Submitted:
17 May 2024
Posted:
20 May 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
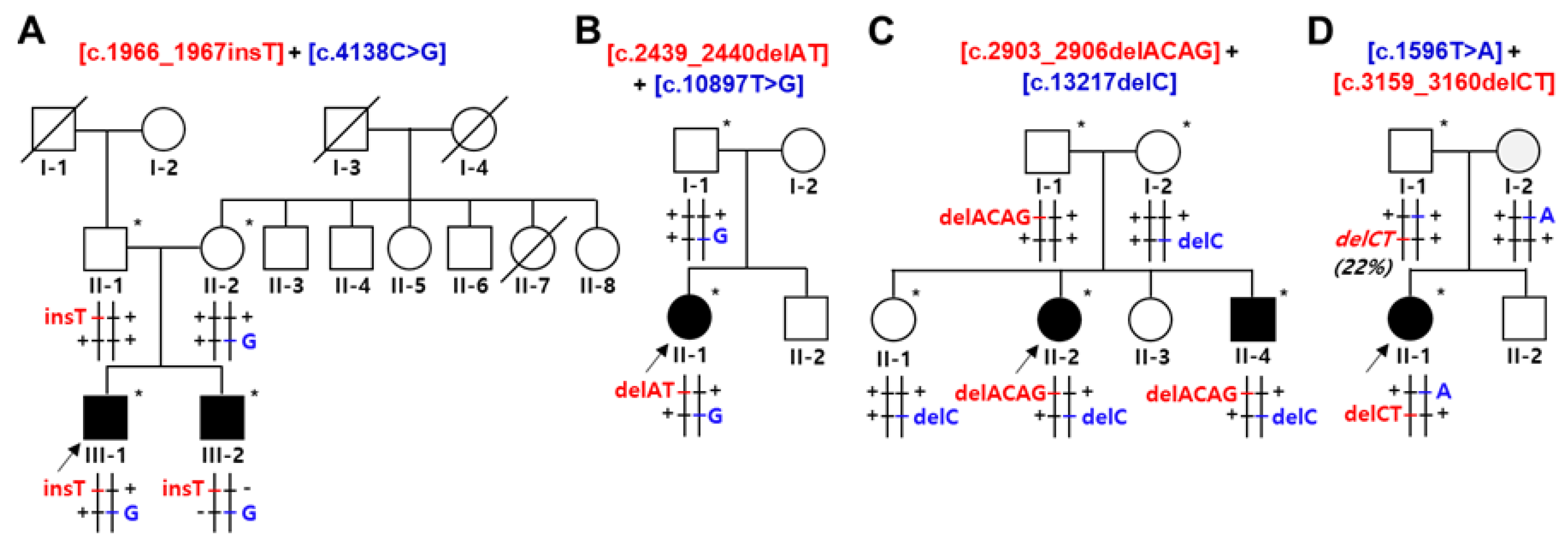
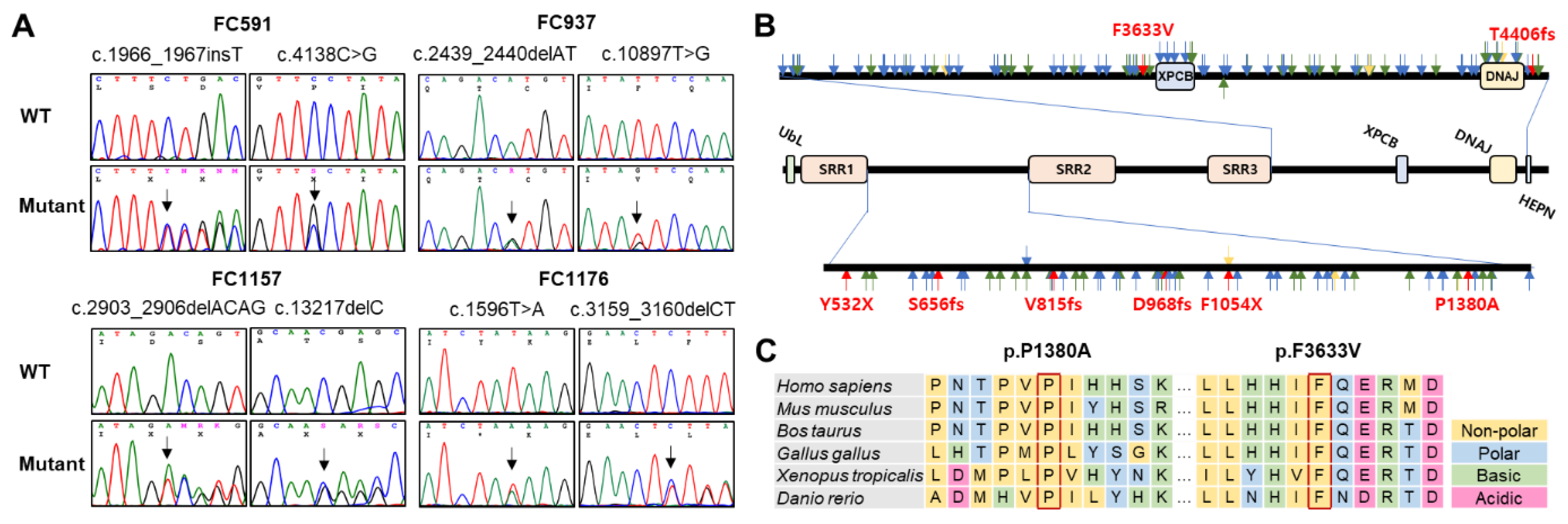
2.1. Identification of Compound Heterozygous SACS Mutations
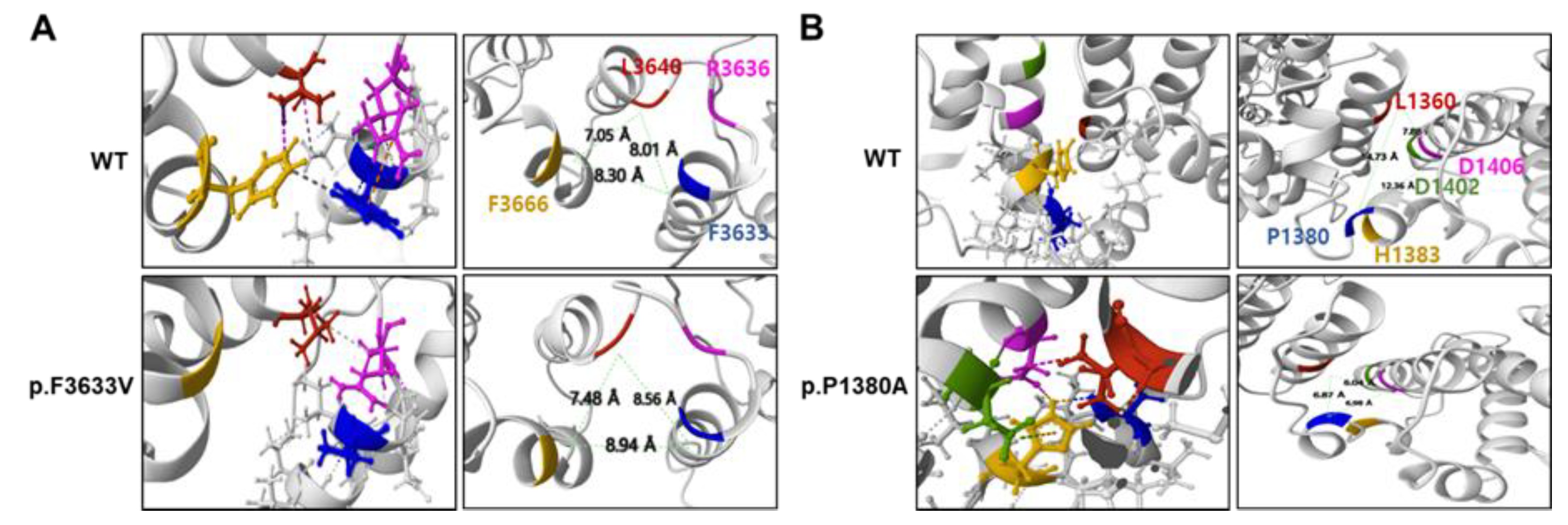
2.2. Prediction of Conformational Changes by Missense Mutations
2.3. Clinical Manifestation
2.4. Electrophysiological Findings
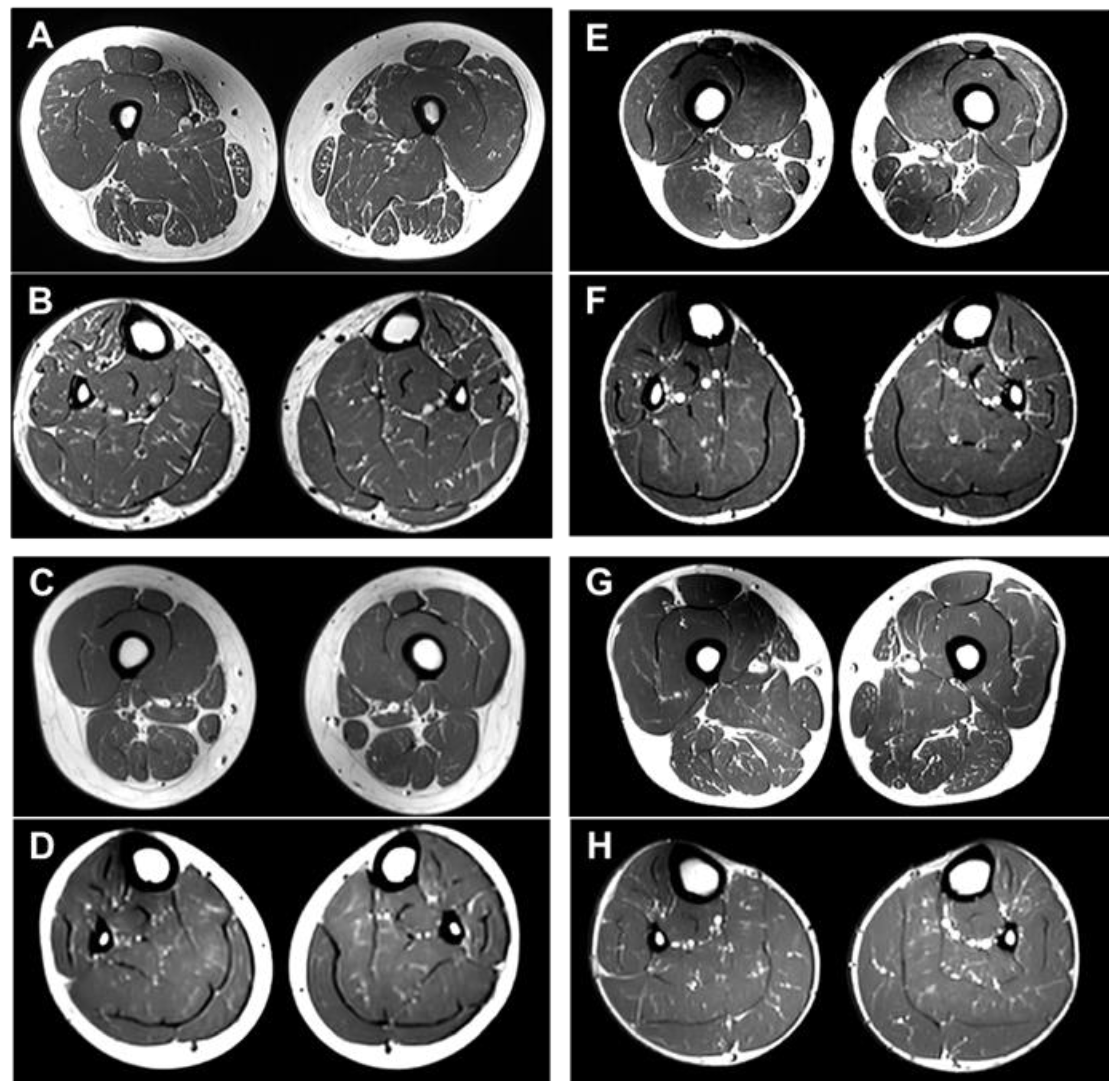
2.5. MRI Features of the Brain and Lower Extremity
3. Discussion
4. Materials and Methods
4.1. Patients
4.2. Molecular Genetic Analysis
4.3. Conservation, Conformational Change, and In Silico Prediction of Mutant Proteins
4.4. Clinical Assessment
4.5. Electrophysiological Examination
4.6. Brain and Lower Extremity MRI
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pipis, M.; Rossor, A.M.; Laura, M.; Reilly, M.M. Next-generation sequencing in Charcot-Marie-Tooth disease: opportunities and challenges. Nat Rev Neurol 2019, 15, 644–656. [Google Scholar] [CrossRef] [PubMed]
- Pareyson, D.; Scaioli, V.; Laura, M. Clinical and electrophysiological aspects of Charcot-Marie-Tooth disease. Neuromolecular Med 2006, 8, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Engert, J.C.; Bérubé, P.; Mercier, J.; Doré, C.; Lepage, P.; Ge, B.; Bouchard, J.P.; Mathieu, J.; Melançon, S.B.; Schalling, M.; et al. ARSACS, a spastic ataxia common in northeastern Québec, is caused by mutations in a new gene encoding an 11.5-kb ORF. Nat Genet 2000, 24, 120–125. [Google Scholar] [CrossRef] [PubMed]
- El Euch-Fayache, G.; Lalani, I.; Amouri, R.; Turki, I.; Ouahchi, K.; Hung, W.Y.; Belal, S.; Siddique, T.; Hentati, F. Phenotypic features and genetic findings in SACSin-related autosomal recessive ataxia in Tunisia. Arch Neurol 2003, 60, 982–988. [Google Scholar] [CrossRef] [PubMed]
