Submitted:
26 February 2024
Posted:
08 March 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. BioGRID
2.2. CYTOSCAPE and Network Topology Analysis
2.3. Evaluation of the HUB-and-Spoke Model
2.4. Cluster Analysis
2.5. GO and KEGG Pathway Analyses
2.6. Network Analyst -- Comprehensive Gene Expression Profiling via Network Visual Analytics: TFs and miRNAs
- tF → TF
- TF → gene
- tF → miRNA
- miRNA → TF
- miRNA → gene
2.7. Protein Intrinsic Disorder and Secondary Structure Prediction
2.8. SARS2-HUMAN Proteome Interaction Database (SHPID)
2.9. Highlighting the Nodes of a STRING Network Involved in the Same Biological Process (GO)
2.10. Comparison between GO Pairs in Enriched Networks
3. Results
3.1. Source of the Data
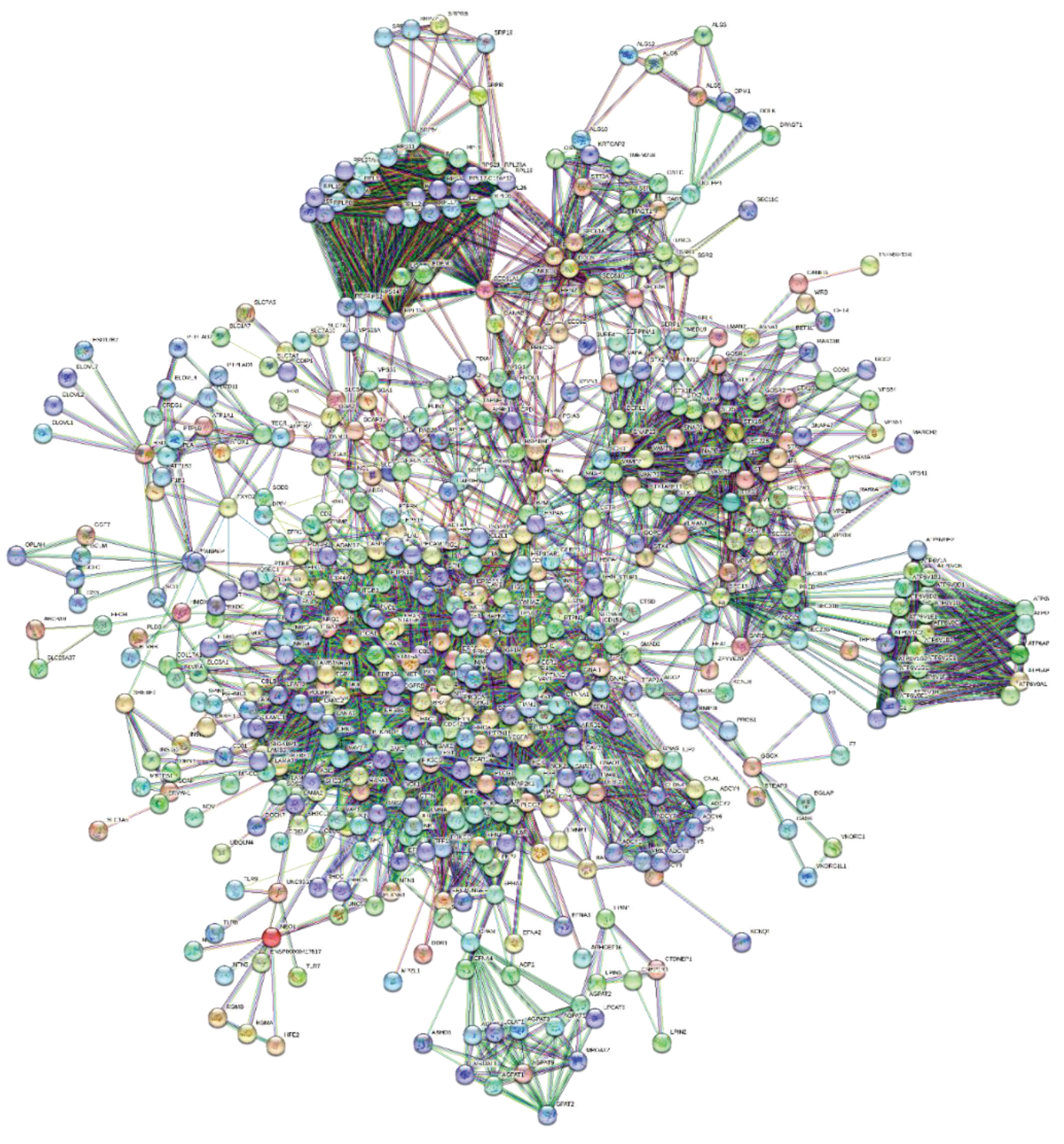
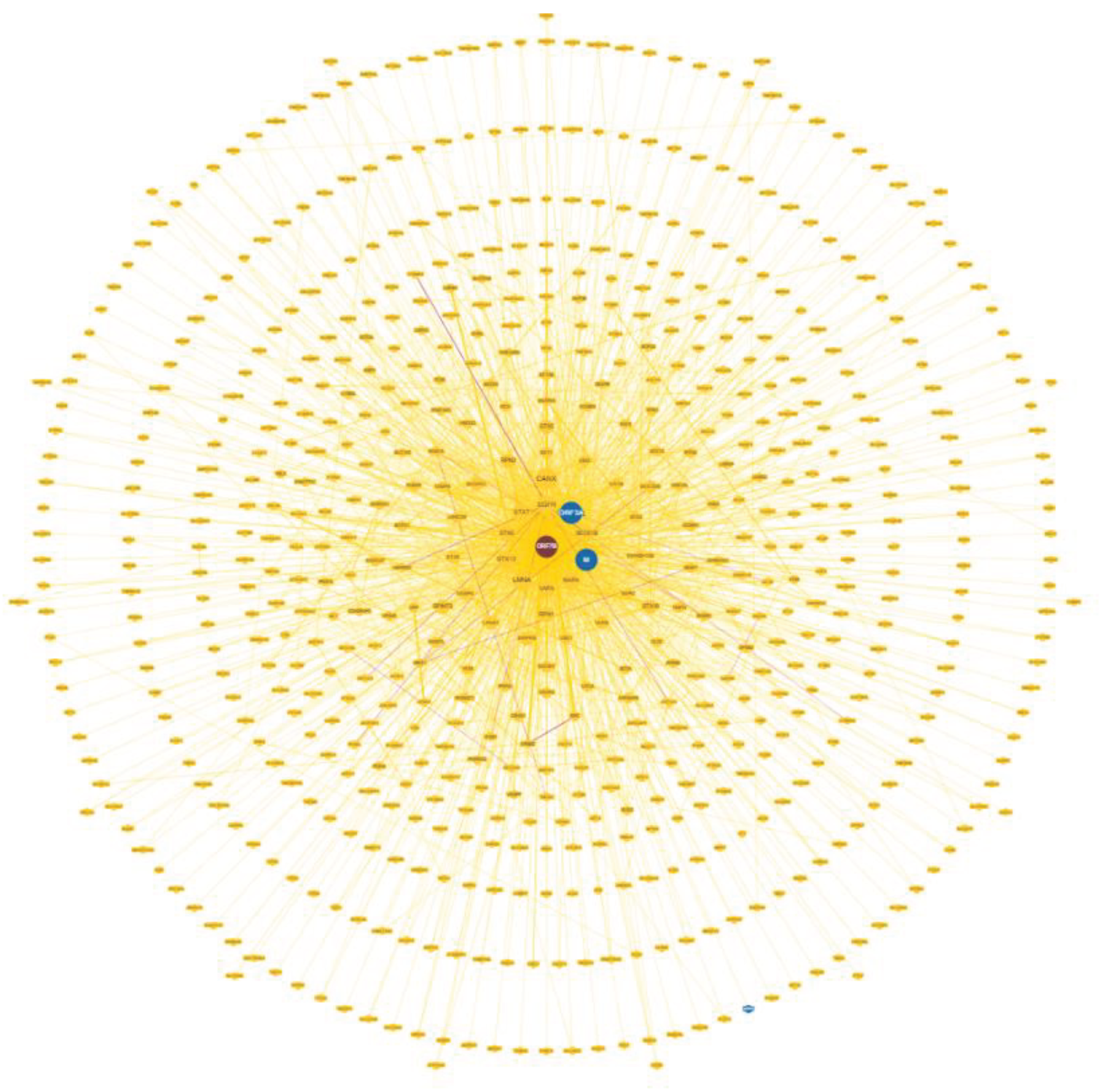
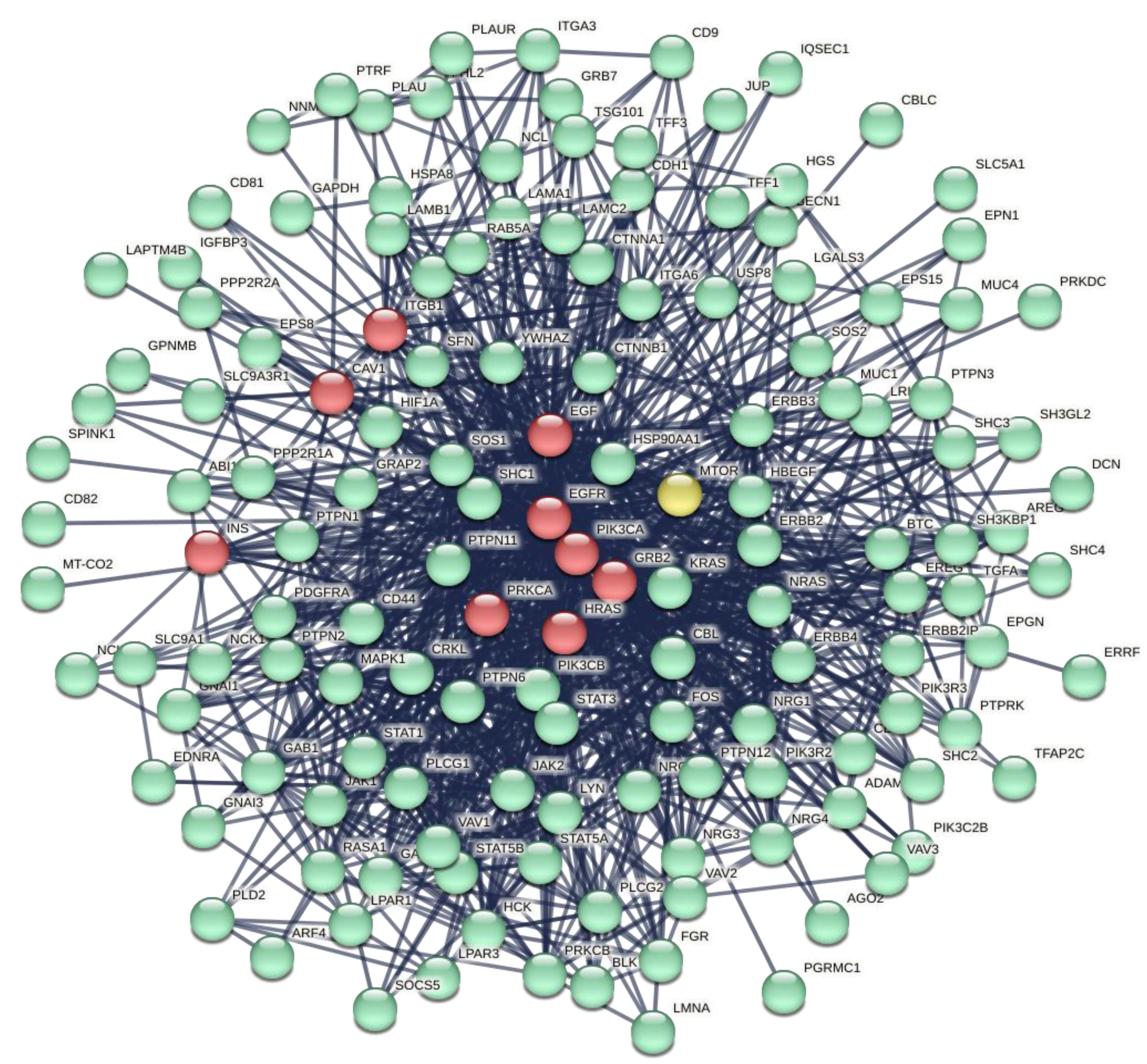
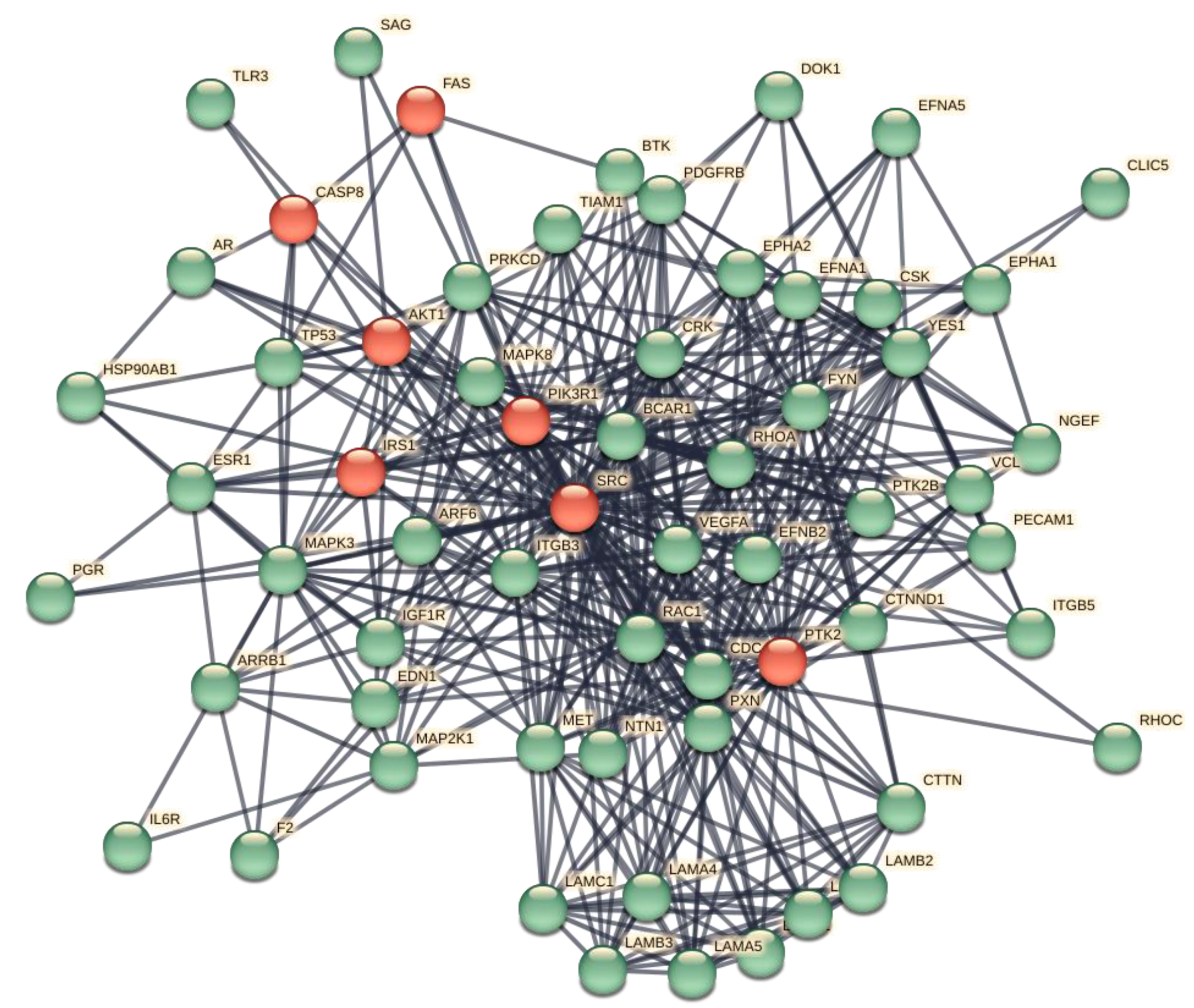
3.2. The Representation of ORF7b Data Using Interactomes
3.3. Principal Characteristics of the Interactome
| Summary Statistics of network* | Notes | |
|---|---|---|
| Number of nodes | 551 | |
