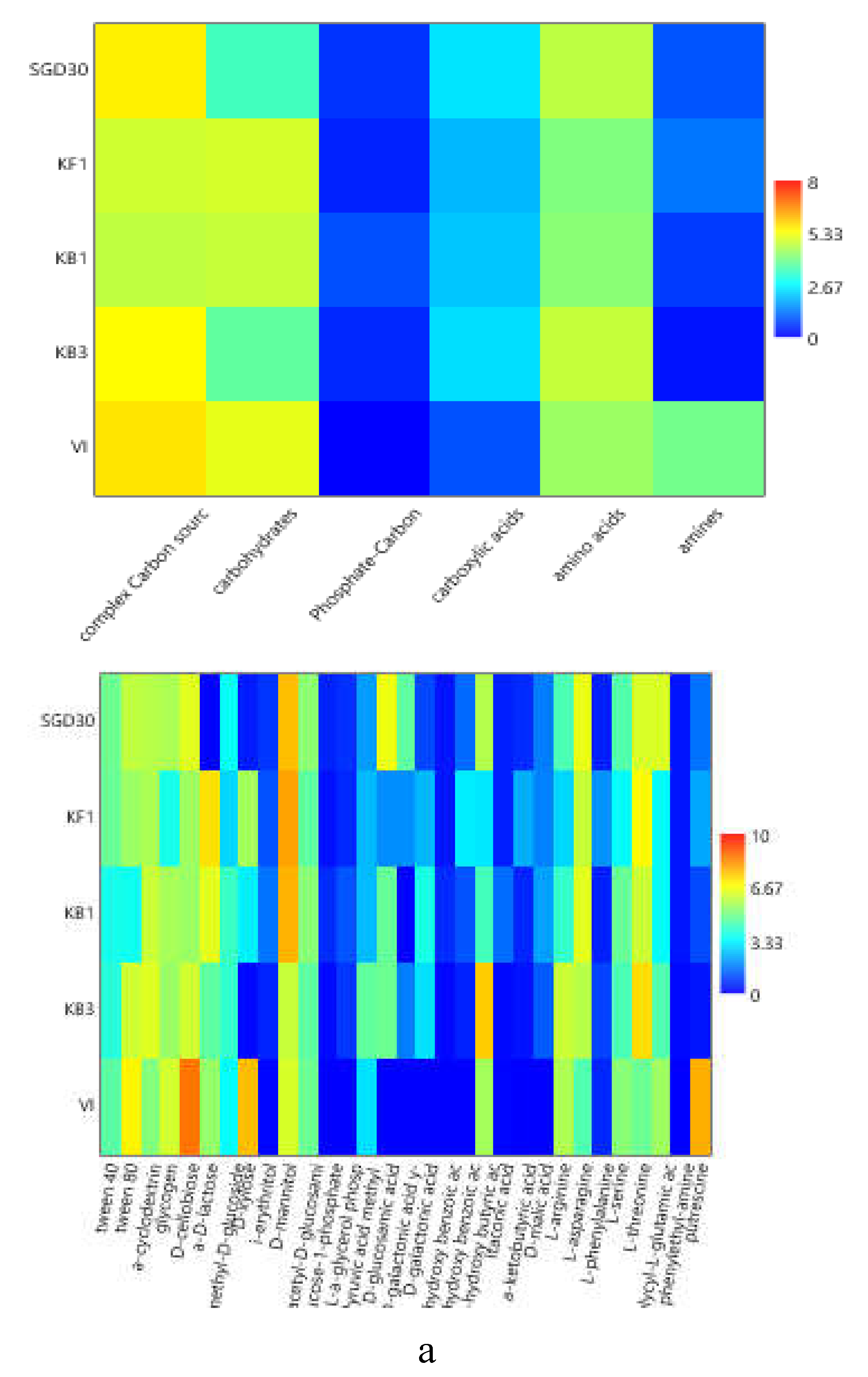
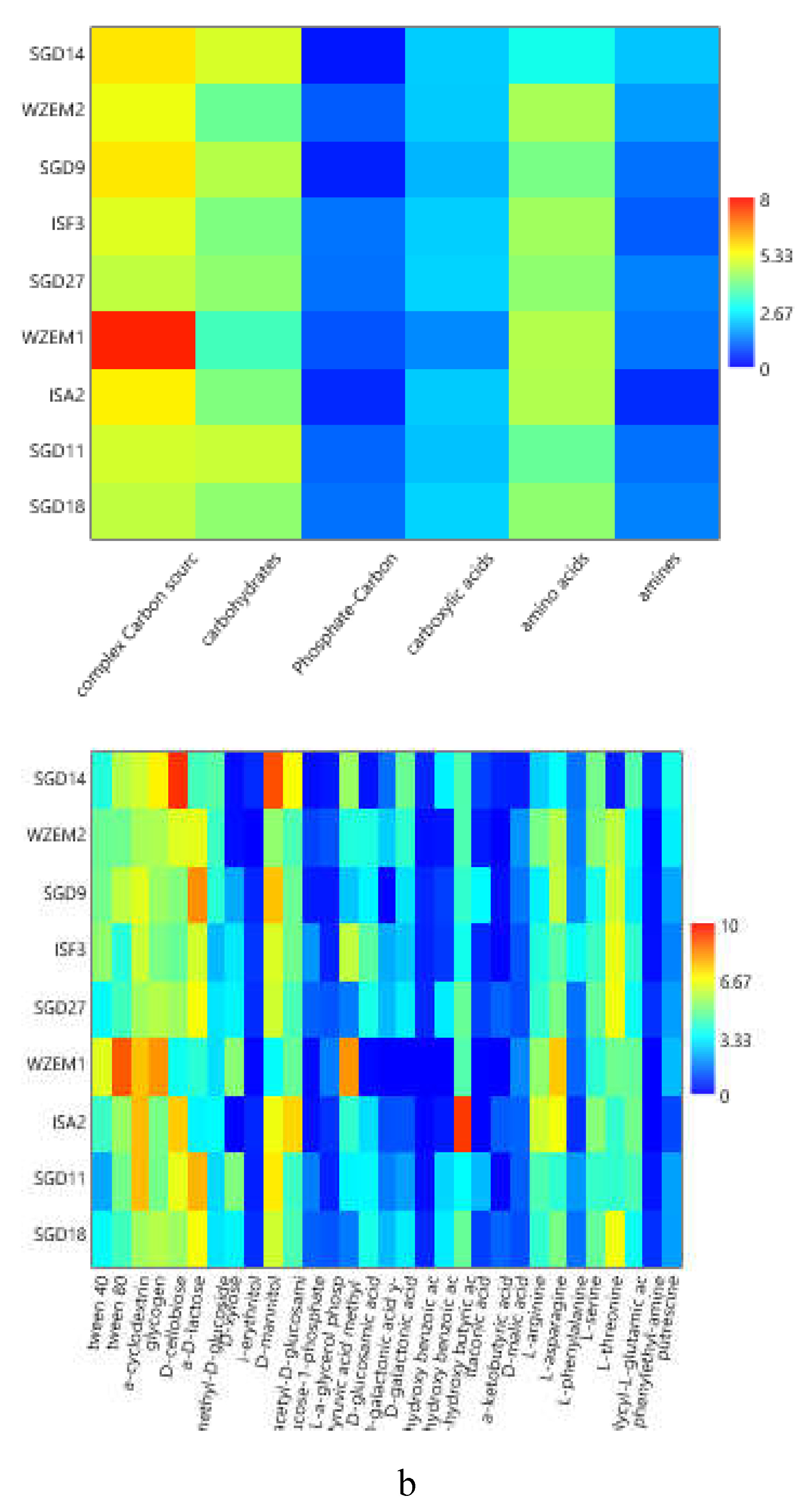
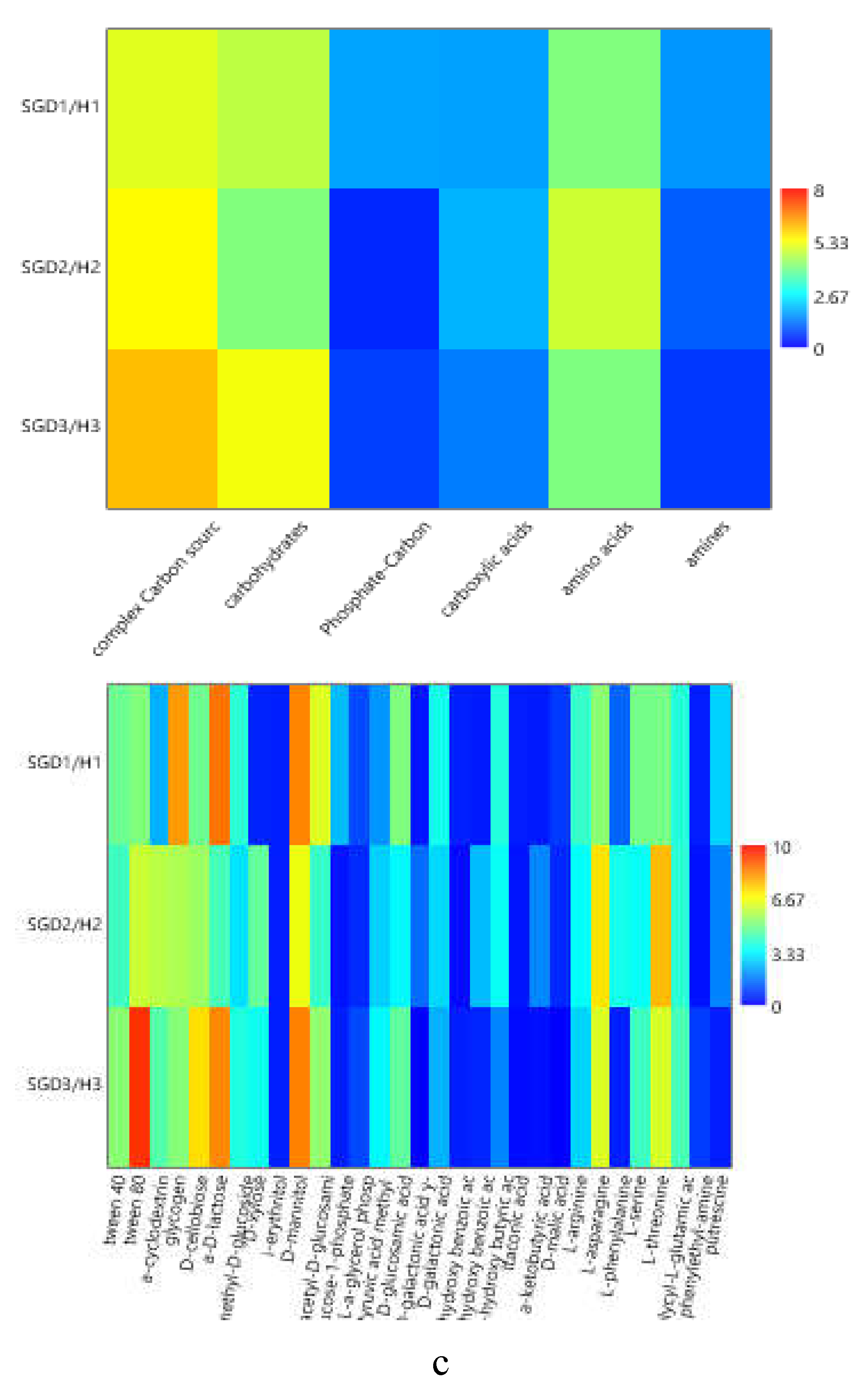
The heatmap graphs (
Figure 6a–c) highlighted the occurrence of peaks in the utilization of complex C sources at stations WZEM1, SGD3/H3 and VI, while of carbohydrates at stations SGD3/H3 and VI. Amino acids utilization was moderate in all the three areas, peaking at stations SGD2/H2 and KB3. Carboxylic acids were metabolized at stations SGD30 and KB3, everywhere in the central area (Isfjorden) and at station SGD2/H2. Amines were also utilized with high efficiency at stations VI and SGD14, while a limited utilization of Phosphate-Carbon compounds was recorded, mostly at stations SGD1/H1, ISF3 and SGD27.
At a single substrate level, within complex Carbon sources, tween 80 was the preferentially metabolized substrate, with peaks of utilization at stations VI, WZEM1 and SGD3/H3 in the northern, central and southern areas respectively. Within carbohydrates, D-cellobiose was utilized mostly at stations VI and SGD14, while a-D-lactose at station SGD1/H1. With respect to amino acids, L-threonine was the amino acid preferentially utilized at stations KB3 and SGD2/H2 in the northern and southern areas, respectively, while L-asparagine was the one utilised in the central area at station WZEM1 Within the phosphate-Carbon compounds, Glucose-1-phosphate was the most metabolised substrate, especially at stations WZEM1 