Submitted:
09 April 2024
Posted:
10 April 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Data Sets
2.2. Phylogeny
2.3. Conserved Domain Analysis
2.4. Plant Growth Conditions
2.5. Genetic Constructs
2.6. Transient Transformation
2.7. Nuclei Staining
2.8. Fluorescence Microscopy
2.9. Molecular Modeling
2.10. Molecular Dynamic Simulations
2.11. Analysis of Promoter cis-Acting Regulatory Elements in Grasses
3. Results
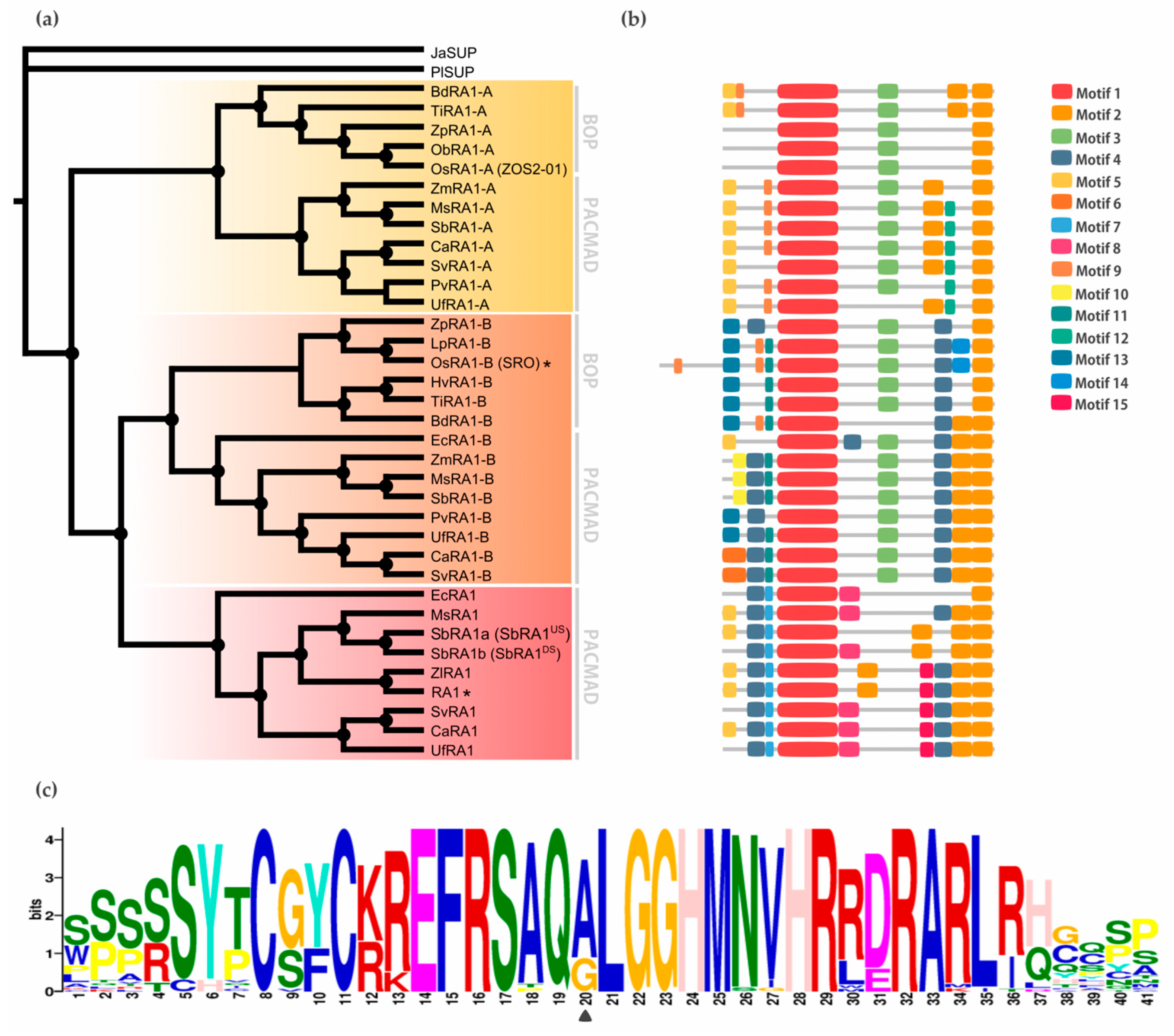
3.1. Phylogenetic Analysis
3.1.1. SUP Evolution and the Origin of RA1
3.1.2. Evolution of RA1 in Grasses
3.2. Conserved Motifs Analysis among RA1 and RA1-Like Coding Sequences
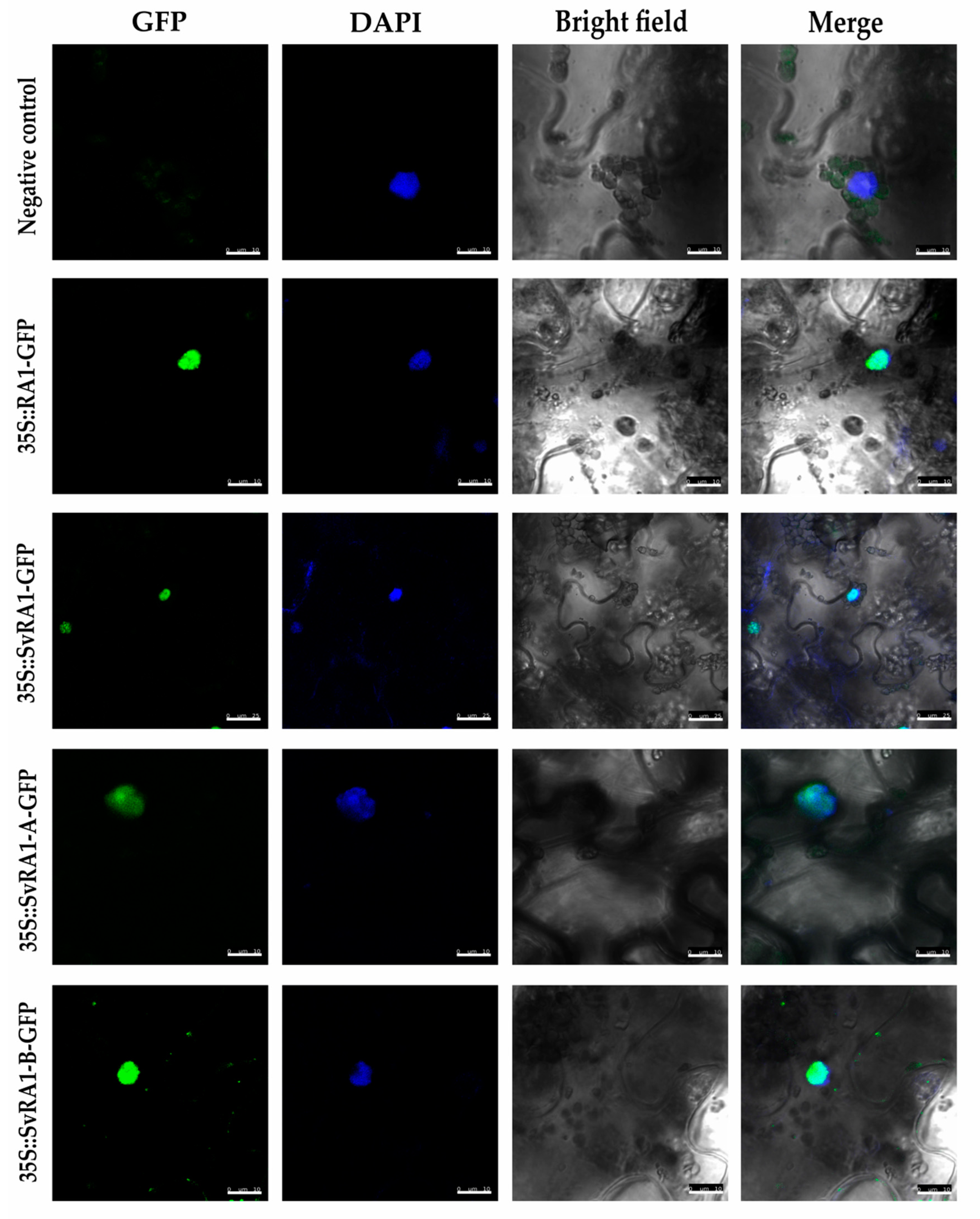
3.3. Subcellular Localization
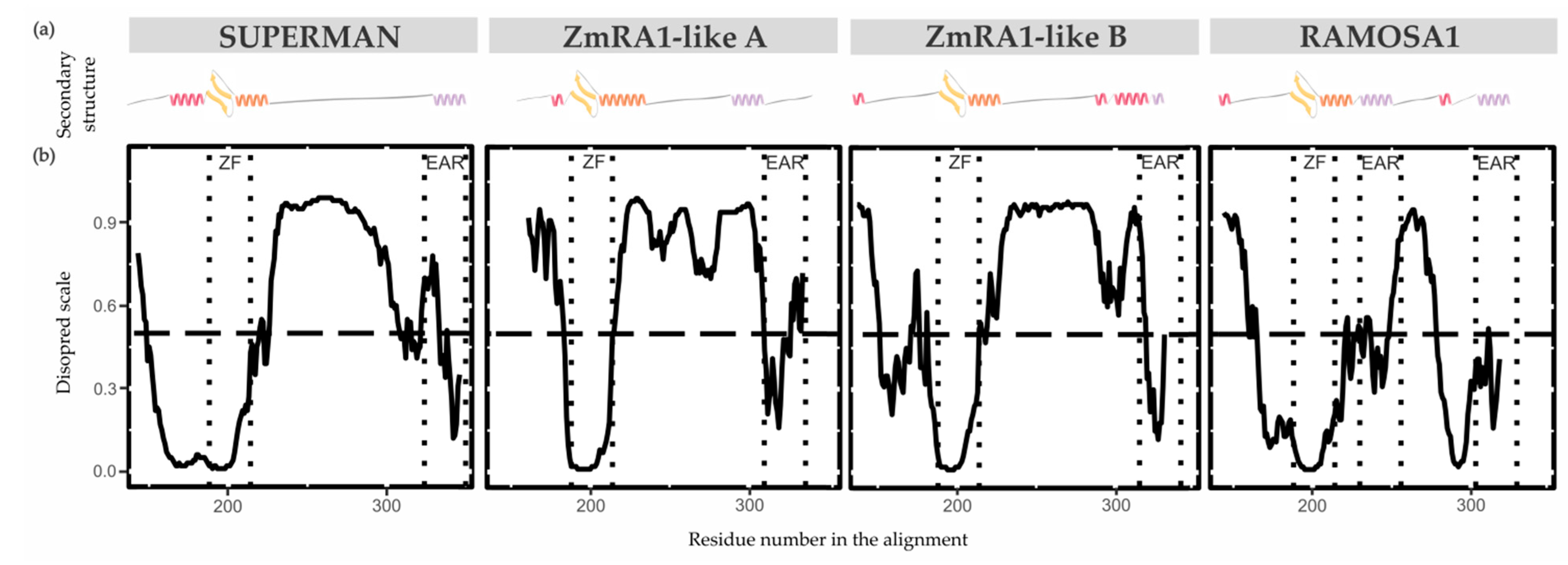
3.4. Molecular Modeling
3.4.1. Proteins Secondary Structure and Conformational Plasticity
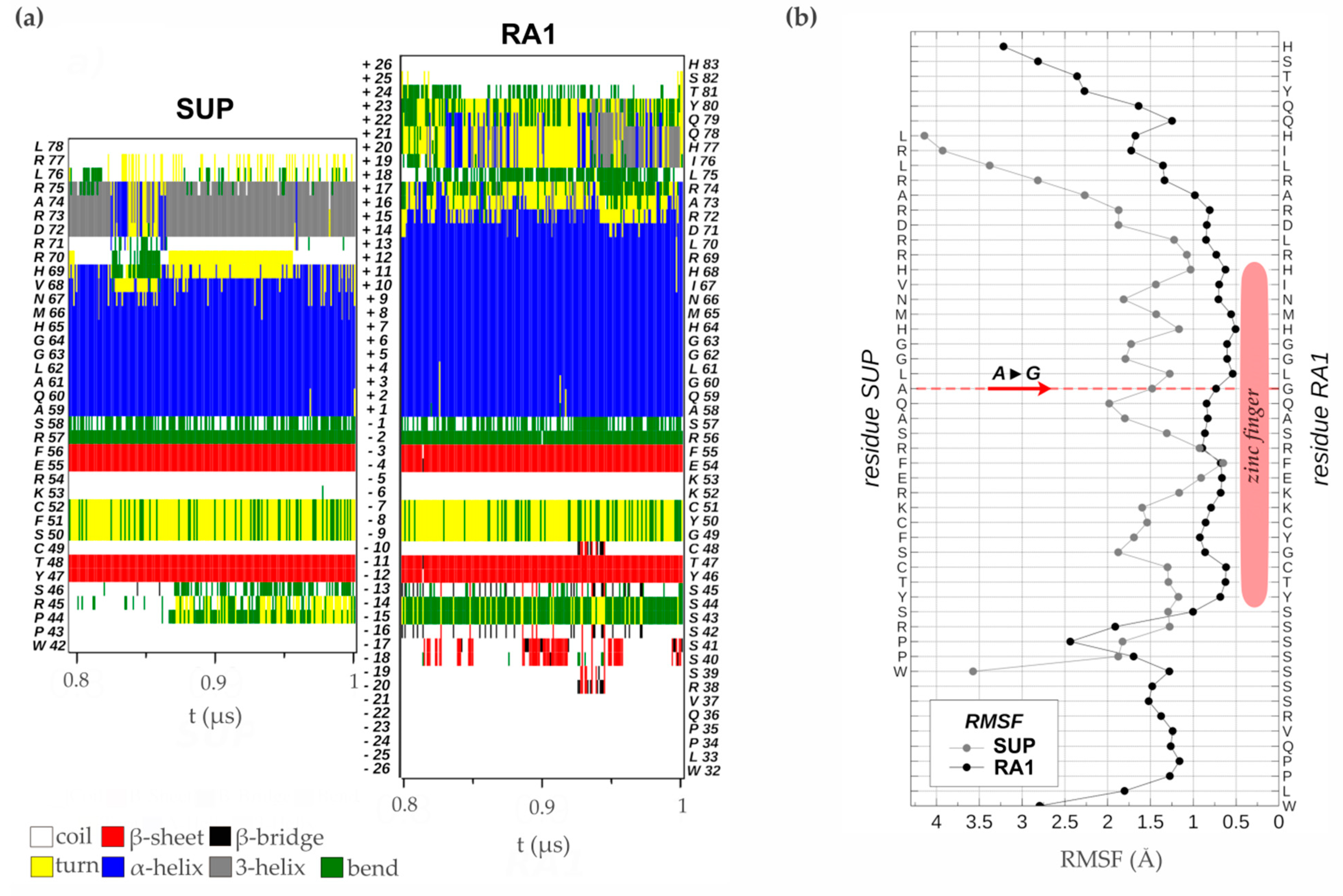
3.4.2. Zinc Finger Molecular Dynamics Simulation
3.5. Cis-Acting Regulatory Elements Analysis among RA1 and RA1-Like Promoter Sequences
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Martienssen, R.; Vollbrecht, E. Nucleotide Sequences Enconding Ramosa 1 Gene and Methods of Use for Same. WO2001090343A2. 2001. [Google Scholar]
- Vollbrecht, E.; Springer, P.S.; Goh, L.; Buckler IV, E.S.; Martienssen, R. Architecture of Floral Branch Systems in Maize and Related Grasses. Nature 2005, 436, 1119–1126. [Google Scholar] [CrossRef] [PubMed]
- Yang, X. Study of RAMOSA1 Function during Maize Inflorescence Development. Ph.D. Thesis, Iowa State University, Ames, IA, USA, 2011. [Google Scholar]
- Gallavotti, A.; Long, J.A.; Stanfield, S.; Yang, X.; Jackson, D.; Vollbrecht, E.; Schmidt, R.J. The Control of Axillary Meristem Fate in the Maize Ramosa Pathway. Developmen, 2010, 137, 2849–2856. [Google Scholar]
- Eveland, A.L.; Goldshmidt, A.; Pautler, M.; Morohashi, K.; Liseron-Monfils, C.; Lewis, M.W.; Kumari, S.; Hirag, S.; Yang, F.; Unger-Wallace, E.; et al. Regulatory Modules Controlling Maize Inflorescence Architecture. Genome Res. 2014, 24, 431–443. [Google Scholar] [CrossRef] [PubMed]
- Strable, J.; Unger-Wallace, E.; Raygoza, A.A.; Briggs, S.; Vollbrecht, E. Interspecies Transfer of RAMOSA1 Orthologs and Promoter Cis Sequences Impacts Maize Inflorescence Architecture. Plant Physiol. 2023, 191, 1084–1101. [Google Scholar] [CrossRef] [PubMed]
- Sigmon, B.; Vollbrecht, E. Evidence of Selection at the Ramosa1 Locus during Maize Domestication. Mol. Ecol. 2010, 19, 1296–1311. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.L.; Sakai, H.; Jack, T.; Weigel, D.; Mayer, U.; Meyerowitz, E.M. SUPERMAN, a Regulator of Floral Homeotic Genes in Arabidopsis. Development 1992, 114, 599–615. [Google Scholar] [CrossRef]
