Submitted:
28 February 2024
Posted:
29 February 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Basics Molecular Dynamics Simulations
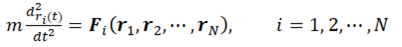
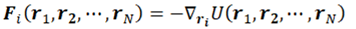
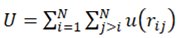
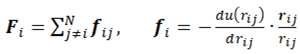
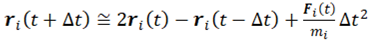
3. Interactions of Waters with Biopolymers
4. Interactions of Nanostructures with Biopolymers
4.1. Interaction of Carbon-Based Nanomaterials with Proteins
4.1.1. Fullerenes and Fullerene Derivatives
4.1.2. Carbon Nanotubes
4.1.3. Graphene and Graphene Derivatives
4.2. Interaction of Noble Metal-Based Nanomaterials with Proteins
4.2.1. Gold Nanomaterials
4.2.2. Silver Nanomaterials
4.2.3. Palladium Nanomaterials
4.3. Interactions of Hydroxyapatite Materials with Proteins
5. Basics of QSAR for Biopolymer Ligand Binding
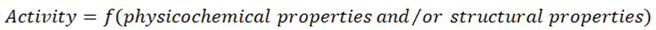
5.1. Human Serum Albumin Promiscuity
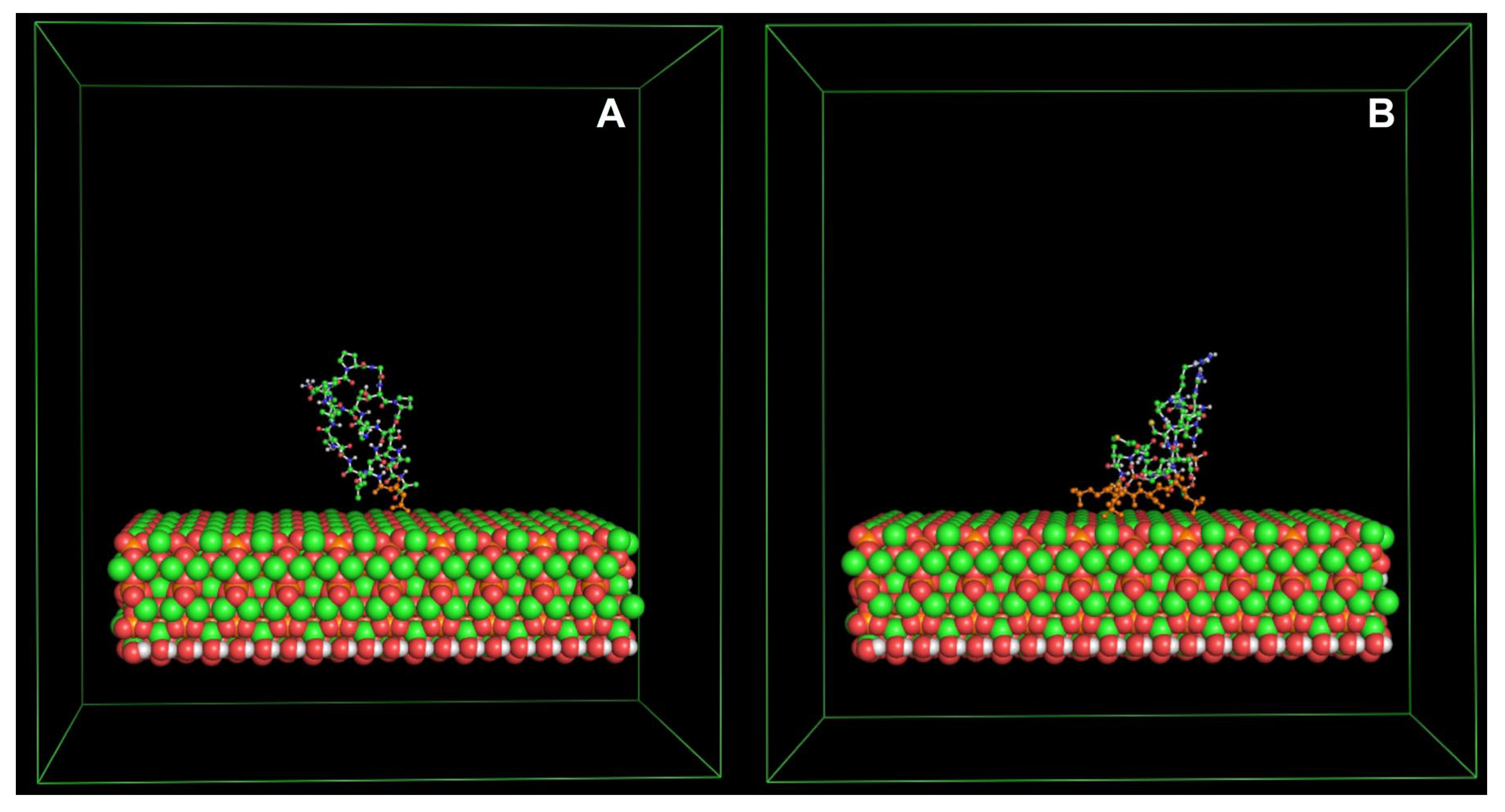
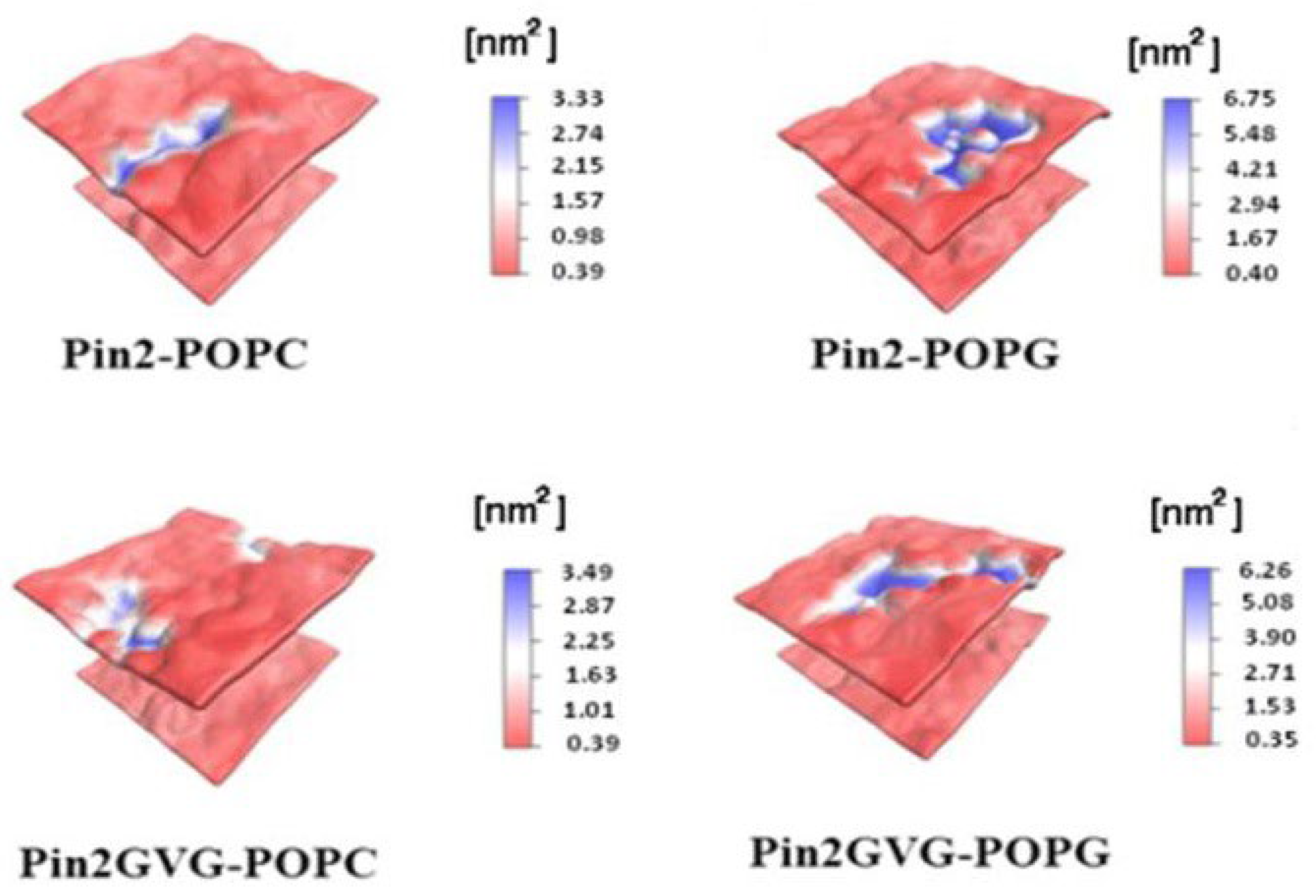