and ISF3 in the central area and SGD1/H1 in the southern one. Within the carboxylic acids, y-hydroxy butyric acid was actively used at stations KB3 and ISA2 in the northern and central areas, respectively. Putrescine was the most metabolized amine at station VI in the northern area.
The diversity indices calculated from CLPP (
Table 2) suggested that microbial communities in the sediment had a good functionality, being able to utilize more than 29 carbon sources in the northern area (in Krossfjorden and Kongsfiorden station KB1), as well as in the inner and outer parts of the central area (Isfjorden stations SGD27, SGD18 and ISF3). High nutritional versatility was observed at the inner station of the southern Hornsund area (station SGD2/H2). The Shannon-Weaver diversity index reflected this trend too.
3.5. Antibiotic Susceptibility Profiles of Bacterial isolates
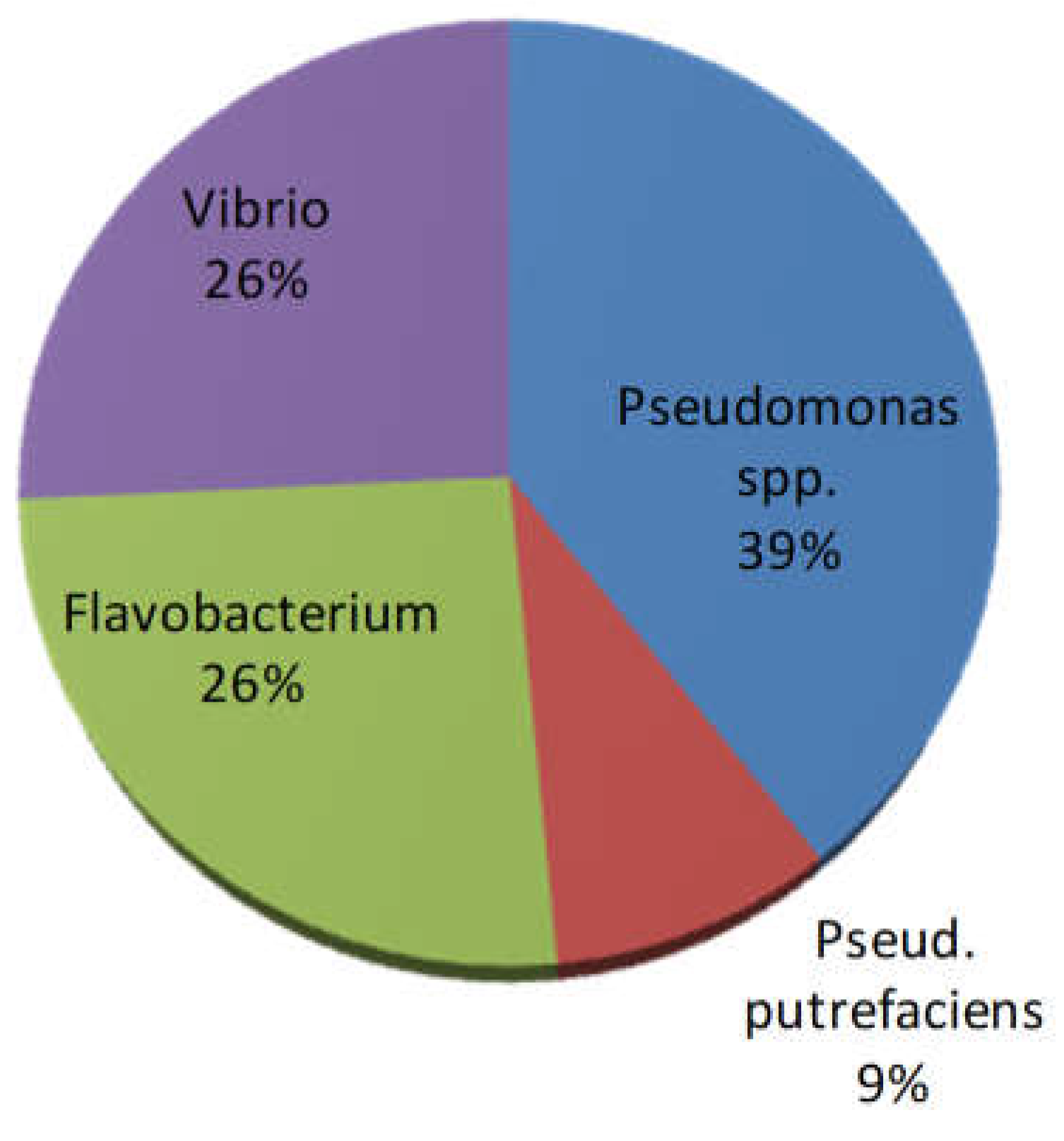
Phenotypic and biochemical tests of bacterial isolates (n=66 in total) pointed out that bacterial isolates were mostly oxidase and catalase positive microorganisms. Glucose fermenting strains were also found (accounting for <30% of the total of isolates). The majority belonged to
Pseudomonas spp. (glucose not fermenting bacteria), followed by
Flavobacterium spp. (pigmented bacteria) and
Vibrio spp. (glucose fermenting bacteria); only 6 strains produced hydrogen sulphide and were identified as
Pseudomonas putrefaciens (
Figure 7).