- Souza, P.V.S.; Bortholin, T.; Naylor, F.G.M.; Pinto, W.B.V.R.; Oliveira, A.S.B. Early-onset axonal Charcot-Marie-Tooth disease due to SACS mutation. Neuromuscul Disord 2018, 28, 169–172. [Google Scholar] [CrossRef] [PubMed]
- Zaman, Q.; Khan, M.A.; Sahar, K.; Rehman, G.; Khan, H.; Rehman, M.; Najumuddin; Ahmad, I. ; Tariq, M.; Muthaffar, O.Y.; et al. Novel variants in MPV17, PRX, GJB1, and SACS cause Charcot-Marie-Tooth and spastic ataxia of Charlevoix-Saguenay type diseases. Genes 2023, 14, 328. [Google Scholar] [CrossRef] [PubMed]
- Feely, S.M.; Laura, M.; Siskind, C.E.; Sottile, S; Davis, M. ; Gibbons, V.S.; Reilly, M.M.; Shy, M.E. MFN2 mutations cause severe phenotypes in most patients with CMT2A. Neurology 2011, 76, 1690–1696. [Google Scholar] [CrossRef] [PubMed]
- Kumar, K.R.; Blair, N.F.; Sue, C.M. An update on the hereditary spastic paraplegias: new genes and new disease models. Mov Disord Clin 2015, Pract 2, 213–223. [Google Scholar] [CrossRef]
- Toft, A.; Birk, S.; Ballegaard, M.; Duno, M.; Hjermind, L.E.; Nielsen, J.E.; Svenstrup, K. Peripheral neuropathy in hereditary spastic paraplegia caused by REEP1 variants. J Neurol 2019, 266, 735–744. [Google Scholar] [CrossRef]
- Bouchard, J.P.; Barbeau, A.; Bouchard, R.; Bouchard, R.W. Autosomal recessive spastic ataxia of Charlevoix-Saguenay. Can J Neurol Sci 1978, 5, 61–69. [Google Scholar] [CrossRef]
- Engert, J.C.; Dore, C.; Mercier, J.; Ge, B.; Betard, C.; Rioux, J.D.; Owen, C.; Berube, P.; Devon, K.; Birren, B.; et al. Autosomal recessive spastic ataxia of Charlevoix-Saguenay (ARSACS): high-resolution physical and transcript map of the candidate region in chromosome region 13q11. Genomics 1999, 62, 156–164. [Google Scholar] [CrossRef] [PubMed]
- Aly, K.A.; Moutaoufik, M.T.; Zilocchi, M.; Phanse, S.; Babu, M. Insights into SACS pathological attributes in autosomal recessive spastic ataxia of Charlevoix-Saguenay (ARSACS). Curr Opin Chem Biol 2022, 71, 102211. [Google Scholar] [CrossRef] [PubMed]
