1. Introduction
The process of aneuploidy that occurs due to chromosome mis-segregation during meiosis is a major cause of birth defects, infertility, and spontaneous miscarriages [
1]. Most constitutional aneuploidies cause embryonic lethality, the most notable exception in humans being trisomy 21, also known as Down syndrome (DS) [
2].
Monosomies are relatively common events in early embryonic development, but survival beyond fetal period is exceptional [
3]. The embryonic and fetal survival of 45,X concepts although extremely low (0.01%) can result in live births, which can be explained by the fact that it is a monosomy of sex chromosomes.
Turner syndrome is due to a partial or total loss of the second sexual chromosome, resulting in the development of highly variable clinical features, affecting 1 in 2,500 female live births [
4,
5]. It is a unique chromosome aneuploidy observed in newborns [
4,
5,
6]. Monosomy of X chromosome is the cytogenetic hallmark for TS [
4], but a wide variety of other anomalies of X or Y chromosome have been found, including chromosomal mosaicism, with whole or part of a sex chromosome [
4,
5,
6]. At least 15-25% of TS patients have mosaicism [
4,
5].
It has been suggested that 45,X lineages have a prolonged cell cycle that can affect the differentiation of tissues and organs [
7]. These preliminary data indicate that there could be an association between a prolonged cell cycle and the phenotypic features in TS [
3].
TS phenotype is a result not only of the genomic imbalance from deleted genes, especially those that escape X inactivation or are in the pseudoautosomal region, but also from epigenetic influences, as X-chromosome and autosomal DNA-methylation and gene expression profiles from TS cells differ from normal female 46,XX cells [
8,
9]. Nonetheless, chromosome instability (CIN) could also play a role in TS complex pathogenesis and clinical variability [
8].
Chromosome instability (CIN) is characterized by a high rate of loss and gain of whole chromosome segments per cell cycle resulting in cell-to-cell variability [10, 11]. It is a hallmark in tumorigenesis and is also common during early human embryogenesis [
11]. CIN can be associated with monosomy-trisomy of a whole chromosome and other genomic and chromosomal variations [
3].
A higher incidence of sister chromatid exchange (SCE) can be associated with CIN, as is seen in some cancers and congenital syndromes [
12,
13,
14,
15,
16]. Therefore, SCE has been used as a sensitive indicator of spontaneous CIN [
12,
17,
18,
19,
20]. Previous research has reported high SCE incidences in patients with Bloom syndrome, Werner syndrome, and various types of neoplasia [
12,
13,
20].
SCE represents the interchange of DNA between replication products at apparently homologous loci, and at low levels constitutes a normal occurrence. Although the molecular basis for SCE formation is not completely understood, it presumably involves DNA damage and repair [
21,
22].
On the other hand, there are few reports comparing cell-cycle kinetics and SCE frequencies in mosaic TS patients until the present date. According to Melaragno et al [
18], mosaic patients are useful for SCE evaluation because the different cell lineages grow under the influence of the same exogenous and endogenous factors. Previously, Bortolai & Melaragno [
23] analyzed by SCE one 45,X/46,XX patient in a small study series. The “mosaic” study design approach not only removes the confounding effects of interindividual differences due to total genetic make-up, but also controls for the effects of environmental influences, since both cell lines in individuals with mosaicism share identical exposure histories [
24].
The main aim of the present cross-sectional study was to analyze the proliferation dynamics and SCE frequencies among 45,X and 46,XN lineages from a moderately large cohort of mosaic TS participants.
2. Materials and Methods
A total of 17 mosaic Turner syndrome (TS) participants were included in the present study from a cohort of circa 124 TS females diagnosed by the Cytogenetic Laboratory of Institute of Pediatrics and Childcare Martagão Gesteira (IPPMG) and followed by the Endocrinology Department of Clementino Fraga Filho University Hospital (HUCFF), Federal University of Rio de Janeiro (UFRJ), Brazil, from 1989 to 2022. The study was approved by the Ethical Review Board of IPPMG, UFRJ (Ethical Review Board protocol n. 2.768.383). All participants or their parents gave their written consent.
Peripheral blood cultures were performed using standard protocols modified from Moorhead et al. [
25]. Peripheral blood from each participant was added to 5 mL RPMI 1640 medium (Gibco, Grand Island, NY, USA) supplemented with 20% fetal bovine serum (Cultilab, Campinas, SP, Brazil) and phytohemagglutinin P (Cultilab).
Cultures were incubated for 48, 72 and 96 hours. Cultures were harvested by standard procedures after one hour in MAS (Mitotic Arresting Solution, Genial Genetics, Chester, UK), treated with a hypotonic solution (0.075 M KCl) and fixed in 3:1 methanol: acetic acid. Cytological preparation and slide mounting were performed according to routine methods.
For bromodeoxyuridine (BrdU) incorporation, 50 µg/mL BrdU (Sigma-Aldrich, Barueri, SP, Brazil) was added in the beginning of the cultures, which were incubated for 72 hours. Differentiation staining was performed using a modified standard technique from previous studies [
26,
27].
Proliferation index (PI) after BrdU labeling was estimated by scoring fifty metaphases from each cell lineage. A total of 100 metaphase cells (50 metaphases from 45,X and 50 from 46,XN cell lineages) from each participant were scored for the determination of the PI, calculated according to the following equation: PI = (M1+ 2M2 + 3M3)/N, where M1, M2, and M3 represent the number of metaphases undergoing first, second and third division and N is the total number of cells scored.
The frequencies of 45,X and 46,XN lineages, and evaluation of in vitro selection were performed in 48, 72 and 96-hour cultures, counting a total of 100 metaphases (50 cells from 45,X lineage and 50 cells from 46,XN lineage).
To appraise chromosomal instability, sister chromatid exchange (SCE) frequencies among 45,X and 46,XN lineages were compared. Twenty well-spread second-division metaphases from each cell lineage were analyzed and the amount of SCE was computed for each one cell.
The frequencies of SCE were compared between 45,X and 46,XN lineages in an intraindividual analysis, comparing the lineages of the same individual participant, and in an interindividual analysis, considering the pool of monosomic and disomic cells in the group of participants.
Fluorescence in situ hybridization (FISH) was performed using the following commercial probes specific for chromosomes X and Y, according to manufacturers’ instructions: DXZ1/DYZ1 (centromeric regions, Xp11.1-q11.1 and Yq12; Vysis/Abbott, Abbott Park, IL, USA or Zytovision, Bremerhaven, Germany), whole chromosome painting (wcp) for X and Y (Cytocell, Milton, Cambridge, UK), SHOX (Xp22.33 and Yp11.3; Cytocell, Milton, Cambridge, UK), XIST (Xq13.2; Vysis/Abbott, Abbott Park, IL, USA or Cytocell, Milton, Cambridge, UK), DYZ3 (centromeric region, Yp11.1-q11.1; Zytovision, Bremerhaven, Germany), ENXY (pericentromeric regions, Xq11.1 and Yq11.22.1; Lexel, Buenos Aires, Argentina), and SRY (Yp11.31; Vysis/Abbott, Abbott Park, IL, USA).
For statistical analysis, the Tukey’s and Šídák multiple comparisons tests were applied to compare interindividual (group) comparisons of the frequencies of lineages 45,X and 46,XN, when considering different cultivation times of the same lineage, or both lineages of each participant at different cultivation times, respectively. The χ2 test or Fisher's exact test were used to compare the frequencies of the 45,X and 46,XN lineages after different cultivation times in the same participant (intraindividual analysis). The two-tailed unpaired t-test was used for the intraindividual comparisons of the SCE frequencies of the 45,X and 46,XN lineages, while the χ2 test or Fisher's exact test was used for the PI. One-way ANOVA with Dunnett's post hoc multiple comparison test was used for the comparisons of SCE frequencies between 45,X and several 46,XN karyotype groups. The means for PI and for SCE frequencies of lineages 45,X and 46,XN were compared by two-tailed paired-t test. Pearson’s correlation coefficient (r) and linear regression were employed to measure the association of PI and of the SCE frequencies between 45,X and 46,XN lineages. Statistical analyses were performed using Prism (GraphPad Software, Boston, MA, USA) and Stata (StataCorp, College Station, TX, USA) statistics software.