| Number of edges | 4648 | ** |
| Avg. Number of neighbors | 16,871 | Average connectivity of the nodes |
| Network diameter | 9 | |
| Characteristic path length | 3.666 | |
| Clustering coefficient | 0.549 | 0 ≤ C ≤ 1 |
| Network density | 0.031 | |
| Network heterogeneity | 1.057 | Tendency to contain hub nodes |
| Network centralization | 0.259 | The extent to which certain nodes are far more central than others |
| Connected Component | 1 | *** |
3.4. Quantitative Evaluation of the Biological Functionalities in the Interactome
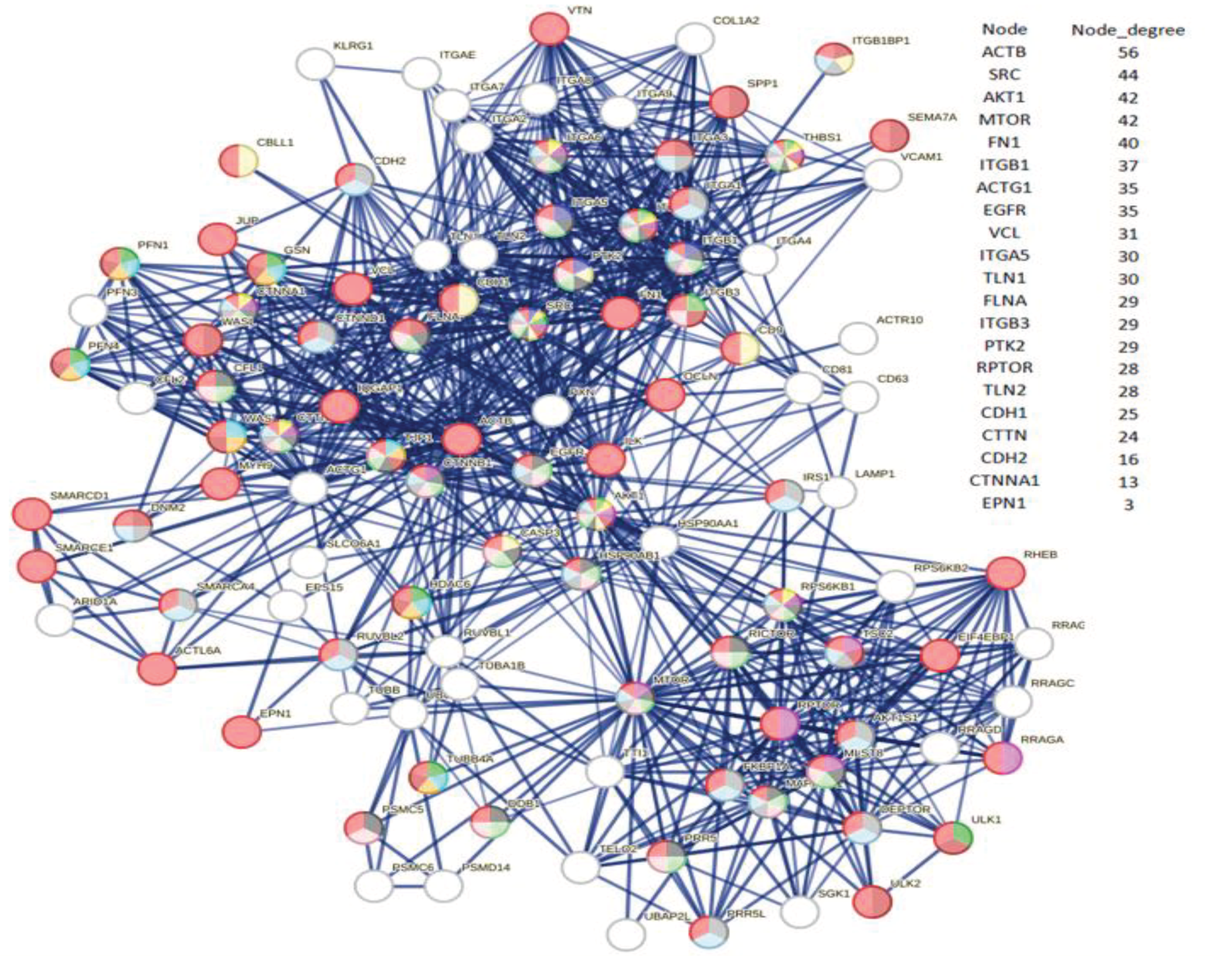
3.5. Exploring the Physical Basis of Cytoskeletal Alterations Caused by ORF7b
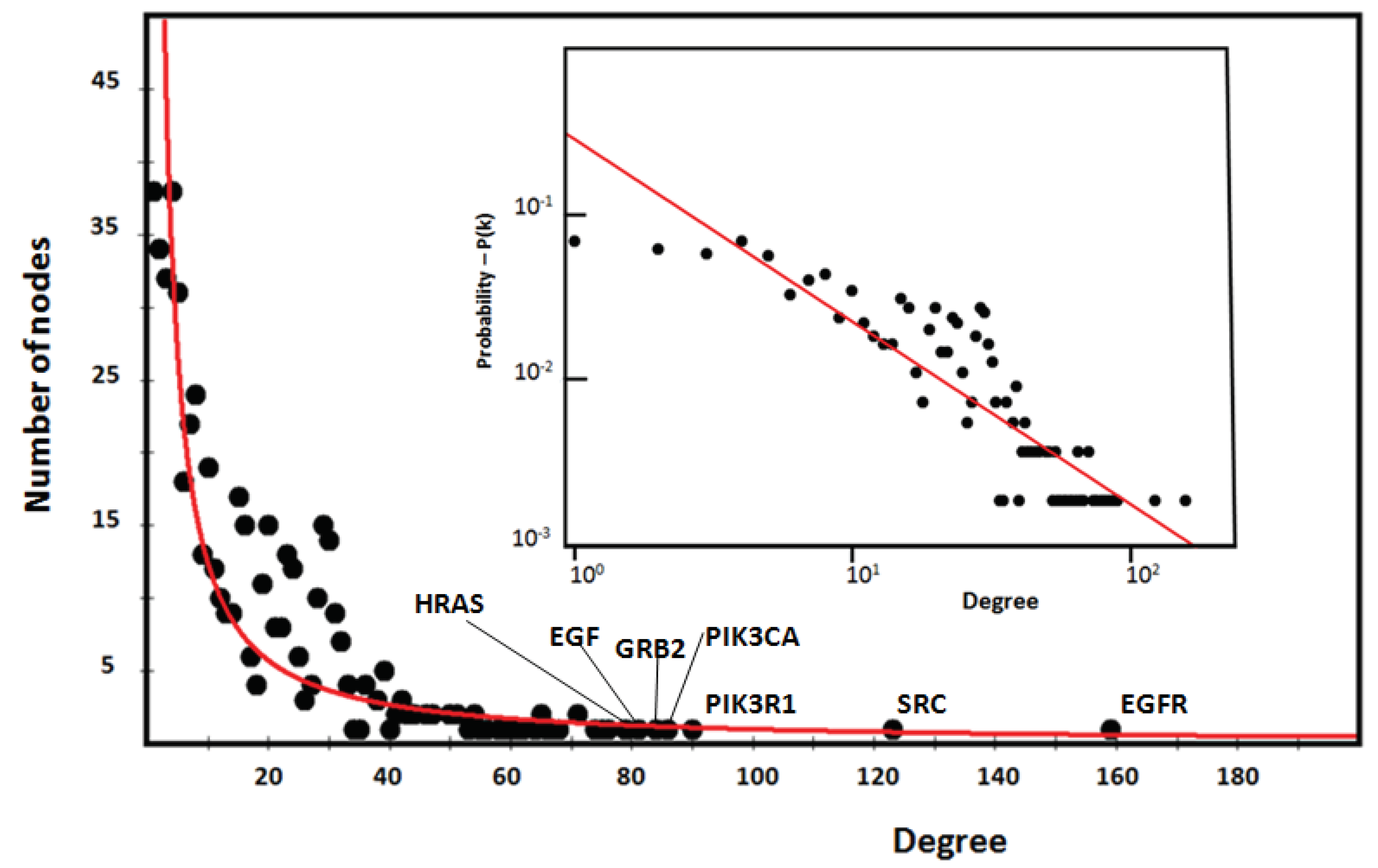
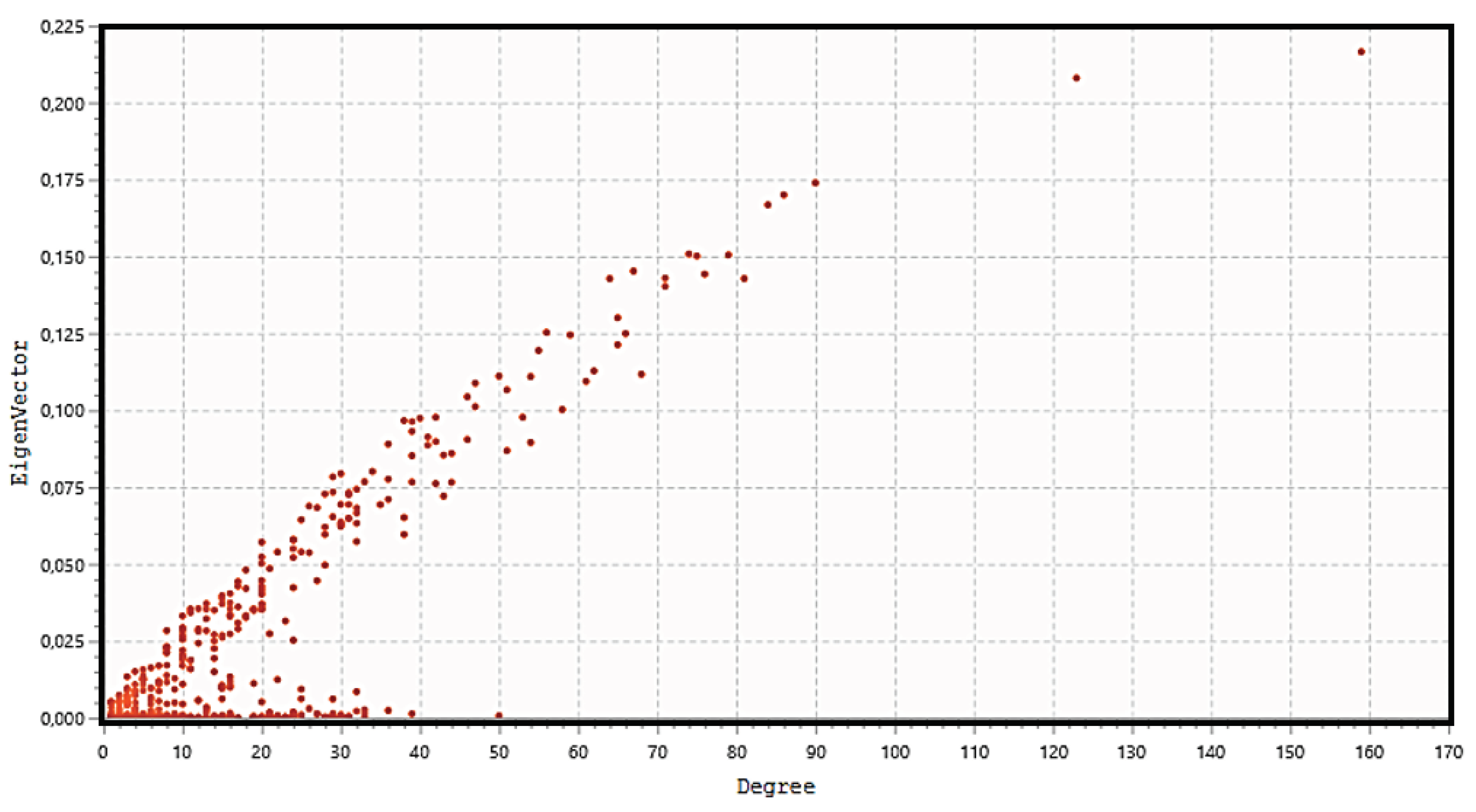
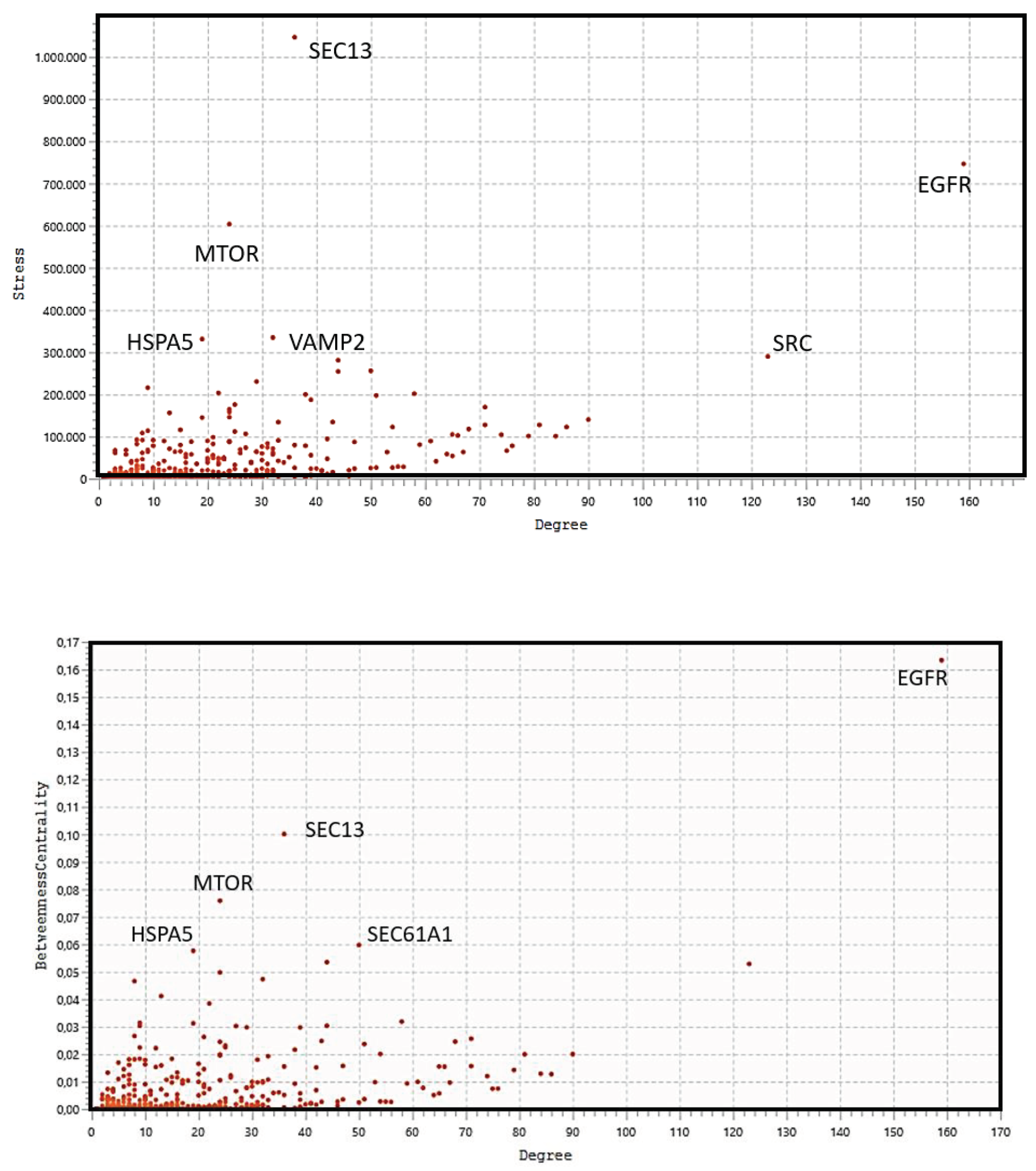
3.6. Topological Analysis
3.7. The Functional Effects Depend Not Only on ORF7b but Also on the Integrated Action of Several Viral Proteins
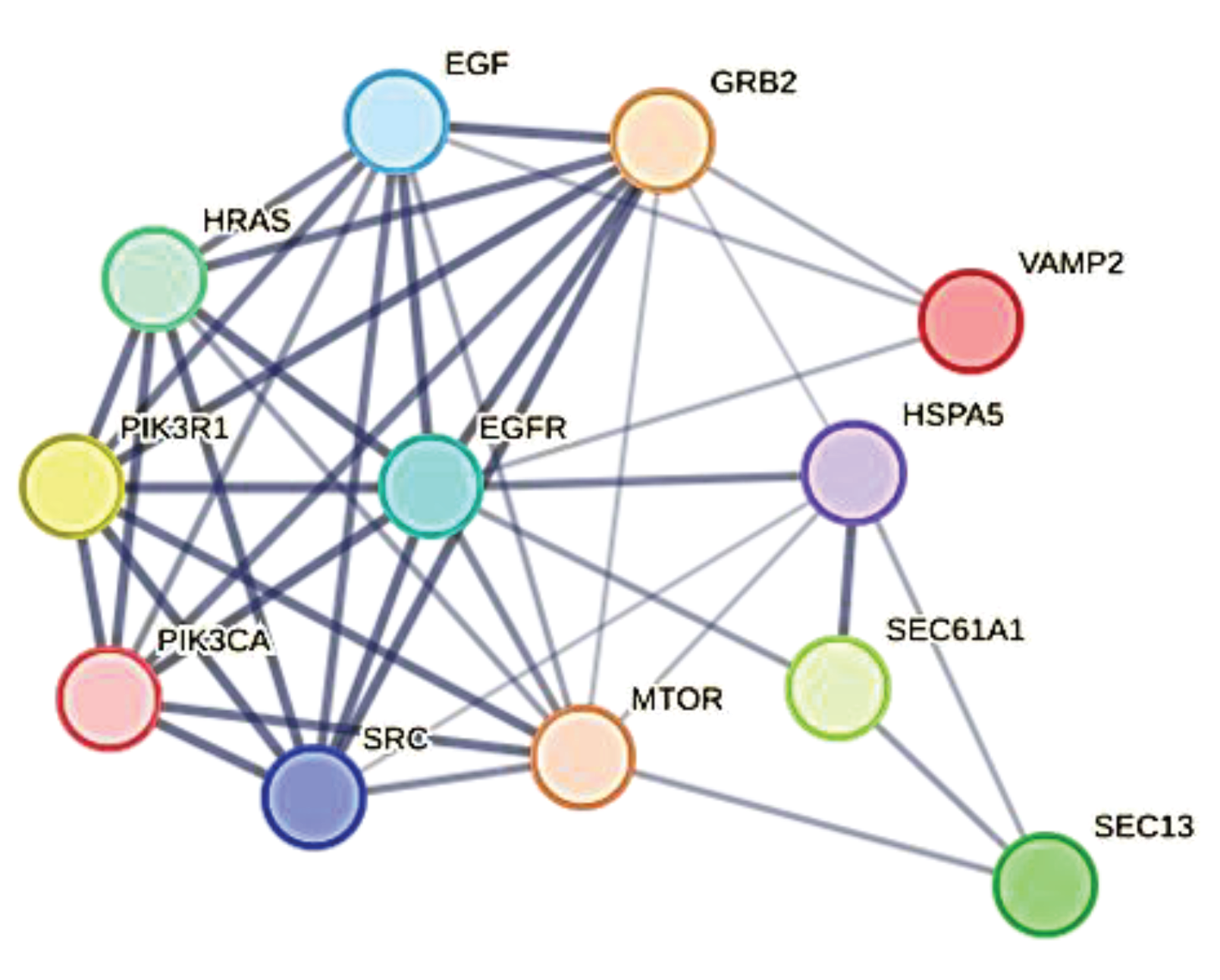
3.8. The Peculiar Case of GRB2, a Protein in the Service of ORF7b
3.9. The Role of ORF7b
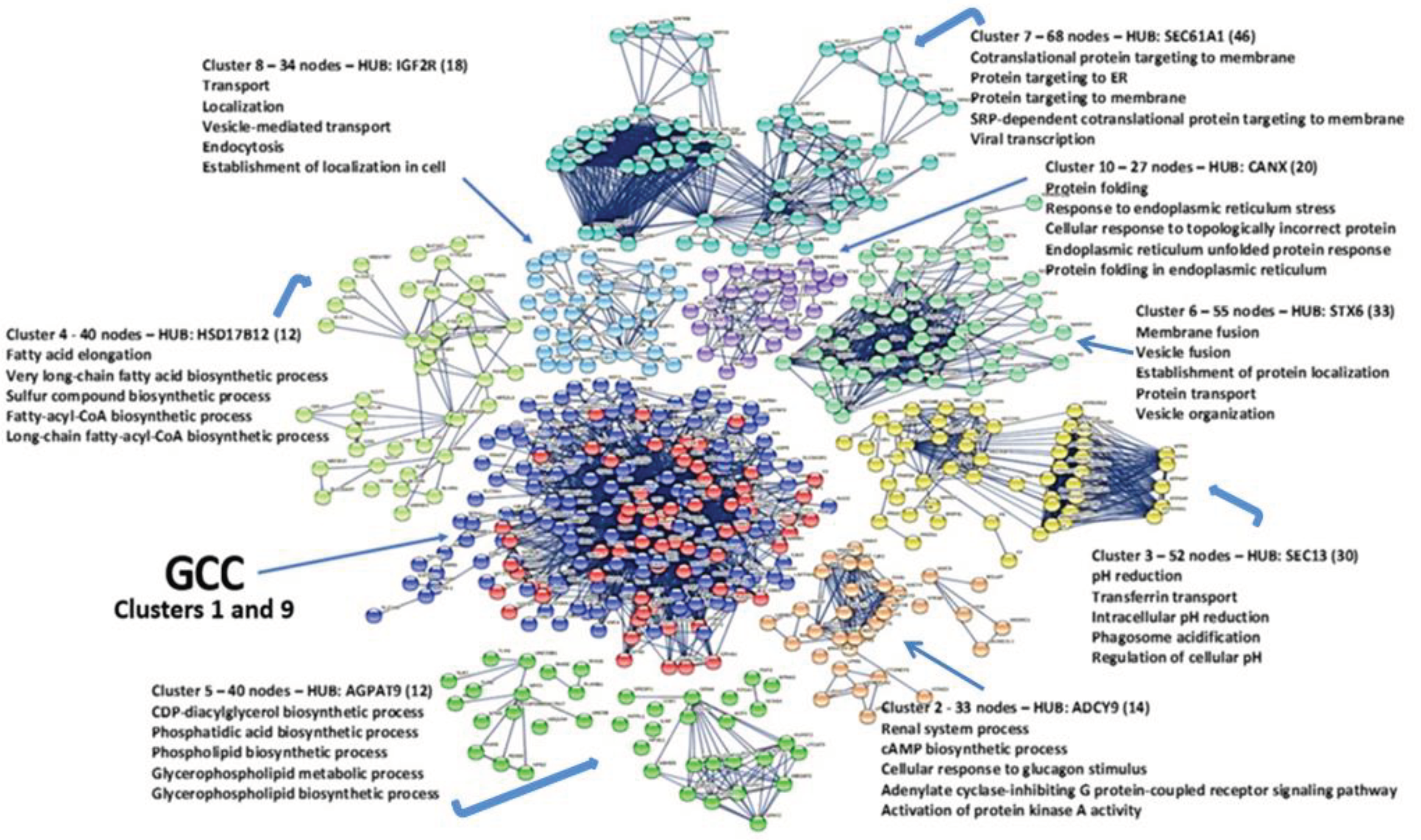