- Bagaria, J.; Bagyinszky, E.; An, S.S.A. Genetics of autosomal recessive spastic ataxia of Charlevoix-Saguenay (ARSACS) and role of SACSin in neurodegeneration. Int J Mol Sci, 2022, 23, 552. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Li, X.B.; Zi, X.H.; Shen, L.; Hu, ZhM. ; Huang, ShX.; Yu, D.L.; Li, H.B.; Xia, K.; Tang, B.S.; et al. A novel hemizygous SACS mutation identified by whole exome sequencing and SNP array analysis in a Chinese ARSACS patient. J Neurol Sci 2016, 362, 111–114. [Google Scholar] [CrossRef] [PubMed]
- Parfitt, D.A.; Michael, G.J.; Vermeulen, E.G.; Prodromou, N.V.; Webb, T.R.; Gallo, J.M.; Cheetham, M.E.; Nicoll, W.S.; Blatch, G.L.; Chapple, J.P. The ataxia protein SACSin is a functional co-chaperone that protects against polyglutamine-expanded ataxin-1. Hum Mol Genet 2009, 18, 1556–1565. [Google Scholar] [CrossRef] [PubMed]
- Lariviere, R.; Gaudet, R.; Gentil, B.J.; Girard, M.; Conte, T.C.; Minotti, S.; Leclerc-Desaulniers, K.; Gehring, K.; McKinney, R.A.; Shoubridge, E.A.; et al. SACS knockout mice present pathophysiological defects underlying autosomal recessive spastic ataxia of Charlevoix-Saguenay. Hum Mol Genet 2015, 24, 727–739. [Google Scholar] [CrossRef] [PubMed]
- Murtinheira, F.; Migueis, M.; Letra-Vilela, R.; Diallo, M.; Quezada, A.; Valente, C.A.; Oliva, A.; Rodriguez, C.; Martin, V.; Herrera, F. SACSin deletion induces aggregation of glial intermediate filaments. Cells 2022, 11, 299. [Google Scholar] [CrossRef] [PubMed]
- Girard, M.; Lariviere, R.; Parfitt, D.A.; Deane, E.C.; Gaudet, R.; Nossova, N.; Blondeau, F.; Prenosil, G.; Vermeulen, E.G.; Duchen, M.R.; et al. Mitochondrial dysfunction and Purkinje cell loss in autosomal recessive spastic ataxia of Charlevoix-Saguenay (ARSACS). Proc Natl Acad Sci USA 2012, 109, 1661–1666. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Gehring, K. Structural studies of parkin and SACSin: Mitochondrial dynamics in neurodegenerative diseases. Mov Disord 2015, 30, 1610–1619. [Google Scholar] [CrossRef]