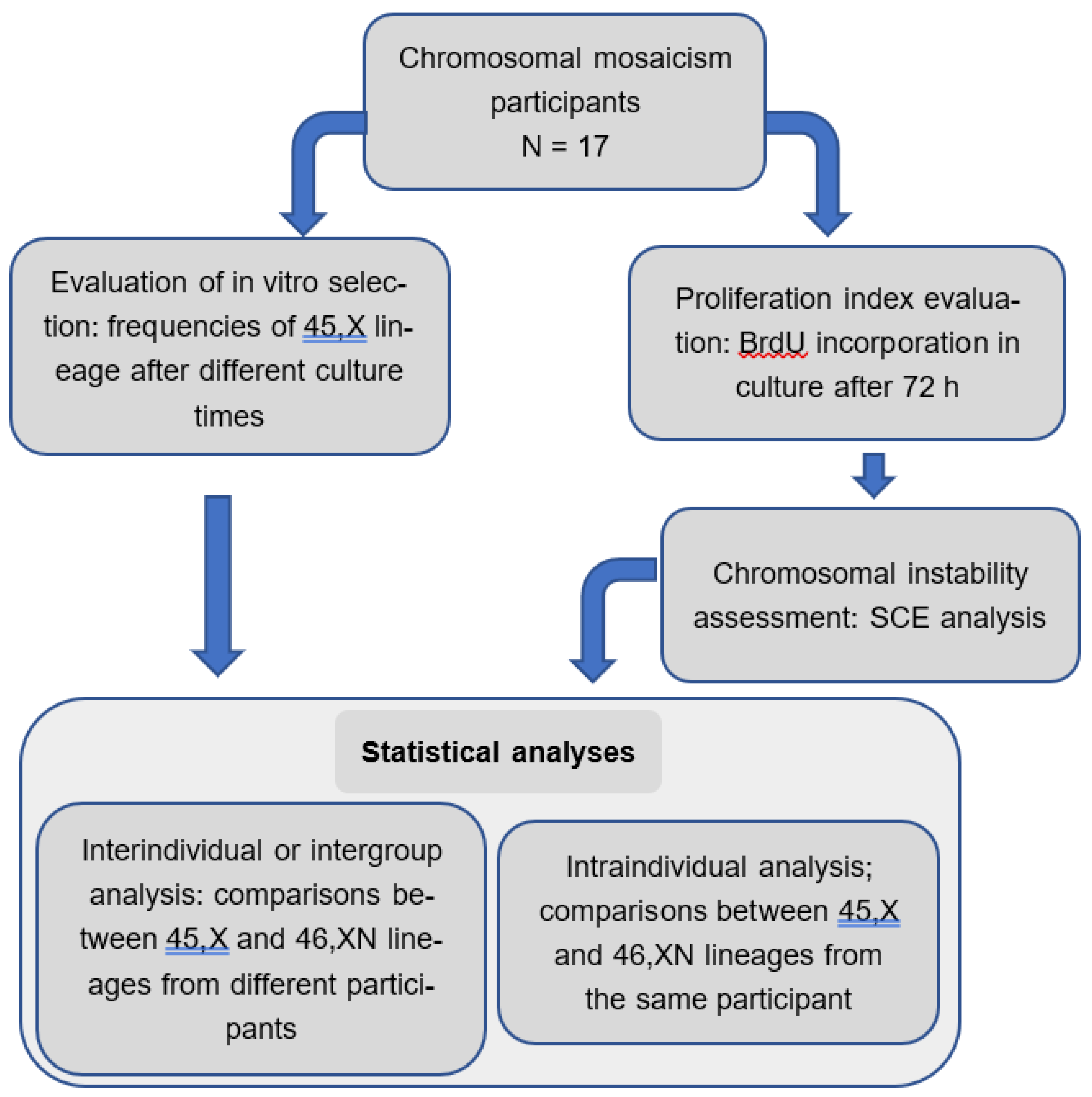
The workflow below summarizes the methods employed in this study (
Figure 1):
3. Results
The participants had a diagnosis of TS verified by karyotype and were selected from the medical records of the Laboratory of Genetics of IPPMG and the Pediatric Endocrinology Service spanning over more than 30 years of healthcare. From this cohort of circa 124 participants, 27 were included by convenience sampling and agreed to participate in a larger study constituting the PhD thesis of one of the authors (MBG). A subset of 17 participants with mosaic TS comprises the sample of the present study. Their age ranged from 1.7 to 42 years (
Table 1).
Karyotype was confirmed using standard cytogenetic technique and GTG-banding for chromosomal analysis of participants 1, 2, 9, and 15, which showed besides 45,X, a typical 46,XX (1, 2, 15) or 46,XY (9) lineages. Anomalous chromosomes found in 13 participants were characterized by FISH (
Figures S1 to S13).
FISH analysis showed that small supernumerary marker chromosomes (sSMC) were derived from the X chromosome in participants 5-8, 10-13, and 16 (
Figures S3-S6, S7-S10, and S12). In two participants (4 and 14,
Figures S2 and S11), sSMC were derived from the Y chromosome. A ring X chromosome was present in one participant (3,
Figure S1) and another participant (17) presented an isochromosome of the short arm of Y chromosome
Figure S13).
One copy of XIST (Xq13.2) was present in sSMC of six participants (5, 6, 10, 11, 13 and 16;
Figures S3, S4, S7, S8, S10, and S12) and two copies in two participants, 3 within a ring chromosome (
Figure S1), and 12 within an sSMC (
Figure S9), both derived from X chromosome. In one participant (7,
Figure S5), the XIST gene was not detected by hybridization on the sSMC.
The sSMC of participant 4 was positive for Y whole chromosome painting, and presented two copies of SRY and SHOX, suggesting that it was an isochromosome for the short arm of the Y chromosome (
Figure S2). On the other hand, the sSMC of participant 14 presented two copies of ENY, suggesting that it was a dicentric sSMC derived from the Y chromosome (
Figure S11). Participant 17 presented an isochromosome for the short arm of the Y chromosome with two copies of ENY, SRY, SHOX and DYZ3 (
Figure S13).
3.1. In Vitro Selection
The occurrence of in vitro cell selection was verified by the amount of cells of 45,X and 46,XN lineages in cultures after 48, 72 and 96 h of incubation. No significant effect of incubation time was found in the proportion of the two lineages for most participants, with two exceptions:
participant 3: there was a decrease in the number of metaphases in the 45,X lineage and a simultaneous increase in the number of metaphases of lineage 46,X+r(X ) over the period (p = 0.0274).
participant 6: there was a decrease in the number of metaphases in the 45,X lineage and an increase in the number of metaphases in the 46,X+mar(X) lineage when comparing 48h and 72h incubation (p = 0.0234).
The frequencies of 45,X and 46,XN lineages in the cultures incubated for 48, 72 and 96 h is shown in
Figure S14.
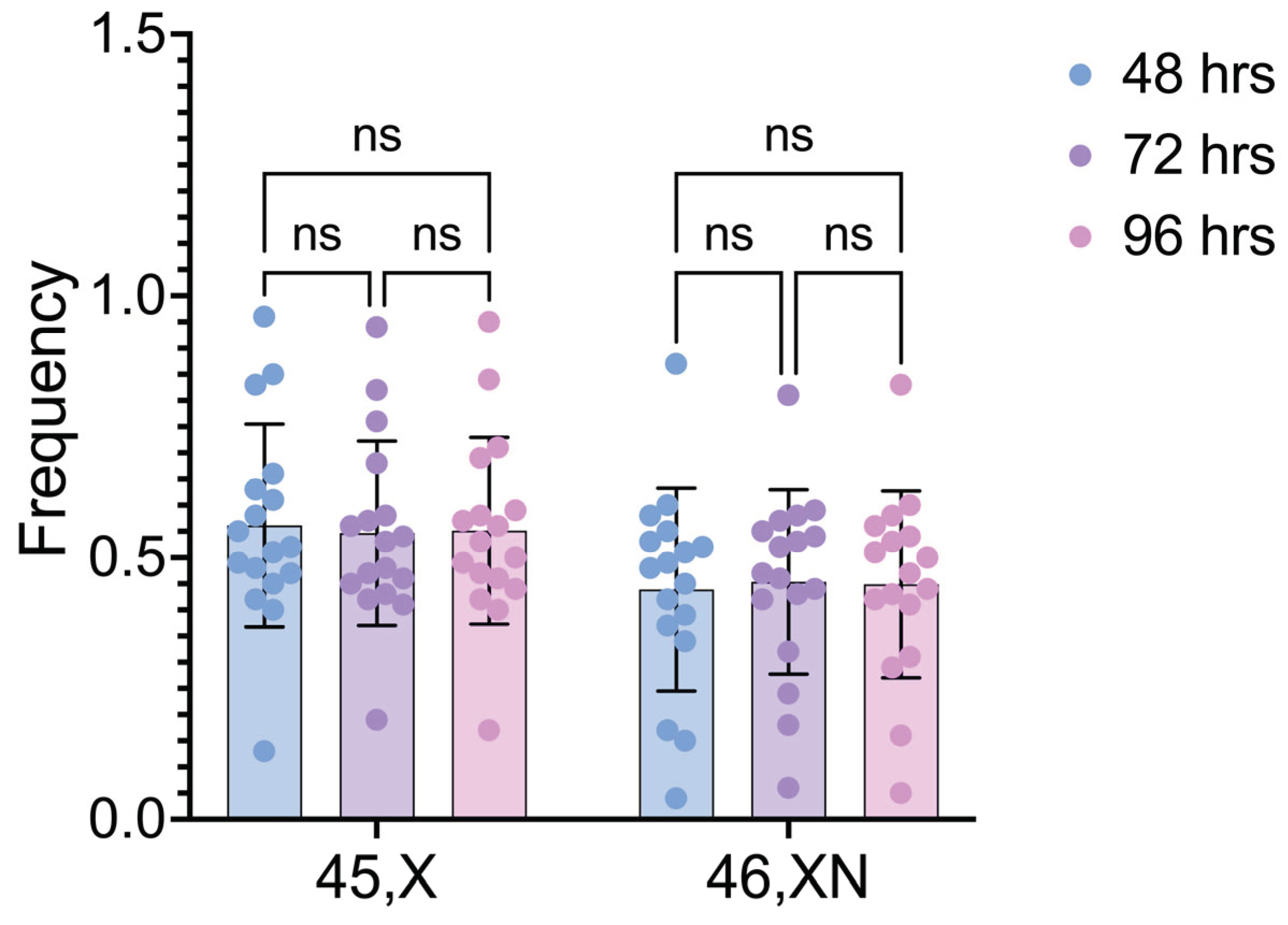
The interindividual comparisons of the frequencies of 45,X and 46,XN cell lineages after different cultivation times revealed no significant difference within both 45,X and 46,XN lineages when 48h vs. 72h, 48h vs. 96h, and 72h vs. 96h were compared (p values= 0.6625, 0.8490 and 0.8803, respectively) (
Figure 2).