3.10. Cluster Analysis
3.11. Analysis of GCC Core
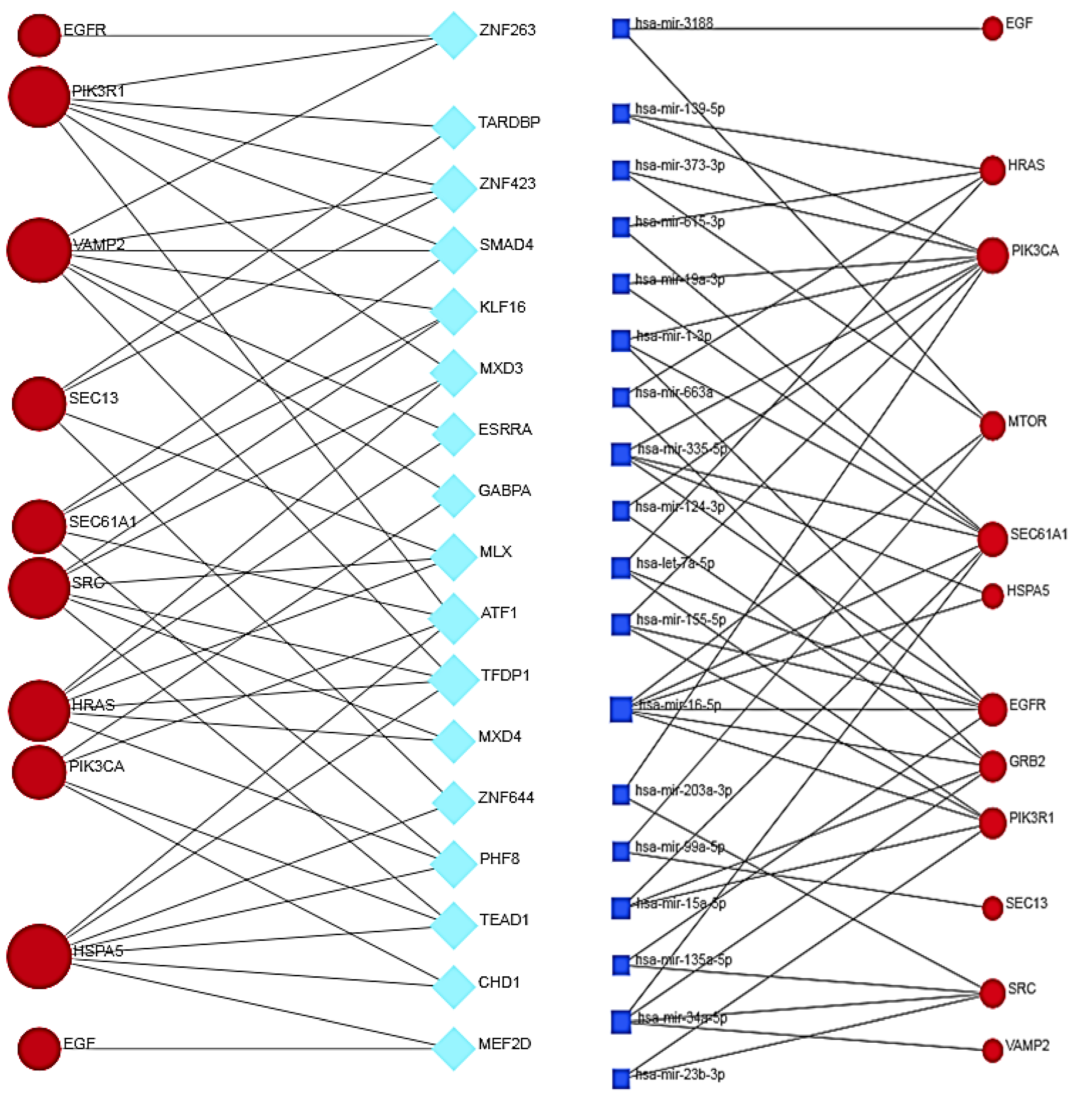
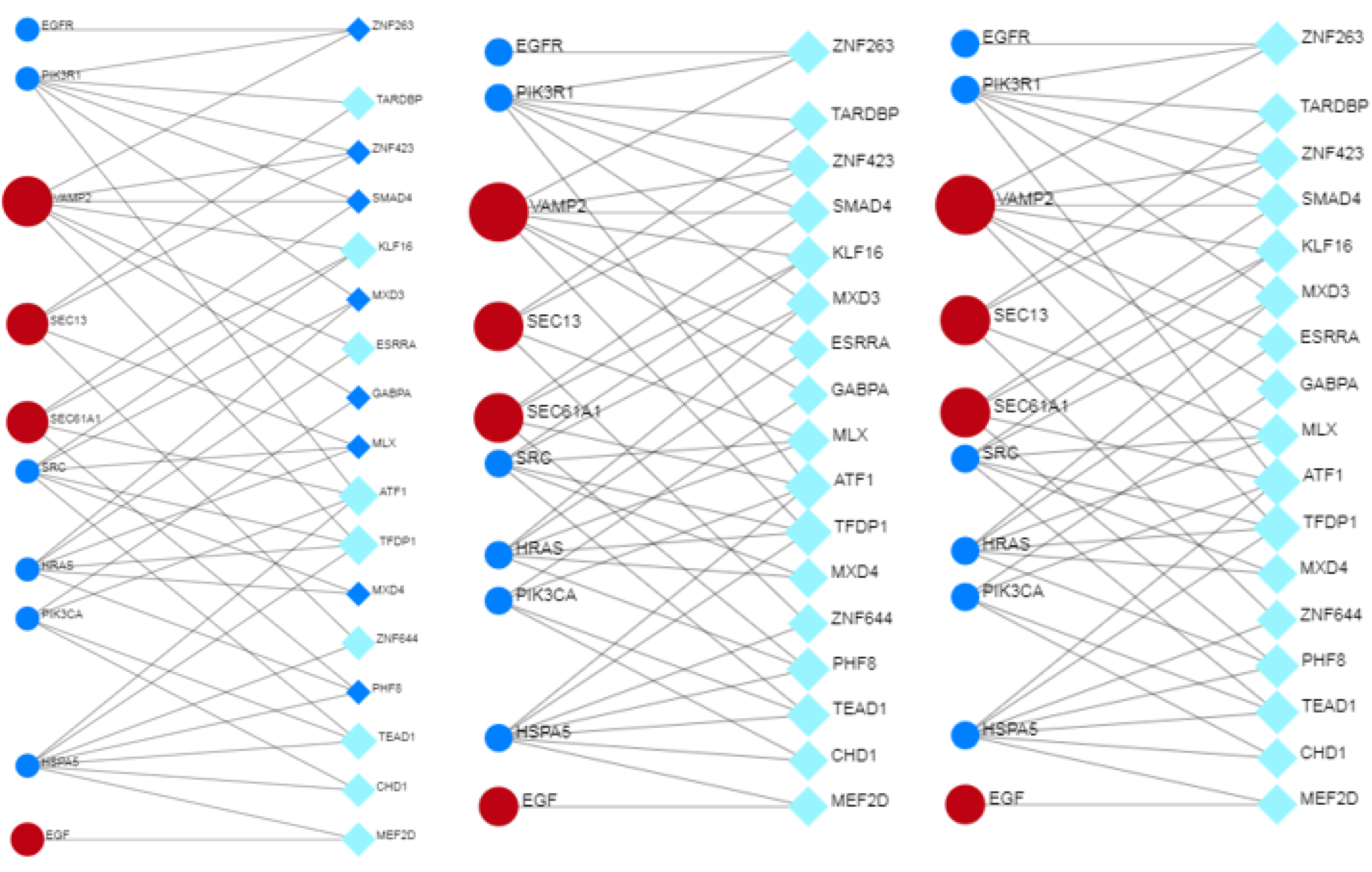
3.12. Co-Regulation between Hub and Bottleneck Proteins, Transcription Factors and miRNAs
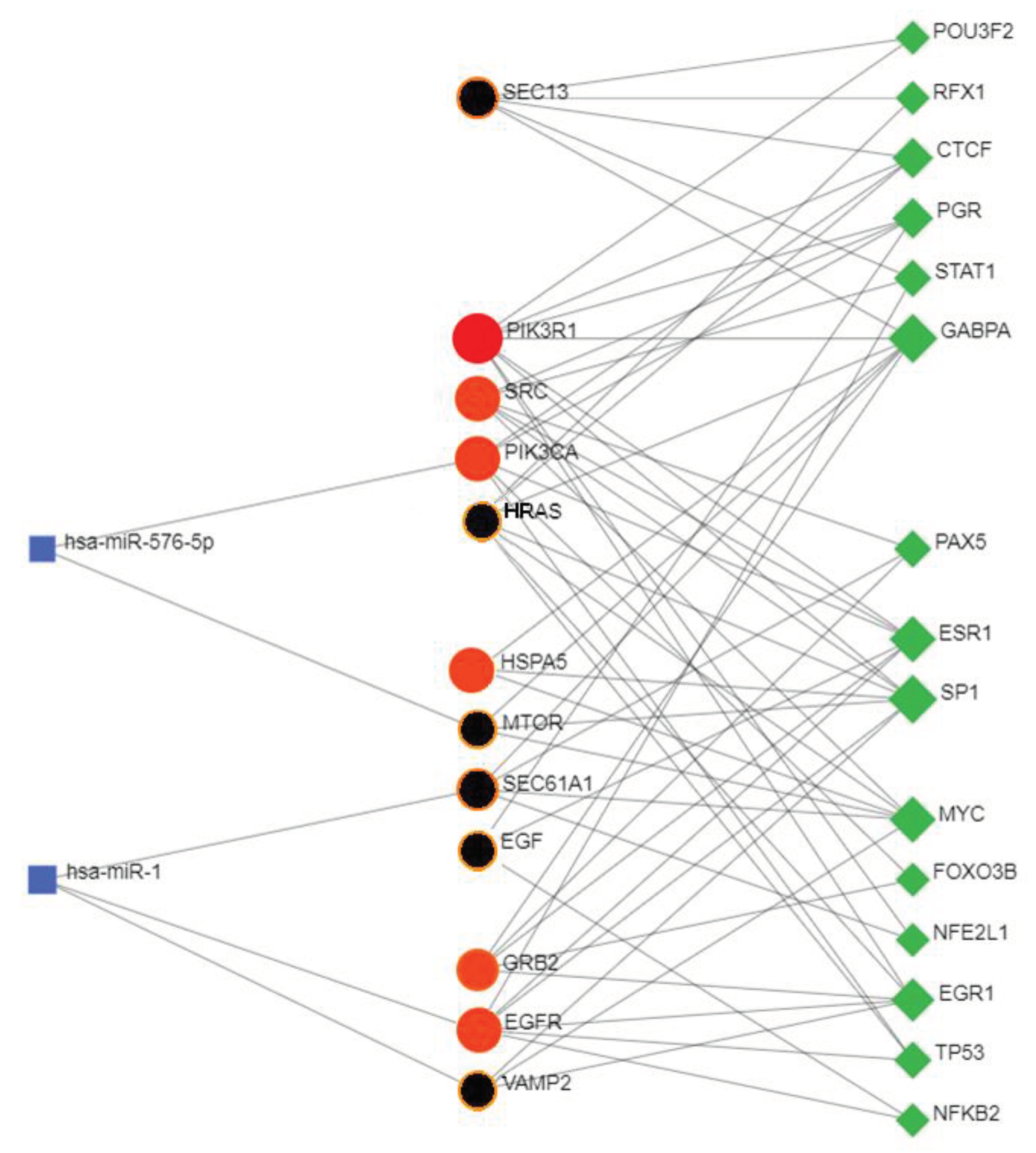
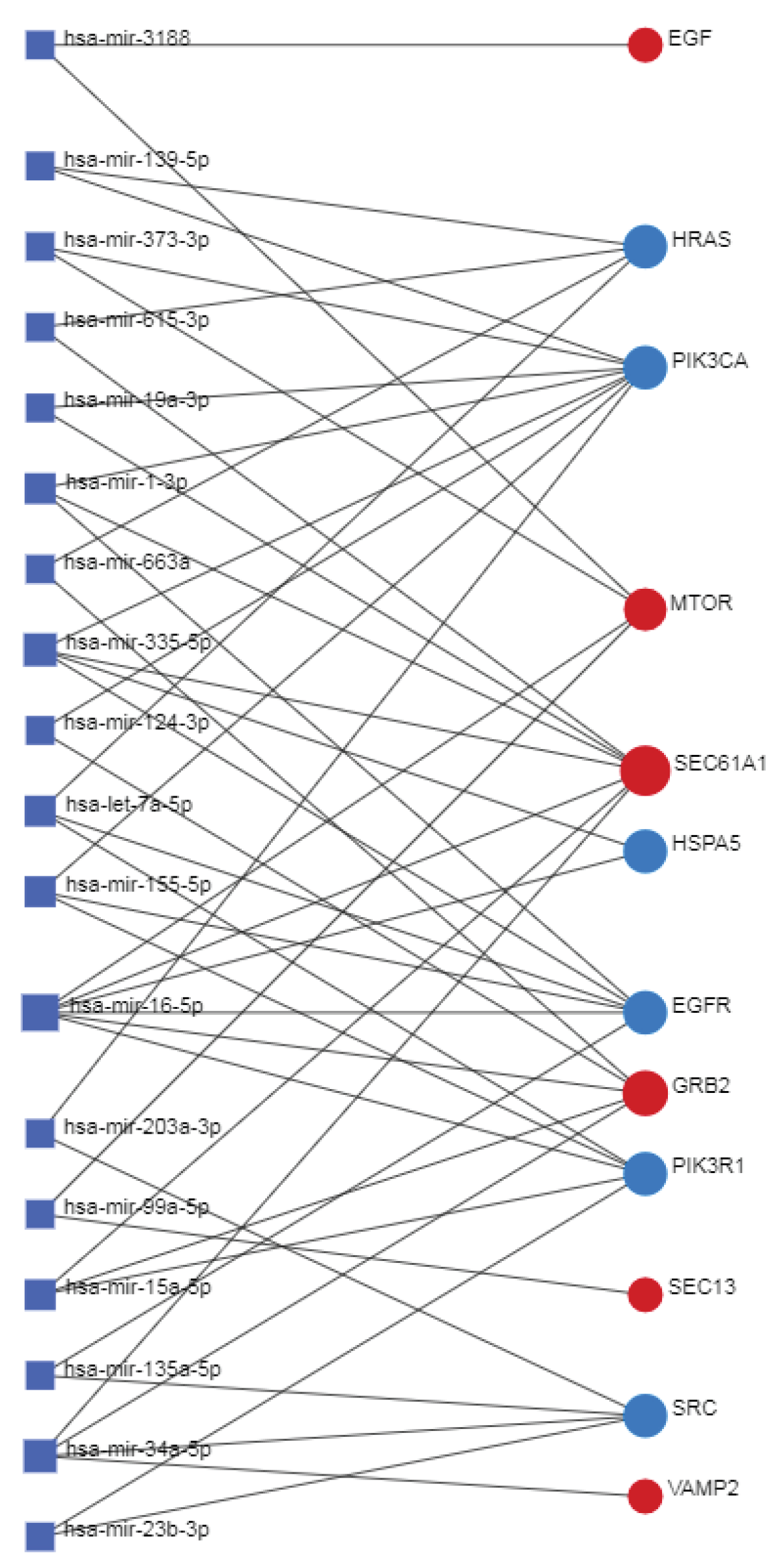
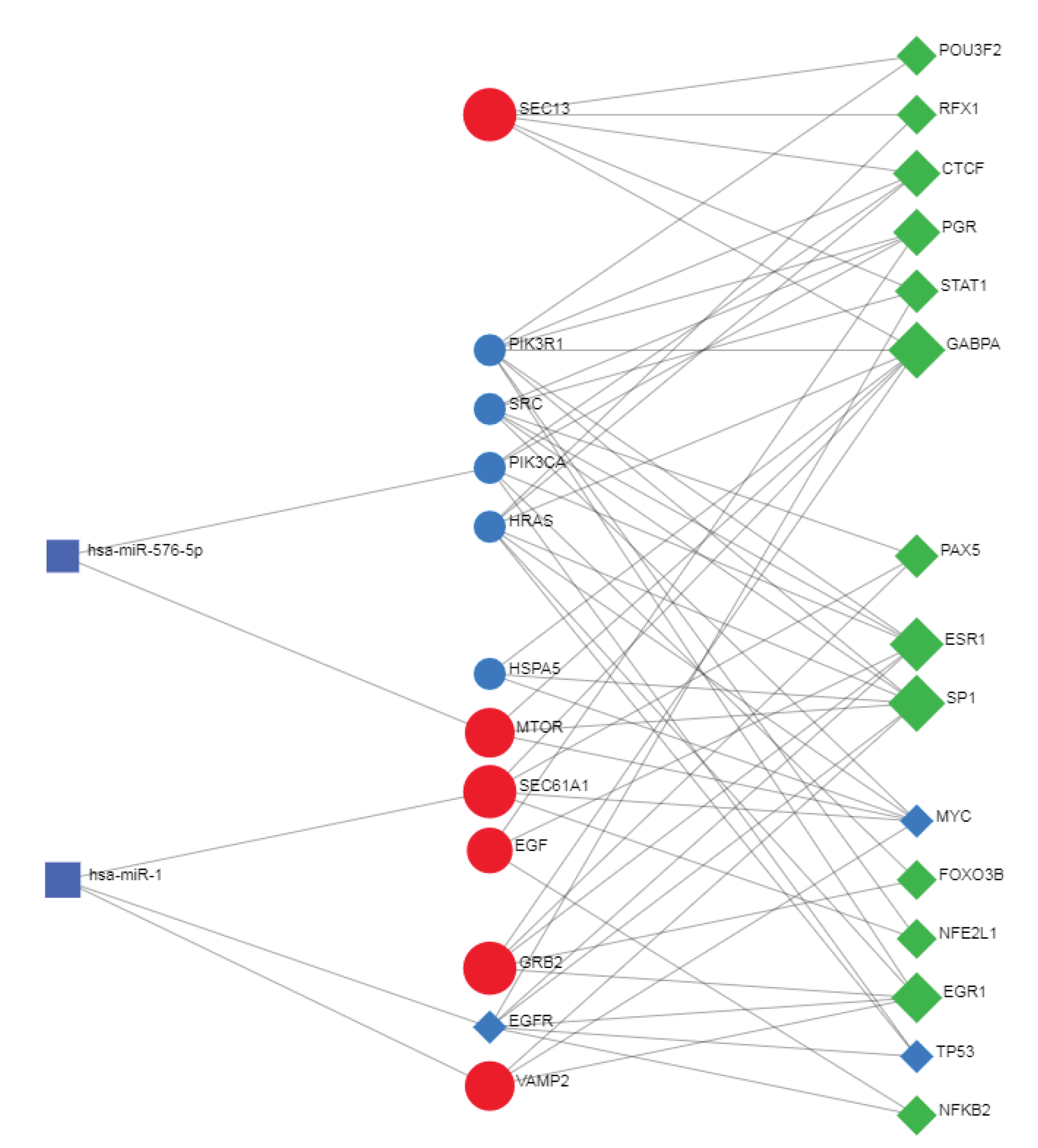
3.13. Transcriptional and Post-Transcriptional Regulatory Networks
3.14. Co-Regulatory Network
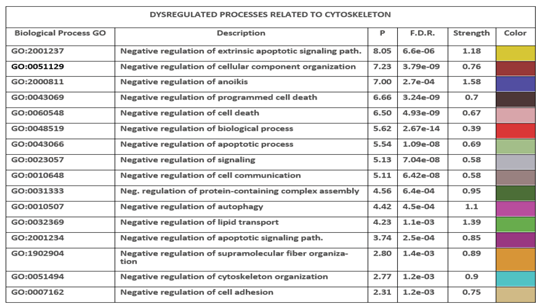
3.15. Comparative Analysis of Negative Regulations According to the GO
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bar-On, Y.M.; et al. SARS-CoV-2 [COVID-19] by the numbers. eLife 2020, 9, e57309. [Google Scholar] [CrossRef]
- Sender, R.; et al. The total number and mass of SARS-CoV-2 virions. PNAS 2021, 118, e2024815118. [Google Scholar] [CrossRef]
- Milo, R. What is the total number of protein molecules per cell volume? A call to rethink some published values. Bioessays. 2013, 35, 1050–1055. [Google Scholar] [CrossRef] [PubMed]