- Cassani, E.; Landoni, M.; Pilu, R. Characterization of the Ra1 Maize Gene Involved in Inflorescence Architecture. Sex. Plant Reprod. 2006, 19, 145–150. [Google Scholar] [CrossRef]
- Landoni, M.; Cassani, E.; Pilu, R. Arabidopsis Thaliana Plants Overexpressing Ramosa1 Maize Gene Show an Increase in Organ Size Due to Cell Expansion. Sex. Plant Reprod. 2007, 20, 191–198. [Google Scholar] [CrossRef]
- Estep, M.C.; Vela Diaz, D.M.; Zhong, J.; Kellogg, E.A. Eleven Diverse Nuclear-Encoded Phylogenetic Markers for the Subfamily Panicoideae (Poaceae). Am. J. Bot. 2012, 99, 443–446. [Google Scholar] [CrossRef]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A Comparative Platform for Green Plant Genomics. Nucleic Acids Res. 2012, 40, 1178–1186. [Google Scholar] [CrossRef] [PubMed]
- Sayers, E.W.; Bolton, E.E.; Brister, J.R.; Canese, K.; Chan, J.; Comeau, D.C.; Connor, R.; Funk, K.; Kelly, C.; Kim, S.; et al. Database Resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2022, 50, D20–D26. [Google Scholar] [CrossRef] [PubMed]
- Tello-Ruiz, M.K.; Jaiswal, P.; Ware, D. Gramene: A Resource for Comparative Analysis of Plants Genomes and Pathways; Springer US: New York, NY, USA, 2022. [Google Scholar]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE : Tools for Motif Discovery and Searching. Nucleic Acids Research 2009, 37, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT Online Service: Multiple Sequence Alignment, Interactive Sequence Choice and Visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. Mrbayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice across a Large Model Space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. Creating the CIPRES Science Gateway for Inference of Large Phylogenetic Trees. In Proceedings of the Gateway Computing Environments Workshop, New Orleans, LA, USA, 14 November 2010; Volume 14, pp. 1–6. [Google Scholar]
- Maddison, W.P.; Maddison, D.R. Mesquite: A Modular System for Evolutionary Analysis. Evolution 2008, 62, 1103–1118. [Google Scholar]
- Lovell, J.T.; Sreedasyam, A.; Schranz, M.E.; Wilson, M.; Carlson, J.W.; Harkess, A.; Emms, D.; Goodstein, D.M. GENESPACE Tracks Regions of Interest and Gene Copy Number Variation across Multiple Genomes. eLife 2022, 11, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.; Elkan, C. Fitting a Mixture Model by Expectation Maximization to Discover Motifs in Biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Hartley, J.L.; Temple, G.F.; Brasch, M.A. DNA Cloning Using in Vitro Site-Specific Recombination. Genome Res, 2000, 10, 1788–1795. [Google Scholar] [CrossRef]