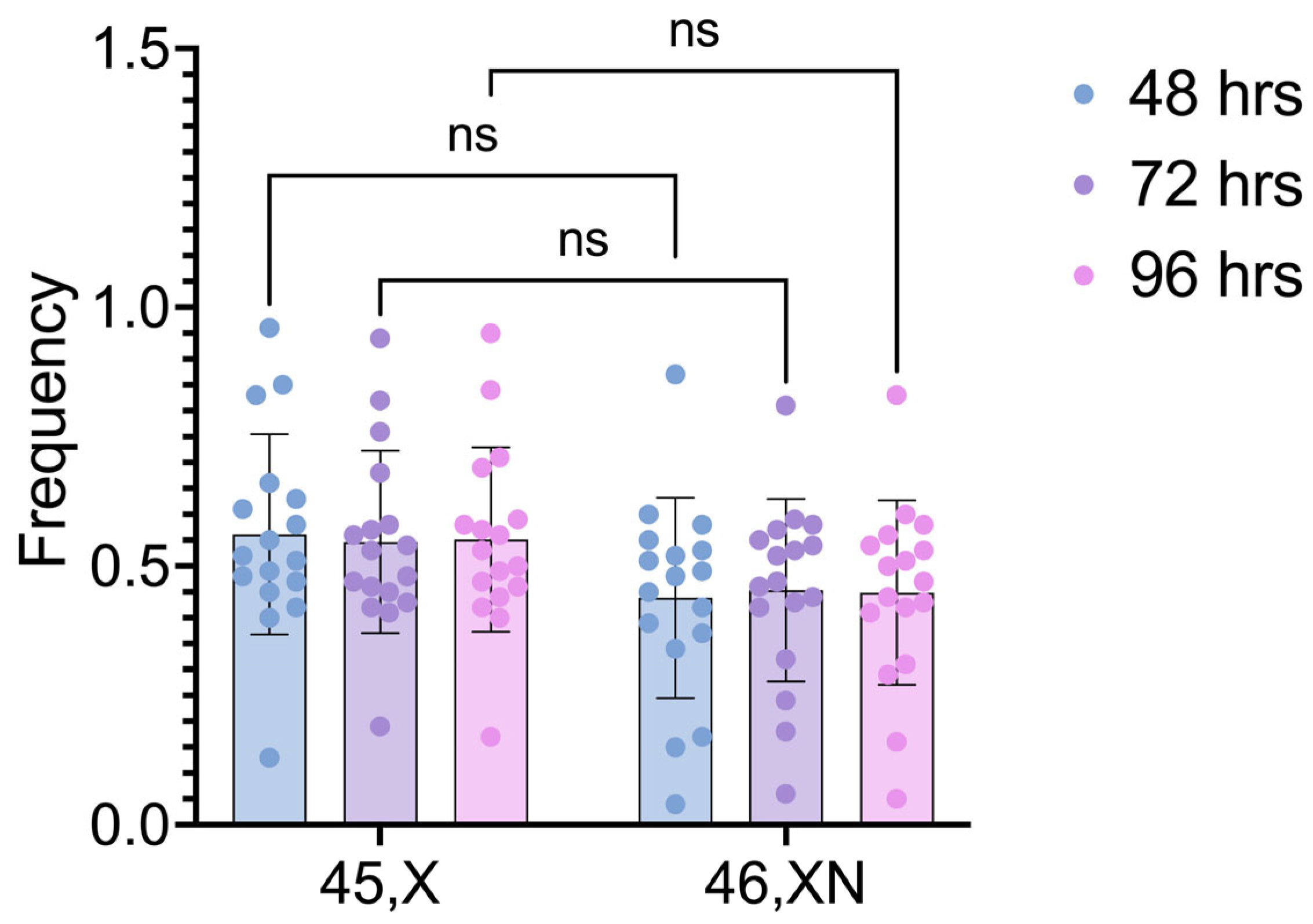
The comparisons between the two cell lineages after different cultivation times also did not detect any statistically significant differences (p values = 0.2082, 0.3511, and 0.2810, for 48h, 72h, and 96h, respectively) (
Figure 3).
3.2. Proliferation Index (PI)
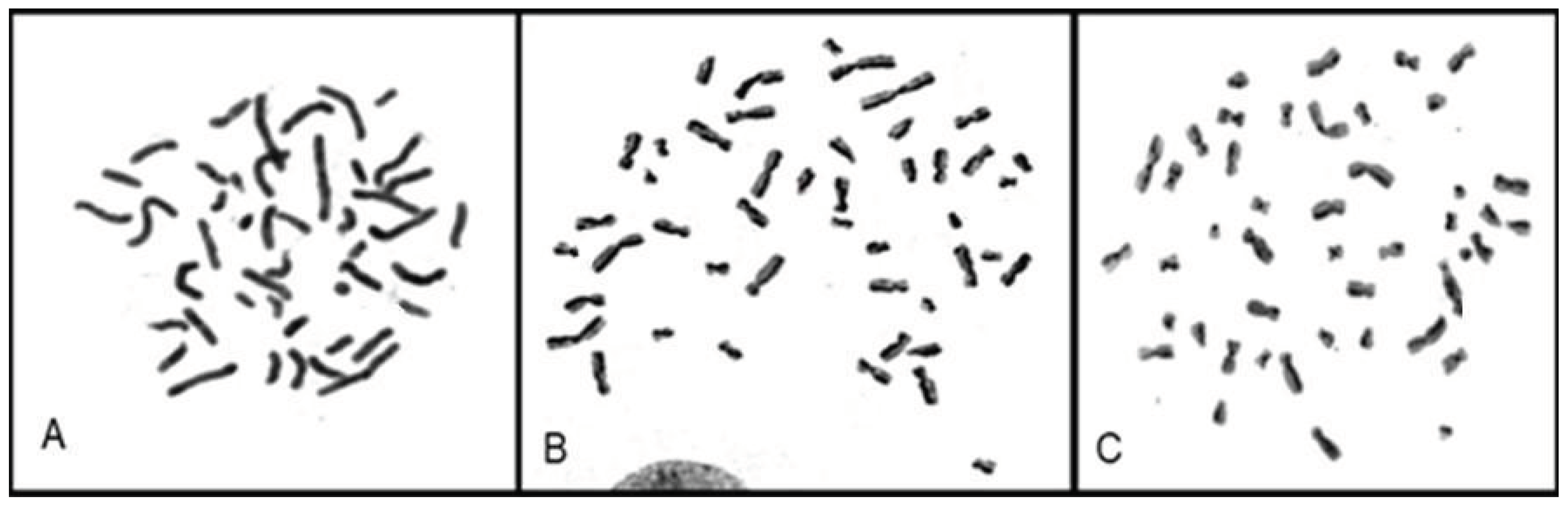
The proliferation index (PI) of each cell lineage was calculated individually by evaluating the number of cells in first, second and third cell division, according to the BrdU incorporation pattern (
Figure 4).
The intraindividual comparisons of PI between 45,X and 46,XN lineages showed no significant differences (p >0.5) in participants with Turner syndrome and chromosomal mosaicism included in this study. However, the intergroup comparison of the PI means showed a significant difference between 45,X and 46,XN lineages (p = 0.0338) (
Table 2).