5.2. Antimicrobial Peptides
5.3. Human Vasopressin V1a Receptor
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bronzino, J. 1 - BIOMEDICAL ENGINEERING: A HISTORICAL PERSPECTIVE. In Introduction to Biomedical Engineering (Second Edition), Enderle, J.D., Blanchard, S.M., Bronzino, J.D., Eds.; Academic Press: Boston, 2005; pp. 1-29. [CrossRef]
- Stal, L.J. Biopolymer. In Encyclopedia of Astrobiology, Gargaud, M., Amils, R., Quintanilla, J.C., Cleaves, H.J., Irvine, W.M., Pinti, D.L., Viso, M., Eds.; Springer Berlin Heidelberg: Berlin, Heidelberg, 2011; pp. 199-200. [CrossRef]
- Stupp, S.I.; Braun, P.V. Molecular Manipulation of Microstructures: Biomaterials, Ceramics, and Semiconductors. Science 1997, 277, 1242–1248. [Google Scholar] [CrossRef]
- Ratner, B.D.; Hoffman, A.S.; Schoen, F.J.; Lemons, J.E. Introduction - Biomaterials Science: An Evolving, Multidisciplinary Endeavor. In Biomaterials Science (Third Edition), Ratner, B.D., Hoffman, A.S., Schoen, F.J., Lemons, J.E., Eds.; Academic Press: 2013; pp. xxv-xxxix.