With respect to the spatial distribution of bacterial species, lower diversity in species composition was observed in Krossfjorden compared to other sites (
Table 3).
The antibiotic susceptibility profiles of the bacterial isolates are shown in
Table 4 and
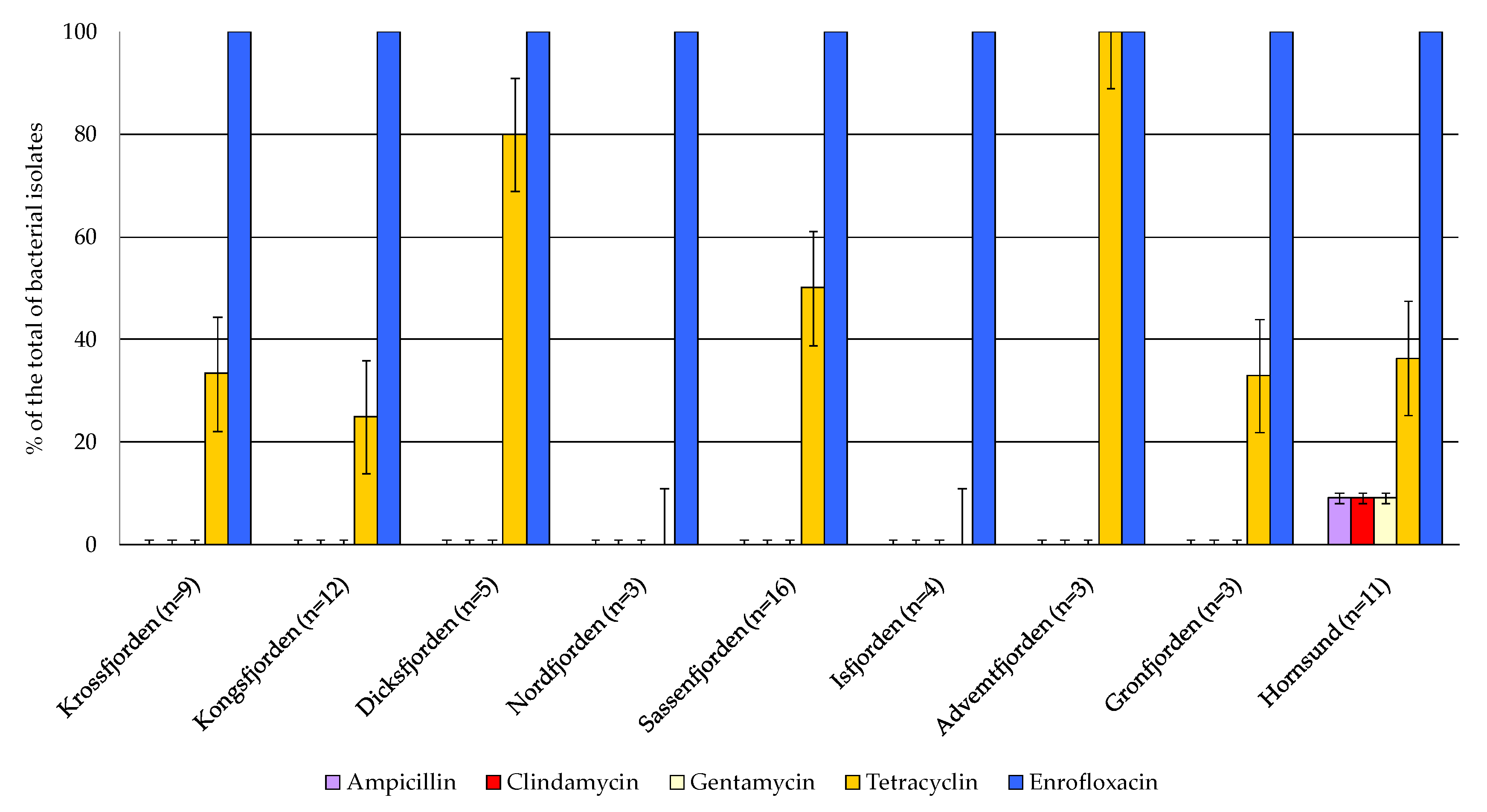
Figure 8, where 0 indicates that the strain was resistant to the assayed antibiotic, while 1 indicates that it was sensitive. All the isolates were sensitive to enrofloxacin, while resistance to tetracycline was observed in a percentage of 60.60% of the total. The majority of the strains were resistant to ampicillin, clindamycin and gentamycin. Regarding the geographical distribution of antibiotic susceptibility, the highest occurrence of strains sensitive to tetracycline was observed in the central area, particularly in Adventfjorden (100% of the total isolates), followed by Dicksonfjorden (80%), Sassenfjorden (50%), Hornsund (36%). Multiple antibiotic resistance to ampicillin, clindamycin and gentamycin was detected in the bacterial strains isolated from the sediments collected in the northern and in the central areas.
4. Discussion
This study contributed to depict on several different fjords of the west Spitsbergen a comprehensive picture of the health status of the sediments. To this end, a dual ecological and biochemical approach was used, looking at the metabolic activity rates and physiological profiles of the benthic microbial community and at the presence and distribution of antibiotic-resistant bacteria as potential bioindicators of pollution, respectively. The microflora hosted into the sediments is driven by complex processes affecting both the community composition and structure [
23]. Besides ocean currents [
24], important factors in shaping benthic microbial community are the sediment age and its diagenetic state [
3] and terrestrial inputs [
25]. The role of marine sediments as an organic carbon reservoir is crucial in fjords with active glacier discharges, which act as hot spots for organic matter burial in the benthic domain and subsequent microbial mineralization [
26]. Bacterial communities associated with Arctic sediments showed different physiological properties and metabolic functions driven by substrates’ availability [
27,
28]. Through the microbial extracellular enzymatic activity measurements and the carbon substrates utilization (by Biolog Ecoplates) new quantitative and qualitative insights on the metabolism of the benthic microbial community were obtained in our research.
4.1. Culturable Heterotrophic Bacterial Abundance and Organic Matter Decompostiion by Enzyme Activities
A quite homogeneous abundance of viable heterotrophic bacteria was observed in the fjord sediments examined in this study, suggesting that fjords hosted a heterotrophic component capable of utilizing a broad range of organic substrates. The abundance of viable heterotrophic bacteria measured in our sediment samples was in the same magnitude order as that found within the Kongsfjorden by Conte et al. [
7], with a mean value of 60.7 × 10
5 CFU/g and ranging from 1.1 to 184.8 × 10
5 CFU/g. The increase of non-marine bacterial abundance towards the inner of Krossfjorden and Kongsfjorden suggested the presence of allochthonous organic matter as a substrate for these microorganisms.