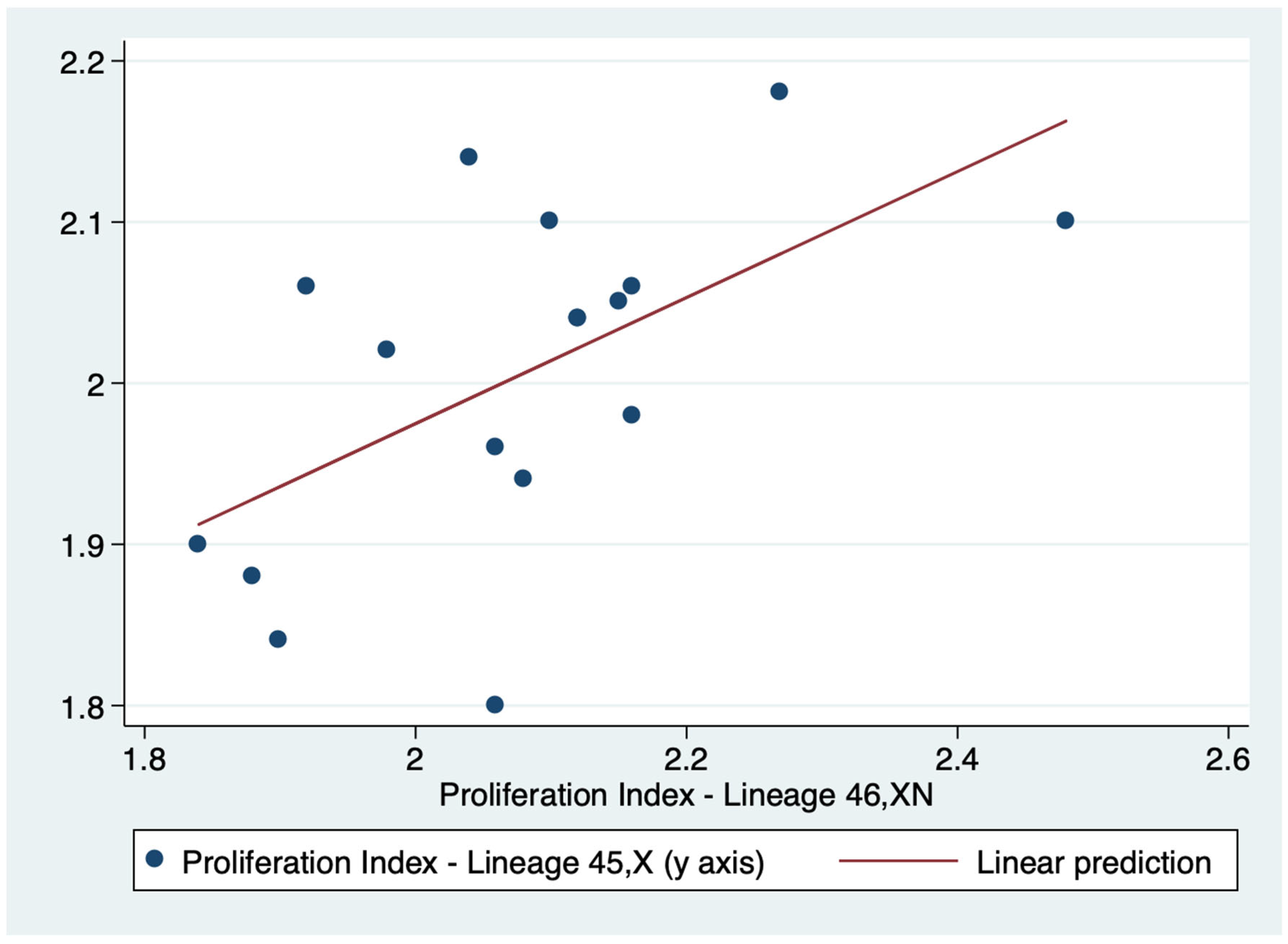
There was a positive correlation between the PI of both lineages (r=0.5716). Consequently the PI of lineages 45,X and 46,XN tended to move in the same direction: as the PI value of one lineage increased or decreased so did the value of the other lineage (
Table 3). Indeed, this correlation was confirmed by linear regression (p = 0.017) (
Figure 5).
3.3. Sister Chromatid Exchange (SCE) Frequency
Chromosomal instability was assessed by calculating the SCE frequencies in 20 second division metaphases of each cell lineage in 72-hour cultures after BrdU labeling. The pictures of second division metaphases of the 17 participants, highlighting SCEs, along with first and third division metaphases, which are employed in PI calculation, are depicted in
supplementary Figures S15-S17.
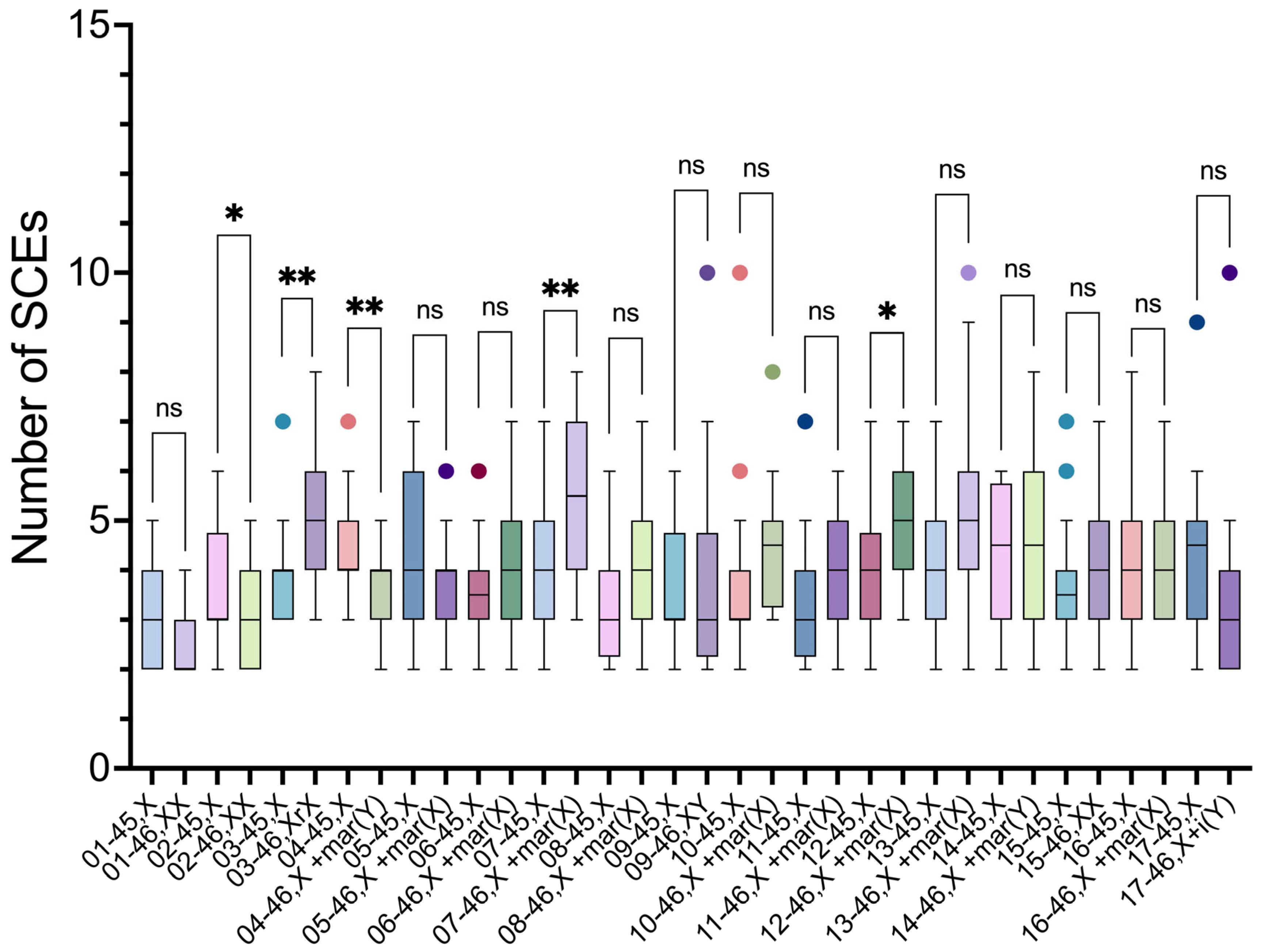
Intraindividual comparisons of SCE frequencies using the unpaired t-test showed a significant difference between the two cell lineages in five participants (2, 3, 4, 7 and 12) (
Table 4).
In 3 participants (3, 7 and 12), the SCE frequencies were higher in 46,XN lineage, compared to 45,X lineage. These 46,XN lineages presented anomalous chromosomes: two participants had an sSMC derived from the X chromosome (46,X+mar(X)), and the third a ring chromosome derived from X chromosome (46,Xr(X)).
In two participants, the SCE frequencies were higher in 45,X lineage. In participant 2, it was higher in relation to 46,XX lineage, and in participant 4, it was higher in relation to the lineage with a sSMC derived from Y chromosome (46,X+mar(Y)) (
Figure 6).
Regarding interindividual comparisons of SCE frequencies, the analysis showed no significant differences in the means of the 45,X and 46,XN lineages (p = 0.2871) (
Table 5).