- Kaplan, D.L. Introduction to Biopolymers from Renewable Resources. In Biopolymers from Renewable Resources, Kaplan, D.L., Ed.; Springer Berlin Heidelberg: Berlin, Heidelberg, 1998; pp. 1-29. [CrossRef]
- Grunwald, I.; Rischka, K.; Kast, S.M.; Scheibel, T.; Bargel, H. Mimicking biopolymers on a molecular scale: nano(bio)technology based on engineered proteins. Philosophical Transactions of the Royal Society A: Mathematical, Physical and Engineering Sciences 2009, 367, 1727–1747. [Google Scholar] [CrossRef] [PubMed]
- Costa, D.; Garrain, P.A.; Baaden, M. Understanding small biomolecule-biomaterial interactions: A review of fundamental theoretical and experimental approaches for biomolecule interactions with inorganic surfaces. Journal of Biomedical Materials Research Part A 2013, 101A, 1210–1222. [Google Scholar] [CrossRef] [PubMed]
- Vogler, E.A. Protein adsorption in three dimensions. Biomaterials 2012, 33, 1201–1237. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.-X.; Burton, S.D.; Xu, Y.S.; Buchko, G.W.; Shaw, W.J. The flexible structure of the K24S28 region of Leucine-Rich Amelogenin Protein (LRAP) bound to apatites as a function of surface type, calcium, mutation, and ionic strength. Front. Physiol. 2014, 5, 254–254. [Google Scholar] [CrossRef] [PubMed]
- Corni, S.; Kokh, D.B.; Ozboyaci, M.; Wade, R.C. Modeling and simulation of protein–surface interactions: achievements and challenges. Quarterly Reviews of Biophysics 2016, 49, e4. [Google Scholar] [CrossRef]
- Halliwell, C.M. Nanoanalytical measurement of protein orientation on conductive sensor surfaces. Analyst 2004, 129, 1166–1170. [Google Scholar] [CrossRef] [PubMed]
- Ostuni, E.; Chapman, R.G.; Holmlin, R.E.; Takayama, S.; Whitesides, G.M. A Survey of Structure−Property Relationships of Surfaces that Resist the Adsorption of Protein. Langmuir : the ACS journal of surfaces and colloids 2001, 17, 5605–5620. [Google Scholar] [CrossRef]
- Rabe, M.; Verdes, D.; Seeger, S. Understanding protein adsorption phenomena at solid surfaces. Advances in Colloid and Interface Science 2011, 162, 87–106. [Google Scholar] [CrossRef]
- Jamadagni, S.N.; Godawat, R.; Garde, S. How Surface Wettability Affects the Binding, Folding, and Dynamics of Hydrophobic Polymers at Interfaces. Langmuir : the ACS journal of surfaces and colloids 2009, 25, 13092–13099. [Google Scholar] [CrossRef]
- Chen, X.; Weber, I.; Harrison, R.W. Hydration Water and Bulk Water in Proteins Have Distinct Properties in Radial Distributions Calculated from 105 Atomic Resolution Crystal Structures. The Journal of Physical Chemistry B 2008, 112, 12073–12080. [Google Scholar] [CrossRef] [PubMed]
- Fogarty, A.C.; Laage, D. Water Dynamics in Protein Hydration Shells: The Molecular Origins of the Dynamical Perturbation. The Journal of Physical Chemistry B 2014, 118, 7715–7729. [Google Scholar] [CrossRef] [PubMed]
- Laage, D.; Elsaesser, T.; Hynes, J.T. Water Dynamics in the Hydration Shells of Biomolecules. Chemical Reviews 2017, 117, 10694–10725. [Google Scholar] [CrossRef] [PubMed]
- Alder, B.J.; Wainwright, T.E. Studies in Molecular Dynamics. I. General Method. The Journal of Chemical Physics 1959, 31, 459–466. [Google Scholar] [CrossRef]
- Bereau, T. Computational compound screening of biomolecules and soft materials by molecular simulations. Modelling and Simulation in Materials Science and Engineering 2021, 29, 023001. [Google Scholar] [CrossRef]
- Dhabal, D.; Jiang, Z.; Pallath, A.; Patel, A.J. Characterizing the Interplay between Polymer Solvation and Conformation. The Journal of Physical Chemistry B 2021, 125, 5434–5442. [Google Scholar] [CrossRef] [PubMed]
- Abkevich, V.I.; Gutin, A.M.; Shakhnovich, E.I. How the first biopolymers could have evolved. Proceedings of the National Academy of Sciences 1996, 93, 839–844. [Google Scholar] [CrossRef] [PubMed]
- van der Giessen, E.; Schultz, P.A.; Bertin, N.; Bulatov, V.V.; Cai, W.; Csányi, G.; Foiles, S.M.; Geers, M.G.D.; González, C.; Hütter, M.; et al. Roadmap on multiscale materials modeling. Modelling and Simulation in Materials Science and Engineering 2020, 28, 043001. [Google Scholar] [CrossRef]
- Salahub, D.R. Multiscale molecular modelling: from electronic structure to dynamics of nanosystems and beyond. Physical Chemistry Chemical Physics 2022, 24, 9051–9081. [Google Scholar] [CrossRef]
- Lazim, R.; Suh, D.; Choi, S. Advances in Molecular Dynamics Simulations and Enhanced Sampling Methods for the Study of Protein Systems. International Journal of Molecular Sciences 2020, 21, 6339. [Google Scholar] [CrossRef]
- Badar, M.S.; Shamsi, S.; Ahmed, J.; Alam, M.A. Molecular Dynamics Simulations: Concept, Methods, and Applications. In Transdisciplinarity, Rezaei, N., Ed.; Springer International Publishing: Cham, 2022; pp. 131-151. [CrossRef]
- Tuckerman, M.E. Statistical Mechanics: Theory and Molecular Simulation; Oxford University Press: 2023.