- Andreas-David Brunner et al., Ultra-high sensitivity mass spectrometry quantifies single-cell proteome changes upon perturbation. Mol Syst Biol 2022, 18, e10798. [CrossRef]
- Brandon Ho, et al. Unification of Protein Abundance Datasets Yields a Quantitative Saccharomyces cerevisiae Proteome. Cell Systems 2018, 6, 1–14. [CrossRef]
- Colonna, G. A Tiny Viral Protein, SARS-CoV-2-ORF7b: Structural Features. Preprints 2023, 2023040522. [Google Scholar] [CrossRef]
- Zhang, J.; et al. A systemic and molecular study of subcellular localization of SARS-CoV-2 proteins. Sig Transduct Target Ther 2020, 5, 269. [Google Scholar] [CrossRef]
- Oughtred R, et al.; et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 2020, 30, 187–200. [Google Scholar] [CrossRef]
- S. Boccaletti, et al. Hwang Complex networks: structure and dynamics. Phys Rep 2006, 424, 175–308. [CrossRef]
- Chris Stark, et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Research 2006, 34, D535–D539. [CrossRef] [PubMed]
- Chen Z, et al. Comprehensive analysis of the host-virus interactome of SARS-CoV-2 (2021 Pre-Publication) Status: Preliminary Report. 2021.