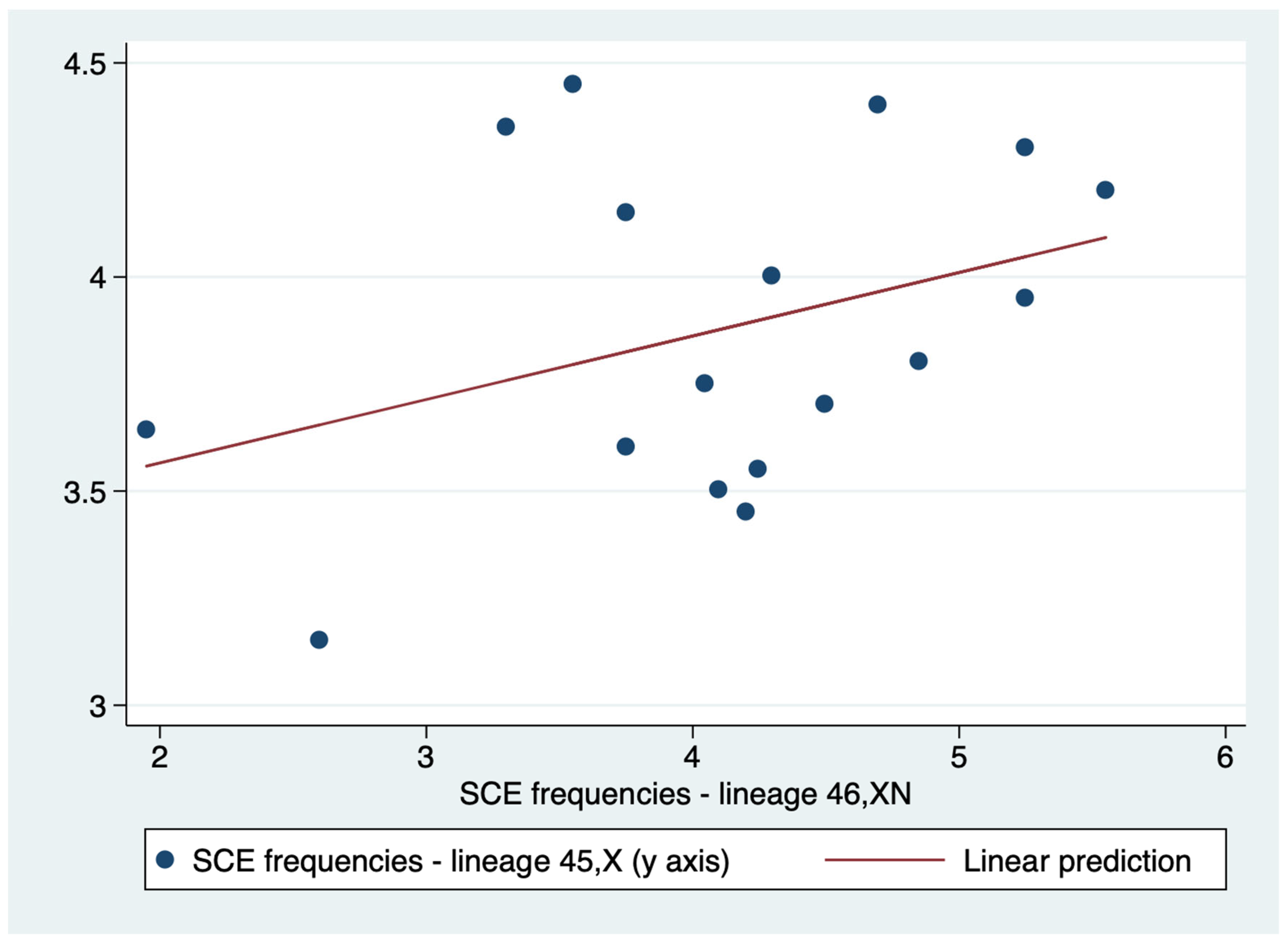
A weak correlation (r=0.3615) was demonstrated between 45,X and 46,XN lineages SCE frequencies. Therefore, unlike PI, the positive association between the SCE frequencies of the two lineages was weak, and it is not possible to affirm with certainty that higher values of the SCE frequency of the 45,X lineage are associated with correspondingly higher values of the SCE frequency of the 46,XN lineage (
Table 6). Additionally, linear regression indicated that the relationship of SCE frequencies of the two lineages are not statistically significant (p = 0.154) (
Figure 7).
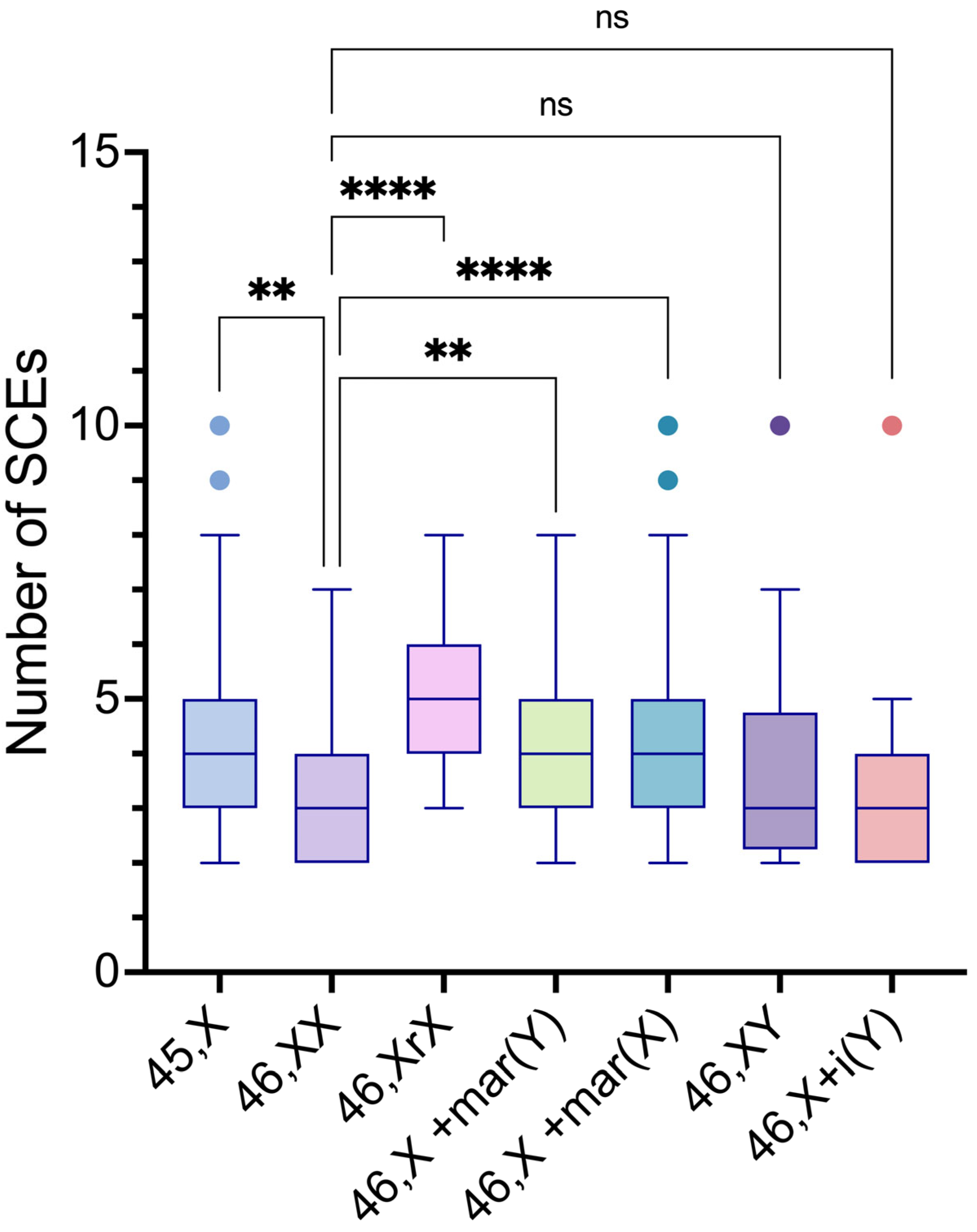
The intergroup comparisons of SCE frequencies among the 45,X and the distinct 46,XN lineages showed statistically significant differences between the 45,X lineage and 46,XX, 46,X,r(X), 46,X,+mar(X) lineages (p values= 0.0155, 0.0007 and <0.0001, respectively); between the 46,XX lineage and 46,Xr(X), 46,X,+mar(Y), 46,X,+mar(X) lineages (p values= <0.0001, 0.0328 and <0.0001, respectively); between the 46,X,r(X) lineage and 46,XY, 46,X,i(Yp) lineages (p values= <0.0198 and 0.0004. respectively); and the 46,X,+mar(X) lineage and 46,X,i(Yp) lineage (p value= 0.0060) (
Table 7 and
Figure 8).
4. Discussion
The present study evaluated in vitro selection, PI and SCE frequencies of seventeen mosaic TS patients. So far in the literature, only few studies have evaluated cell proliferation of 45,X cells and of cells having other sex or autosomal chromosome anomalies [
12,
13,
20,
24,
28,
29,
30,
31,
32]. TS individuals who carry structurally abnormal chromosomes are a unique group, and they have provided opportunities to appraise the cellular consequences of chromosomal imbalance.
Evidence of in vitro selection was observed in only two of 17 participants. Participant 3, displaying a ring X chromosome, and 6, displaying an sSMC derived from X chromosome, both of them with a decrease in monosomic cells after increased cultivation times, suggesting that there was a selective disadvantage in vitro of 45,X lineage in these participants. However, interindividual comparisons of the frequencies of 45,X and 46,XN cell lineages revealed no significant differences in the proportions of 45,X and 46,XN lineages after cultivation over time. Summarizing, results from most of the participants in this study did not reveal differences in 45,X and 46,XX cell lineages proportions in culture over time. However, differences were found eventually in some (12%) mosaic TS individuals.