- Petrenko, R.; Meller, J. Molecular Dynamics. In Encyclopedia of Life Sciences. [CrossRef]
- Spreiter, Q.; Walter, M. Classical Molecular Dynamics Simulation with the Velocity Verlet Algorithm at Strong External Magnetic Fields. Journal of Computational Physics 1999, 152, 102–119. [Google Scholar] [CrossRef]
- Jones, D.; Allen, J.E.; Yang, Y.; Drew Bennett, W.F.; Gokhale, M.; Moshiri, N.; Rosing, T.S. Accelerators for Classical Molecular Dynamics Simulations of Biomolecules. Journal of Chemical Theory and Computation 2022, 18, 4047–4069. [Google Scholar] [CrossRef]
- Singh, N.; Li, W. Recent Advances in Coarse-Grained Models for Biomolecules and Their Applications. International Journal of Molecular Sciences 2019, 20, 3774. [Google Scholar] [CrossRef]
- Cerutti, D.S.; Freddolino, P.L.; Duke, R.E., Jr.; Case, D.A. Simulations of a Protein Crystal with a High Resolution X-ray Structure: Evaluation of Force Fields and Water Models. The Journal of Physical Chemistry B 2010, 114, 12811–12824. [Google Scholar] [CrossRef]
- Iwamoto, N. Water Effects in Polymers Through Molecular Dynamics. In Proceedings of the 2018 IEEE 68th Electronic Components and Technology Conference (ECTC), 29 May-1 June 2018, 2018; pp. 1659-1667. [CrossRef]
- Bellissent-Funel, M.-C.; Hassanali, A.; Havenith, M.; Henchman, R.; Pohl, P.; Sterpone, F.; van der Spoel, D.; Xu, Y.; Garcia, A.E. Water Determines the Structure and Dynamics of Proteins. Chemical Reviews 2016, 116, 7673–7697. [Google Scholar] [CrossRef]
- Carugo, O. Protein hydration: Investigation of globular protein crystal structures. International Journal of Biological Macromolecules 2017, 99, 160–165. [Google Scholar] [CrossRef] [PubMed]
- Scoppola, E.; Sodo, A.; McLain, Sylvia E.; Ricci, Maria A.; Bruni, F. Water-Peptide Site-Specific Interactions: A Structural Study on the Hydration of Glutathione. Biophysical Journal 2014, 106, 1701-1709. [CrossRef]
- Qin, Y.; Wang, L.; Zhong, D. Dynamics and mechanism of ultrafast water–protein interactions. Proceedings of the National Academy of Sciences 2016, 113, 8424–8429. [Google Scholar] [CrossRef] [PubMed]
- Conti Nibali, V.; Havenith, M. New Insights into the Role of Water in Biological Function: Studying Solvated Biomolecules Using Terahertz Absorption Spectroscopy in Conjunction with Molecular Dynamics Simulations. Journal of the American Chemical Society 2014, 136, 12800–12807. [Google Scholar] [CrossRef] [PubMed]
- Dielmann-Gessner, J.; Grossman, M.; Conti Nibali, V.; Born, B.; Solomonov, I.; Fields, G.B.; Havenith, M.; Sagi, I. Enzymatic turnover of macromolecules generates long-lasting protein–water-coupled motions beyond reaction steady state. Proceedings of the National Academy of Sciences 2014, 111, 17857–17862. [Google Scholar] [CrossRef] [PubMed]
- Raschke, T.M. Water structure and interactions with protein surfaces. Current Opinion in Structural Biology 2006, 16, 152–159. [Google Scholar] [CrossRef] [PubMed]
- Lopes, P.E.M.; Guvench, O.; MacKerell, A.D. Current Status of Protein Force Fields for Molecular Dynamics Simulations. In Molecular Modeling of Proteins, Kukol, A., Ed.; Springer New York: New York, NY, 2015; pp. 47-71. [CrossRef]
- Pattni, V.; Vasilevskaya, T.; Thiel, W.; Heyden, M. Distinct Protein Hydration Water Species Defined by Spatially Resolved Spectra of Intermolecular Vibrations. The Journal of Physical Chemistry B 2017, 121, 7431–7442. [Google Scholar] [CrossRef] [PubMed]
- Persson, F.; Söderhjelm, P.; Halle, B.; D., K.I.; W., K. The spatial range of protein hydration. The Journal of Chemical Physics 2018, 148, 215104. [CrossRef]