Svalbard fjord sediments are affected by the different physical-chemical properties (temperature, nutrient content, light and dissolved oxygen) and hydrodynamic patterns of the overlaying water column, which in turn affect the species composition of the bacterial community [
29,
30]. In the water column of Hornsund fjord, higher bacterial numbers and biomass than in Kongsfjorden were reported [
31], suggesting a better adaptation of Arctic bacteria to the low water temperatures and availability of organic substrates. At a sediment level, the microbial community hosted in Hornsund was considered mostly of Arctic origin [
32], compared to the one inhabiting the Kongsfjorden sediments, more affected by the Atlantic influence [
2,
33].
Benthic microbial communities play a critical role in driving the biogeochemical processes, including carbon, nitrogen, phosphorus, and sulfur cycles [
34,
35]. Therefore, measurements of the organic matter turnover mediated by hydrolytic enzymes, although in terms of potential activities, are a key step for the understanding of organic matter processing carried out by the microbial assemblage [
1]. The fjord sediments examined in the present study were characterized by the predominance of LAP and AP compared to GLU. The spatial enzyme activity patterns recorded in the sediments suggested that in the central area, especially at the inner Isfjorden (stations ISF3 and SGD9) the organic matter pool was rich in proteins and mucopolysaccharides, as shown by high LAP and GLU activity rates. Inner areas of Arctic fjords were found to be characterized by high dissolved organic matter (DOM) and the prevalence of Bacteroidetes [
1,
36]; this microbial group, especially with Cytophaga-Flavobacteria members, is known to contribute effectively to the turnover of detritus and more generally in carbon cycling within aquatic ecosystems [
37]. Diversified spectra of polysaccharide hydrolases were detected in Svalbard sediments by Teske et al. [
38], who attributed the occurrence of these enzyme activities to members of the Chloroflexi genus. Conversely, low GLU activities (around 0.01 mmol ml
-1 d
-1 ) were measured in surface sediments from the LTER Hausgarten station, in the Fram Strait [
39].
Organic phosphate esters were more abundant in the northern (Krossfjorden station SGD30) and central areas. The southern Hornsund area and the outer stations of the Isfjorden (SDG11 and SDG18) were characterized by low activity rates for all the three examined enzymes; this could indicate that sediment organic pool was quite refractory or hardly prone to active microbial decomposition, as also suggested by the low heterotrophic bacterial abundance found in this area.
4.2. Microbial Community Metabolism (CLPP)
While extracellular enzyme activities provide information on the microbial ability to hydrolyse complex molecules, Biolog Ecoplates give additional information on the utilization of single monomers or oligomers (i.e. amino acids, sugars, complex carbon substrates) at a community level, providing an index of the metabolic complexity of the microbial community [
17,
18]. Complex Carbon sources, carbohydrates and amino acids were the main categories of organic substrates preferentially metabolized by the benthic microbial assemblage in our fjord sediments. Distinct CLPP were detected, with complex Carbon sources actively metabolized in all the three areas, particularly in the central and southern areas, amino acids in the northern and southern areas and carbohydrates in the southern Hornsund fjord. Variable metabolic patterns suggested the active role played by the microbial community in driving carbon and nutrient cycling and also the quality of the benthic organic matter pool. Indeed, in Arctic fjords, terrestrial runoff, water mass exchanges and glaciers’ melting waters coexist, creating contrasting environmental conditions, and their interplay is complex especially during the summer season, when the biological activities are more enhanced and detritus released from melting ice accumulates into deep matrices [
1,
12,
40]. Furthermore, on a spatial scale the metabolic diversification probably reflected a variable community structure, like what reported in other Svalbard fjord sediments, strongly affected by changing glacial, marine and terrestrial inputs [
7].
The diversity indices suggested that in the sediments of the central (Isfjorden, stations SGD27 and SGD18) and in the southern area (Hornsund fjord, station SGD2/H2) the microbial community was able to metabolise the highest number of carbon substrates. The presence of a muddy sediment was a common trait of these three stations; nevertheless, even where clay sediments were found (i.e. in Krossfjorden, at station KB3 in Kongsfjorden and at station SGD9 in Isfjorden), CLPP showed the presence of a metabolically active microbial community in sediments unless of their granulometry, in agreement with Hargrave et al. [
41]. Moreover, functional spectra of the benthic microbial community did not seem to relate to a glacio-marine transition pattern; indeed, the functional diversity of sediments collected from outer fjord stations did not differ significantly from that of inner ones.
The use of a wide range of organic compounds as carbon sources - as shown by the Biolog Ecoplates data - confirmed the metabolic complexity and versatility of the microbial community; on the other hand, this latter is closely linked to the synthesis of extracellular enzymes. From an ecological point of view, the presence of a complex metabolism indicates the high nutritional versatility of the microbial community and of its large adaptation to diversified ecological niches; conversely, a simple metabolism suggests that microorganisms have a limited capacity of adaptation and thus can colonize only specific ecological niches [
42].
Looking at the single substrates metabolized by the microbial community, within complex Carbon sources, the widespread utilization of tween 80 was related to the presence of specific enzymes like lipases or esterases; lipases are, after proteinases, one of the most spread enzymes in marine ecosystems [
43]. Within carbohydrates, α-D-glucose was preferentially used by the microbial community; this result was not surprising, since this sugar is the principal substrate utilized to obtain pyruvate [
42]. Other well metabolized carbohydrates were mannitol and a-D-lactose.
Within carboxylic acids, γ-hydroxybutyric acid is involved in the metabolism of fatty acids and amino acids [
10]. These organic substrates are normal constituents of hydrophobic molecules. With respect to amino acids, L-threonine and L-asparagine were the most metabolised molecules; both amino acids play a key role in the cell metabolism.
The diversity indices (both richness and Shannon-Weaver) suggested that the microbial assemblage was metabolically well adapted to cope with changing conditions such as variable nutrient inputs coming from terrestrial runoff and glacier melting.
4.3. Antibiotic Resistant Bacteria
Marine sediments represent natural reservoirs of pollutants originating from surface runoff, oceanic transport, sea ice transport and atmospheric deposition. Most of organic and inorganic contaminants can also undergo transformation processes along the trophic web, accumulating into organisms. Previous studies from Svalbard and Bjørnøya documented in sediment cores the pollutants’ presence more than 100 years back in time [
44,
45]. A better understanding of the contaminant distribution within fjord environment is important since global changes are affecting the strength and directions of contaminant transport processes. Due to melting of glaciers, ice cover, and permafrost, contaminants accumulated on glacier surfaces for many decades contaminants can be discharged to marine ecosystem.
Despite their recalcitrant nature, organic contaminants such as polychlorinated biphenils (PCBs) can be transformed into chemical substances by different microbial metabolic pathways, both aerobic and anaerobic, facilitating further metabolization. Bacteria can also adsorb, accumulate and transform heavy metals; heavy metals-resistance has been reported for a number of different bacterial genera and sometimes the co-occurrence of antibiotic and metal resistances has been shown [
46].
Extreme environments represent important pristine ecosystems to study the evolution of antibiotic-resistance genes (ARGs) and the spread of antibiotic-resistant bacteria (ARB) in the presence/absence of pollution from antibiotics [
47]. Several studies have reported the occurrence in polar environments of both ARB and ARGs; this has been related to the mobilization of antibiotic residues as chemical contaminants accumulated in sediments or sea ice as a consequence of climate warming [
48,
49]. Moreover, plastic debris, frequently sinking into sediments, may act as a reservoir of ARB and ARGs [
50]. Although antibiotic resistance is a global problem, the diversity and spread of ARB and ARGs and generally of the aquatic resistome in Arctic environments are strongly understudied [
51]. Even though Norway is characterised by a lower use of antibiotics for humans' health, animal husbandry and food production compared with many other countries [
52,
53], there is still a gap of current data on the usage of antimicrobial agents in humans and animals [
54] making not possible to predict the effects of these pollutants on the environment and the local wildlife.
Contaminants may be transported in the Arctic from several remote and local sources across different pathways, including biological transport driven by migrating animals, in addition to the sea ice, riverine, groundwater discharge and atmospheric transport pathways [
55]
, but data on the presence and impact of contaminants in polar regions are still fragmentary.