- Stukalov A, Girault V, et al. Multilevel proteomics reveals host perturbations by SARS-CoV-2 and SARS-CoV. Nature 2021, 594, 246–252. [CrossRef]
- Estelle, M.N. Laurent, Yorgos Sofianatos, et al. Global BioID-based SARS-CoV-2 proteins proximal interactome unveils novel ties between viral polypeptides and host factors involved in multiple COVID19-associated mechanisms (2020 Pre-Publication). Status: Preliminary Report.
- Samavarchi-Tehrani P, et al. A SARS-CoV-2 - host proximity interactome (2020 Pre-Publication) Status: Preliminary Report.
- St-Germain JR, Astori A, et al., A SARS-CoV-2 BioID-based virus-host membrane protein interactome and virus peptide compendium: new proteomics resources for COVID-19 research (2020 Pre-Publication) Status: Preliminary Report.
- Bamberger TC, Pankow S, et al. The Host Interactome of Spike Expands the Tropism of SARS-CoV-2 (2021 Pre-Publication). Status: Preliminary Report.
- Liu X, Huuskonen S, et al. SARS-CoV-2-host proteome interactions for antiviral drug discovery. Mol Syst Biol. 2020, 17, e10396. [CrossRef]
- Snider J, et al. Fundamentals of protein interaction network mapping. Mol Syst Biol. 2015, 11, 848. [CrossRef]
- O’Connor, D.J. Emergent Properties. In Old and New Questions in Physics, Cosmology, Philosophy, and Theoretical Biology; van der Merwe, A., Ed.; Springer: Boston, MA, 1983. [Google Scholar] [CrossRef]
- Boojari, M.A. Investigating the Evolution and Development of Biological Systems from the Perspective of Thermo-Kinetics and Systems Theory. Orig Life Evol Biosph 2020, 50, 121–143. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk D, et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015, 43, D447–52. [CrossRef]
- Machicao J, et al. On the Use of Topological Features of Metabolic Networks for the Classification of Cancer Samples. Curr Genomics. 2021, 22, 88–97. [CrossRef] [PubMed]
- Xueming Liu, et al. Controllability of giant connected components in a directed network. PHYSICAL REVIEW E 2017, 95, 042318. [CrossRef]
- Wagner, How the global structure of protein interaction networks evolves. Proceedings of the Royal Society of London. Series B 2003, 270, 457–466. [CrossRef]
- Shannon P, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–504. [CrossRef]
- Assenov Y, Ramírez F, Schelhorn SE, Lengauer T, Albrecht M. Computing topological parameters of biological networks. Bioinformatics. 2008, 24, 282–4. [CrossRef] [PubMed]
- Doncheva, N., et al. Topological analysis and interactive visualization of biological networks and protein structures. Nat Protoc 2012, 7, 670–685. [CrossRef] [PubMed]
- Dong, J., Horvath, S. Understanding network concepts in modules. BMC Syst Biol 2007, 1, 24. [CrossRef]
- Kumar M, Saini S, Gayen K. Elementary mode analysis reveals that Clostridium acetobutylicum modulates its metabolic strategy under external stress. Mol Biosyst. 2014, 10, 2090–105. [CrossRef] [PubMed]
- Barabási, A.L., Oltvai, Z.N.: Network biology: understanding the cell's functional organization. Nat Rev Genet 2004, 5, 101–113. [CrossRef] [PubMed]
- Wagner, How the global structure of protein interaction networks evolves. Proc.R.Soc.Lond.B 2003, 270, 457–466. [CrossRef]
- M. Gerlach, E. G. Altmann, Testing statistical laws in complex systems. Phys. Rev. Lett. 2019, 122, 168301. [CrossRef]
- Jose C Nacher and Tatsuya Akutsu, Dominating scale-free networks with variable scaling exponent: heterogeneous networks are not difficult to control. New J. Phys. 2012, 14, 073005. [CrossRef]
- Winterbach W, et al. Topology of molecular interaction networks. BMC Syst Biol. 2013, 7, 90. [CrossRef]
- Yoon TY, Munson M. SNARE complex assembly and disassembly. Curr Biol. 2018, 28, R397–R401. [CrossRef] [PubMed]
- Kádková A, Radecke J, Sørensen JB. The SNAP-25 Protein Family. Neuroscience 2019, 420, 50–71. [CrossRef] [PubMed]
- Grabowski P, et al.,. "Expression of neuroendocrine markers: a signature of human undifferentiated carcinoma of the colon and rectum". Virchows Archiv. 2022, 441, 256–263. [CrossRef]
- Xi Z, et al. "Association of Alpha-Soluble NSF Attachment Protein with Epileptic Seizure". Journal of Molecular Neuroscience 2015, 57, 3): 417–425. [CrossRef]
- Morton AJ, Faull RL, Edwardson JM. "Abnormalities in the synaptic vesicle fusion machinery in Huntington's disease". Brain Research Bulletin 2001, 56, 111–117. [CrossRef] [PubMed]
- albandian A, et al. Post-acute COVID-19 syndrome. Nat Med. 2021, 27, 601–615. [CrossRef]
- Silva Andrade B, et al. Long-COVID and Post-COVID Health Complications: An Up-to-Date Review on Clinical Conditions and Their Possible Molecular Mechanisms. Viruses. 2021, 13, 700. [CrossRef]
- Vadivalagan C, et al. Exosomal mediated signal transduction through artificial microRNA (amiRNA): A potential target for inhibition of SARS-CoV-2. Cell Signal. 2022, 95, 110334. [CrossRef]
- Krachmarova E, et al. Insights into the SARS-CoV-2 ORF6 Mechanism of Action. Int J Mol Sci. 2023, 24, 11589. [CrossRef]
- Rozen EJ, Shohet JM. Systematic review of the receptor tyrosine kinase superfamily in neuroblastoma pathophysiology. Cancer Metastasis Rev. 2022, 41, 33–52. [CrossRef]
- Wang, Z. ErbB Receptors and Cancer. Methods Mol Biol. 2017, 1652, 3–35. [Google Scholar] [CrossRef] [PubMed]
- Hadid T, et al.,. Coagulation and anticoagulation in COVID-19. Blood Rev. 2021, 47, 100761. [CrossRef] [PubMed]
- Khunti K, et al. COVID-19, Hyperglycemia, and New-Onset Diabetes. Diabetes Care. 2021, 44, 2645–2655. [CrossRef]
- Loughrey D, Dahlman JE. Non-liver mRNA Delivery. Acc Chem Res. 2022, 55, 13–23. [CrossRef] [PubMed]
- Peiris S, et al. Pathological findings in organs and tissues of patients with COVID-19: A systematic review. PLoS One. 2021, 16, e0250708. [CrossRef]
- Koçak Tufan Z, et al. COVID-19 and Sepsis. Turk J Med Sci. 2021, 51, 3301–3311. [CrossRef]
- Kopańska M, et al. Effects of SARS-CoV-2 Inflammation on Selected Organ Systems of the Human Body. Int J Mol Sci. 2022, 23, 4178. [CrossRef]
- Zeng C, et al. SARS-CoV-2 spreads through cell-to-cell transmission. Proc Natl Acad Sci U S A. 2022, 119, e2111400119. [CrossRef]
- Cifuentes-Munoz N, et al. Viral cell-to-cell spread: Conventional and non-conventional ways. Adv Virus Res. 2020, 108, 85–125. [CrossRef]
- Tiwari V, et al. Role of Tunneling Nanotubes in Viral Infection, Neurodegenerative Disease, and Cancer. Front Immunol. 2021, 12, 680891. [CrossRef] [PubMed]
- Ermagun, A., Tajik, N. & Mahmassani, H. Uncertainty in vulnerability of networks under attack. Sci Rep 2023, 13, 3179. [CrossRef]
- Masoomy, H. and al., Relation between the degree and betweenness centrality distribution in complex networks. Phys. Rev. E 2023, 107, 044303. [CrossRef]
- Vistain LF, Tay S. Single-Cell Proteomics. Trends Biochem Sci. 2021, 46, 661–672. [CrossRef] [PubMed]
- Rosa SS, et al. mRNA vaccines manufacturing: Challenges and bottlenecks. Vaccine. 2021, 39, 2190–2200. [CrossRef] [PubMed]
- Yu H, Kim PM, Sprecher E, Trifonov V, Gerstein M. The importance of bottlenecks in protein networks: correlation with gene essentiality and expression dynamics. PLoS Comput Biol. 2007, 3, e59. [CrossRef] [PubMed]
- Di Silvestre D, et al. Network Topological Analysis for the Identification of Novel Hubs in Plant Nutrition. Front Plant Sci. 2021, 12, 629013. [CrossRef] [PubMed]
- Kei Yura, et al. Alternative splicing in human transcriptome: Functional and structural influence on proteins. Gene 2006, 380, 63–71. [CrossRef]
- Chandramohan Nithya, et al. Dissection of hubs and bottlenecks in a protein-protein interaction network. Computational Biology and Chemistry 2023, 102, 107802. [CrossRef]
- Manjari Kiran et al., Global versus Local Hubs in Human Protein–Protein Interaction Network, J. Proteome Res. 2013, 12, 12, 5436–5446. [CrossRef]
- Uversky, V. N., Oldfield, C. J. & Dunker, A. K. Intrinsically disordered proteins in human diseases: introducing the D2 concept. Annu Rev Biophys 2008, 37, 215–246.
- Lazaros, K. Lazaros K.Gallos, et al. Stability and Topology of Scale-Free Networks under Attack and Defense Strategies. Physical Review Letters 2005, 94, 188701. [Google Scholar] [CrossRef]
- S. Perera, et al. "Structural characteristics of complex supply chain networks," 2017 Moratuwa Engineering Research Conference (MERCon), Moratuwa, Sri Lanka. 2017; 135–140. [CrossRef]
- Barabási, AL., Oltvai, Z. Network biology: understanding the cell's functional organization. Nat Rev Genet 2004, 5, 101–113. [CrossRef]
- Barabási, A.-L. Network Science, 1st ed.; Cambridge University Press: Cambridge, 2016. [Google Scholar]
- Dietz KJ, et al. Hubs and bottlenecks in plant molecular signalling networks. New Phytol. 2010, 188, 919–38. [CrossRef] [PubMed]
- Ermagun, A., et al. Uncertainty in vulnerability of networks under attack. Sci Rep 2023, 13, 3179. [CrossRef]
- McCabe ERB. Metabolite flux: A dynamic concept for inherited metabolic disorders as complex traits. Mol Genet Metab. 2019, 128, 14–18. [CrossRef] [PubMed]
- Newman, Mark. "The physics of networks.". Physics today 2008, 61, 33–38. [CrossRef]
- Bader G.D. and Hogue, C.W. Analyzing yeast protein–protein interaction data obtained from different sources. Nat. Biotechnol. 2002, 20, 991–997. [CrossRef] [PubMed]
- J. Wang, et al. Low protein diet up-regulate intramuscular lipogenic gene expression and down-regulate lipolytic gene expression in growth finishing pigs. Livestock Science 2012, 148, 119–128. [CrossRef]
- Cowley, M., et al., Intra- and inter-individual genetic differences in gene expression. Nat Prec 2008. [CrossRef]
- Sosa-Acosta, P., et al. Proteomics of ZIKV infected amniotic fluids of microcephalic fetuses reveals extracellular matrix and immune system dysregulation. Proteomics Clinical Applications 2022, 16, e2100041. [CrossRef]
- Davis, H.E., et al. Long COVID: major findings, mechanisms and recommendations. Nat Rev Microbiol 2023, 21, 133–146. [CrossRef] [PubMed]
- Zandi M, et al. The role of SARS-CoV-2 accessory proteins in immune evasion. Biomed Pharmacother. 2022, 156, 113889. [CrossRef]
- Ascenzi D, et al. Stereodynamical Effects by Anisotropic Intermolecular Forces. Front Chem. 2019, 7, 390. [CrossRef]
- Malaney, P., et al. Intrinsic Disorder in PTEN and its Interactome Confers Structural Plasticity and Functional Versatility. Sci Rep 2013, 3, 2035. [CrossRef] [PubMed]
- Uversky, V. N., et al. Intrinsically disordered proteins in human diseases: introducing the D2 concept. Annu Rev Biophys 2008, 37, 215–246. [CrossRef]
- Sharma A, Colonna G. System-Wide Pollution of Biomedical Data: Consequence of the Search for Hub Genes of Hepatocellular Carcinoma Without Spatiotemporal Consideration. Mol Diagn Ther. 2021, 25, 9–27. [CrossRef] [PubMed]
- Flynn RA, et al. Discovery and functional interrogation of SARS-CoV-2 RNA-host protein interactions. Cell. 2021, 184, 2394–2411e16. [CrossRef] [PubMed]
- Charitou, T. Charitou, T., Bryan, K. & Lynn, D.J. Using biological networks to integrate, visualize and analyze genomics data. Genet Sel Evol 2016, 48, 27. [Google Scholar] [CrossRef]
- Breuer K, Foroushani AK, Laird MR, Chen C, Sribnaia A, Lo R, et al. InnateDB: systems biology of innate immunity and beyond–recent updates and continuing curation. Nucleic Acids Res. 2013, 41, D1228–33. [CrossRef]
- Hong G, et al. Separate enrichment analysis of pathways for up- and downregulated genes. J R Soc Interface. 2013, 11, 20130950. [CrossRef] [PubMed]
- Wu D, Lim E, et al. ROAST: rotation gene set tests for complex microarray experiments. Bioinformatics 2010, 26, 2176–2182. [CrossRef]
- Tavleeva MM, et al. Effects of Antioxidant Gene Overexpression on Stress Resistance and Malignization In Vitro and In Vivo: A Review. Antioxidants 2022, 11, 2316. [CrossRef]
- Sharifi-Rad Mehdi, et al. Lifestyle, Oxidative Stress, and Antioxidants: Back and Forth in the Pathophysiology of Chronic Diseases. Frontiers in Physiology, Review 2020, 11. [CrossRef]
- Lu T, et al. Gene regulation and DNA damage in the ageing human brain. Nature. 2004, 429, 883–91. [CrossRef] [PubMed]
- Jolly MK, et al. EMT and MET: necessary or permissive for metastasis? Mol Oncol. 2017, 11, 755–769. [CrossRef] [PubMed]
- Kupsco A, Schlenk D. Oxidative stress, unfolded protein response, and apoptosis in developmental toxicity. Int Rev Cell Mol Biol. 2015, 317, 1–66. [CrossRef] [PubMed]
- Priya Wadgaonkar, Fei Chen, Connections between endoplasmic reticulum stress-associated unfolded protein response, mitochondria, and autophagy in arsenic-induced carcinogenesis. Seminars in Cancer Biology 2021, 76, 258–266, ISSN 1044. [CrossRef]
- Bokai Song et al. Intercellular communication within the virus microenvironment affects the susceptibility of cells to secondary viral infections. Sci. Adv. 2023, 9, eadg3433. [CrossRef] [PubMed]
- Zeng C, et al. SARS-CoV-2 spreads through cell-to-cell transmission. Proc Natl Acad Sci U S A. 2022, 119, e2111400119. [CrossRef]
- K. P. Sinaga and M. -S. Yang, "Unsupervised K-Means Clustering Algorithm,". IEEE Access 2020, 8, 80716–80727. [CrossRef]
- M. P. Rombach, et al. Core-periphery structure in networks. SIAM Journal on Applied Mathematics 2014, 74, 167–190. [CrossRef]
- Laughlin RB, et al. The middle way. Proc. Natl. Acad. Sci. USA. 2000, 97, 32–37. [CrossRef]
- Sear RP, Pagonabarraga I, Flaus A. Life at the mesoscale: The self-organised cytoplasm and nucleoplasm. BMC Biophys. 2015, 8, 4.