- Hellens, R.P.; Anne Edwards, E.; Leyland, N.R.; Bean, S.; Mullineaux, P.M. PGreen: A Versatile and Flexible Binary Ti Vector for Agrobacterium-Mediated Plant Transformation. Plant Mol. Biol. 2000, 42, 819–832. [Google Scholar] [CrossRef] [PubMed]
- Voinnet, O.; Rivas, S.; Mestre, P.; Baulcombe, D. An Enhanced Transient Expression System in Plants Based on Suppression of Gene Silencing by the P19 Protein of Tomato Bushy Stunt Virus. Plant J. 2003, 33, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Belda-Palazón, B.; Ruiz, L.; Martí, E.; Tárraga, S.; Tiburcio, A.F.; Culiáñez, F.; Farràs, R.; Carrasco, P.; Ferrando, A. Aminopropyltransferases Involved in Polyamine Biosynthesis Localize Preferentially in the Nucleus of Plant Cells. PLoS ONE 2012, 7, e46907. [Google Scholar] [CrossRef] [PubMed]
- Felippes, F.F. De; Weigel, D. Transient Assays for the Analysis of MiRNA Processing and Function. In Plant MicroRNAs; Meyers, B.C., Green, P.J., Eds.; Humana Press: Totowa, NJ, USA, 2009. [Google Scholar]
- Jones, D.T. Protein Secondary Structure Prediction Based on Position-Specific Scoring Matrices. J. Mol. Biol. 1999, 292, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Cozzetto, D. DISOPRED3: Precise Disordered Region Predictions with Annotated Protein-Binding Activity. Bioinformatics 2015, 31, 857–863. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindah, E. Gromacs: High Performance Molecular Simulations through Multi-Level Parallelism from Laptops to Supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J.L.; Dror, R.O.; Shaw, D.E. Improved Side-Chain Torsion Potentials for the Amber Ff99SB Protein Force Field. Proteins Struct. Funct. Bioinform. 2010, 78, 1950–1958. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A Smooth Particle Mesh Ewald Method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A Linear Constraint Solver for Molecular Simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Miyamoto, S.; Kollman, P.A. Settle: An Analytical Version of the SHAKE and RATTLE Algorithm for Rigid Water Models. J. Comput. Chem. 1992, 13, 952–962. [Google Scholar] [CrossRef]
- Solovyev, V.V.; Shahmuradov, I.A.; Salamov, A.A. Computational Biology of Transcription Factor Binding; Humana Press: Totowa, NJ, USA, 2010; Volume 674. [Google Scholar]
- Castro-Mondragon, J.A.; Riudavets-Puig, R.; Rauluseviciute, I.; Berhanu Lemma, R.; Turchi, L.; Blanc-Mathieu, R.; Lucas, J.; Boddie, P.; Khan, A.; Perez, N.M.; et al. JASPAR 2022: The 9th Release of the Open-Access Database of Transcription Factor Binding Profiles. Nucleic Acids Res. 2021, 50, D165–D173. [Google Scholar] [CrossRef] [PubMed]