- Baets, J.; Deconinck, T.; Smets, K.; Goossens, D.; Van den Bergh, P.; Dahan, K.; Schmedding, E.; Santens, P.; Rasic, V.M.; Van Damme, P.; et al. Mutations in SACS cause atypical and late-onset forms of ARSACS. Neurology 2010, 75, 1181–1188. [Google Scholar] [CrossRef]
- Synofzik, M.; Soehn, A.S.; Gburek-Augustat, J.; Schicks, J.; Karle, K.N.; Schule, R.; Haack, T.B.; Schoning, M.; Biskup, S.; Rudnik-Schoneborn, S.; et al. Autosomal recessive spastic ataxia of Charlevoix Saguenay (ARSACS): expanding the genetic, clinical and imaging spectrum. Orphanet J Rare Dis 2013, 8, 41. [Google Scholar] [CrossRef] [PubMed]
- Miyatake, S.; Miyake, N.; Doi, H.; Saitsu, H.; Ogata, K.; Kawai, M.; Matsumoto, N. A novel SACS mutation in an atypical case with autosomal recessive spastic ataxia of Charlevoix-Saguenay (ARSACS). Intern Med 2012, 51, 2221–2226. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, F.; Doi, H.; Kunii, M. Autosomal recessive spinocerebellar ataxias in Japan. Rinsho Shinkeigaku 2016, 56, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Aida, I.; Ozawa, T.; Fujinaka, H.; Goto, K.; Ohta, K.; Nakajima, T. Autosomal recessive spastic ataxia of Charlevoix-Saguenay without spasticity. Intern Med 2021, 60, 3963–3967. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.; Tang, J.G.; Yang, Y.F.; Tan, Z.P.; Tan, J.Q. A novel homozygous SACS mutation identified by whole-exome sequencing in a consanguineous family with autosomal recessive spastic ataxia of Charlevoix-Saguenay. Cytogenet Genome Res 2017, 152, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Lu, X.; Jin, Y.; Li, D.; Ye, X.; Tao, C.; Zhou, M.; Jiang, H.; Yu, H. A novel SACS variant identified in a Chinese patient: case report and review of the literature. Front Neurol 2022, 13, 845318. [Google Scholar] [CrossRef] [PubMed]
