Submitted:
21 February 2024
Posted:
22 February 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Results
2. ITS sequencing of 62 accessions of bananas
2. Classification of different subgroups of bananas using the “420 bp region” of ITS
2. Identification of hybrid bananas using the “420 bp region” of ITS
Discussion
Material and methods
4. Materials
4. Rapid extraction of banana genomic DNA
4. PCR, sequencing, and analysis
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Simmonds, N.W. The evolution of the bananas. Longmans Green 1962, 170. [Google Scholar]
- Simmonds, N.W.; Shepherd, K. Taxonomy and origins of cultivated bananas. Bot. J. Linn. Soc 1955, 55, 302–312. [Google Scholar] [CrossRef]
- Kaemmer, D.; Afza, R.; Weising, K.; Kahl, G.; Novak, F. Oligonucleotide and amplification fingerprinting of wild species and cultivars of banana (Musa spp.). Nat. Biotechnol. 1992, 10, 1030–1035. [Google Scholar] [CrossRef]
- Risterucci, AM.; Hippolyte, I.; Perrier, X.; Xia, L.; Caig, V.; Evers, M.; Huttner, E.; Kilian, A. Development and assessment of diversity arrays technology for high-throughput DNA analyses in Musa. Theor. Appl. Genet. 2009, 119, 1093–1103. [Google Scholar] [CrossRef]
- Hinge, V.; Shaikh, I.; Chavhan, R.; Deshmukh, A.; Shelake, R.; Ghuge, S.; Dethe, A.; Penna, S.; Kadam, U. Assessment of genetic diversity and volatile content of commercially grown banana (Musa spp.) cultivars. Sci. Rep-UK 2022, 12, 1–16. [Google Scholar] [CrossRef]
- Ravishankar, K.; Vidhya, L.; Cyriac, A.; Rekha, A.; Goel, R.; Sharma, N.K. Development of SSR markers based on a survey of genomic sequences and their molecular analysis in banana (Musa spp.). J. Hortic. Sci. Biotech. 2012, 87, 84–88. [Google Scholar] [CrossRef]