Since a first detection of multidrug-resistant bacteria in Arctic [
56] reports on the spread of this phenomenon have exponentially increased. Bacterial isolates showing multiple antibiotic resistance patterns were detected in the Pasvik river (Arctic Norway [
20]). Kalinowska et al. [
57] evidenced the role of wastewater discharge in the dissemination of antibiotic-resistant
Enterococcus spp. in two Arctic lakes (a water supply system affected by birds and a wastewater close to the Polish Polar Station) in the West Spitsbergen. In ice, water and sediments samples from Longyearbyen and Adventfjorden, the most populated area of Svalbard, class 1 integrons – as markers of anthropogenic pollution- were detected in ARB isolates. Antibiotic resistance and virulence genes were found in the variable regions of integrons; the relative abundance of intI1 genes was higher in the wastewater outflow site and correlated also with the abundance of the heavy metals genes, suggesting that Svalbard resistome was affected by both natural (melting glaciers) and human (wastewater discharge) drivers. The presence of ARB and integrons in the marine environment may potentially cause a threat due to the potential transfer of ARBs and integrons to humans and the environment.
5. Conclusions
The metabolic potential of microbial communities is a helpful criterion to understand their role in complex environments like Arctic marine sediments. CLPP determined in our research indicate a nutritional versatility of the benthic microbial community to adapt to changing scenarios, and this characteristic is of paramount importance to enable microbial survival under extreme conditions. Although the functional spectra of sediment bacterial communities occurring in the fjords of the Svalbard archipelago did not seem to clearly reflect a glacio-marine transition, on a spatial scale, community metabolism was more diversified in the inner central Isfjorden area, probably in response to the availability of organic detritus made available by glacier melting and terrestrial inputs.
Multiple resistance to ampicillin, clindamycin and gentamycin was detected in all the sampled sediments, excepting in the southern Hornsund area, stressing the role of sediments as a potential reservoir of chemical wastes ascribable to antibiotic residuals, although no evident environmental risks were pointed out from this preliminary screening, given the widespread occurrence of a resistome in many natural environments.
Overall, microbial functional patterns and antibiotic susceptibility profiles varied differently in the various northern, central and southern areas of the West Spitsbergen, suggesting the suitability of these microbial characteristics as potential descriptors of the health status of the fjord sediments.
Author Contributions
Conceptualization, G.C. and A.Z.; methodology, G.C., G.M., A.C.R.; software, G.C., A.C.R.; validation, G.C., G.M., G.Z.; formal analysis, G.Z.; investigation, A.Z.; resources, G.C. and A.Z.; data curation, G.C.; writing—original draft preparation, G.C. and G.M.; writing—review and editing, G.C., G.M., A.Z.; visualization, G.Z.; supervision, G.C.; project administration, A.Z.; funding acquisition, A.Z. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by statutory IOPAN funds: theme II.2. The APC was funded by G.C. Check carefully that the details given are accurate and use the standard spelling of funding agency names at https://search.crossref.org/funding.
Data Availability Statement
Dataset available on request from the authors.
Acknowledgments
In this section, you can acknowledge any support given which is not covered by the author contribution or funding sections. This may include administrative and technical support, or donations in kind (e.g., materials used for experiments).
Conflicts of Interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
References
- Caruso:, G.; Madonia, A.; Bonamano, S.; Miserocchi, S.; Giglio, F.; Maimone, G.; Azzaro, F.; Decembrini, F.; La Ferla, R.; Piermattei, V.; Piazzolla, D.; Marcelli, M.; Azzaro, M. Microbial Abundance and Enzyme Activity Patterns: Response to Changing Environmental Characteristics along a Transect in Kongsfjorden (Svalbard Islands). J. Mar. Sci. Eng 2020, 8, 824. [Google Scholar] [CrossRef]
- Svendsen, H.; Beszczynska-Møller, A.; Hagen, J.O.; Lefauconnier, B.; Tverberg, V.; Gerland, S.; Ørbaek, J.B.; Bischof, K.; Papucci, C.; Zajaczkowski, M.; Azzolini, R.; Bruland, O.; Wiencke, C. The physical environment of Kongsfjorden–Krossfjorden, an Arctic fjord system in Svalbard. Polar Res. 2002, 21(1), 133–166. [Google Scholar] [CrossRef]
- Pelikan, C.; Jaussi, M.; Wasmund, K.; Seidenkrantz, M.-S.; Pearce, C.; Kuzyk, Z.Z.A.; Herbold, C.W.; Røy, H.; Kjeldsen, K.U.; Loy, A. Glacial runoff promotes deep burial of sulfur cycling-associated microorganisms in marine sediments. Front. Microbiol. 2019, 10, 2558. [Google Scholar] [CrossRef] [PubMed]
- Tobias-Hünefeldt, S.P.; Wing, S.R.; Baltar, F.; Morales, S.E. Changes in microbial community phylogeny and metabolic activity along the water column uncouple at near sediment aphotic layers in fjords. Sci Rep. 2021, 11, 19303. [Google Scholar] [CrossRef] [PubMed]
- Hedges, J.I.; Keil, R.G. Sedimentary organic matter preservation: an assessment and speculative synthesis. <italic>Mar. Chem</italic>. <bold>1995</bold>, <italic>49(2-3)</italic>, 81-115. [CrossRef]
- Caruso, G.; Genovese, L.; Mancuso, M.; Modica, A. Effects of fish farming on microbial enzyme activities and densities: comparison between three Mediterranean sites. Lett. Appl. Microbiol. 2003, 37(4), 324–328. [Google Scholar] [CrossRef] [PubMed]
- Conte, A.; Papale, M.; Amalfitano, S.; Mikkonen, A.; Rizzo, C.; De Domenico, E.; Michaud, L.; Lo Giudice, A. Bacterial community structure along the subtidal sandy sediment belt of a high Arctic fjord (Kongsfjorden, Svalbard Islands). Sci. Tot. Environ. 2018, 619-620, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Granberg, M.; Winberg von Friesen, L.; Bach, L.; Collard, F.; Strand, J.; Gabrielsen, G.W. Anthropogenic microlitter in wastewater and marine samples from Ny-Ålesund, Barentsburg and Signehamna, Svalbard. IVL Swedish Environmental Research Institute 2019, Stockholm, Sweden, Report number C 373, 1-28, ISBN 978-91-7883-020-6.
- Caruso, G.; Maimone, G.; Rappazzo, A.C.; Dell’Acqua, O.; Laganà, P.; Azzaro, M. Microbial Biofilm Colonizing Plastic Substrates in the Ross Sea (Antarctica): First Overview of Community-Level Physiological Profiles. J. Mar. Sci. Eng. 2023, 11, 1317. [Google Scholar] [CrossRef]
- Papale, M.; Caruso, G.; Maimone, G.; La Ferla, R.; Lo Giudice, A.; Rappazzo, A.C.; Cosenza, A.; Azzaro, F.; Ferretti, R.; Paranhos, R.; et al. Microbial Community Abundance and Metabolism Close to the Ice-Water Interface of the Blomstrandbreen Glacier (Kongsfjorden, Svalbard): A Sampling Survey Using an Unmanned Autonomous Vehicle. Water 2023, 15, 556. [Google Scholar] [CrossRef]
- Wan, L.; Caruso, G.; Cao, X.; Song, C.; Maimone, G.; Rappazzo, A.C.; Laganà, P.; Zhou, Y. Microbial Response to Coastal-Offshore Gradients in Taiwan Straits: Community Metabolism and Total Prokaryotic Abundance as Potential Proxies. Micro.b Ecol.. 2023, 85, 1253–1264. [Google Scholar] [CrossRef]
- Piquet, A.M.-T.; Scheepens, J.F.; Bolhuis, H.; Wiencke, C.; Buma, A.G.J. Variability of protistan and bacterial communities in two Arctic fjords (Spitsbergen). Polar Biol. 2010, 33, 1521–1536. [Google Scholar] [CrossRef]
- Skogseth, R.; Olivier, L.L.A.; Nilsen, F.; Falck, E.; Fraser, N.; Tverberg, V.; Ledang, A.B.; Vader, A.; Jonassen, M.O.; Søreide, J.; Cottier, F.; Berge, J.; Ivanov, B.V.; Falk-Petersen, S. Variability and decadal trends in the Isfjorden (Svalbard) ocean climate and circulation – An indicator for climate change in the European Arctic. Progr. Oceanogr. 2020, 187, 102394. [Google Scholar] [CrossRef]
- Görlich, K.; Weslawski, J.M.; Zajaczkowski, M. Suspension settling effect on macrobenthos biomass distribution in the Hornsund fjord, Spitsbergen. Polar Res. 1987, 5(2), 175–192. [Google Scholar] [CrossRef]
- Hoppe, H.G. Use of fluorogenic model substrates for extracellular enzyme activity (EEA) measurement of bacteria. In Handbook of methods in aquatic microbial ecology; Kemp, P.F., Sherr, B.F., Sherr, E.B., Cole, J.J., Eds.; Lewis Publisher, London, UK, 1993, pp. 423-431.