- Vlachakis, D.; Bencurova, E.; Papangelopoulos, N.; Kossida, S. Chapter Seven - Current State-of-the-Art Molecular Dynamics Methods and Applications. In Advances in Protein Chemistry and Structural Biology, Donev, R., Ed.; Academic Press: 2014; Volume 94, pp. 269-313. [CrossRef]
- Zhou, J.; Ranjith, P.G. Insights into interfacial behaviours of surfactant and polymer: A molecular dynamics simulation. Journal of Molecular Liquids 2022, 346, 117865. [Google Scholar] [CrossRef]
- Yin, Q.; Zhang, L.; Jiang, B.; Yin, Q.; Du, K. Effect of water in amorphous polyvinyl formal: insights from molecular dynamics simulation. Journal of Molecular Modeling 2015, 21, 2. [Google Scholar] [CrossRef]
- Zhou, R. Modeling of Nanotoxicity: Molecular Interactions of Nanomaterials with Bionanomachines; Springer International Publishing. [CrossRef]
- Liping, T.; Paul, T.; Wenjing, H. Surface Chemistry Influences Implant Biocompatibility. Current Topics in Medicinal Chemistry 2008, 8, 270–280. [Google Scholar] [CrossRef]
- Firkowska-Boden, I.; Zhang, X.; Jandt, K.D. Controlling Protein Adsorption through Nanostructured Polymeric Surfaces. Advanced Healthcare Materials 2018, 7, 1700995. [Google Scholar] [CrossRef]
- Zhou, R. Introduction. In Modeling of Nanotoxicity: Molecular Interactions of Nanomaterials with Bionanomachines; Springer International Publishing: Cham, 2015; pp. 1–15. [Google Scholar] [CrossRef]
- Qun, W.; Meng-hao, W.; Ke-feng, W.; Yaling, L.; Hong-ping, Z.; Xiong, L.; Xing-dong, Z. Computer simulation of biomolecule–biomaterial interactions at surfaces and interfaces. Biomedical Materials 2015, 10, 032001. [Google Scholar] [CrossRef]
- Debouck, C. The HIV-1 Protease as a Therapeutic Target for AIDS. AIDS Research and Human Retroviruses 1992, 8, 153–164. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.H.; DeCamp, D.L.; Sijbesma, R.P.; Srdanov, G.; Wudl, F.; Kenyon, G.L. Inhibition of the HIV-1 protease by fullerene derivatives: model building studies and experimental verification. Journal of the American Chemical Society 1993, 115, 6506–6509. [Google Scholar] [CrossRef]
- Chen, B.X.; Wilson, S.R.; Das, M.; Coughlin, D.J.; Erlanger, B.F. Antigenicity of fullerenes: Antibodies specific for fullerenes and their characteristics. Proceedings of the National Academy of Sciences 1998, 95, 10809–10813. [Google Scholar] [CrossRef]
- Braden, B.C.; Goldbaum, F.A.; Chen, B.-X.; Kirschner, A.N.; Wilson, S.R.; Erlanger, B.F. X-ray crystal structure of an anti-Buckminsterfullerene antibody Fab fragment: Biomolecular recognition of C60. Proceedings of the National Academy of Sciences 2000, 97, 12193–12197. [Google Scholar] [CrossRef]
- Noon, W.H.; Kong, Y.; Ma, J. Molecular dynamics analysis of a buckyball–antibody complex. Proceedings of the National Academy of Sciences 2002, 99, 6466–6470. [Google Scholar] [CrossRef]
- Kang, S.-g.; Zhou, G.; Yang, P.; Liu, Y.; Sun, B.; Huynh, T.; Meng, H.; Zhao, L.; Xing, G.; Chen, C.; et al. Molecular mechanism of pancreatic tumor metastasis inhibition by Gd@C82(OH)22 and its implication for de novo design of nanomedicine. Proceedings of the National Academy of Sciences 2012, 109, 15431–15436. [Google Scholar] [CrossRef]
- Kang, S.-g.; Huynh, T.; Zhou, R. Non-destructive Inhibition of Metallofullerenol Gd@C82(OH)22 on WW domain: Implication on Signal Transduction Pathway. Scientific Reports 2012, 2, 957. [Google Scholar] [CrossRef]
- Kang, S.-g.; Huynh, T.; Zhou, R. Metallofullerenol Gd@C82(OH)22 distracts the proline-rich-motif from putative binding on the SH3 domain. Nanoscale 2013, 5, 2703–2712. [Google Scholar] [CrossRef]
- Prato, M.; Kostarelos, K.; Bianco, A. Functionalized Carbon Nanotubes in Drug Design and Discovery. Accounts of Chemical Research 2008, 41, 60–68. [Google Scholar] [CrossRef]