Bortolai and Melaragno [
23] also did not observe statistically significant differences between the proliferation of aneuploid and normal cells of a mosaic 45,X/46,XX TS patient after 48, 72 and 96 h cultivation times. We did not detect in vitro selective disadvantage of 45,X lineage in three 45,X/46XX and one 45,X/46XY participants in this study (
Figure S14). It is possible that cultures for periods longer than 96 h are required to detect the in vitro selective disadvantage of 45,X cell lineages.
Although the intraindividual comparisons showed no significant differences between 45,X and 46,XN lineages, the intergroup comparison of the PI means of the 17 participants evaluated in this study showed a significant difference between both lineages (p = 0.0338). PI of 46,XN lineage was slightly higher than the 45,X lineage, suggesting 45,X lineages had a proliferative disadvantage in relation to 46XN lineages. A previous study with skin fibroblast cultures comparing differences in cell cycle of aneuploid cell lines (45,X) to cell lines with structural anomalies of the X chromosome, and normal diploid cell lines, 46,XX and 46,XY, showed the cell cycle of 45,X cell lines was significantly longer compared to euploid controls [
3]. A possible explanation for this phenomenon would be a longer S phase, resulting in a prolonged inter-mitotic period in 45,X cells that could lead to a decrease in their quantity during organogenesis, which could contribute to important phenotypical features in TS [
3,
7].
This differential PI between 45,X and 46,XN lineages could explain the increase in the proportion of diploid cells with age in TS individuals [
33], contrasting with the increase of aneuploid cells due to a loss of X chromosome in unaffected women with age [
34,
35].
SCE are considered to be cytological manifestations of homologous recombination. They occur naturally as events associated with normal DNA replication, with estimates being 3-4 exchanges per-cell per replication cycle [
21,
36]. In Bloom's syndrome, a very high SCE frequency is found, well above that of cells from unaffected individuals or individuals with any other genetic disorder. SCE can also reflect genomic instability and may serve as a preclinical biomarker for early detection of genomic instability [
12,
20,
22]. Nonetheless, in this study, the amount of SCE in the participants was not remarkable, and the average was similar to the baseline.
SCE frequencies were significantly different between 45,X and 46,XN cell lineages in five participants. In three participants (3, 7 and 12), the SCE frequencies were higher in the lineage with a structurally anomalous sex chromosome compared to 45,X lineage. In participant 2 the SCE frequency was higher in the 45,X lineage compared to the 46,XX lineage, and in participant 4 the SCE frequency of 45,X lineage was higher than that of a lineage with sSMC derived from Y chromosome. However, interindividual comparisons of SCE frequencies revealed no significant differences in the means of 45,X and 46,XN lineages (p value = 0.2871).
Iqbal, Martin & Simpson demonstrated significantly increased mean SCE frequencies in all fibroblast cell lines with abnormalities of the X chromosome: 45,X; 46,X,del(X)(q13); 46,X,del(X)(p11); and 46,X,i(Xq) compared to normal 46,XX cell lines [
37]. In our study, more than 70% of the participants presented a lineage with anomalous chromosomes and only four were mosaics with a normal female (46,XX) or male (46,XY) cell lineage. Consequently, most of the intraindividual comparisons were between cell lineages with a certain amount of genomic instability. However, we employed primary lymphocyte cultures and not established fibroblast cell lines, therefore our results may reflect more closely what is happening in mosaic TS tissues and organs.
The frequency of the Y chromosome in TS is variable [16, 38] usually being found in 3 to 12% of individuals [38-41]. The presence of Y sequences in our study was 23.5% (4/17). Probably, this higher frequency in our study is due to the significant number of individuals with marker chromosomes (sSMC). From the 4 participants with intact or anomalous chromosomes derived from the Y chromosome, only one showed significant differences in SCE frequencies between the two lineages. Surprisingly, 45,X lineage from this participant showed higher SCE frequency than the lineage with a sSMC derived from Y chromosome cells.
As far as we know, the number of articles that have studied genomic instability in aneuploidies is limited. Yet, all data suggest that cells from Down syndrome, Edward syndrome, Turner syndrome and Patau syndrome patients may be karyotypically less stable than cells from unaffected individuals [
2]. Our cross-sectional study with a sample of mosaic TS selected by convenience, with a significant number of participants with sSMC, showed some evidence that the 45,X cell lineage may have a selective disadvantage over time, although there was a significant variation concerning the differences in SCE frequencies between 45,X and 46,XN lineages. Therefore, the presence of chromosomal structural abnormalities, especially the presence of sSMC, could have been responsible for the variation of 46,XN lineages instability observed in our study.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org, Figure S1: FISH results for participant 3; partial metaphases showing the presence of a ring chromosome derived from chromosome X. Figure S2: FISH results for participant 4; partial metaphases showing sSMC derived from Y chromosome. Figure S3: FISH results for participant 5; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S4: FISH results for participant 5; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S5: FISH results for participant 7; interphase cells showing an sSMC derived from chromosome X. Figure S6: FISH results for participant 8; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S7: FISH results for participant 10; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S8: FISH results for participant 11; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S9: FISH results for participant 12; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S10: FISH results for participant 13; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S11: FISH results for participant 14; partial metaphases showing the presence of an sSMC derived from Y chromosome. Figure S12: FISH results for participant 16; partial metaphases showing the presence of an sSMC derived from X chromosome. Figure S13: FISH results for participant 17; partial metaphases showing isochromosome for short arm of Y chromosome. Figure S14: Frequencies of 45,X and 46,XN cell lineages after different incubation times. Figure S15: Micrographs of metaphases in second division of participants 1 to 6 displaying sister chromatid exchanges. Figure S16: Micrographs of metaphases in second division of participants 7 to 12 displaying sister chromatid exchanges. Figure S17: Micrographs of metaphases in second division of participants 13 to 17 displaying sister chromatid exchanges.
Author Contributions
MBG performed experiments, interpreted the results and drafted the initial manuscript. EVN performed the statistical analysis, interpreted the results, drafted the initial manuscript, and participated in its design and coordination. DRNG drafted and edited the initial manuscript. ISP, MMG, KF, NCKS and MGR accomplished participant clinical diagnosis and evaluation. LSB executed technical support. AFF performed G-banding analysis. MCMR reviewed all laboratory results, participated in its design and coordination. MGR participated in its design and coordination. All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by Rio de Janeiro State Research Foundation – FAPERJ (Jovem Cientista do Nosso Estado RJ 2008/ MGR; E-26/103.116/2008 FAPERJ) and National Council for Scientific and Technological Development – CNPq.
Institutional Review Board Statement
This study was performed in accordance with the Guidelines and Standards for Research in Human Beings, established by Brazilian National Health Council and with the guidelines of the Declaration of Helsinki. Ethics approval was obtained from the Research Ethics Review Board of Institute of Childcare and Pediatrics Martagão Gesteira (IPPMG), Federal University of Rio de Janeiro (UFRJ) (protocol code: 2.768.383; date of approval: 12 July 2018).
Informed Consent Statement
Informed consent was obtained from all participants, or their parents involved in the study.
Data Availability Statement
The data presented in this study are available by the corresponding author upon request.