- Garland, J.L.; Mills, A.L. Classification and characterization of heterotrophic microbial communities on the basis of patterns of community-level sole-carbon-source utilization. Appl. Environ. Microbiol 1991, 57, 2351–2359. [Google Scholar] [CrossRef] [PubMed]
- Sala, M.M.; Arin, L.; Balagué, V.; Felipe, J.; Guadayol, O.; Vaqué, D. Functional diversity of bacterioplankton assemblages in Western Antarctic seawaters during late spring. Mar. Ecol. Progr. Ser. 2005, 292, 13–21. [Google Scholar] [CrossRef]
- Sala, M.M.; Estrada, M.; Gasol, J.M. Seasonal changes in the functional diversity of bacterioplankton in contrasting coastal environments of the NW Mediterranean. Aquat. Microb. Ecol. 2006, 44, 1–9. [Google Scholar] [CrossRef]
- Holt, J.G.; Krieg, N.R.; Sneath, P.H.A.; Staley, J.T.; Williams, S.T. <italic>Bergey’s Manual of Determinative Bacteriology</italic>, 9th ed.; Williams & Wilkins, Baltimore, USA, 1994, pp. 1-787.
- Laganà, P.; Votano, L.; Caruso, G.; Azzaro, M.; Lo Giudice, A.; Delia, S. Bacterial isolates from the Arctic region (Pasvik River, Norway): assessment of biofilm production and antibiotic susceptibility profiles. Environ. Sci. Pollut. Res. 2018, 25, 1089–1102. [Google Scholar] [CrossRef]
- Bauer, A.W.; Kirby, W.M.; Sherris, J.C.; Turk, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [PubMed]
- EUCAST (European Committee on Antimicrobial Susceptibility Testing). Breakpoint tables for interpretation of MICs and zone diameters. Version 12.0, 2022. Available at http://www.eucast.org.
- Petro, C.; Starnawski, P.; Schramm, A.; Kjeldsen, K.U. Microbial community assembly in marine sediments. Aquat. Microb. Ecol. 2017, 79, 177–195. [Google Scholar] [CrossRef]
- Hamdan, L.J.; Coffin, R.B.; Sikaroodi, M.; Greinert, J.; Treude, T.; Gillevet, P.M. Ocean currents shape the microbiome of Arctic marine sediments. ISME J. 2013, 7, 685–696. [Google Scholar] [CrossRef]
- Delpech, L.M.; Vonnahme, T.R.; McGovern, M.; Gradinger, R.; Præbel, K.; Poste, A.E. Terrestrial inputs shape coastal bacterial and archaea communities in a high Arctic fjord (Isfjorden, Svalbard). Front. Microbiol. 2021, 12, 614634. [Google Scholar] [CrossRef]
- Jørgensen, B.B.; Laufer, K.; Michaud, A.B. , Wehrmann, L.M. Biogeochemistry and microbiology of high Arctic marine sediment ecosystems—Case study of Svalbard fjords. Limnol. Oceanogr. 2020, 66, S273–S292. [Google Scholar] [CrossRef]
- Thomas, F.A.; Mohan, M.; Krishnan, K.P. Bacterial diversity and their metabolic profiles in the sedimentary environments of Ny-Ålesund, Arctic. Antonie Van Leeuwenhoek 2021, 114, 1339–1360. [Google Scholar] [CrossRef] [PubMed]
- Vishnupriya, S.; Jabir, T.; Krishnan, K.P.; Mohamed Hatha, A.A. Bacterial community structure and functional profiling of high Arctic fjord sediments. World J. Microbiol. Biotechnol. 2021, 37, 133. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.M.; Zhang, T.; Li, J.; Wang, N.-F.; Wang, Z.; Yu, L.-Y. Bacterial community pattern along the sediment seafloor of the Arctic fjorden (Kongsfjorden, Svalbard). Antonie van Leeuwenhoek 2019, 112, 1121–1136. [Google Scholar] [CrossRef] [PubMed]
- Venkatachalam, S.; Gopinath, A.; Krishnan, K.P. Fjords of the western and northern regions of Svalbard harbour distinct bacterioplankton community structures. World J. Microbiol. Biotechnol. 2023, 39, 57. [Google Scholar] [CrossRef] [PubMed]
- Kalinowska, A.; Ameryk, A.; Jankowska, K. Microbiological survey in two Arctic fjords: Total bacterial number and biomass comparison of Hornsund and Kongsfjorden. In <italic>Impact of Climate Changes on Marine Environments, GeoPlanet: Earth and Planetary Sciences</italic>, Zielinski, T.; Weslawski, M.; Kuliński, K., Eds.; Springer International Publishing, Switzerland, 2015, pp. 115-126. [CrossRef]
- Børsheim, K.Y.; Drinkwater, K.F. Different temperature adaptation in Arctic and Atlantic heterotrophic bacteria in the Barents Sea Polar Front region. J. Mar. Syst. 2014, 130, 160–166. [Google Scholar] [CrossRef]
- Beszczynska-Moller, A. : Weslawski, J.M.; Walczowski, W.; Zajaczkowski, M. Estimation of glacial meltwater discharge into Svalbard coastal waters. Oceanologia 1997, 39(3), 289–299. [Google Scholar]
- Falkowski, P.G.; Fenchel, T.; Delong, E.F. The microbial engines that drive Earth’s biogeochemical cycles. Science 2008, 320, 1034–1039. [Google Scholar] [CrossRef]
- Gray, J.S.; Elliott, M. Ecology of marine sediments: from science to management. Oxford University Press, 2009, pp-1-216. [CrossRef]
- Brakstad, O.G.; Nonstad, I.; Faksness, LG.; Brandvik, P.J. Responses of Microbial Communities in Arctic Sea Ice After Contamination by Crude Petroleum Oil. Microb. Ecol. 2008, 55, 540–552. [Google Scholar] [CrossRef]
- Cottrell, M.T.; Kirchman, D.L. Natural assemblages of marine proteobacteria and members of the Cytophaga-Flavobacter cluster consuming low- and high-molecular-weight dissolved organic matter. Appl. Environ. Microbiol. 2000, 66(4), 1692–1697. [Google Scholar] [CrossRef]
- Teske, A.; Durbin, A.; Ziervogel, K.; Cox, C.; Arnosti, C. Microbial community composition and function in permanently cold seawater and sediments from an Arctic fjord of Svalbard. Appl. Environ. Microbiol. 2011, 77, 2008–2018. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, K.; Hassenrück, C.; Salman-Carvalho, V.; Holtappels, M.; Bienhold, C. Response of bacterial communities to different detritus compositions in arctic deep-sea sediments. Front. Microbiol. 2017, 8, 266. [Google Scholar] [CrossRef] [PubMed]
- Bourgeois, S.; Kerhervé, P.; Calleja, M.Ll.; Many, G.; Morata, N. Glacier inputs influence organic matter composition and prokaryotic distribution in a high Arctic fjord (Kongsfjorden, Svalbard). J. Mar. Syst. 2016, 164, 112–127. [Google Scholar] [CrossRef]
- Hargrave, B.T.; Holmer, M.; Newcombe, C.P. Towards a classification of organic enrichment in marine sediments based on biogeochemical indicators. Mar. Pollut. Bull. 2008, 56(5), 810–824. [Google Scholar] [CrossRef] [PubMed]
- Fenice, M.; Gallo, A.; Juarez-Jimenez, B.; Gonzalez-Lopez, J. Screening for extracellular enzyme activities by bacteria isolated from samples collected in the Tyrrhenian Sea. Ann. Microbiol. 2007, 57, 93–99. [Google Scholar] [CrossRef]
- Davey, K.E.; Kirby, R.R.; Turley, C.M.; Weightman, A.J.; Fry, J.C. Depth variation of bacterial extracellular enzyme activity and population diversity in the northeastern North Atlantic Ocean. Deep Sea Res. Part II 2001, 48, 1003–1017. [Google Scholar] [CrossRef]
- Zaborska, A.; Beszczyńska-Moller, A.; Włodarska-Kowalczuk, M. History of heavy metal accumulation in the Svalbard area: Distribution, origin and transport pathways. Environ. Poll. 2017, 231, 437–450. [Google Scholar] [CrossRef] [PubMed]
- Evenset, A.; Christensen, G.N.; Carroll, J.; Zaborska, A.; Berger, U.; Herzke, D.; Gregor, D. Historical trends in persistent organic pollutants and metals recorded in sediment from Lake Ellasjøen, Bjørnøya, Norwegian Arctic. Environ. Pollut. 2007, 146(1), 196–205. [Google Scholar] [CrossRef] [PubMed]
- Li, L.-G.; Xia, Y.; Zhang, T. Co-occurrence of antibiotic and metal resistance genes revealed in complete genome collection. ISME J. 2017, 11, 651–662. [Google Scholar] [CrossRef]
- Scott, L.C.; Lee, N.; Aw, T.G. Antibiotic resistance in minimally human-impacted environments. Int. J. Environ. Res. Public Health 2020, 17, 3939. [Google Scholar] [CrossRef]
- Mohamed Hatha, A.A.; Neethu, S.; Nikhil, S.M.; Mujeeb Rahiman, K.M.; Krishnan, K.P.; Saramma, A.V. Relatively high antibiotic resistance among heterotrophic bacteria from Arctic fjord sediments than water - Evidence towards better selection pressure in the fjord sediments. Polar Sci. 2015, 9, 382–388. [Google Scholar] [CrossRef]
- AMAP (Arctic Monitoring and Assessment Programme). <italic>AMAP Assessment 2018: Biological Effects of Contaminants on Arctic Wildlife and Fish. </italic>Arctic Monitoring and Assessment Programme (AMAP), 2018, Tromsø, Norway. vii+84pp.