- Nepal, D.; Geckeler, K.E. Proteins and Carbon Nanotubes: Close Encounter in Water. Small 2007, 3, 1259–1265. [Google Scholar] [CrossRef] [PubMed]
- Chiu, C.-c.; Maher, M.C.; Dieckmann, G.R.; Nielsen, S.O. Molecular Dynamics Study of a Carbon Nanotube Binding Reversible Cyclic Peptide. ACS Nano 2010, 4, 2539–2546. [Google Scholar] [CrossRef] [PubMed]
- Salvador-Morales, C.; Townsend, P.; Flahaut, E.; Vénien-Bryan, C.; Vlandas, A.; Green, M.L.H.; Sim, R.B. Binding of pulmonary surfactant proteins to carbon nanotubes; potential for damage to lung immune defense mechanisms. Carbon 2007, 45, 607–617. [Google Scholar] [CrossRef]
- Botas, C.; Poulain, F.; Akiyama, J.; Brown, C.; Allen, L.; Goerke, J.; Clements, J.; Carlson, E.; Gillespie, A.M.; Epstein, C.; et al. Altered surfactant homeostasis and alveolar type II cell morphology in mice lacking surfactant protein D. Proceedings of the National Academy of Sciences 1998, 95, 11869–11874. [Google Scholar] [CrossRef] [PubMed]
- Ge, C.; Du, J.; Zhao, L.; Wang, L.; Liu, Y.; Li, D.; Yang, Y.; Zhou, R.; Zhao, Y.; Chai, Z.; et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proceedings of the National Academy of Sciences 2011, 108, 16968–16973. [Google Scholar] [CrossRef] [PubMed]
- Zuo, G.; Huang, Q.; Wei, G.; Zhou, R.; Fang, H. Plugging into Proteins: Poisoning Protein Function by a Hydrophobic Nanoparticle. ACS Nano 2010, 4, 7508–7514. [Google Scholar] [CrossRef]
- Zuo, G.; Gu, W.; Fang, H.; Zhou, R. Carbon Nanotube Wins the Competitive Binding over Proline-Rich Motif Ligand on SH3 Domain. The Journal of Physical Chemistry C 2011, 115, 12322–12328. [Google Scholar] [CrossRef]
- Nel, A.E.; Mädler, L.; Velegol, D.; Xia, T.; Hoek, E.M.V.; Somasundaran, P.; Klaessig, F.; Castranova, V.; Thompson, M. Understanding biophysicochemical interactions at the nano–bio interface. Nature Materials 2009, 8, 543–557. [Google Scholar] [CrossRef]
- Zuo, G.; Zhou, X.; Huang, Q.; Fang, H.; Zhou, R. Adsorption of Villin Headpiece onto Graphene, Carbon Nanotube, and C60: Effect of Contacting Surface Curvatures on Binding Affinity. The Journal of Physical Chemistry C 2011, 115, 23323–23328. [Google Scholar] [CrossRef]
- Rosi, N.L.; Mirkin, C.A. Nanostructures in Biodiagnostics. Chemical Reviews 2005, 105, 1547–1562. [Google Scholar] [CrossRef] [PubMed]
- Colangelo, E.; Comenge, J.; Paramelle, D.; Volk, M.; Chen, Q.; Lévy, R. Characterizing Self-Assembled Monolayers on Gold Nanoparticles. Bioconjugate Chemistry 2017, 28, 11–22. [Google Scholar] [CrossRef]
- Liu, R.; Wang, Y.; Yuan, Q.; An, D.; Li, J.; Gao, X. The Au clusters induce tumor cell apoptosis via specifically targeting thioredoxin reductase 1 (TrxR1) and suppressing its activity. Chemical Communications 2014, 50, 10687–10690. [Google Scholar] [CrossRef]
- An, D.; Su, J.; Weber, J.K.; Gao, X.; Zhou, R.; Li, J. A Peptide-Coated Gold Nanocluster Exhibits Unique Behavior in Protein Activity Inhibition. Journal of the American Chemical Society 2015, 137, 8412–8418. [Google Scholar] [CrossRef]
- Ding, F.; Radic, S.; Chen, R.; Chen, P.; Geitner, N.K.; Brown, J.M.; Ke, P.C. Direct observation of a single nanoparticle–ubiquitin corona formation. Nanoscale 2013, 5, 9162–9169. [Google Scholar] [CrossRef] [PubMed]
- Fang, G.; Li, W.; Shen, X.; Perez-Aguilar, J.M.; Chong, Y.; Gao, X.; Chai, Z.; Chen, C.; Ge, C.; Zhou, R. Differential Pd-nanocrystal facets demonstrate distinct antibacterial activity against Gram-positive and Gram-negative bacteria. Nature Communications 2018, 9, 129. [Google Scholar] [CrossRef]
- Villarreal-Ramirez, E.; Eliezer, D.; Garduño-Juarez, R.; Gericke, A.; Perez-Aguilar, J.M.; Boskey, A. Phosphorylation regulates the secondary structure and function of dentin phosphoprotein peptides. Bone 2017, 95, 65–75. [Google Scholar] [CrossRef]