- Rotchanapreeda, T.; Wongniam, S.; Swangpol, S.; Chareonsap, P.; Sukkaewmanee, N.; Somana, J. Development of SSR markers from Musa balbisiana for genetic diversity analysis among Thai bananas. Plant Syst. Evol. 2016, 302, 739–761. [Google Scholar] [CrossRef]
- Suthanthiram, B.; Arumugam, C.; Uma, S.; Saraswathi, M. MusatransSSRDB (a transcriptome derived SSR database)–An advanced tool for banana improvement. J. Biosciences 2019, 44, 4. [Google Scholar]
- Ahmad, F.; Megia, R.; Poerba, Y.S. Genetic diversity of Musa balbisiana Colla in Indonesia based on AFLP marker. HAYATI J. Biosci. 2014, 21, 39–47. [Google Scholar] [CrossRef]
- Jesus, O.; Silva, S.; Amorim, E.; Ferreira, C.; de, C.; José, M.; Silva, G.; Figueira, A. Genetic diversity and population structure of Musa accessions in ex situ conservation. BMC Plant Biol. 2013, 13, 41. [Google Scholar] [CrossRef] [PubMed]
- Mertens, A.; Bawin, Y.; Vanden Abeele, S.; Kallow, S.; Vu, D.T.; Le, L.; Tuong, V.D.; Swennen, R.; Vandelook, F.; Panis, B.; et al. Genetic diversity and structure of Musa balbisiana populations in Vietnam and its implications for the conservation of banana crop wild relatives. PLOS ONE 2021, 16, e0253255. [Google Scholar] [CrossRef]
- Sardos, J.; Christelová, P.; Cizkova, J.; Paofa, J.; Sachter-Smith, G.; Janssens, S.; Rauka, G.B.; Ruas, M.; Daniells, J.; Dolezel, J.; Roux, N. Collection of new diversity of wild and cultivated bananas (Musa spp.) in the Autonomous Region of Bougainville, Papua New Guinea. Genet. Resour. Crop Ev. 2018, 65, 2267–2286. [Google Scholar] [CrossRef]