- PJ Thul, et al. A subcellular map of the human proteome. Science 2017, 356, eaal3321. [CrossRef]
- S Garg, et al. Characterization of methionine dependence in melanoma cells. Molecular Omics, in press. 2023. [CrossRef]
- Samad AFA, et al. MicroRNA and Transcription Factor: Key Players in Plant Regulatory Network. Front Plant Sci. 2017, 8, 565. [CrossRef]
- Vishnubalaji R, Reciprocal interplays between MicroRNAs and pluripotency transcription factors in dictating stemness features in human cancers. Semin Cancer Biol. 2022, 87, 1–16. [CrossRef] [PubMed]
- Raices, M., D'Angelo, M. Nuclear pore complex composition: a new regulator of tissue-specific and developmental functions. Nat Rev Mol Cell Biol 2012, 13, 687–699. [CrossRef]
- Tang BL, et al. The mammalian homolog of yeast Sec13p is enriched in the intermediate compartment and is essential for protein transport from the endoplasmic reticulum to the Golgi apparatus. Mol Cell Biol. 1997, 17, 256–66. [CrossRef] [PubMed]
- Betel D, et al. The microRNA.org resource: targets and expressio. Nucleic Acids Res. 2008, 36, D149–153.
- Isakova A, et al. A mouse tissue atlas of small noncoding RNA. Proc Natl Acad Sci U S A. 2020, 117, 25634–25645. [CrossRef] [PubMed]
- Pérez-Galarza, J. Immune Response to SARS-CoV-2 Infection in Obesity and T2D: Literature Review. Vaccines 2021, 9, 102. [Google Scholar] [CrossRef] [PubMed]
- Claudia Pinacchio, et al. Analysis of serum microRNAs and rs2910164 GC single-nucleotide polymorphism of miRNA-146a in COVID-19 patients. Journal of Immunoassay and Immunochemistry 2022, 43, 347–364. [CrossRef]
- Wang, Y., et al. Decreased inhibition of exosomal miRNAs on SARS-CoV-2 replication underlies poor outcomes in elderly people and diabetic patients. Sig Transduct Target Ther 2021, 6, 300. [CrossRef]
- Hadavi R, et al. Expression of Bioinformatically Candidate miRNAs including, miR-576-5p, miR-501-3p and miR-3143, Targeting PI3K Pathway in Triple-Negative Breast Cancer. Galen Med J. 2019, 8, e1646. [CrossRef] [PubMed]
- Su Li, et al. Identification of candidate biomarkers for epithelial ovarian cancer metastasis using microarray data. Oncology Letters 2017, 14, 3967–3974. [CrossRef] [PubMed]
- Khokhar M, et al. Novel Molecular Networks and Regulatory MicroRNAs in Type 2 Diabetes Mellitus: Multiomics Integration and Interactomics Study. JMIR Bioinform Biotech 2022, 3, e32437. [CrossRef]
- Wang Y, et al. A transcriptional roadmap to the induction of pluripotency in somatic cells. Stem Cell Rev Rep. 2010, 6, 282–96. [CrossRef] [PubMed]
- Aubrey BJ, Strasser A, Kelly GL. Tumor-Suppressor Functions of the TP53 Pathway. Cold Spring Harb Perspect Med. 2016, 6, a026062. [CrossRef] [PubMed]
- Olivier M, Hollstein M, Hainaut P. TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol. 2010, 2, a001008. [CrossRef] [PubMed]
- Lee JD, et al. Differences in syncytia formation by SARS-CoV-2 variants modify host chromatin accessibility and cellular senescence via TP53. bioRxiv, Preprint. 2023. [CrossRef] [PubMed]
- Tiwari R, et al. Structural similarity-based prediction of host factors associated with SARS-CoV-2 infection and pathogenesis. J Biomol Struct Dyn. 2022, 40, 5868–5879. [CrossRef] [PubMed]
- Liu CG, et al. "MicroRNA expression profiling using microarrays". Nature Protocols. 2008, 3, 563–568. [CrossRef] [PubMed]
- Lagendijk AK, et al. "Revealing details: whole mount microRNA in situ hybridization protocol for zebrafish embryos and adult tissues". Biology Open. 2012, 1, 566–569. [CrossRef]
- Shingara J, et al. "An optimized isolation and labeling platform for accurate microRNA expression profiling". RNA. 2005, 11, 1461–1470. [CrossRef] [PubMed]
- Chen Z, et al. Interactomes of SARS-CoV-2 and human coronaviruses reveal host factors potentially affecting pathogenesis. EMBO J. 2021, 40, e107776. [CrossRef] [PubMed]
- Delgado-Chaves FM, et al. Computational Analysis of the Global Effects of Ly6E in the Immune Response to Coronavirus Infection Using Gene Networks. Genes (Basel). 2020, 11, 831. [CrossRef] [PubMed]
- Guzzi PH, et al. Master Regulator Analysis of the SARS-CoV-2/Human Interactome. J Clin Med. 2020, 9, 982. [CrossRef] [PubMed]
- Zaim S, et al. COVID-19 and Multiorgan Response. Curr Probl Cardiol. 2020, 45, 100618. [CrossRef] [PubMed]
- Thakur, V.; et al. Multi-Organ Involvement in COVID-19: Beyond Pulmonary Manifestations. J. Clin. Med. 2021, 10, 446. [Google Scholar] [CrossRef]
- K. Moravcevic, et al. “Conditional Peripheral Membrane Proteins: Facing up to Limited Specificity”. Structure 2012, 20, 15–27. [CrossRef] [PubMed]
- Seaton, B.A. Seaton B.A. and Roberts M.F. Peripheral membrane proteins. In Biological Membranes; Mertz, K., Roux, B., Eds.; Birkhauser: Boston; pp. 355–403.
- A.M. Whited, A. Johs. The interactions of peripheral membrane proteins with biological membranes. Chemistry and Physics of Lipids 2015, 192, 51–59. [CrossRef]
- Félix M. Goñi. Non-permanent proteins in membranes: when proteins come as visitors (Review). Molecular Membrane Biology 2002, 19, 237–245. [CrossRef]
- Puthenveetil, R.; Christenson, E.T.; Vinogradova, O. New Horizons in Structural Biology of Membrane Proteins: Experimental Evaluation of the Role of Conformational Dynamics and Intrinsic Flexibility. Membranes 2022, 12, 227. [Google Scholar] [CrossRef]
- Cho, W. and Stahelin, R.V. "Membrane-protein interactions in cell signaling and membrane trafficking". Annual Review of Biophysics and Biomolecular Structure 2005, 34, 119–151. [CrossRef]
- Robertson RM, et al. A two-helix motif positions the lysophosphatidic acid acyltransferase active site for catalysis within the membrane bilayer. Nat Struct Mol Biol. 2017, 24, 666–671. [CrossRef]
- Seelig J. "Thermodynamics of lipid-peptide interactions". Biochim Biophys Acta 2004, 1666, 40–50. [CrossRef]
- Allen KN, et al. Monotopic Membrane Proteins Join the Fold. Trends Biochem Sci. 2019, 44, 7–20. [CrossRef]
- Dowran R, et al. Reinfection and reactivation of SARS-CoV-2. Future Virol. 2022. [CrossRef]
- Costanzo M, et al. Deciphering the Relationship between SARS-CoV-2 and Cancer. Int J Mol Sci. 2023, 24, 7803. [CrossRef]
- Stingi A, Cirillo L. SARS-CoV-2 infection and cancer: Evidence for and against a role of SARS-CoV-2 in cancer onset. Bioessays 2021, 43, e2000289. [CrossRef]
- Cardozo C.M., Hainaut P. Viral strategies for circumventing p53: the case of severe acute respiratory syndrome coronavirus. Curr Opin Oncol. 2021, 33, 149–158. [PubMed]
- Gómez-Carballa A, et al. Is SARS-CoV-2 an oncogenic virus? J Infect. 2022, 85, 573–607. [CrossRef]
- Jahankhani K, et al. Possible cancer-causing capacity of COVID-19: Is SARS-CoV-2 an oncogenic agent? Biochimie. 2023, 213, 130–138. [CrossRef] [PubMed]
- Carlson CR, et al. Phosphoregulation of phase separation by the SARS-CoV-2 N protein suggests a biophysical basis for its dual functions. Mol. Cell 2020, 80, 1092–1103.e4. [CrossRef] [PubMed]
- Cubuk J, et al. The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nat. Commun. 2021, 12, 1936. [CrossRef] [PubMed]
- Wu C, Qavi AJ, et al. Characterization of SARS-CoV-2 nucleocapsid protein reveals multiple functional consequences of the C-terminal domain. iScience 2021, 24, 102681. [CrossRef]
- Zheng Y, Gao C. Phase Separation: The Robust Modulator of Innate Antiviral Signaling and SARS-CoV-2 Infection. Pathogens. 2023, 12, 243. [CrossRef]
- Jack A, et al. SARS-CoV-2 nucleocapsid protein forms condensates with viral genomic RNA. PLOS Biol 2021, 19, e3001425. [CrossRef] [PubMed]
- Lu S, et al. The SARS-CoV-2 nucleocapsid phosphoprotein formsmutually exclusive condensates with RNA and the membrane-associatedMprotein. Nat.Commun. 2021, 12, 502. [CrossRef] [PubMed]
- Risso-Ballester J., et al. A condensate-hardening drug blocks RSV replication in vivo. Nature 2021, 595, 596–599. [CrossRef] [PubMed]
- Li H., et al. Phase separation in viral infections. Trends Microbiol. 2022, 30, 1217–1231. [CrossRef]
- Wang B., et al. Liquid-liquid phase separation in human health and diseases. Signal Transduct. Target. Ther. 2021, 6, 290. [CrossRef]
- Li P, et al. Phase transitions in the assembly of multivalent signalling proteins. Nature 2012, 483, 336–340. [CrossRef]
- Szklarczyk D, et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [CrossRef]
- Szklarczyk D, et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023, 51, D638–D646. [CrossRef]
- Doncheva NT, et al. Cytoscape StringApp: Network Analysis and Visualization of Proteomics Data. J Proteome Res. 2019, 18, 623–632. [CrossRef]
- Chung F, Lu L, Dewey TG, Galas DJ. Duplication models for biological networks. J Comput Biol. 2003, 10, 677–87. [CrossRef] [PubMed]
- Scardoni G, Tosadori G, Faizan M, Spoto F, Fabbri F, Laudanna C. Biological network analysis with CentiScaPe: centralities and experimental dataset integration. F1000Res. 2014, 3, 139. [CrossRef]
- Perera, S.; Perera, H.N.; Kasthurirathna, D. "Structural characteristics of complex supply chain networks," 2017 Moratuwa Engineering Research Conference (MERCon), Moratuwa, Sri Lanka, 2017. 135–140. [CrossRef]
- Barabási, A.-L. Network Science, 1st ed.; Cambridge University Press: Cambridge, 2016. [Google Scholar]
- Syakur, M A et al. Integration K-Means Clustering Method and Elbow Method For Identification of The Best Customer Profile Cluster - IOP Conference Series. Materials Science and Engineering; Bristol Vol. 336, Fasc. 1, (Apr 2018). [CrossRef]
- Zhou, G., et al. "NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis". Nucleic Acids Research 2019, 47, W234–W241. [CrossRef] [PubMed]
- Xia J, et al. NetworkAnalyst--integrative approaches for protein-protein interaction network analysis and visual exploration. Nucleic Acids Res. 2014, 42, W167–74. [CrossRef] [PubMed]
- Zhi-Ping Liu, et al. (2015). RegNetwork: an integrated database of transcriptional and posttranscriptional regulatory networks in human and mouse. Database 2015. [CrossRef]
- Gábor Erdős, Zsuzsanna Dosztányi. Analyzing Protein Disorder with IUPred2A. Current Protocols in Bioinformatics 2020, 70, e99.
- Drozdetskiy A, et al. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 2015, 43, W389–94. [CrossRef] [PubMed]
- Yuan Ren et al., Force redistribution in clathrin-mediated endocytosis revealed by coiled-coil force sensors. Sci. Adv. 2023, 9, eadi1535. [CrossRef] [PubMed]
- M. Skruzny, E. Pohl, S. Gnoth, G. Malengo, V. Sourjik, The protein architecture of the endocytic coat analyzed by FRET microscopy. Mol. Syst. Biol. 2020, 16, e9009.
- Mahapatra A, et al. The Mechanics and Thermodynamics of Tubule Formation in Biological Membranes. J Membr Biol. 2021, 254, 273–291. [CrossRef] [PubMed]
- R. C. Aguilar, et al. The yeast epsin Ent1 is recruited to membranes through multiple independent interactions. J. Biol. Chem. 2003, 278, 10737–10743. [CrossRef]
- Å. E. Y. Engqvist-Goldstein, et al. The actin-binding protein Hip1R associates with clathrin during early stages of endocytosis and promotes clathrin assembly in vitro. J. Cell Biol. 2001, 154, 1209–1224. [CrossRef] [PubMed]
- J. D. Wilbur, et al. Actin binding by Hip1 (Huntingtin-interacting protein 1) and Hip1R (Hip1-related protein) is regulated by clathrin light chain. J. Biol. Chem. 2008, 283, 32870–32879. [CrossRef] [PubMed]
- Skruzny, M. The endocytic protein machinery as an actin-driven membrane-remodeling machine. Eur J Cell Biol. 2022, 101, 151267. [Google Scholar] [CrossRef] [PubMed]
- Shin N, et al. SNX9 regulates tubular invagination of the plasma membrane through interaction with actin cytoskeleton and dynamin 2. J Cell Sci. 2008, 121, 1252–63. [CrossRef] [PubMed]
- Hsu PP, et al. The mTOR-regulated phosphoproteome reveals a mechanism of mTORC1-mediated inhibition of growth factor signaling. Science. 2011, 332, 1317–22. [CrossRef]
- Ekim B, et al. mTOR kinase domain phosphorylation promotes mTORC1 signaling, cell growth, and cell cycle progression. Mol Cell Biol. 2011, 31, 2787–801. [CrossRef]
- Jacinto E, et al. Mammalian TOR complex 2 controls the actin cytoskeleton and is rapamycin insensitive. Nat Cell Biol. 2004, 6, 1122–8. [CrossRef] [PubMed]
- Ma G, et al. DEPTOR as a novel prognostic marker inhibits the proliferation via deactivating mTOR signaling pathway in gastric cancer cells. Exp Cell Res. 2023, 427, 113598. [CrossRef] [PubMed]
- Information about DEPTOR is also from NCBI – USA. Available online: https://www.ncbi.nlm.nih.gov/gene/64798.