- Boskey, A.L.; Villarreal-Ramirez, E. Intrinsically disordered proteins and biomineralization. Matrix Biology 2016, 52-54, 43–59. [Google Scholar] [CrossRef] [PubMed]
- Pancsa, R.; Schad, E.; Tantos, A.; Tompa, P. Emergent functions of proteins in non-stoichiometric supramolecular assemblies. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 2019, 1867, 970–979. [Google Scholar] [CrossRef] [PubMed]
- Hunter, G.K.; O'Young, J.; Grohe, B.; Karttunen, M.; Goldberg, H.A. The Flexible Polyelectrolyte Hypothesis of Protein−Biomineral Interaction. Langmuir : the ACS journal of surfaces and colloids 2010, 26, 18639–18646. [Google Scholar] [CrossRef]
- Kim, J. Systematic approach to characterize the dynamics of protein adsorption on the surface of biomaterials using proteomics. Colloids and Surfaces B: Biointerfaces 2020, 188, 110756. [Google Scholar] [CrossRef] [PubMed]
- Shaw, W.J.; Ferris, K. Structure, Orientation, and Dynamics of the C-Terminal Hexapeptide of LRAP Determined Using Solid-State NMR. The Journal of Physical Chemistry B 2008, 112, 16975–16981. [Google Scholar] [CrossRef]
- Chadha, R.; Bhandari, S. Drug–excipient compatibility screening—Role of thermoanalytical and spectroscopic techniques. Journal of Pharmaceutical and Biomedical Analysis 2014, 87, 82–97. [Google Scholar] [CrossRef]
- Zheng, H.; Handing, K.B.; Zimmerman, M.D.; Shabalin, I.G.; Almo, S.C.; Minor, W. X-ray crystallography over the past decade for novel drug discovery – where are we heading next? Expert Opinion on Drug Discovery 2015, 10, 975–989. [Google Scholar] [CrossRef]
- Mahapatra, M.K.; Karuppasamy, M. Chapter 2 - Fundamental considerations in drug design. In Computer Aided Drug Design (CADD): From Ligand-Based Methods to Structure-Based Approaches, Rudrapal, M., Egbuna, C., Eds.; Elsevier: 2022; pp. 17-55. [CrossRef]
- McGee, T.D.; Edwards, J.; Roitberg, A.E. Preliminary Molecular Dynamic Simulations of the Estrogen Receptor Alpha Ligand Binding Domain from Antagonist to Apo. International Journal of Environmental Research and Public Health 2008, 5, 111–114. [Google Scholar] [CrossRef]
- Zhao, H.; Caflisch, A. Molecular dynamics in drug design. European Journal of Medicinal Chemistry 2015, 91, 4–14. [Google Scholar] [CrossRef]
- Malathi, K.; Ramaiah, S. Bioinformatics approaches for new drug discovery: a review. Biotechnology and Genetic Engineering Reviews 2018, 34, 243–260. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-Galicia, G.; Garduño-Juárez, R.; Correa-Basurto, J.; Deeb, O. Exploring QSARs for inhibitory effect of a set of heterocyclic thrombin inhibitors by multilinear regression refined by artificial neural network and molecular docking simulations. Journal of Enzyme Inhibition and Medicinal Chemistry 2012, 27, 174–186. [Google Scholar] [CrossRef]
- Macalino, S.J.Y.; Gosu, V.; Hong, S.; Choi, S. Role of computer-aided drug design in modern drug discovery. Archives of Pharmacal Research 2015, 38, 1686–1701. [Google Scholar] [CrossRef]
- Ishikawa, T.; Hirano, H.; Saito, H.; Sano, K.; Ikegami, Y.; Yamaotsu, N.; Hirono, S. Quantitative Structure-Activity Relationship (QSAR) Analysis to Predict Drug-Drug Interactions of ABC Transporter ABCG2. Mini-Reviews in Medicinal Chemistry 2012, 12, 505–514. [Google Scholar] [CrossRef]
- Martínez-Archundia, M.; Colín-Astudillo, B.; Moreno-Vargas, L.M.; Ramírez-Galicia, G.; Garduño-Juárez, R.; Deeb, O.; Contreras-Romo, M.C.; Quintanar-Stephano, A.; Abarca-Rojano, E.; Correa-Basurto, J. Ligand recognition properties of the vasopressin V2 receptor studied under QSAR and molecular modeling strategies. Chemical Biology & Drug Design 2017, 90, 840–853. [Google Scholar] [CrossRef]