- Kwon, K.Y.; Huh, K.; Eun, B.L.; Yoo, H.W.; Kamsteeg, E.J.; Scheffer, H.; Koh, S.B. A probable Korean case of autosomal recessive spastic ataxia of Charlevoix-Saguenay. Can J Neurol Sci 2015, 42, 271–273. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.; Lyoo, C.H.; Park, S.E.; Seo, Y.; Han, S.H.; Han, J. Optical coherence tomography findings facilitate the diagnosis of autosomal recessive spastic ataxia of Charlevoix-Saguenay. Korean J Ophthalmol 2021, 35, 330–331. [Google Scholar] [CrossRef] [PubMed]
- Bong, J.B.; Kim, S.W.; Lee, S.-T.; Choi, J.R.; Shin, H.Y. Autosomal recessive spastic ataxia of Charlevoix-Saguenay. J Korean Neurol Assoc 2019, 37, 69–72. [Google Scholar] [CrossRef]
- Hammer, M.B.; Eleuch-Fayache, G.; Gibbs, J.R.; Arepalli, S.K.; Chong, S.B.; Sassi, C.; Bouhlal, Y.; Hentati, F.; Amouri, R.; Singleton, A.B. Exome sequencing: an efficient diagnostic tool for complex neurodegenerative disorders. Eur J Neurol 2013, 20, 486–492. [Google Scholar] [CrossRef]
- Shakya, S.; Kumari, R.; Suroliya, V.; Tyagi, N.; Joshi, A.; Garg, A.; Singh, I.; Kalikavil Puthanveedu, D.; Cherian, A.; Mukerji, M.; et al. Whole exome and targeted gene sequencing to detect pathogenic recessive variants in early onset cerebellar ataxia. Clin Genet 2019, 96, 566–574. [Google Scholar] [CrossRef] [PubMed]
- Panwala, T.F.; Garcia-Santibanez, R.; Vizcarra, J.A.; Garcia, A.G.; Verma, S. Childhood-onset hereditary spastic paraplegia (HSP): a case series and review of literature. Pediatr Neurol 2022, 130, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Karakaya, M.; Storbeck, M.; Strathmann, E.A.; Delle Vedove, A.; Holker, I.; Altmueller, J.; Naghiyeva, L.; Schmitz-Steinkruger, L.; Vezyroglou, K.; Motameny, S.; et al. Targeted sequencing with expanded gene profile enables high diagnostic yield in non-5q-spinal muscular atrophies. Hum Mutat 2018, 39, 1284–1298. [Google Scholar] [CrossRef] [PubMed]