- Wu, W., et al. The SARS-CoV-2 nucleocapsid protein: its role in the viral life cycle, structure and functions, and use as a potential target in the development of vaccines and diagnostics. Virol J 2023, 20, 6. [CrossRef] [PubMed]
- Kloc, M., et al. Virus interactions with the actin cytoskeleton—what we know and do not know about SARS-CoV-2. Arch Virol 2022, 167, 737–749. [CrossRef]
- Zhong P., et al. Trasmissione dei virus da cellula a cellula. Curr. Opinion. Virolo. 2013, 3, 44–50.
- Dale B. M., et al. Cell-to-cell transfer of HIV-1 via virological synapses leads to endosomal virion maturation that activates viral membrane fusion. Cell Host Microbe 2011, 10, 551–562. [CrossRef]
- Kruglova N, Siniavin A, Gushchin V, Mazurov D. Different Neutralization Sensitivity of SARS-CoV-2 Cell-to-Cell and Cell-Free Modes of Infection to Convalescent Sera. Viruses. 2021, 13, 1133. [CrossRef]
- Zeng C, et al. SARS-CoV-2 spreads through cell-to-cell transmission. Proc Natl Acad Sci U S A. 2022, 119, e2111400119. [CrossRef]
- Carlson CR, et al. Phosphoregulation of phase separation by the SARS-CoV-2 N protein suggests a biophysical basis for its dual functions. Mol Cell. 2020, 80, 1092–103e4. [CrossRef]
- Zhao M, et al. GCG inhibits SARS-CoV-2 replication by disrupting the liquid phase condensation of its nucleocapsid protein. Nat Commun. 2021, 12, 2114. [CrossRef] [PubMed]
- Zhao D, et al. Understanding the phase separation characteristics of nucleocapsid protein provides a new therapeutic opportunity against SARS-CoV-2. Protein Cell. 2021, 12, 734–40. [CrossRef]
- Alizon S, et al. Virulence evolution and the trade-off hypothesis: History, current state of affairs and the future. J Evol Biol. 2009, 22, 245–259 pmid:19196383. [CrossRef]
- Miller MR, et al. The evolution of parasites in response to tolerance in their hosts: The good, the bad, and apparent commensalism. Evolution. 2006, 60, 945. [PubMed]
- Best A, et al. The coevolutionary implications of host tolerance. Evolution. 2014, 68, 1426–1435. [CrossRef] [PubMed]
- Randolph HE, et al. Holy Immune Tolerance. Batman! Immunity. 2018, 48, 1074–1076. [CrossRef] [PubMed]
- Hsieh, Min-Kang, and Jeffery B. Klauda. "Multiscale Molecular Dynamics Simulations of the Homodimer Accessory Protein ORF7b of SARS-CoV-2.". The Journal of Physical Chemistry B 2023, 128, 150–162, Virtual Special Issue “Gregory A. Voth Festschrift”.
- Chen WS et al. Functional independence of the epidermal growth factor receptor from a domain required for ligand-induced internalization and calcium regulation. Cell. 1989, 59, 33–43. [CrossRef] [PubMed]
- Zhang, C.; et al. Transmembrane and coiled-coil domain family 1 is a novel protein of the endoplasmic reticulum. PLoS One. 2014, 9, e85206. [Google Scholar] [CrossRef]
- Principe, S.; et al. In-depth proteomic analyzes of exosomes isolated from expressed prostatic secretions in urine. Proteomics. 2013, 13, 1667–1671. [Google Scholar] [CrossRef]
- Scardoni G, Tosadori G, Faizan M, Spoto F, Fabbri F, Laudanna C. Biological network analysis with CentiScaPe: centralities and experimental dataset integration. F1000Res. 2014, 3, 139. [CrossRef]
- Appel, E. A. et al. Supramolecular cross-linked networks via host-guest complexation with cucurbituril. J. Am. Chem. Soc. 2010, 132, 14251–14260. [CrossRef]
- Tan, C. S. Y. et al. Distinguishing relaxation dynamics in transiently crosslinked polymeric networks. Polym. Chem. 2017, 8, 5336–5343. [CrossRef]
- Anna, V. Ignatenko, et al. Chapter Seven - Nonlinear Chaotic Dynamics of Quantum Systems: Molecules in an Electromagnetic Field. In Advances in Quantum Chemistry; Samantha Jenkins, et al., Eds.; Academic Press, 2019; Volume 78, pp. 149–170. ISBN 9780128160848. ISSN 0065-3276. Available online: https://www.sciencedirect.com/science/article/pii/S0065327618300303. [CrossRef]
- Yu AC, et al. Physical networks from entropy-driven non-covalent interactions. Nat Commun. 2021, 12, 746. [CrossRef] [PubMed]
- VanHulle, M M. Entropy driven artificial neuronal networks and sensorial representation; A proposal. Journal of Parallel and Distributed Computing 1989, 6, 2. ISSN 0743–7315.
- Bernaschi, M., et al. The Fitness-Corrected Block Model, or how to create maximum-entropy data-driven spatial social networks. Sci Rep 2022, 12, 18206. [CrossRef] [PubMed]
- Yongxin Li, et al. Entropy driven circuit as an emerging molecular tool for biological sensing: A review. TrAC Trends in Analytical Chemistry 2021, 134, 116142. ISSN 0165–9936. [CrossRef]
- Vladimir N. Uversky. Dancing Protein Clouds: The Strange Biology and Chaotic Physics of Intrinsically Disordered Proteins. Journal of Biological Chemistry 2016, 291, 6689–6695. [CrossRef]
- Tanaka, F. Tanaka, F. & Edwards, S. F. Viscoelastic properties of physically cross-linked networks. 2. Dynamic mechanical moduli. J. Non-Newton. Fluid Mech. 1992, 43, 273–288. [Google Scholar]
- Lun XK, Zanotelli VR, et al. Influence of node abundance on signaling network state and dynamics analyzed by mass cytometry. Nat Biotechnol. 2017, 35, 164–172. [CrossRef] [PubMed]
- Chen, Y., Li, Y., Xiong, J. et al. Role of PRKDC in cancer initiation, progression, and treatment. Cancer Cell Int 2021, 21, 563. [CrossRef]
- V.N. Uversky, C.J. Oldfield, A.K. Dunker. Intrinsically disordered proteins in human diseases: Introducing the D 2 concept. Annu Rev Biophys 2008, 37, 215–246. [CrossRef] [PubMed]
- E. Koshland. Application of a Theory of Enzyme Specificity to Protein Synthesis. Proc Natl Acad Sci 1958, 44, 98–104. [CrossRef] [PubMed]

| Action | Enriched terms |
|---|---|
| Biological Process (Gene Ontology): | 1690 GO-terms |
| Molecular Function (Gene Ontology): | 166 GO-terms |
| Cellular Component (Gene Ontology): | 267 GO-terms |
| Reference publications (PubMed): | >10,000 publications |
| Local network cluster (STRING): | 137 clusters |
| KEGG Pathways: | 195 pathways |
| Reactome Pathways: | 494 pathways |
| WikiPathways: | 259 pathways |
| Disease-gene associations (DISEASES): | 112 diseases |
| Tissue expression (TISSUES): | 186 tissues |
| Subcellular localization (compartments): | 249 compartments significantly |
| Human Phenotype (Monarch): | 1002 phenotypes |
| Annotated Keywords (UniProt): | 99 keywords |
| Protein Domains (Pfam): | 63 domains |
| Protein Domains and Features (InterPro): | 118 domains |
| Protein Domains (SMART): | 20 domains |
| All enriched terms (without PubMed): | 5,057 enriched terms in 15 categories |
| GO Term ID | Term description | Number of involved proteins | p-value |
|---|---|---|---|
| GO:0051179 | Localization | 378 | 2.01e-77 |
| GO:0006810 | Transport | 320 | 3.04e-67 |
| GO:0007169 | Transmembrane receptor protein tyrosine kinase signaling pathway | 124 | 1.23e-66 |
| GO:0051234 | Establishment of localization | 322 | 7.72e-66 |
| GO:0015833 | Peptide transport | 187 | 1.09e-62 |
| GO:0051649 | Establishment of localization in cell | 230 | 3.37e-62 |
| GO:0051641 | Cellular localization | 254 | 1.29e-60 |
| GO:0015031 | Protein transport | 181 | 7.86e-60 |
| GO:0007167 | Enzyme linked receptor protein signaling pathway | 131 | 7.95e-59 |
| GO:0008104 | Protein localization | 213 | 1.46e-58 |
| GO:0045184 | Establishment of protein localization | 183 | 2.85e-58 |
| GO:0016192 | Vesicle-mediated transport | 189 | 1.18e-53 |
| GO:0032879 | Regulation of localization | 229 | 2.95e-51 |
| GO:0009987 | Cellular process | 546 | 4.49e-51 |
| GO:0046907 | Intracellular transport | 168 | 1.19e-49 |
| GO Term ID | COMPARTMENT | Number of involved proteins | p-value |
|---|---|---|---|
| GOCC:0016020 | Membrane | 399 | 2.58e-92 |
| GOCC:0012505 | Endomembrane system | 302 | 1.36e-91 |
| GOCC:0031090 | Organelle membrane | 243 | 2.07e-77 |
| GOCC:0098796 | Membrane protein complex | 189 | 7.35e-74 |
| GOCC:0005737 | Cytoplasm | 437 | 1.41e-73 |
| GOCC:0031982 | Vesicle | 213 | 5.13e-62 |
| GOCC:0098588 | Bounding membrane of organelle | 174 | 3.17e-58 |
| GOCC:0005783 | Endoplasmic reticulum | 133 | 6.29e-55 |
| GOCC:0098805 | Whole membrane | 156 | 1.76e-53 |
| GOCC:0110165 | Cellular anatomical entity | 531 | 2.59e-51 |
| GOCC:0005789 | Endoplasmic reticulum membrane | 105 | 4.93e-51 |
| GOCC:0042175 | Nuclear outer membrane-ER membrane network | 106 | 1.79e-50 |
| GOCC:0031410 | Cytoplasmic vesicle | 177 | 1.29e-49 |
| GOCC:0032991 | Protein-containing complex | 306 | 4.76e-44 |
| GOCC:0043226 | Organelle | 437 | 1.20e-41 |
| GOCC:0043227 | Membrane-bounded organelle | 406 | 5.80e-41 |
| GOCC:0005622 | Intracellular | 462 | 8.33e-38 |
| GOCC:0043229 | Intracellular organelle | 407 | 4.82e-34 |
| GOCC:0005829 | Cytosol | 201 | 2.18e-32 |
| GOCC:0005886 | Plasma membrane | 220 | 3.75e-30 |
| GOCC:0031201 | SNARE complex | 34 | 3.79e-30 |
| GOCC:0043231 | Intracellular membrane-bounded organelle | 349 | 6.22e-30 |
| Term ID | Molecular Mechanism | Number of involved proteins | p-value |
|---|---|---|---|
| HSA-9006934 | Signaling by Receptor Tyrosine Kinases | 140 | 4.44e-84 |
| HSA-1643685 | Disease | 189 | 2.66e-63 |
| HSA-422475 | Axon guidance | 101 | 6.79e-45 |
| HSA-9675108 | Nervous system development | 103 | 6.79e-45 |
| HSA-168256 | Immune System | 176 | 6.06e-41 |
| HSA-5663205 | Infectious disease | 115 | 6.30e-41 |
| HSA-162582 | Signal Transduction | 204 | 2.84e-37 |
| HSA-5653656 | Vesicle-mediated transport | 95 | 2.70e-34 |
| HSA-199991 | Membrane Trafficking | 92 | 5.34e-34 |
| HSA-392499 | Metabolism of proteins | 163 | 3.25e-33 |
| HSA-109582 | Hemostasis | 89 | 1.02e-32 |
| HSA-1799339 | SRP-dependent cotranslational protein targeting to membrane | 45 | 2.19e-31 |
| HSA-168249 | Innate Immune System | 111 | 1.33e-30 |
| HSA-1227986 | Signaling by ERBB2 | 35 | 2.43e-30 |
| HSA-74752 | Signaling by Insulin receptor | 38 | 5.77e-29 |
| HSA-177929 | Signaling by EGFR | 33 | 5.40e-28 |
| HSA-4420097 | VEGFA-VEGFR2 Pathway | 39 | 5.35e-27 |
| HSA-202733 | Cell surface interactions at the vascular wall | 42 | 3.19e-25 |
| HSA-76002 | Platelet activation, signaling and aggregation | 52 | 4.89e-24 |
| HSA-6811558 | PI5P, PP2A and IER3 Regulate PI3K/AKT Signaling | 37 | 5.16e-24 |
| HSA-5683057 | MAPK family signaling cascades | 52 | 1.19e-20 |
| HSA-5684996 | MAPK1/MAPK3 signaling | 49 | 1.37e-20 |
| HSA-77387 | Insulin receptor recycling | 21 | 1.07e-19 |
| HSA-192823 | Viral mRNA Translation | 27 | 1.54e-16 |
| HSA-1500931 | Cell-Cell communication | 30 | 1.91e-15 |
| TERM ID | HUMAN TISSUES INVOLVED WITH ORF7b | Number of involved proteins | p-value |
|---|---|---|---|
| BTO:0000345 | Digestive gland | 233 | 4.73e-56 |