- Deeb, O.; Martínez-Pachecho, H.; Ramírez-Galicia, G.; Garduño-Juárez, R. Application of Docking Methodologies in QSAR-Based Studies. Pharmaceutical Sciences: Breakthroughs in Research and Practice 2017, 850-876. Pharmaceutical Sciences. [CrossRef]
- Vilar, S.; Costanzi, S. Predicting the Biological Activities Through QSAR Analysis and Docking-Based Scoring. In Membrane Protein Structure and Dynamics: Methods and Protocols, Vaidehi, N., Klein-Seetharaman, J., Eds.; Humana Press: Totowa, NJ, 2012; pp. 271-284. [CrossRef]
- Cherkasov, A.; Muratov, E.N.; Fourches, D.; Varnek, A.; Baskin, I.I.; Cronin, M.; Dearden, J.; Gramatica, P.; Martin, Y.C.; Todeschini, R.; et al. QSAR Modeling: Where Have You Been? Where Are You Going To? Journal of Medicinal Chemistry 2014, 57, 4977–5010. [Google Scholar] [CrossRef] [PubMed]
- Leach, A.R. Molecular modelling : principles and applications, 2nd ed. ed.; Harlow etc. Prentice Hall: Harlow [etc., 2001.
- Rasulev, B. Recent Developments in 3D QSAR and Molecular Docking Studies of Organic and Nanostructures. In Handbook of Computational Chemistry, Leszczynski, J., Kaczmarek-Kedziera, A., Puzyn, T., G. Papadopoulos, M., Reis, H., K. Shukla, M., Eds.; Springer International Publishing: Cham, 2017; pp. 2133-2161. [CrossRef]
- Veerasamy, R.; Rajak, H.; Jain, A.; Sivadasan, S.; Christapher, P.V.; Agrawal, R. Validation of QSAR Models - Strategies and Importance. Int J Drug Design and Discov 2011, 2, 511–519. [Google Scholar]
- Uversky, V.N. Chapter One - Protein intrinsic disorder and structure-function continuum. In Progress in Molecular Biology and Translational Science, Uversky, V.N., Ed.; Academic Press: 2019; Volume 166, pp. 1-17. [CrossRef]
- Nam, K.; Wolf-Watz, M. Protein dynamics: The future is bright and complicated! Structural Dynamics 2023, 10, 014301. [Google Scholar] [CrossRef]
- Putri, R.M.; Zulfikri, H.; Fredy, J.W.; Juan, A.; Tananchayakul, P.; Cornelissen, J.J.L.M.; Koay, M.S.T.; Filippi, C.; Katsonis, N. Photoprogramming Allostery in Human Serum Albumin. Bioconjugate Chemistry 2018, 29, 2215–2224. [Google Scholar] [CrossRef]
- Deeb, O.; Rosales-Hernández, M.C.; Gómez-Castro, C.; Garduño-Juárez, R.; Correa-Basurto, J. Exploration of human serum albumin binding sites by docking and molecular dynamics flexible ligand–protein interactions. Biopolymers 2010, 93, 161–170. [Google Scholar] [CrossRef]
- Lei, J.; Sun, L.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q. The antimicrobial peptides and their potential clinical applications. American journal of translational research 2019, 11, 3919–3931. [Google Scholar] [PubMed]
- Velasco-Bolom, J.-L.; Corzo, G.; Garduño-Juárez, R. Molecular dynamics simulation of the membrane binding and disruption mechanisms by antimicrobial scorpion venom-derived peptides. Journal of Biomolecular Structure and Dynamics 2018, 36, 2070–2084. [Google Scholar] [CrossRef] [PubMed]
- Velasco-Bolom, J.-L.; Corzo, G.; Garduño-Juárez, R. Folding profiles of antimicrobial scorpion venom-derived peptides on hydrophobic surfaces: a molecular dynamics study. Journal of Biomolecular Structure and Dynamics 2020, 38, 2928–2938. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Romo, M.C.; Martínez-Archundia, M.; Deeb, O.; Ślusarz, M.J.; Ramírez-Salinas, G.; Garduño-Juárez, R.; Quintanar-Stephano, A.; Ramírez-Galicia, G.; Correa-Basurto, J. Exploring the Ligand Recognition Properties of the Human Vasopressin V1a Receptor Using QSAR and Molecular Modeling Studies. Chemical Biology & Drug Design 2014, 83, 207–223. [Google Scholar] [CrossRef]
- Yoshimura, M.; Conway-Campbell, B.; Ueta, Y. Arginine vasopressin: Direct and indirect action on metabolism. Peptides 2021, 142, 170555. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