- Canal, W.M. SNP identification in crop plants. Curr. Opin. Plant Biol. 2009, 12, 211–217. [Google Scholar]
- Alberto, C.; Julie, S.; Yann, H.; Guillaume, M.; Catherine, B.; Nicolas, R.; Rony, S.; Sebastien, C.C.; Mathieu, R. Unravelling the complex story of intergenomic recombination in ABB allotriploid bananas. Ann Bot-London 2021, 127, 7–20. [Google Scholar]
- Sardos, J.; Rouard, M.; Hueber, Y.; Cenci, A.; Hyma, K.; Houwe, I.; Hribova, E.; Courtois, B.; Roux, N. A genome-wide association study on the seedless phenotype in banana (Musa spp.) reveals the potential of a selected panel to detect candidate genes in a vegetatively propagated crop. PLOS ONE 2016, 11, e0154448. [Google Scholar] [CrossRef] [PubMed]
- Gardoce, R.; Manohar, A.N.; Mendoza, JV.; Tejano, M.; Nocum, J.D.; Lachica, G.; Gueco, L.; Cueva, F.; Lantican, D. A novel SNP panel developed for targeted genotyping-by-sequencing (GBS) reveals genetic diversity and population structure of Musa spp. germplasm collection. Mol. Genet. Genomics 2023, 298, 857–869. [Google Scholar] [CrossRef] [PubMed]
- Nwakanma, D.C.; Pillay, M.; Okoli, B.E.; Tenkouano, A. PCR-RFLP of the ribosomal DNA internal transcribed spacers (ITS) provides markers for the A and B genomes in Musa L. Theor. Appl. Genet., 2003, 108, 154–159. [Google Scholar] [CrossRef] [PubMed]
- Dita, M.A.; Waalwijk, C.; Buddenhagen, I.W.; Souza, M.T.; Kema, H.J. A molecular diagnostic for tropical race 4 of the banana fusarium wilt pathogen. Plant Pathol. 2010, 59, 348–357. [Google Scholar] [CrossRef]
- Teo, C.; Tan, S.; Ho, C.; Faridah, Q.; Othman, Y.; Heslop-Harrison, J.; Kalendar, R.; Schulman, A. Genomic constitution and classification using retrotransposon-based markers in the orphan crop banana. J. Plant Biol. 2005, 48, 96–105. [Google Scholar] [CrossRef]