- Grant, C.E.; Bailey, T.L. XSTREME: Comprehensive Motif Analysis of Biological Sequence Datasets. bioRxiv, 4587. [Google Scholar]
- Ohta, M.; Matsui, K.; Hiratsu, K.; Shinshi, H.; Ohme-Takagi, M. Repression Domains of Class II ERF Transcriptional Repressors Share an Essential Motif for Active Repression. Plant Cell 2001, 13, 1959–1968. [Google Scholar] [CrossRef] [PubMed]
- Hiratsu, K.; Ohta, M.; Matsui, K.; Ohme-Takagi, M. The SUPERMAN Protein Is an Active Repressor Whose Carboxy-Terminal Repression Domain Is Required for the Development of Normal Flowers. FEBS Lett. 2002, 514, 351–354. [Google Scholar] [CrossRef] [PubMed]
- Hiratsu, K.; Mitsuda, N.; Matsui, K.; Ohme-Takagi, M. Identification of the Minimal Repression Domain of SUPERMAN Shows That the DLELRL Hexapeptide Is Both Necessary and Sufficient for Repression of Transcription in Arabidopsis. Biochem. Biophys. Res. Commun. 2004, 321, 172–178. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, S.B.; Hagen, G.; Guilfoyle, T.J. Aux/IAA Proteins Contain a Potent Transcriptional Repression Domain. Plant Cell 2004, 16, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Ke, J.; Ma, H.; Gu, X.; Thelen, A.; Brunzelle, J.S.; Li, J.; Xu, H.E.; Melcher, K. Structural Basis for Recognition of Diverse Transcriptional Repressors by the TOPLESS Family of Corepressors. Sci. Adv. 2015, 1, e1500107. [Google Scholar] [CrossRef] [PubMed]
- Dathan, N.; Zaccaro, L.; Esposito, S.; Insernia, C.; Omichinski, J.G.; Riccio, A.; Pedone, C.; Di Blasio, B.; Fattorusso, R.; Pedone, P.V. The Arabidopsis SUPERMAN Protein Is Able to Specifically Bind DNA through Its Single Cys2-His2 Zinc Finger Motif. Nucleic Acids Res. 2002, 30, 4945–4951. [Google Scholar] [CrossRef]
- Isernia, C.; Bucci, E.; Leone, M.; Zaccaro, L.; Di Lello, P.; Digilio, G.; Esposito, S.; Saviano, M.; Di Blasio, B.; Pedone, C.; et al. NMR Structure of the Single QALGGH Zinc Finger Domain from the Arabidopsis Thaliana SUPERMAN Protein. ChemBioChem 2003, 4, 171–180. [Google Scholar] [CrossRef]
- Buchan, D.W.A.; Jones, D.T. The PSIPRED Protein Analysis Workbench: 20 Years On. Nucleic Acids Res. 2019, 47, W402–W407. [Google Scholar] [CrossRef]
- Sakai, H.; Medrano, L.J.; Meyerowitz, E.M. Role of SUPERMAN in Maintaining Araidopsis Floral Whorl Boundaries. Nature 1995, 378, 199–202. [Google Scholar] [CrossRef]
- Bennetzen, J.L. Patterns in Grass Genome Evolution. Curr. Opin. Plant Biol., 2007, 10, 176–181. [Google Scholar] [CrossRef] [PubMed]