- Naz, S.; Imtiaz, A.; Mujtaba, G.; Maqsood, A.; Bashir, R.; Bukhari, I.; Khan, M.R.; Ramzan, M.; Fatima, A.; Rehman, A.U.; et al. Genetic causes of moderate to severe hearing loss point to modifiers. Clin Genet 2017, 91, 589–598. [Google Scholar] [CrossRef] [PubMed]
- Kanwal, S.; Choi, Y.J.; Lim, S.O.; Choi, H.J.; Park, J.H.; Nuzhat, R.; Khan, A.; Perveen, S.; Choi, B.O.; Chung, K.W. Novel homozygous mutations in Pakistani families with Charcot-Marie-Tooth disease. BMC Med Genomics 2021, 14, 174. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.J.; Kanwal, S.; Hameed, R.; Tamanna, N.; Perveen, S.; Mahreen, H.; Son, W.; Lee, K.S.; Chung, K.W. Biallelic mutations in pakistani families with autosomal recessive prelingual nonsyndromic hearing loss. Genes Genomics 2023, 45, 145–156. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Jung, S.C.; Hong, Y.B.; Yoo, J.H.; Koo, H.; Lee, J.H.; Hong, H.D.; Kim, S.B.; Chung, K.W.; Choi, B.O. Recessive optic atrophy, sensorimotor neuropathy and cataract associated with novel compound heterozygous mutations in OPA1. Mol Med Rep 2016, 14, 33–40. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.J.; Nam, S.H.; Park, J.M.; Kanwal, S.; Choi, Y.J.; Lee, H.J.; Lee, K.S.; Lee, J.E.; Park, J.S.; Choi, B.O.; et al. ; Compound heterozygous mutations of SH3TC2 in Charcot-Marie-Tooth disease type 4C patients. J Hum Genet 2019, 64, 961–965. [Google Scholar] [CrossRef] [PubMed]
- Paternostro-Sluga, T.; Grim-Stieger, M.; Posch, M.; Schuhfried, O.; Vacariu, G.; Mittermaier, C.; Bittner, C.; Fialka-Moser, V. Reliability and validity of the Medical Research Council (MRC) scale and a modified scale for testing muscle strength in patients with radial palsy. J Rehabil Med 2008, 40, 665–671. [Google Scholar] [CrossRef]