Acknowledgments
We acknowledge Dr. Marcelo Gerardin Poirot Land and Dr. Elaine Sobral da Costa for their support in laboratory maintenance and infrastructure improvements; Sandra Alves Peixoto Pellegrini in memoriam for technical support; and Dr. Karinne Condack Mafort Branco for clinical support.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Radonova L, Svobodova T, Anger M. Regulation of the cell cycle in early mammalian embryos and its clinical implications. The International Journal of Developmental Biology. 2019 Feb 6;63(3-4–5):113–22. [CrossRef]
- Potapova TA, Zhu J, Li R. Aneuploidy and chromosomal instability: a vicious cycle driving cellular evolution and cancer genome chaos. Cancer Metastasis Rev. 2013 Dec;32(3–4):377–89. [CrossRef]
- Álvarez-Nava F, Soto‐Quintana M. The Hypothesis of the Prolonged Cell Cycle in Turner Syndrome. J Dev Biol. 2022 May 11;10(2):16. 11 May. [CrossRef]
- Oliveira RMR de, Verreschi IT do N, Lipay MVN, Eça LP, Guedes AD, Bianco B. Y chromosome in Turner syndrome: review of the literature. Sao Paulo Med J. 2009 Nov;127(6):373–8. [CrossRef]
- Gravholt CH, Viuff MH, Brun S, Stochholm K, Andersen NH. Turner syndrome: mechanisms and management. Nat Rev Endocrinol. 2019 Oct;15(10):601–14. [CrossRef]
- Marzuki NS, Anggaratri HW, Suciati LP, Ambarwati DD, Paramayuda C, Kartapradja H, et al. Diversity of sex chromosome abnormalities in a cohort of 95 Indonesian patients with monosomy X. Molecular Cytogenetics. 2011 Oct 12;4(1):23. [CrossRef]
- Simpson JL, Lebeau MM. Gonadal and statural determinants on the X chromosome and their relationship to in vitro studies showing prolonged cell cycles in 45,X; 46,X,del(X)(p11); 46,X,del(X)(q13); and 46,X,del(X)(q22) fibroblasts. Am J Obstet Gynecol. 1981 Dec 15;141(8):930-40. [CrossRef]
- Álvarez-Nava F, Lanes R. Epigenetics in Turner syndrome. Clin Epigenetics. 2018;10:45. [CrossRef]
- Trolle C, Nielsen MM, Skakkebæk A, Lamy P, Vang S, Hedegaard J, et al. Widespread DNA hypomethylation and differential gene expression in Turner syndrome. Sci Rep. 2016 Sep 30;6:34220. [CrossRef]
- Thompson SL, Bakhoum SF, Compton DA. Mechanisms of chromosomal instability. Curr Biol. 2010 Mar 23;20(6):R285-295. [CrossRef]
- Vanneste E, Voet T, Le Caignec C, Ampe M, Konings P, Melotte C, et al. Chromosome instability is common in human cleavage-stage embryos. Nat Med. 2009 May;15(5):577–83. [CrossRef]
- Salawu A, Wright K, Al-Kathiri A, Wyld L, Reed M, Sisley K. Sister Chromatid Exchange and Genomic Instability in Soft Tissue Sarcomas: Potential Implications for Response to DNA-Damaging Treatments. Sarcoma. 2018;2018:3082526. [CrossRef]
- Cobanoglu H, Cayir A. Assessment of the genotoxic potential of tetrachlorvinphos insecticide by cytokinesis-block micronucleus and sister chromatid exchange assays. Hum Exp Toxicol. 2021 Dec;40(12_suppl):S158–63. [CrossRef]
- Montenegro MM, Quaio CR, Palmeira P, Gasparini Y, Rangel-Santos A, Damasceno J, et al. Gene expression profile suggesting immunological dysregulation in two Brazilian Bloom’s syndrome cases. Mol Genet Genomic Med. 2020 Feb 19;8(4):e1133. [CrossRef]
- Bianco B, Nunes Lipay MV, Guedes AD, Verreschi ITN. Clinical implications of the detection of Y-chromosome mosaicism in Turner’s syndrome: report of 3 cases. Fertility and Sterility. 2008 Oct 1;90(4):1197.e17-1197.e20. [CrossRef]
- Bispo AVS, Burégio-Frota P, Oliveira dos Santos L, Leal GF, Duarte AR, Araújo J, et al. Y chromosome in Turner syndrome: detection of hidden mosaicism and the report of a rare X;Y translocation case. Reprod Fertil Dev. 2014 Oct;26(8):1176–82. [CrossRef]
- Barale R, Chelotti L, Davini T, Del Ry S, Andreassi MG, Ballardin M, et al. Sister chromatid exchange and micronucleus frequency in human lymphocytes of 1,650 subjects in an Italian population: II. Contribution of sex, age, and lifestyle. Environ Mol Mutagen. 1998;31(3):228–42. [CrossRef]
- Melaragno MI, Ribeiro MC, Smith M de A, Andrade JA, Erwenne CM. Sister chromatid exchange frequency in a retinoblastoma mosaic patient with del(13). Cancer Genet Cytogenet. 1988 Jun;32(2):177–81. [CrossRef]
- Oztas S, Gullulu G, Tatar A, Astam N, Akyol I, Karakuzu A, et al. Chromosome and sister chromatid exchange studies in Behcet’s patients. J Dermatol. 2006 Jun;33(6):406–10. [CrossRef]
- Xu J, Liu P, Meng X, Bai J, Fu S, Guan R, et al. Association between sister chromatid exchange and double minute chromosomes in human tumor cells. Molecular Cytogenetics. 2015 Nov 19;8(1):91. [CrossRef]
- Wilson DM, Thompson LH. Molecular mechanisms of sister-chromatid exchange. Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis. 2007 Mar 1;616(1):11–23. [CrossRef]
- Latt SA, Schreck RR. Sister chromatid exchange analysis. Am J Hum Genet. 1980 May;32(3):297–313.
- Bortolai A, Melaragno MI. Cell-cycle kinetics of cell lines from patients with chromosomal mosaicism. Ann Genet. 2001 Apr-Jun;44(2):93-7. [CrossRef]
- Rafferty K, Archer KJ, Turner K, Brown R, Jackson-Cook C. Trisomy 21-associated increases in chromosomal instability are unmasked by comparing isogenic trisomic/disomic leukocytes from people with mosaic Down syndrome. PLoS One. 2021;16(7):e0254806. [CrossRef]
- Moorhead PS, Nowell PC, Mellman WJ, Battips DM, Hungerford DA. Chromosome preparations of leukocytes cultured from human peripheral blood. Experimental Cell Research. 1960 Sep 1;20(3):613–6. [CrossRef]
- Perry P, Wolff S. New Giemsa method for the differential staining of sister chromatids. Nature. 1974 Sep;251(5471):156–8. [CrossRef]
- Korenberg JR, Freedlender EF. Giemsa technique for the detection of sister chromatid exchanges. Chromosoma. 1974;48(4):355–60. [CrossRef]
- Butler MG. Sister Chromatid Exchanges in A Male With A Y/Y Translocation. Trans Nebr Acad Sci Affil Soc. 1981;9:9–11.
- Heidemann A, Schmalenberger B, Zankl H. Sister chromatid exchange and lymphocyte proliferation in a Down syndrome mosaic. Clin Genet. 1983 Feb;23(2):139–42. [CrossRef]
- Santovito A, Gendusa C, Cervella P. Evaluation of baseline frequency of sister chromatid exchanges in an Italian population according to age, sex, smoking habits, and gene polymorphisms. Am J Hum Biol. 2017 Nov;29(6). [CrossRef]
- Landi S, Frenzilli G, Milillo PC, Cocchi L, Sbrana I, Scapoli C, et al. Spontaneous sister chromatid exchange and chromosome aberration frequency in humans: the familial effect. Mutat Res. 1999 Aug 18;444(2):337–45. [CrossRef]
- Laurie CC, Laurie CA, Rice K, Doheny KF, Zelnick LR, McHugh CP, et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat Genet. 2012 May 6;44(6):642–50. [CrossRef]
- Denes AM, Landin-Wilhelmsen K, Wettergren Y, Bryman I, Hanson C. The proportion of diploid 46,XX cells increases with time in women with Turner syndrome--a 10-year follow-up study. Genet Test Mol Biomarkers. 2015 Feb;19(2):82–7. [CrossRef]
- Machiela MJ, Zhou W, Karlins E, Sampson JN, Freedman ND, Yang Q, et al. Female chromosome X mosaicism is age-related and preferentially affects the inactivated X chromosome. Nat Commun. 2016 Jun 13;7(1):11843. [CrossRef]
- Catalán J, Falck GCM, Norppa H. The X Chromosome Frequently Lags Behind in Female Lymphocyte Anaphase. Am J Hum Genet. 2000 Feb;66(2):687–91. [CrossRef]
- Payne M, Hickson ID. Genomic instability and cancer: lessons from analysis of Bloom’s syndrome. Biochem Soc Trans. 2009 Jun;37(Pt 3):553–9. [CrossRef]
- Iqbal MA, Martin AO, Simpson JL. Increased sister chromatid exchanges in human cell lines characterized by monosomy X or structural abnormalities of the X chromosome. Hum Genet. 1984;68(3):205–8. [CrossRef]
- Gravholt CH, Viuff M, Just J, Sandahl K, Brun S, van der Velden J, et al. The Changing Face of Turner Syndrome. Endocr Rev. 2023 Jan 12;44(1):33–69. [CrossRef]
- Thunström S, Landin-Wilhelmsen K, Bryman I, Hanson C. Side differences in the degree of mosaicism of the buccal mucosa in Turner syndrome. Mol Genet Genomic Med. 2019 Aug 29;7(10):e00938. [CrossRef]
- Liehr T, Mrasek K, Hinreiner S, Reich D, Ewers E, Bartels I, et al. Small Supernumerary Marker Chromosomes (sSMC) in Patients with a 45,X/46,X,+mar Karyotype – 17 New Cases and a Review of the Literature. Sexual Development. 2008 Jan 18;1(6):353–62. [CrossRef]
- Wang H, Wang T, Yang N, He Y, Chen L, Hong L, et al. The clinical analysis of small supernumerary marker chromosomes in 17 children with mos 45,X/46,X,+mar karyotype. Oncol Lett. 2017 Jun;13(6):4385–9. [CrossRef]
Figure 1.
Flowchart showing the methods and statistical analyses used in this study.
Figure 1.
Flowchart showing the methods and statistical analyses used in this study.
Figure 2.
Frequencies of 45,X and 46,XN cell lineages after different culture times. Comparisons done within each cell lineage. ns: non-significant.
Figure 2.
Frequencies of 45,X and 46,XN cell lineages after different culture times. Comparisons done within each cell lineage. ns: non-significant.
Figure 3.
Frequencies of 45,X and 46,XN cell lineages after different culture times. Comparisons done between cell lineages. ns: non-significant.
Figure 3.
Frequencies of 45,X and 46,XN cell lineages after different culture times. Comparisons done between cell lineages. ns: non-significant.
Figure 4.