- Schubert, M.; Marcussen, T.; Meseguer, A.S.; Fjellheim, S. The Grass Subfamily Pooideae: Cretaceous–Palaeocene Origin and Climate-Driven Cenozoic Diversification. Glob. Ecol. Biogeogr. 2019, 28, 1168–1182. [Google Scholar] [CrossRef]
- Huang, W.; Zhang, L.; Columbus, J.T.; Hu, Y.; Zhao, Y.; Tang, L.; Guo, Z.; Chen, W.; McKain, M.; Bartlett, M.; et al. A Well-Supported Nuclear Phylogeny of Poaceae and Implications for the Evolution of C4 Photosynthesis. Mol. Plant 2022, 15, 755–777. [Google Scholar] [CrossRef] [PubMed]
- Preston, J.C.; Kellogg, E.A. Reconstructing the Evolutionary History of Paralogous APETALA1/FRUITFULL- like Genes in Grasses (Poaceae). Genetics 2006, 174, 421–437. [Google Scholar] [CrossRef] [PubMed]
- McKain, M.R.; Tang, H.; McNeal, J.R.; Ayyampalayam, S.; Davis, J.I.; Depamphilis, C.W.; Givnish, T.J.; Chris Pires, J.; Stevenson, D.W.; Leebens-Mack, J.H. A Phylogenomic Assessment of Ancient Polyploidy and Genome Evolution across the Poales. Genome Biol. Evol. 2016, 8, 1150–1164. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Sakai, H.; Meyerowitz, E.M. Whorl-Specific Expression of the SUPERMAN Gene of Arabidopsis Is Mediated by Cis Elements in the Transcribed Region. Curr. Biol. 2003, 13, 1524–1530. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Jacobsen, S.E. Locus-Specific Control of Asymmetric and CpNpG Methylation by the DRM and CMT3 Methyltransferase Genes. Proc. Natl. Acad. Sci. USA 2002, 99, 16491–16498. [Google Scholar] [CrossRef] [PubMed]
- Kubo, K.I.; Sakamoto, A.; Kobayashi, A.; Rybka, Z.; Kanno, Y.; Nakagawa, H.; Nishino, T.; Takatsuji, H. CYs2/His2 Zinc-Finger Protein Family of Petunia: Evolution and General Mechanism of Target-Sequence Recognition. Nucleic Acids Res. 1998, 26, 608–615. [Google Scholar] [CrossRef]
- Kagale, S.; Links, M.G.; Rozwadowski, K. Genome-Wide Analysis of Ethylene-Responsive Element Binding Factor-Associated Amphiphilic Repression Motif-Containing Transcriptional Regulators in Arabidopsis. Plant Physiol. 2010, 152, 1109–1134. [Google Scholar] [CrossRef]
- Chow, V.; Kirzinger, M.W.; Kagale, S. Lend Me Your EARs: A Systematic Review of the Broad Functions of EAR Motif-Containing Transcriptional Repressors in Plants. Genes 2023, 14, 270. [Google Scholar] [CrossRef]
- Terao, J. SUPERMAN, the Guardian of Floral Organ Gene Expression in Arabidopsis thaliana. Master’s Thesis, American University, Washington, DC, USA, 2019. [Google Scholar]
- Berg, J.M.; Godwin, H.A. Lessons from zinc-binding peptides. Annu. Rev. Biophys. Biomol. Struct. 1997, 26:1, 357–371.