| BTO:0001491 | Viscus | 322 | 1.59e-54 |
| BTO:0001489 | Whole body | 504 | 2.18e-45 |
| BTO:0000522 | Gland | 356 | 2.76e-45 |
| BTO:0000759 | Liver | 178 | 2.17e-44 |
| BTO:0001488 | Endocrine gland | 323 | 1.15e-37 |
| BTO:0003091 | Urogenital system | 341 | 3.03e-36 |
| BTO:0000227 | Central nervous system | 303 | 1.41e-35 |
| BTO:0001484 | Nervous system | 307 | 3.23e-35 |
| BTO:0000449 | Fetus | 125 | 1.68e-32 |
| BTO:0001078 | Placenta | 119 | 1.40e-30 |
| BTO:0000081 | Reproductive system | 308 | 5.01e-30 |
| BTO:0003099 | Internal female genital organ | 183 | 5.01e-30 |
| BTO:0000174 | Embryonic structure | 159 | 7.75e-28 |
| BTO:0000203 | Respiratory system | 127 | 9.81e-28 |
| BTO:0000083 | Female reproductive system | 292 | 1.71e-27 |
| BTO:0000089 | Blood | 136 | 1.39e-26 |
| BTO:0000570 | Hematopoietic system | 172 | 6.39e-26 |
| BTO:0000763 | Lung | 105 | 3.59e-23 |
| BTO:0000988 | Pancreas | 72 | 6.06e-23 |
| BTO:0000431 | Excretory gland | 106 | 8.44e-21 |
| BTO:0003092 | Urinary system | 97 | 3.43e-19 |
| BTO:0001244 | Urinary tract | 97 | 4.17e-19 |
| BTO:0000671 | Kidney | 86 | 5.49e-19 |
| BTO:0001129 | Prostate gland | 58 | 8.08e-19 |
| BTO:0000132 | Blood platelet | 50 | 2.81e-18 |
| BTO:0000511 | Gastrointestinal tract | 116 | 4.50e-17 |
| BTO:0000131 | Blood plasma | 51 | 1.46e-16 |
| BTO:0000574 | Hematopoietic cell | 77 | 1.21e-14 |
| BTO:0000082 | Male reproductive system | 148 | 3.21e-14 |
| BTO:0000751 | Leukocyte | 72 | 3.31e-14 |
| BTO:0000080 | Male reproductive gland | 138 | 6.39e-13 |
| BTO:0000254 | Female reproductive gland | 145 | 7.61e-12 |
| BTO:0005810 | Immune system | 96 | 3.68e-11 |
| BTO:0003096 | Internal male genital organ | 122 | 4.92e-11 |
| BTO:0000088 | Cardiovascular system | 70 | 4.40e-10 |
| BTO:0000421 | Connective tissue | 63 | 1.19e-09 |
| BTO:0000439 | Eye | 59 | 1.33e-09 |
| BTO:0000706 | Large intestine | 54 | 1.57e-09 |
| BTO:0000202 | Sense organ | 69 | 1.98e-09 |
| BTO:0000855 | Lymph | 25 | 4.56e-09 |
| BTO:0001085 | Vascular system | 38 | 9.90e-09 |
| BTO:0001424 | Uterus | 67 | 1.11e-08 |
| BTO:0000269 | Colon | 46 | 3.05e-08 |
| BTO:0001363 | Testis | 85 | 2.48e-05 |
| Pathway | Description | Number of involved proteins | p-value |
|---|---|---|---|
| hsa04012 | ErbB signaling pathway | 50 | 3.02e-41 |
| hsa04510 | Focal adhesion | 64 | 2.27e-40 |
| hsa01521 | EGFR tyrosine kinase inhibitor resistance | 46 | 4.29e-38 |
| hsa04151 | PI3K-Akt signaling pathway | 74 | 1.16e-36 |
| hsa04141 | Protein processing in ER | 55 | 1.44e-35 |
| hsa04015 | Rap1 signaling pathway | 51 | 3.72e-28 |
| hsa04014 | Ras signaling pathway | 52 | 3.76e-27 |
| hsa05206 | MicroRNAs in cancer | 45 | 1.48e-26 |
| hsa04935 | Growth hormone synthesis, secretion action | 40 | 3.80e-26 |
| hsa04130 | SNARE interactions in vesicular transport | 27 | 1.38e-25 |
| hsa04062 | Chemokine signaling pathway | 45 | 2.35e-24 |
| hsa04145 | Phagosome | 40 | 9.33e-24 |
| hsa04360 | Axon guidance | 43 | 2.15e-23 |
| hsa04072 | Phospholipase D signaling pathway | 39 | 2.11e-22 |
| hsa04917 | Prolactin signaling pathway | 30 | 2.84e-22 |
| hsa04150 | mTOR signaling pathway | 39 | 4.38e-22 |
| hsa04810 | Regulation of actin cytoskeleton | 42 | 3.07e-20 |
| hsa01522 | Endocrine resistance | 31 | 4.10e-20 |
| hsa04915 | Estrogen signaling pathway | 35 | 4.10e-20 |
| hsa04722 | Neurotrophin signaling pathway | 33 | 4.29e-20 |
| hsa04919 | Thyroid hormone signaling pathway | 32 | 9.72e-19 |
| hsa04664 | Fc epsilon RI signaling pathway | 26 | 1.16e-18 |
| hsa04010 | MAPK signaling pathway | 45 | 5.16e-18 |
| hsa04721 | Synaptic vesicle cycle | 26 | 1.05e-17 |
| hsa04660 | T cell receptor signaling pathway | 29 | 1.05e-17 |
| hsa04662 | B cell receptor signaling pathway | 26 | 2.90e-17 |
| hsa04650 | Natural killer cell mediated cytotoxicity | 30 | 7.46e-17 |
| HUB protein | Number of GO Processes | Bottleneck protein | Number of GO Processes |
|---|---|---|---|
| EGFR | 408 | EGFR | 408 |
| PIK3R1 | 328 | HSPA5 | 234 |
| EGF | 646 | MTOR | 413 |
| HRAS | 245 | SEC13 | 83 |
| GRB2 | 233 | SEC61A1 | 63 |
| SRC | 508 | SRC | 508 |
| PIK3CA | 271 | VAMP2 | 143 |
| Viral Protein | Human target | Viral protein features** |
|---|---|---|
| nsp4* | EGFR | Is involved in the assembly of virally induced cytoplasmic double- membrane vesicles necessary for viral replication. |
| M* | EGFR | Component of the viral envelope. |
| ORF3a* | EGFR | Homotetrameric potassium sensitive ion channels (viroporin) and may modulate virus release |
| ORF7b* | EGFR | This paper |
| S | EGFR | Spike or Surface glycoprotein |
| nsp4* | SRC | See above |
| nsp5* | SRC | Is a cysteine protease, essential for the viral life cycle. |
| nsp6* | SRC | Plays a role in the initial induction of auto-phagosomes from host reticulum endoplasmic |
| nsp13* | SRC | Multi-functional helicase with a zinc-binding domain in N-terminus |
| nsp14* | SRC | 3'-5' deoxyribonuclease |
| E* | SRC | Plays a central role in virus morphogenesis and assembly |
| M* | SRC | See above |
| ORF3a* | SRC | See above |
| ORF3b | SRC | Could be involved in immune evasion as interferon agonist (78) |
| ORF6* | SRC | Could be a determinant of virus virulence |
| ORF7a* | SRC | Non-structural protein, which is dispensable for virus replication in cell culture |
| ORF7b* | SRC | See above |
| ORF8 | SRC | Is a viral cytokine regulating immune responses |
| S | SRC | See above |
| M* | PIK3R1 | See above |
| ORF7b* | PIK3R1 | See above |
| ORF3b | PIK3R2 | See above |
| M* | PIK3R3 | See above |
| S | PIK3R3 | See above |
| N* | ORF7b | Responsible for wrapping viral RNA into a symmetric helical structure |
| Function | Strength* | p-value | P | Human Proteins involved in the process |
|---|---|---|---|---|
| Negative regulation of ERBB signaling pathway | 1.22 | 1.38e-18 | 22.13 | HBEGF, EREG, PTPN12, TSG101, CBL, CBLB, EGF, ERBB2, CBLC, EGFR, TGFA, SOCS5, PTPN2, HGS, EPS15, ERRFI1, SNX5, SH3GL2, GRB2, BTC, AREG, SH3KBP1, CDC42, EPN1, EPGN |
| Negative regulation of EGFR signaling pathway | 1.23 | 3.24e-17 | 21.53 | HBEGF, EREG, TSG101, CBL, CBLB, EGF, CBLC, EGFR, TGFA, SOCS5, PTPN2, HGS, EPS15, ERRFI1, SNX5, SH3GL2, GRB2, BTC, AREG, SH3KBP1, CDC42, EPN1, EPGN |
| Negative regulation of anoikis | 1.14 | 5.72e-05 | 6.56 | PIK3CA, ITGA5, BCL2L1, CAV1, PTK2, SRC, ITGB1 |
| Negative regulation of extrinsic apoptotic signaling pathway | 0.76 | 9.26e-07 | 6.05 | GCLC, LGALS3, BCL2L1, IGF1, CTNNA1, UNC5B, FYN, FAS, CASP8, LMNA, GCLM, SRC, AR, CTTN, NRG1, ITGA6, AKT1 |
| Negative regulation of protein tyrosine kinase activity | 0.99 | 9.13e-05 | 5.90 | TSG101, CBL, CBLB, CBLC, SOCS5, PTPN2, CAV1, ERRFI1 |
| Negative regulation of epidermal growth factor-activated receptor activity | 1.18 | 1.7e-04 | 4.99 | TSG101, CBL, CBLB, CBLC, SOCS5, ERRFI1 |
| Negative regulation of interleukin-6 production | 0.81 | 5.0e-05 | 4.61 | CSK, SOCS5, GAS6, TLR9, VIMP, PTPN6, ARRB1, ENSP00000417517 |
| Negative regulation of peptidyl-tyrosine phosphorylation | 0.88 | 1.50e-05 | 4.55 | TSG101, CBL, CBLB, CBLC, SPINK1, SOCS5, PTPN2, CAV1, ERRFI1, PRKCD, PTPN6 |
| Negative regulation of PERK-mediated unfolded protein response | 1.33 | 9.2e-03 | 4.12 | NCK2, PTPN1, NCK1 |
| Negative regulation of endoplasmic reticulum unfolded protein response | 1.04 | 8.9e-03 | 4.11 | NCK2, HSPA5, PTPN1, NCK1 |
| Negative regulation of blood-brain barrier permeability | 1.55 | 3.13e-02 | 3.88 | SH3GL2, VEGFA |
| Negative regulation of response to oxidative stress | 0.81 | 4.1e-04 | 3.74 | SLC7A11, MET, GGT7, CTNNB1, FYN, NFE2L2, INS, HIF1A, AKT1 |
| Negative regulation of protein tyrosine phosphatase activity | 1.39 | 4.67e-02 | 3.71 | LGALS3, GNAI2 |
| Negative regulation of mesenchymal to epithelial transition | 1.38 | 4.77e-02 | 3.69 | CTNNB1, STAT1 |
| Negative regulation of blood coagulation | 0.83 | 2.8e-04 | 3.69 | PROC, PDGFRA, F2, PLAUR, PLAU, EDN1, CD9, PROS1, PRKCD |
| Negative regulation of primary miRNA processing | 1.38 | 4.67e-02 | 3.68 | STAT3, IL6 |
| Negative regulation of lipid transport | 0.79 | 1.74e-02 | 1.77 | EGF, PTPN11, SREBF2, AKT1, ITGB3 |
| COMPARTMENT | PROTEINS* | Protein number |
|---|---|---|
| EXTRACELLULAR | AREG, BTC, CD81, CD9, EGF, EGFR, ERBB3, EREG, HBEGF, HSPA8, INS, LAMA1, LAMB1, MUC1, NRG1, NRG3, PLAU, SFN, TGFA, TSG101 | 20 |
| CYTOSKELETON | CTNNA1, CTNNB1, GNAI1, GNAI3, LMNA, MAPK1, PPP2R1A, PTPN3 | 8 |
| PLASMA MEMBRANE | ADAM17, ARF4, BTC, CAV1, CAV2, CD44, CD81, CD82, CDH1, CTNNA1, CTNNB1, EDNRA, EGFR, EPS15, ERBB2, ERBB2IP, ERBB3, ERBB4, EREG, GAB2, GNAI1, GNAI3, HBEGF, HCK, HRAS, ITGA3, ITGB1, ITGB4, JUP, KRAS, LAPTM4B, LPAR1, LPAR3, LYN, MUC1, NRG1, NRG3, PDGFRA, PIK3C2B, PLCG1, PLCG2, PPP2R1A, PRKCA, PRKCB, PTPN2, PTPN3, PTPRK, PTRF, SHC1, SLC9A1, SLC9A3R1, TGFA, TSG101, USP8 | 54 |
| CYTOSOL | PIK3C2B, GRB7, ARF4, PPP2R1A, PLCG1, HCK, USP8, PRKCA, MAPK1, RAB5A, FOS, HSPA8, CTNNB1, HIF1A, GAPDH | 15 |
| MITOCHONDRION | PPP2R1A, MAPK1, HSP90AA1,LGALS3, ERBB4, PTRF, MT-CO2 | 7 |
| GOLGI | CAV2, CBL, CDH1, HRAS, LYN, MAPK1 | 5 |
| ER | FOS, NCK1, PTPN2 | 3 |
| PEROXISOME | No level 5 protein | - |
| ENDOSOME | CDH1, EGFR, CAV1, ERBB2, PTPN1, RAB5A, MAPK1, TSG101, GRB2, HGS, USP8, LPAR1, LAPTM4B, GRAP2 | 14 |
| LYSOSOME | LAPTM4B, HSPA8, MTOR, HCK | 4 |
| NUCLEUS | CAV2,CTNNB1, EGFR, ERBB2, ERBB2IP, ERBB4, FOS, GRAP2, GRB2, HIF1A, HRAS, HSPA8, IGFBP3, JAK2, LGALS3, LMNA, LYN, MAPK1, MUC1, NCK1, NCL, NRG1, PLCG1, PPP2R1A, PRKCB, PRKDC, PTPN11, PTPN2, PTPN6, PTRF, STAT1, STAT3, STAT5B, TFAP2C | 34 |
| COMPARTMENT | PROTEINS* | Protein number |
|---|---|---|
| EXTRACELLULAR | EDN1, F2, FAS, HSP90AB1, LAMA5, LAMC1, MET, NTN1, VEGFA | 9 |
| CYTOSKELETON | CDC42, CTNNB1, CTTN, LMNA, MAPK3, PTK2, PXN, YES1 | 8 |
| PLASMA MEMBRANE | AKT1, ARF6, CASP8, CAV1, CDC42, CDH1, CDH2, CTNNB1, CTNND1, EFNA5, EFNB2, EPHA1, EPHA2, ESR1, FAS, HRAS, IGF1R, ITGB3, MET, NEDD4, PDGFRB, PECAM1, PRKCD, PTK2, PTK2B, PXN, RAC1, RHOA, SRC, TIAM1, TJP1, YES1 | 32 |
| CYTOSOL | AKT1, ARF6, CASP8, CTNNB1, MAPK3, PRKCD, PTK2, RHOA, SRC, YES1, | 10 |
| MITOCHONDRION | GJA1, HSP90AA1, MAP2K1, MAPK3, SRC | 5 |
| GOLGI | CBL, CDH1, ESR1, HRAS, MAP2K1, MAPK3, NEDD4, RAC1, YES1 | 9 |
| ER | PRKCD, MAP2K1 | 2 |
| PEROXISOME | No level 5 protein | - - |
| ENDOSOME | ARF6, CAV1, CDH1, MAP2K1, MAPK3, PRKCD, RAC1, SRC | 8 |
| LYSOSOME | PDGFRB, PRKCD, SRC | 3 |
| NUCLEUS | AKT1, AR, ARRB1, CTNNB1, ESR1, GJA1, HRAS, HSP90AB1, ITGB3, LMNA, MAP2K1, MAPK8, NEDD4, PGR, PRKCD, PTK2, PTK2B, RAC1, STAT3 | 19 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