Metaphases in (A) first division (participant 3), (B) second division (participant 9), and (C) third division (participant 3).
Figure 4.
Metaphases in (A) first division (participant 3), (B) second division (participant 9), and (C) third division (participant 3).
Figure 5.
Linear regression of the proliferation index (PI) of lineages 45,X and 46,XN.
Figure 5.
Linear regression of the proliferation index (PI) of lineages 45,X and 46,XN.
Figure 6.
Intraindividual comparisons of SCE frequencies of 45,X and 46,XN lineages from the 17 participants. There were significant differences between the lineages of participants 2, 3, 4, 7, and 12. ns, non-significant (p>0.05), ∗p≤0.05, ∗∗p≤0.01.
Figure 6.
Intraindividual comparisons of SCE frequencies of 45,X and 46,XN lineages from the 17 participants. There were significant differences between the lineages of participants 2, 3, 4, 7, and 12. ns, non-significant (p>0.05), ∗p≤0.05, ∗∗p≤0.01.
Figure 7.
Linear regression of the sister chromatid exchange (SCE) frequencies of lineages 45,XN and 46,XN.
Figure 7.
Linear regression of the sister chromatid exchange (SCE) frequencies of lineages 45,XN and 46,XN.
Figure 8.
Intergroup comparisons of SCE frequency means among 45,X and the various 46,XN lineages found in the study participants.
Figure 8.
Intergroup comparisons of SCE frequency means among 45,X and the various 46,XN lineages found in the study participants.
Table 1.
Karyotypes of participants with Turner syndrome included in this study.
Table 1.
Karyotypes of participants with Turner syndrome included in this study.
| Participant |
Karyotype by conventional cytogenetics and FISH |
Age (yr.) |
| 1 |
mos45,X/46,XX |
27 |
| 2 |
mos45,X/46,XX |
30 |
| 3 |
mos45,X/46,X,ish.r(X)(DXZ1+,XIST++) |
16 |
| 4 |
mos45,X/46,X,+mar.ish.der(Y)(SHOX++,SRY ++,wcpY+) |
11 |
| 5 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST+) |
19 |
| 6 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST+) |
11 |
| 7 |
mos45,X/46,X,+mar.ish. der(X)(DXZ1+,XIST-) |
1.7 |
| 8 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST++) |
12 |
| 9 |
mos45,X/46,XY |
31 |
| 10 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST+) |
15 |
| 11 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST+) |
5 |
| 12 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST++) |
17 |
| 13 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST+) |
22 |
| 14 |
mos45,X/46,X+mar.ish.der(Y)(ENY++) |
23 |
| 15 |
mos45,X/46,XX |
42 |
| 16 |
mos45,X/46,X,+mar.ish.der(X)(DXZ1+,XIST+) |
19 |
| 17 |
mos45,X/46,X,i(Y)(pter->p10->pter).ish (ENY++,SRY++,SHOX++,DYZ3++) |
14 |
Table 2.
Paired t test on the equality of means for the proliferation index (PI) of lineages 45,X and 46,XN.
Table 2.
Paired t test on the equality of means for the proliferation index (PI) of lineages 45,X and 46,XN.
| PI |
N |
Mean |
Std. Err. |
Std. Dev.(±) |
95% Conf. Interval |
| 45,X lineage |
17 |
2.005 |
0.026 |
0.106 |
1.951 |
2.060 |
| 46,XN lineage |
17 |
2.078 |
0.038 |
0.155 |
1.998 |
2.157 |
|
t =-2.3211 |
p = 0.0338 |
|
Table 3.
Pearson’s correlation for the proliferation index (PI) of lineages 45,X and 46,XN.
Table 3.
Pearson’s correlation for the proliferation index (PI) of lineages 45,X and 46,XN.
| PI |
Mean |
Std. Dev.(±) |
Min |
Max |
45,X PI |
46,XN PI |
| 45,X lineage |
2.005 |
0.106 |
1.80 |
2.18 |
1.000 |
|
| 46,XN lineage |
2.078 |
0.155 |
1.84 |
2.48 |
0.5716 |
1.000 |
Table 4.
SCE frequencies and PI of the two cell lineages of the participants.
Table 4.
SCE frequencies and PI of the two cell lineages of the participants.
| Participants |
Lineages |
SCE mean |
p value |
PI |
χ2
|
p value |
| 1 |
45,X
46,XX |
3.15
2.6 |
0.0551 |
2.05
2.15 |
0.9387 |
0.6254 |
| 2 |
45,X
46,XX |
3.65
2.95 |
0.0454* |
2.18
2.27 |
0.9859 |
0.6108 |
| 3 |
45,X
46,X,+r(X) |
3.95
5.25 |
0.0046** |
2.06
2.16 |
1.016 |
0.6017 |
| 4 |
45,X
46,X,+mar(Y) |
4.45
3.55 |
0.0046** |
2.10
2.10 |
0.1640 |
0.9213 |
| 5 |
45,X
46,X,+mar(X) |
4.15
3.75 |
0.3583 |
2.02
1.98 |
0.7311 |
0.6938 |
| 6 |
45,X
46,X,+mar(X) |
3.55
4.25 |
0.0779 |
2.04
2.12 |
0.4380 |
0.8033 |
| 7 |
45,X
46,X,+mar(X) |
4.2
5.55 |
0.0100** |
2.10
2.10 |
0,1829 |
0.9126 |
| 8 |
45,X
46,X,+mar(X) |
3.45
4.2 |
0.0714 |
1.88
1.88 |
0.1814 |
0.9133 |
| 9 |
45,X
46,XY |
3.6
3.75 |
0.7805 |
2.06
1.92 |
2,123 |
0.3460 |
| 10 |
45,X
46,X,+mar(X) |
3.7
4.5 |
0.1183 |
1.98
2.16 |
3,187 |
0.2032 |
| 11 |
45,X
46,X,+mar(X) |
3.5
4.1 |
0.1544 |
2.04
2.12 |
0.4600 |
0.7945 |
| 12 |
45,X
46,X,+mar(X) |
3.8
4.85 |
0.0105* |
1.94
2.08 |
1.432 |
0.4886 |
| 13 |
45,X
46,X+mar(X) |
4.3
5.25 |
0.0745 |
1.84
1.90 |
0.2741 |
0.8719 |
| 14 |
45,X
46,X+mar(Y) |
4.4
4.7 |
0.5248 |
2.14
2.04 |
0.7435 |
0.6895 |
| 15 |
45,X
46,XX |
3.75
4.05 |
0.4647 |
1.80
2.06 |
4.399 |
0.1108 |
| 16 |
45,X
46,X,+mar(X) |
4
4.3 |
0.5228 |
1.90
1.90 |
0.1673 |
0.9198 |
| 17 |
45,X
46,X,i(Yp) |
4.35
3.3 |
0.0703 |
1.96
2.06 |
1.000 |
0.6065 |
Table 5.
Paired t test on the equality of means for the sister chromatid exchange (SCE) frequencies of lineages 45,X and 46,XN.
Table 5.
Paired t test on the equality of means for the sister chromatid exchange (SCE) frequencies of lineages 45,X and 46,XN.
| SCE |
N |
Mean |
Std. Err. |
Std. Dev. (±) |
95% Conf. Interval |
| 45,X |
17 |
3.879 |
0.093 |
0.383 |
3.682 |
4.076 |
| 46,XN |
17 |
4.112 |
0.227 |
0.934 |
3.631 |
4.592 |
|
t = -1.1012 |
p=0.2871 |
|
|
|
|
|
Table 6.
Pearson’s correlation for the sister chromatid exchange (SCE) frequencies of the lineages 45,X and 46,XN.
Table 6.
Pearson’s correlation for the sister chromatid exchange (SCE) frequencies of the lineages 45,X and 46,XN.
| SCE |
Mean |
Std. Dev (±) |
Min |
Max |
SCE 45,X |
SCE 46,XN |
| 45,X |
3.879 |
0.383 |
3.15 |
4.45 |
1.000 |
|
| 46,XN |
4.112 |
0.934 |
1.95 |
5.55 |
0.3615 |
1.000 |
Table 7.
p-values of Šídák's multiple comparisons test to support intergroup differences of SCE frequencies among 45,X and 46,XN lineages of mosaic Turner syndrome participants.
Table 7.
p-values of Šídák's multiple comparisons test to support intergroup differences of SCE frequencies among 45,X and 46,XN lineages of mosaic Turner syndrome participants.
| Karyotype |
46,XX |
46,Xr(X) |
46,X+mar(Y) |
46,X+mar(X) |
46,XY |
46,X+i(Yp) |
| 45,X |
0.0151* |
0.0007* |
0.9995 |
<0.0001* |
>0.9999 |
0.8204 |
| 46,XX |
|
<0.0001* |
0.0328* |
<0.0001* |
0.9542 |
>0.9999 |
| 46,Xr(X) |
|
|
0.0841 |
0.4991 |
<0.0198* |
0.0004* |
| 46,X+mar(Y) |
|
|
|
0.9079 |
0.9998 |
0,5310 |
| 46,X+mar(X) |
|
|
|
|
0.3630 |
0.0060* |
| 46,XY |
|
|
|
|
|
0.9997 |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).