- Takatsuji, H. Zinc-Finger Transcription Factors in Plants. Cell. Mol. Life Sci. 1998, 54, 582–596. [Google Scholar] [CrossRef] [PubMed]
- Takatsuji, H. Zinc-Finger Proteins: The Classical Zinc Finger Emerges in Contemporary Plant Science. Plant Mol. Biol. 1999, 39, 1073–1078. [Google Scholar] [CrossRef]
- Lee, M.I.N.S.; Gippert, G.P.; Soman, K.V.; Case, D.A.; Wright, P.E. Three-Dimensional Solution Structure of a Single Zinc Finger DNA-Binding Domain. Science 1989, 245, 635–637. [Google Scholar] [CrossRef]
- Kochoyan, M.; Keutmannt, H.T.; Weiss, M.A. Architectural Rules of the Zinc-Finger Motif: Comparative Two-Dimensional NMR Studies of Native and “Aromatic-Swap” Domains Define. Proc. Natl. Acad. Sci. USA 1991, 88, 8455–8459. [Google Scholar] [CrossRef]
- Michael, S.F.; Kilfoil, V.J.; Schmidt, M.H.; Amann, B.T.; Berg, J.M. Metal Binding and Folding Properties of a Minimalist Cys2His2 Zinc Finger Peptide. Proc. Natl. Acad. Sci. USA 1992, 89, 4796–4800. [Google Scholar] [CrossRef] [PubMed]
- Fairall, L.; Schwabe, J.W.R.; Chapman, L.; Finch, J.T.; Rhodes, D. The Crystal Structure of a Two Zinc-Finger Peptide Reveals an Extension to the Rules for Zinc-Finger/DNA Recognition. Nature 1993, 366, 483–487. [Google Scholar] [CrossRef] [PubMed]
- Elrod-Erickson, M.; Rould, M.A.; Nekludova, L.; Pabo, C.O. Zif268 Protein-DNA Complex Refined at 1.6 Å: A Model System for Understanding Zinc Finger-DNA Interactions. Structure 1996, 4, 1171–1180. [Google Scholar] [CrossRef]
- Houbaviy, H.B.; Usheva, A.; Shenk, T.; Burley, S.K. Cocrystal Structure of YY1 Bound to the Adeno-Associated Virus P5 Initiator. Proc. Natl. Acad. Sci. USA 1996, 93, 13577–13582. [Google Scholar] [CrossRef]
- Omichinski, J.G.; Pedone, P.V.; Felsenfeld, G.; Gronenborn, A.M.; Clore, G.M. The Solution Structure of a Specific GAGA Factor-DNA Complex Reveals a Modular Binding Mode. Nat. Struct. Biol. 1997, 4, 122–132. [Google Scholar] [CrossRef]
- Bowers, P.M.; Schaufler, L.E.; Klevit, R.E. A Folding Transition and Novel Zinc Finger Accessory Domain in the Transcription Factor ADR1. Nat. Struct. Biol. 1999, 6, 478–485. [Google Scholar] [PubMed]
- Berg, J.M. Zinc Finger Domains: From Predictions to Design. Acc. Chem. Res. 1995, 28, 14–19. [Google Scholar] [CrossRef]
- Pavletich, N.P.; Pabo, C. Zinc Finger-DNA Recognition: Crystal Structure of a Zif268-DNA Complex at 2.1 A. Science 1991, 252, 809–817. [Google Scholar] [CrossRef]
- Jacobs, G.H. Determination of the Base Recognition Positions of Zinc Fingers from Sequence Analysis. EMBO J. 1992, 11, 4507–4517. [Google Scholar] [CrossRef]
- Omichinski, J.G.; Clore, G.M.; Schaad, O.; Felsenfeld, G.; Trainor, C.; Appella, E.; Stahl, S.J.; Gronenborn, A.M. NMR Structure of a Specific DNA Complex of Zn-Containing DNA Binding Domain of GATA-1. Science 1993, 261, 438–446. [Google Scholar] [CrossRef]
- Takatsuji, H.; Nakamura, N.; Katsumoto, Y. A New Family of Zinc Finger Proteins in Petunia: Structure, DNA Sequence Recognition, and Floral Organ-Specific Expression. Plant Cell 1994, 6, 947–958. [Google Scholar] [PubMed]
- Ciftci-Yilmaz, S.; Mittler, R. The Zinc Finger Network of Plants. Cell. Mol. Life Sci. 2008, 65, 1150–1160. [Google Scholar] [CrossRef] [PubMed]
- Takatsuji, H. A Single Amino Acid Determines the Specificity for the Target Sequence of Two Zinc-Finger Proteins in Plants The EPF Family Is a Group of DNA-Binding Proteins with Two Canonical Cys 2 / His 2 Zinc- Expressed in a Floral Organ-Specific Manner That Is Corre. Biochem. Biophys. Res. Commun. 1996, 223, 219–223. [Google Scholar] [CrossRef]
- Turquetti-Moraes, D.K.; Moharana, K.C.; Almeida-Silva, F.; Pedrosa-Silva, F.; Venancio, T.M. Integrating Omics Approaches to Discover and Prioritize Candidate Genes Involved in Oil Biosynthesis in Soybean. Gene 2022, 808. [Google Scholar] [CrossRef]
- Finkelstein, R.R. Mutations at Two New Arabidopsis ABA Response Loci Are Similar to the Abi3 Mutations. Plant J. 1994, 5, 765–771. [Google Scholar] [CrossRef]