- Yang, Y.; Liu, G.; Song, W.; Ye, C.; Lin, H.; Li, Z.; Liu, W. Plastics in the marine environment are réservoirs for antibiotic and metal resistance genes. Environ. Int. 2019, 123, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Makowska-Zawierucha, N.; Mokracka, J.; Małecka, M.; Balazy, P.; Chełchowski, M.; Ignatiuk, D.; Zawierucha, K. Quantification of class 1 integrons and characterization of the associated gene cassettes in the high Arctic – Interplay of humans and glaciers in shaping the aquatic resistome. Ecol. Indic. 2022, 145, 109633. [Google Scholar] [CrossRef]
- Astrup, E. Antibiotic resistance in Norway- NIPH. Public Health Report, 2014. Available at https://www.fhi.no/en/op/public-health-report-2014/health--disease/antibiotic-resistance-in-norway---p/.
- Norwegian Ministries. National Strategy against Antibiotic Resistance 2015-2020. Norwegian Ministry of Health and Care Services, 2015, pp. 1-36.
- NORM/NORM-VET 2021. Usage of Antimicrobial Agents and Occurrence of Antimicrobial Resistance in Norway. Tromsø, Norway, September 2022, pp. 1-64, Available at www.vetinst.no.
- Pouch, A.; Zaborska, A. Climate change influence on migration of contaminants in the Arctic marine environment. GeoPlanet Earth Planet. Sci. 2015, 22, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Sjölund, M.; Bonnedahl, J.; Hernandez, J.; Bengtsson, S.; Cederbrant, G.; Pinhassi, J.; Kahlmeter, G.; Olsen, B. Dissemination of multidrug-resistant bacteria into the Arctic. Emerg. Infect. Dis. 2008, 14(1), 70–72. [Google Scholar] [CrossRef]
- Kalinowska, A.; Jankowska, K.; Fudala-Ksiazek, S.; Pierpaoli, M.; Luczkiewicz, A. The microbial community, its biochemical potential, and the antimicrobial resistance of <italic>Enterococcus</italic> spp. in Arctic lakes under natural and anthropogenic impact (West Spitsbergen). Sci. Tot Environ. 2021, 763, 142998. [Google Scholar] [CrossRef]
Figure 1.
Location of the sampling sites.
Figure 1.
Location of the sampling sites.
Figure 2.
Marine (MA) and non-marine (TSA) bacterial concentrations (in Colony-Forming Units per g of sediment) recorded in the sediments collected in northern, central and southern Svalbard regions separately.
Figure 2.
Marine (MA) and non-marine (TSA) bacterial concentrations (in Colony-Forming Units per g of sediment) recorded in the sediments collected in northern, central and southern Svalbard regions separately.
Figure 3.
(a-c). Leucine aminopeptidase (LAP, a), beta-glucosidase (GLU, b) and alkaline phosphatase (AP, c) activity rates measured in the sampled sediments.
Figure 3.
(a-c). Leucine aminopeptidase (LAP, a), beta-glucosidase (GLU, b) and alkaline phosphatase (AP, c) activity rates measured in the sampled sediments.
Figure 4.
(a-c). Trend of the Average Well Color Development (as Absorbance values) recorded during the incubation period in the sediments collected from the Northern (a), Central (b) and Southern (c) areas of the Svalbard archipelago.
Figure 4.
(a-c). Trend of the Average Well Color Development (as Absorbance values) recorded during the incubation period in the sediments collected from the Northern (a), Central (b) and Southern (c) areas of the Svalbard archipelago.
Figure 5.
(a-c). Community-level physiological profiles (as % of the total) of the sediments collected from the Northern (a), Central (b) and Southern areas (c) of the Svalbard archipelago.
Figure 5.
(a-c). Community-level physiological profiles (as % of the total) of the sediments collected from the Northern (a), Central (b) and Southern areas (c) of the Svalbard archipelago.
Figure 6.
(a-c). Heatmaps showing the utilization patterns of the different organic substrates (carbohydrates, phosphate carbon sources, carboxylic and acetic acids, amino acids and amines) (left) and of each substrate assayed by the Biolog Ecoplates (right) in the sediments of the Northern (a), Central (b) and Southern areas ( c) of the Svalbard archipelago.
Figure 6.
(a-c). Heatmaps showing the utilization patterns of the different organic substrates (carbohydrates, phosphate carbon sources, carboxylic and acetic acids, amino acids and amines) (left) and of each substrate assayed by the Biolog Ecoplates (right) in the sediments of the Northern (a), Central (b) and Southern areas ( c) of the Svalbard archipelago.
Figure 7.
Presumptive identification of retrieved bacterial isolates (n=66).
Figure 7.
Presumptive identification of retrieved bacterial isolates (n=66).
Figure 8.
Antibiotic susceptibility test of bacterial isolates; the strains whose growth was inhibited by the antibiotic were scored as negative.
Figure 8.
Antibiotic susceptibility test of bacterial isolates; the strains whose growth was inhibited by the antibiotic were scored as negative.
Table 1.
Geographical coordinates of the sampled stations and characteristics of the sediments.
Table 1.
Geographical coordinates of the sampled stations and characteristics of the sediments.
| Station |
Area |
Latitude |
Longitude |
Depth (m) |
sediment type |
sediment colour |
| KL2 |
Krossfjorden inner |
79,3282 |
11,6341 |
174 |
clay |
grey |
| SGD30 |
Krossfjorden |
79,2015 |
11,7395 |
282 |
clay |
grey |
| KF1 |
Krossfjorden outer |
79,1261 |
11,8177 |
60 |
clay |
grey |
| KB1 |
Kongsfjorden outer |
79,0112 |
11,4058 |
369 |
mud |
noir |
| KB3 |
Kongsfjorden |
78,965 |
11,8994 |
347 |
clay |
grey |
| VI |
Konsfjorden inner |
78,8933 |
12,4722 |
74 |
mud |
brown |
| SGD14 |
Dicksonfjorden |
78,8097 |
15,3842 |
16 |
mud |
brown |
| WZEM2 |
Nordfjorden |
78,5152 |
14,9392 |
144 |
mud |
noir |
| SGD9 |
Billefjorden |
78,609 |
16,4805 |
141 |
clay |
grey |
| ISF3 |
Sassenfjorden |
78,4479 |
16,078 |
94 |
mud |
noir |
| SGD27 |
Sassenfjorden |
78,4003 |
16,3756 |
31 |
mud |
noir |
| WZEM1 |
Isfjorden |
78,395 |
15,5663 |
202 |
mud |
noir |
| ISA2 |
Isfjorden |
78,2657 |
15,1257 |
248 |
mud |
noir |
| SGD11 |
Adventfjorden |
78,2372 |
15,636 |
58 |
mud |
noir |
| SGD18 |
Gronfjorden |
78,0873 |
14,1221 |
122 |
mud |
noir |
| SGD1/H1 |
Hornsund West |
77,0021 |
16,4646 |
125 |
mud |
noir |
| SGD2/H2 |
Hornsund inner |
77,0019 |
16,0846 |
84 |
mud |
noir |
| SGD3/H3 |
Hornsund East |
76,9664 |
15,7297 |
242 |
mud |
noir |
Table 2.