- Goutallier, D.; Postel, J.M.; Bernageau, J.; Lavau, L.; Voisin, M.C. Fatty muscle degeneration in cuff ruptures: pre- and postoperative evaluation by CT scan. Clin Orthop Relat Res 1994, 304, 78–83. [Google Scholar] [CrossRef]
| Family ID | Mutations | Clinical phenotype | ACMG/AMP | |
|---|---|---|---|---|
| Nucleotide 1 | Amino acid 1 | |||
| FC591 | [c.1966_1967insT] + [c.4138C>G] | [p.S656Ffs*1] + [p.P1380A] | CMT, cerebellar ataxia, spasticity, and HL | P p |
| FC937 | [c.2439_2440delAT] + [c.10897T>G] | [p.V815Gfs*2] + [p.F3633V] | CMT, cerebellar ataxia, and spasticity, HL | P LP |
| FC1157 | [c.2903_2906delACAG] + [c.13217delC] | [p.D968Vfs*13] + [p.T4406Rfs*45] | CMT, cerebellar ataxia, spasticity, and HL | P P |
| FC1176 | [c.1596T>A] + [c.3159_3160delCT] | [p.Y532X] + [p.F1054X] | CMT, cerebellar ataxia, spasticity, and HL | P P |
| Variants 1 | dbSNP Acc. No. | Mutant allele frequencies | GERP | In silico analyses 2 | References | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| IGSR | gnomAD | KRGDB | MutT | REVEL | PP2 | MU | ||||
| p.Y532X | rs2137720760 | - | - | - | -2.77 | 200/0* | - | - | - | |
| p.S656Ffs*1 | - | - | - | - | 5.14 | 200/0* | - | - | - | |
| p.V815Gfs*2 | rs775059063 | - | 0.0000142 | - | -1.5 | 200/0* | - | - | - | 30-32 |
| p.D968Vfs*13 | rs1259615333 | - | 0.0000099 | - | 3.71 | 199/1* | - | - | - | 33 |
| p.F1054X | rs2137637877 | - | - | - | 5.09 | 200/0* | - | - | - | |
| p.P1380A | - | - | - | - | 6.06 | 66/34* | 0.637* | 0.965* | -0.757* | |
| p.F3633V | rs1382541188 | - | 0.000013 | - | 6.04 | 72/28* | 0.626* | 0.999* | -0.292* | |
| p.T4406Rfs*45 | - | - | - | - | 5.85 | 198/2* | - | - | - | |
| Mutation | Protein stability prediction tools 1 | ||
|---|---|---|---|
| PremPS | MAESTROweb | DynaMut2 | |