- Bäumlein, H.; Miséra, S.; Luerßen, H.; Kölle, K.; Horstmann, C.; Wobus, U.; Müller, A.J. The FUS3 Gene of Arabidopsis Thaliana Is a Regulator of Gene Expression during Late Embryogenesis. Plant J. 1994, 6, 379–387. [Google Scholar] [CrossRef]
- Monfared, M.M.; Simon, M.K.; Meister, R.J.; Roig-Villanova, I.; Kooiker, M.; Colombo, L.; Fletcher, J.C.; Gasser, C.S. Overlapping and Antagonistic Activities of BASIC PENTACYSTEINE Genes Affect a Range of Developmental Processes in Arabidopsis. Plant J. 2011, 66, 1020–1031. [Google Scholar] [CrossRef] [PubMed]
- Burnham, C.R.; Yagyu, P. Linkage Relations of Teosinte Branched. Maize Genet. Coop. Newsl. 1961. [Google Scholar]
- Pekker, I.; Alvarez, J.P.; Eshed, Y. Auxin Response Factors Mediate Arabidopsis Organ Asymmetry via Modulation of KANADI Activity. Plant Cell 2005, 17, 2899–2910. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Wang, L.J.; Mao, Y.B.; Cai, W.J.; Xue, H.W.; Chen, X.Y. Control of Root Cap Formation by MicroRNA-Targeted Auxin Response Factors in Arabidopsis. Plant Cell 2005, 2005 17, 2204–2216. [Google Scholar] [CrossRef]
- Murmu, J.; Bush, M.J.; de Long, C.; Li, S.; Xu, M.; Khan, M.; Malcolmson, C.; Fobert, P.R.; Zachgo, S.; Hepworth, S.R. Arabidopsis Basic Leucine-Zipper Transcription Factors TGA9 and TGA10 Interact with Floral Glutaredoxins ROXY1 and ROXY2 and Are Redundantly Required for Anther Development. Plant Physiol. 2010, 154, 1492–1504. [Google Scholar] [CrossRef] [PubMed]
- Ito, S.; Song, Y.H.; Josephson-Day, A.R.; Miller, R.J.; Breton, G.; Olmstead, R.G.; Imaizumi, T. FLOWERING BHLH Transcriptional Activators Control Expression of the Photoperiodic Flowering Regulator CONSTANS in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 3582–3587. [Google Scholar] [CrossRef]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 Promotes Panicle Branching and Higher Grain Productivity in Rice. Nat. Genet. 2010, 42, 545–549. [Google Scholar] [CrossRef] [PubMed]
- Son, O.; Hur, Y.S.; Kim, Y.K.; Lee, H.J.; Kim, S.; Kim, M.R.; Nam, K.H.; Lee, M.S.; Kim, B.Y.; Park, J.; et al. ATHB12, an ABA-Inducible Homeodomain-Leucine Zipper (HD-Zip) Protein of Arabidopsis, Negatively Regulates the Growth of the Inflorescence Stem by Decreasing the Expression of a Gibberellin 20-Oxidase Gene. Plant Cell Physiol. 2010, 51, 1537–1547. [Google Scholar] [CrossRef]
- Scortecci, K.C.; Michaels, S.D.; Amasino, R.M. Identification of a MADS-Box Gene, FLOWERING LOCUS M, That Represses Flowering. Plant J. 2001, 26, 229–236. [Google Scholar] [CrossRef]
- Dorca-Fornell, C.; Gregis, V.; Grandi, V.; Coupland, G.; Colombo, L.; Kater, M.M. The Arabidopsis SOC1-like Genes AGL42, AGL71 and AGL72 Promote Flowering in the Shoot Apical and Axillary Meristems. Plant J. 2011, 67, 1006–1017. [Google Scholar] [CrossRef] [PubMed]
- Fornara, F.; Panigrahi, K.C.S.; Gissot, L.; Sauerbrunn, N.; Rühl, M.; Jarillo, J.A.; Coupland, G. Arabidopsis DOF Transcription Factors Act Redundantly to Reduce CONSTANS Expression and Are Essential for a Photoperiodic Flowering Response. Dev. Cell 2009, 17, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Palatnik, J.F.; Allen, E.; Wu, X.; Schommer, C.; Schwab, R.; Carrington, J.C.; Weigel, D. Control of Leaf Morphogenesis by MicroRNAs. Nature 2003, 425, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Zhang, L.; Wang, W.; Tian, P.; Wang, W.; Wang, K.; Gao, Z.; Liu, S.; Zhang, Y.; Irish, V.F.; et al. TCP5 Controls Leaf Margin Development by Regulating KNOX and BEL-like Transcription Factors in Arabidopsis. J. Exp. Bot. 2021, 72, 1809–1821. [Google Scholar] [CrossRef] [PubMed]
- Omidbakhshfard, M.A.; Fujikura, U.; Olas, J.J.; Xue, G.P.; Balazadeh, S.; Mueller-Roeber, B. GROWTH-REGULATING FACTOR 9 Negatively Regulates Arabidopsis Leaf Growth by Controlling ORG3 and Restricting Cell Proliferation in Leaf Primordia. PLoS Genet. 2018, 14, e1007484. [Google Scholar] [CrossRef] [PubMed]