Diversity indices.
Table 2.
Diversity indices.
| |
Richness |
Shannon-Weaver |
| Northern area |
|
|
| SGD30 |
28 |
3.054 |
| KF1 |
28 |
3.171 |
| KB1 |
29 |
3.162 |
| KB3 |
24 |
3.012 |
| VI |
19 |
2.875 |
| Central area |
|
|
| SGD14 |
23 |
2.965 |
| WZEM2 |
25 |
3.108 |
| SGD9 |
28 |
3.127 |
| ISF3 |
29 |
3.191 |
| SGD27 |
31 |
3.251 |
| WZEM1 |
21 |
2.935 |
| ISA2 |
26 |
3.001 |
| SGD11 |
28 |
3.180 |
| SGD18 |
31 |
3.251 |
| Southern area |
|
|
| SGD1/H1 |
26 |
3.048 |
| SGD2/H2 |
31 |
3.177 |
| SGD3/H3 |
27 |
2.999 |
Table 3.
Presumptive identification of the bacterial strains isolated per each sampling area according to the phenotypic and biochemical tests.
Table 3.
Presumptive identification of the bacterial strains isolated per each sampling area according to the phenotypic and biochemical tests.
| |
|
Pseudomonas spp. |
Pseud. putrefaciens |
Flavobacterium spp. |
Vibrio spp. |
Total of isolates |
| Northern area |
Krossfjorden |
4 |
|
5 |
|
9 |
| |
Kongsfjorden |
3 |
5 |
1 |
3 |
12 |
| Central area |
Isfjorden |
17 |
|
4 |
13 |
34 |
| Southern area |
Hornsund |
2 |
1 |
7 |
1 |
11 |
| Total of isolated strains |
|
26 |
6 |
17 |
17 |
66 |
Table 4.
Profiles of susceptibility to the assayed antibiotics of the bacterial strains isolated in this study.
Table 4.
Profiles of susceptibility to the assayed antibiotics of the bacterial strains isolated in this study.
| |
|
Susceptibility to: |
| |
Station |
Ampicillin |
Clindamycin |
Gentamycin |
Tetracycline |
Enrofloxacin |
| Northern area |
|
|
|
|
|
|
| Krossfjorden |
SGD 30 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 30 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 30 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 30 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 30 |
0 |
0 |
0 |
0 |
1 |
| |
KF1 |
0 |
0 |
0 |
0 |
1 |
| |
KF1 |
0 |
0 |
0 |
1 |
1 |
| |
KF1 |
0 |
0 |
0 |
1 |
1 |
| |
KF1 |
0 |
0 |
0 |
0 |
1 |
| Total |
n=9 |
0 |
0 |
0 |
3 |
9 |
| Kongsfjordem |
KB1 |
0 |
0 |
0 |
0 |
1 |
| |
KB1 |
0 |
0 |
0 |
0 |
1 |
| |
KB1 |
0 |
0 |
0 |
0 |
1 |
| |
KB3 |
0 |
0 |
0 |
0 |
1 |
| |
KB3 |
0 |
0 |
0 |
0 |
1 |
| |
KB3 |
0 |
0 |
0 |
0 |
1 |
| |
KB3 |
0 |
0 |
0 |
0 |
1 |
| |
KB3 |
0 |
0 |
0 |
0 |
1 |
| |
VI |
0 |
0 |
0 |
0 |
1 |
| |
VI |
0 |
0 |
0 |
1 |
1 |
| |
VI |
0 |
0 |
0 |
1 |
1 |
| |
VI |
0 |
0 |
0 |
1 |
1 |
| Total |
n=12 |
0 |
0 |
0 |
3 |
12 |
| Northern area |
n=21 |
0 (0.00%) |
0 (0.00%) |
0 (0.00%) |
6 (28.60%) |
21 (100.00%) |
| Central area |
|
|
|
|
|
|
| Dicksonfjorden |
SGD 14 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 14 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 14 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 14 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 14 |
0 |
0 |
0 |
1 |
1 |
| Total |
n=5 |
0 |
0 |
0 |
4 |
5 |
| Nordfjorden |
WZEM2 |
0 |
0 |
0 |
0 |
1 |
| |
WZEM2 |
0 |
0 |
0 |
0 |
1 |
| |
WZEM2 |
0 |
0 |
0 |
0 |
1 |
| Total |
n=3 |
0 |
0 |
0 |
0 |
3 |
| Sassenfjorden |
SGD9 |
0 |
0 |
0 |
0 |
1 |
| |
SGD9 |
0 |
0 |
0 |
1 |
1 |
| |
SGD9 |
0 |
0 |
0 |
0 |
1 |
| |
ISF 3 |
0 |
0 |
0 |
0 |
1 |
| |
ISF 3 |
0 |
0 |
0 |
0 |
1 |
| |
ISF 3 |
0 |
0 |
0 |
0 |
1 |
| |
ISF 3 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 27 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 27 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 27 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 27 |
0 |
0 |
0 |
1 |
1 |
| |
SGD 27 |
0 |
0 |
0 |
1 |
1 |
| |
WZEM1 |
0 |
0 |
0 |
1 |
1 |
| |
WZEM1 |
0 |
0 |
0 |
1 |
1 |
| |
WZEM1 |
0 |
0 |
0 |
1 |
1 |
| |
WZEM1 |
0 |
0 |
0 |
0 |
1 |
| Total |
n=16 |
0 |
0 |
0 |
8 |
16 |
| Isfjorden |
ISA 2 |
0 |
0 |
0 |
0 |
1 |
| |
ISA 2 |
0 |
0 |
0 |
0 |
1 |
| |
ISA 2 |
0 |
0 |
0 |
0 |
1 |
| |
ISA 2 |
0 |
0 |
0 |
0 |
1 |
| Total |
n=4 |
0 |
0 |
0 |
0 |
4 |
| Advemtfjorden |
SDG11 |
0 |
0 |
0 |
1 |
1 |
| |
SDG11 |
0 |
0 |
0 |
1 |
1 |
| |
SDG11 |
0 |
0 |
0 |
1 |
1 |
| Total |
n=3 |
0 |
0 |
0 |
3 |
3 |
| Gronfjorden |
SGD 18 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 18 |
0 |
0 |
0 |
0 |
1 |
| |
SGD 18 |
0 |
0 |
0 |
1 |
1 |
| Total |
n=3 |
0 |
0 |
0 |
1 |
3 |
| Central area |
n=34 |
0 (0.00%) |
0 (0.00%) |
0 (0.00%) |
16 (47.06%) |
34 (100.00%) |
| Southern area |
|
|
|
|
|
|
| Hornsund |
SGD1/H1 |
0 |
0 |
0 |
0 |
1 |
| |
SGD1/H1 |
0 |
0 |
0 |
0 |
1 |
| |
SGD1/H1 |
0 |
0 |
0 |
0 |
1 |
| |
SGD1/H1 |
1 |
1 |
1 |
1 |
1 |
| |
SGD1/H1 |
0 |
0 |
0 |
0 |
1 |
| |
SGD2/H2 |
0 |
0 |
0 |
0 |
1 |
| |
SGD2/H2 |
0 |
0 |
0 |
0 |
1 |
| |
SGD3/H3 |
0 |
0 |
0 |
0 |
1 |
| |
SGD3/H3 |
0 |
0 |
0 |
1 |
1 |
| |
SGD3/H3 |
0 |
0 |
0 |
1 |
1 |
| |
SGD3/H3 |
0 |
0 |
0 |
1 |
1 |
| Southern area |
n=11 |
1 (0.09%) |
1 (0.09%) |
1 (0.09%) |
4 (36.40%) |
11 (100%) |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).