| p.F3633V | 0.68* | 0.283* | -0.73* |
| p.P1380A | 0.91* | 0.121* | -1.10* |
| Family ID | FC591 | FC937 | FC1157 | FC1176 | ||
|---|---|---|---|---|---|---|
| Patients | III-1 | III-2 | II-1 | II-2 | II-4 | II-1 |
| Mutations | p.S656Ffs*1 + p.P1380A |
p.V815Gfs*2 + p.F3633V |
p.D968Vfs*13 + p.T4406Rfs*45 |
p.Y532X+ p.F1054X | ||
| Sex | M | M | F | F | M | M |
| Examined age (yrs) | 35 | 33 | 27 | 26 | 25 | 22 |
| Onset age (yrs) | 4 | 5 | 15 | 17 | 15 | 3 |
| Muscle weakness | ||||||
| Upper limb (MRC) | ||||||
| Proximal (Rt/Lt) 1 | 4+/4+ | 4+/4+ | 4/4 | 4+/4+ | 4+/4+ | 4/4 |
| Distal (Rt/Lt) 2 | 4+/4+ | 4+/4+ | 4/4 | 4+/4+ | 4+/4+ | 4/4 |
| Lower limb (MRC) | ||||||
| Proximal (Rt/Lt) 3 | 4/4 | 4+/4+ | 4-/4- | 4+/4+ | 4/4 | 4/4 |
| Distal (Rt/Lt) 4 | 4/4 | 4+/4+ | 4-/4- | 4+/4+ | 4/4 | 4/4 |
| Muscle atrophy | Mild | Minimal | Moderate | Minimal | Mild | Mild |
| Sensory disturbance | Yes | Yes | Yes | Yes | Yes | Yes |
| DTR | ||||||
| Biceps jerk reflex | ++ | ++ | +++ | +++ | +++ | +++ |
| Knee jerk reflex | +++ | +++ | +++ | +++ | +++ | +++ |
| Disability score | ||||||
| FDS | 2 | 2 | 4 | 2 | 4 | 4 |
| CMTNSv2 | 13 | 11 | 24 | 17 | 22 | 23 |
| Pyramidal sign | Yes | Yes | Yes | Yes | Yes | Yes |
| Spasticity | Yes | Yes | Yes | Yes | Yes | Yes |
| Cerebellar ataxia | Yes | Yes | Yes | Yes | Yes | Yes |
| Dysarthria | No | No | Yes | Yes | No | Yes |
| Nystagmus | Yes | Yes | Yes | Yes | Yes | Yes |
| Foot deformity | Yes | Yes | Yes | Yes | Yes | Yes |
| Intellectual disability | No | No | No | No | No | No |
| Hearing loss | Yes | Yes | Yes | Yes | Yes | Yes |
| Electrophysiology | ||||||
| Nerve conduction | HMSN | HMSN | HMSN | HMSN | HMSN | HMSN |
| BAEP | NA | NA | Abnormal | Abnormal | NA | Abnormal |
| Brain MRI | NA | NA | NA | Cerebellar atrophy | Cerebellar atrophy | Cerebellar atrophy |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