- Kubo, H.; Peeters, A.J.M.; Aarts, M.G.M.; Pereira, A.; Koornneef, M. ANTHOCYANINLESS2, a Homeobox Gene Affecting Anthocyanin Distribution and Root Development in Arabidopsis. Plant Cell 1999, 11, 1217–1226. [Google Scholar] [CrossRef] [PubMed]
- Aida, M.; Beis, D.; Heidstra, R.; Willemsen, V.; Blilou, I.; Galinha, C.; Nussaume, L.; Noh, Y.S.; Amasino, R.; Scheres, B. The PLETHORA Genes Mediate Patterning of the Arabidopsis Root Stem Cell Niche. Cell 2004, 119, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yang, H.; Wu, C.; Feng, J.; Liu, X.; Qin, H.; Wang, D. Overexpressing HRS1 Confers Hypersensitivity to Low Phosphate-Elicited Inhibition of Primary Root Growth in Arabidopsis Thaliana. J. Integr. Plant Biol. 2009, 51, 382–392. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Kang, J.Y.; Cho, D.I.; Park, J.H.; Soo, Y.K. ABF2, an ABRE-Binding BZIP Factor, Is an Essential Component of Glucose Signaling and Its Overexpression Affects Multiple Stress Tolerance. Plant J. 2004, 40, 75–87. [Google Scholar] [CrossRef]
- Kang, H.G.; Singh, K.B. Characterization of Salicylic Acid-Responsive, Arabidopsis Dof Domain Proteins: Overexpression of OBP3 Leads to Growth Defects. Plant J. 2000, 21, 329–339. [Google Scholar] [CrossRef]
- Akhtar, M.; Jaiswal, A.; Taj, G.; Jaiswal, J.P.; Qureshi, M.I.; Singh, N.K. DREB1/CBF Transcription Factors: Their Structure, Function and Role in Abiotic Stress Tolerance in Plants. J. Genet. 2012, 91, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Cao, F.Y.; DeFalco, T.A.; Moeder, W.; Li, B.; Gong, Y.; Liu, X.M.; Taniguchi, M.; Lumba, S.; Toh, S.; Shan, L.; et al. Arabidopsis ETHYLENE RESPONSE FACTOR 8 (ERF8) Has Dual Functions in ABA Signaling and Immunity. BMC Plant Biol. 2018, 18, 211. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Sheerin, D.J.; von Roepenack-Lahaye, E.; Stahl, M.; Hiltbrunner, A. The Phytochrome Interacting Proteins ERF55 and ERF58 Repress Light-Induced Seed Germination in Arabidopsis Thaliana. Nat. Commun. 2022, 13, 1656. [Google Scholar] [CrossRef]
- Heyman, J.; Cools, T.; Canher, B.; Shavialenka, S.; Traas, J.; Vercauteren, I.; Van Den Daele, H.; Persiau, G.; De Jaeger, G.; Sugimoto, K.; et al. The Heterodimeric Transcription Factor Complex ERF115-PAT1 Grants Regeneration Competence. Nat. Plants 2016, 2016 2, 16165. [Google Scholar] [CrossRef]
- Hao, Y.J.; Wei, W.; Song, Q.X.; Chen, H.W.; Zhang, Y.Q.; Wang, F.; Zou, H.F.; Lei, G.; Tian, A.G.; Zhang, W.K.; et al. Soybean NAC Transcription Factors Promote Abiotic Stress Tolerance and Lateral Root Formation in Transgenic Plants. Plant J. 2011, 68, 302–313. [Google Scholar] [CrossRef]
- Zheng, X.; Chen, B.; Lu, G.; Han, B. Overexpression of a NAC Transcription Factor Enhances Rice Drought and Salt Tolerance. Biochem. Biophys. Res. Commun. 2009, 379, 985–989. [Google Scholar] [CrossRef]
- Balazadeh, S.; Siddiqui, H.; Allu, A.D.; Matallana-Ramirez, L.P.; Caldana, C.; Mehrnia, M.; Zanor, M.I.; Köhler, B.; Mueller-Roeber, B. A Gene Regulatory Network Controlled by the NAC Transcription Factor ANAC092/AtNAC2/ORE1 during Salt-Promoted Senescence. Plant J. 2010, 62, 250–264. [Google Scholar] [CrossRef]
- Takagi, H.; Tamiru, M.; Abe, A.; Yoshida, K.; Uemura, A.; Yaegashi, H.; Obara, T.; Oikawa, K.; Utsushi, H.; Kanzaki, E.; et al. MutMap Accelerates Breeding of a Salt-Tolerant Rice Cultivar. Nat. Biotechnol. 2015, 33, 445–449. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.C.; Lai, Z.; Fan, B.; Chen, Z. Arabidopsis WRKY38 and WRKY62 Transcription Factors Interact with Histone Deacetylase 19 in Basal Defense. Plant Cell 2008, 20, 2357–2371. [Google Scholar] [CrossRef]
- Chen, X.; Liu, J.; Lin, G.; Wang, A.; Wang, Z.; Lu, G. Overexpression of AtWRKY28 and AtWRKY75 in Arabidopsis Enhances Resistance to Oxalic Acid and Sclerotinia Sclerotiorum. Plant Cell Rep. 2013, 32, 1589–1599. [Google Scholar] [CrossRef]
- Jia, J.; XinG, J. hong; Dong, J. gao; Han, J. min; Liu, J. sheng. Functional Analysis of MYB73 of Arabidopsis Thaliana Against Bipolaris Oryzae. Agric. Sci. China 2011, 10, 721–727. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