- Nair, A.S.; Teo, C.H.; Schwarzacher, T.; Harrison, P.H. Genomic classification of banana cultivars from south India using IRAP markers. Euphytica 2005, 144, 285–290. [Google Scholar] [CrossRef]
- Li, L.; Häkkinen, M.; Yuan, Y.; Hao, G.; Ge, X. Molecular phylogeny and systematics of the banana family (Musaceae) inferred from multiple nuclear and chloroplast DNA fragments, with a special reference to the genus Musa. Mol. Phylogenet. Evol. 2010, 57, 1–10. [Google Scholar] [CrossRef]
- MORÁN, J.F. Improvement of cavendish banana cultivars through conventional breeding. Acta Hortic. 2013, 986, 205–208. [Google Scholar] [CrossRef]
- Biswas, M.K.; Yi, G. Genes and markers: Application in banana crop improvement. In Banana: Genomics and transgenic approaches for genetic improvement; Mohandas, S., Ravishankar, K., Eds.; Springer Press: Singapore, Singapore, 2016; pp. 35–50. [Google Scholar]
- Saraswathi, M.S.; Uma, S.; Vadivel, E.; Durai, P.; Siva, S.A.; Rajagopal, G.; Sathiamoorthy, S. Diversity analysis in Indian cooking bananas (Musa, ABB) through morphotaxonomic and molecular characterization. Acta Hortic. 2011.
- Christelová, P.; Langhe, E.; Hribova, E.; Cizkova, J.; Sardos, J.; Hušáková, M.; Houwe, I.; Sutanto, A.; Kepler, A.; Swennen, R.; et al. Molecular and cytological characterization of the global Musa germplasm collection provides insights into the treasure of banana diversity. Biodivers. Conserv. 2017, 26, 801–824. [Google Scholar] [CrossRef]
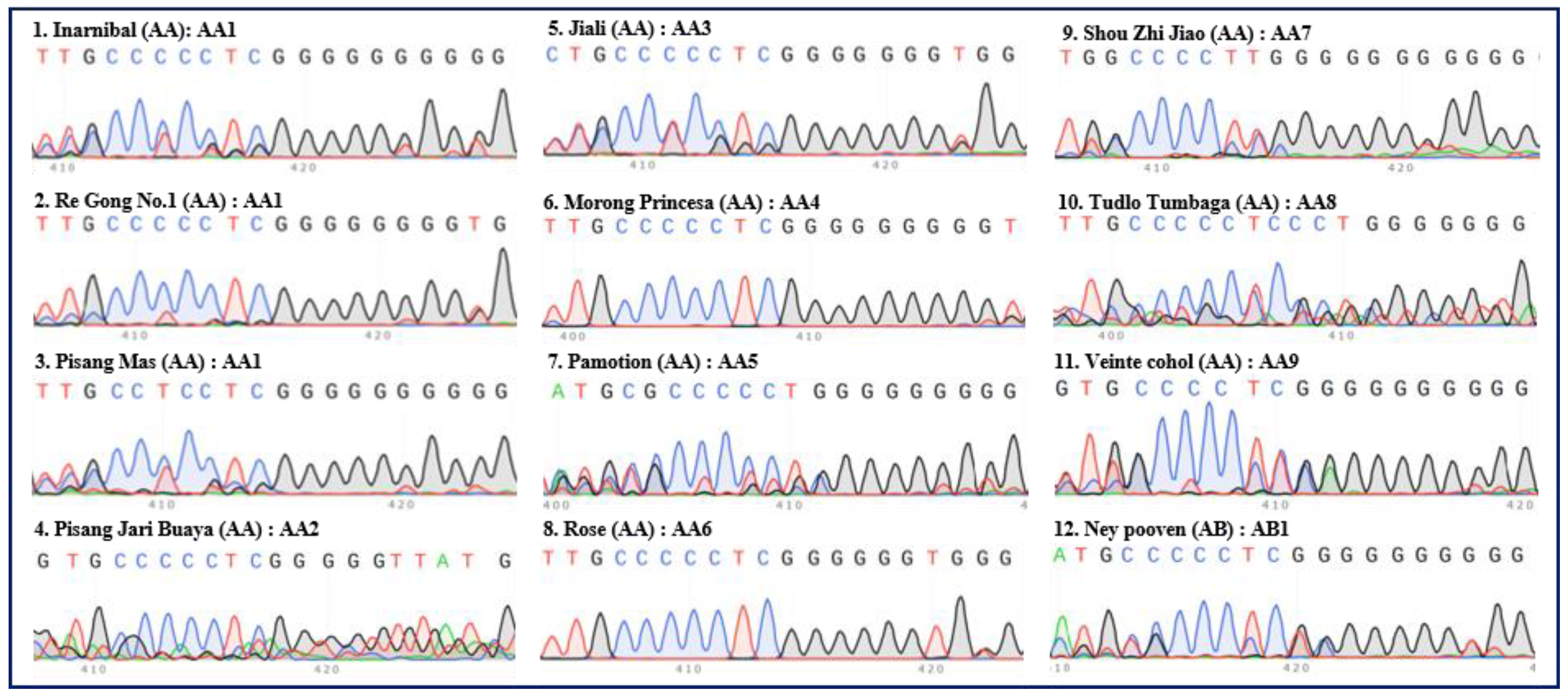
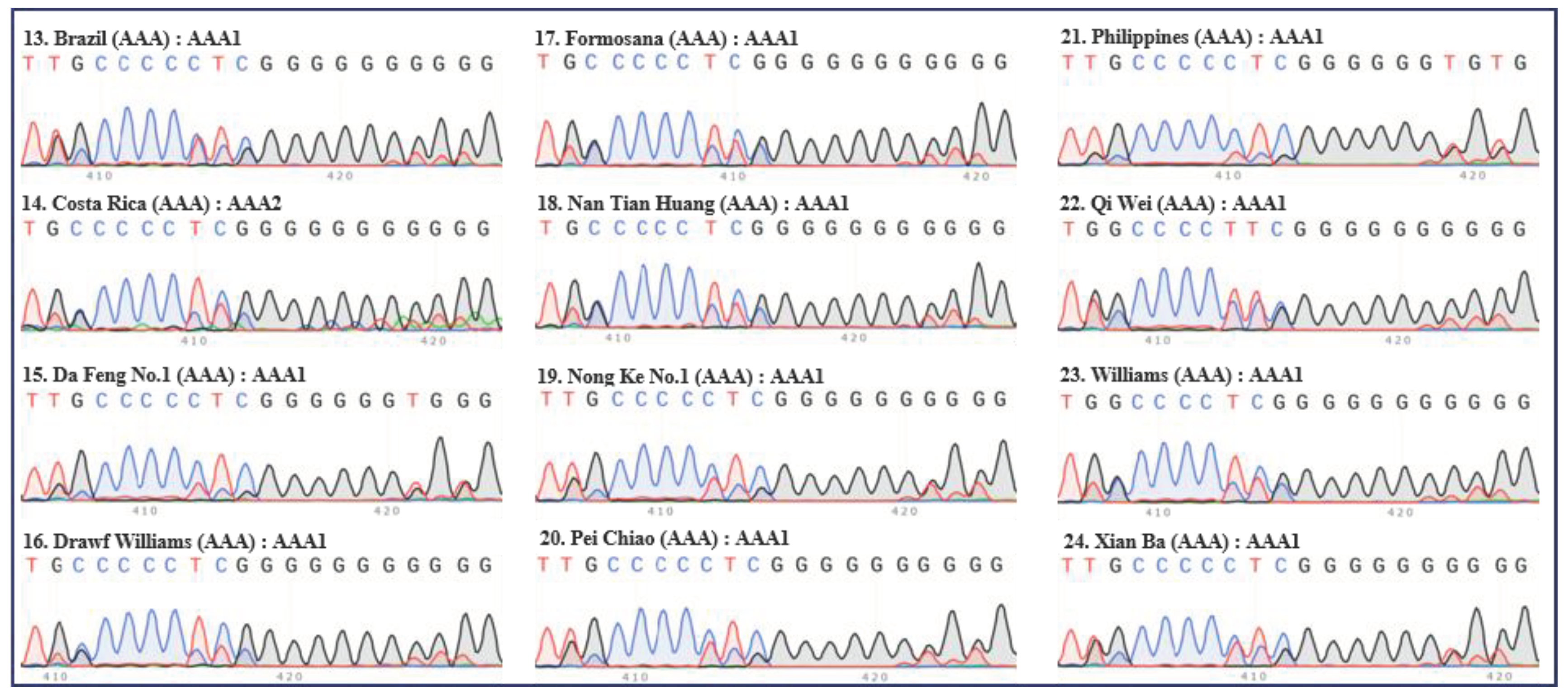
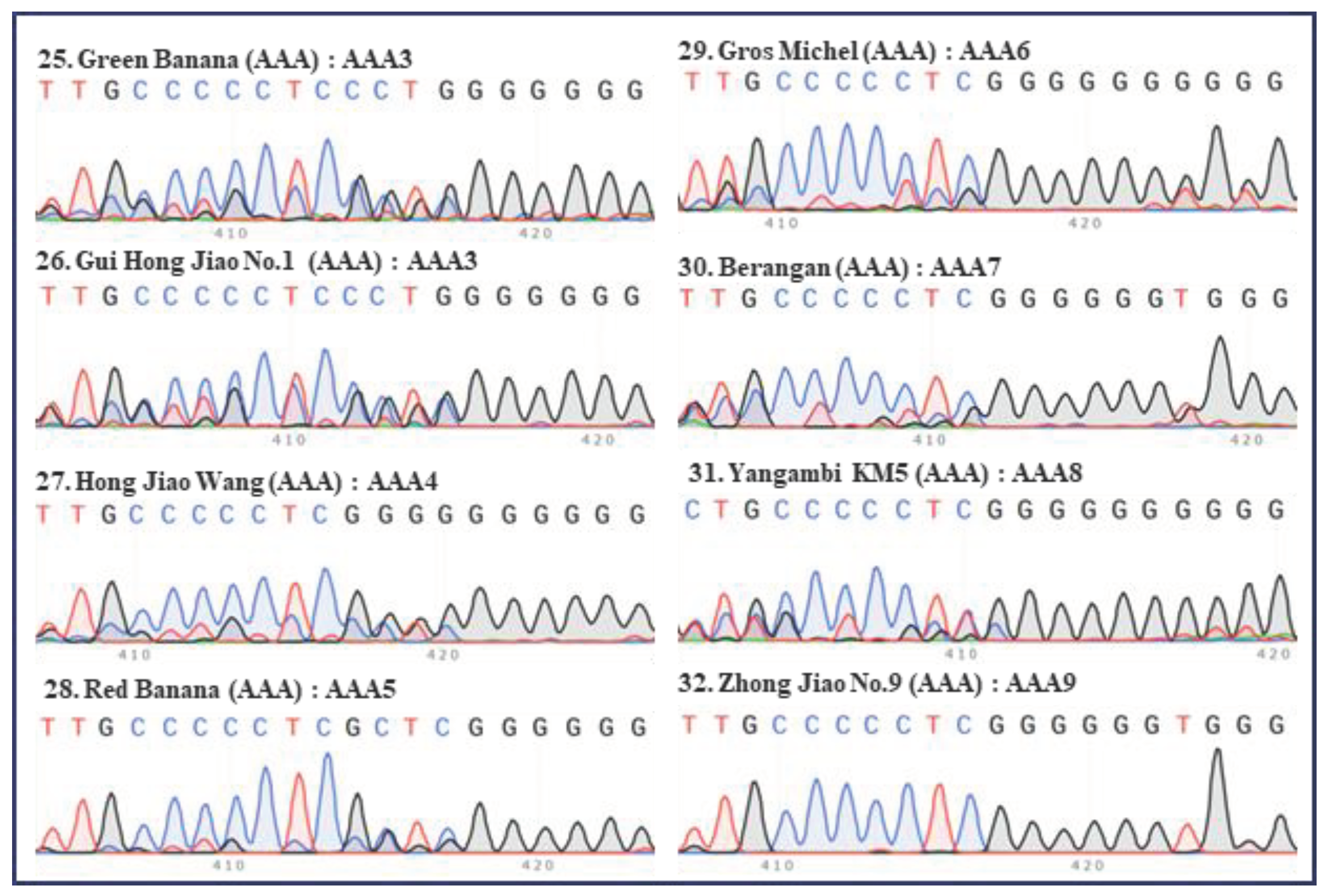
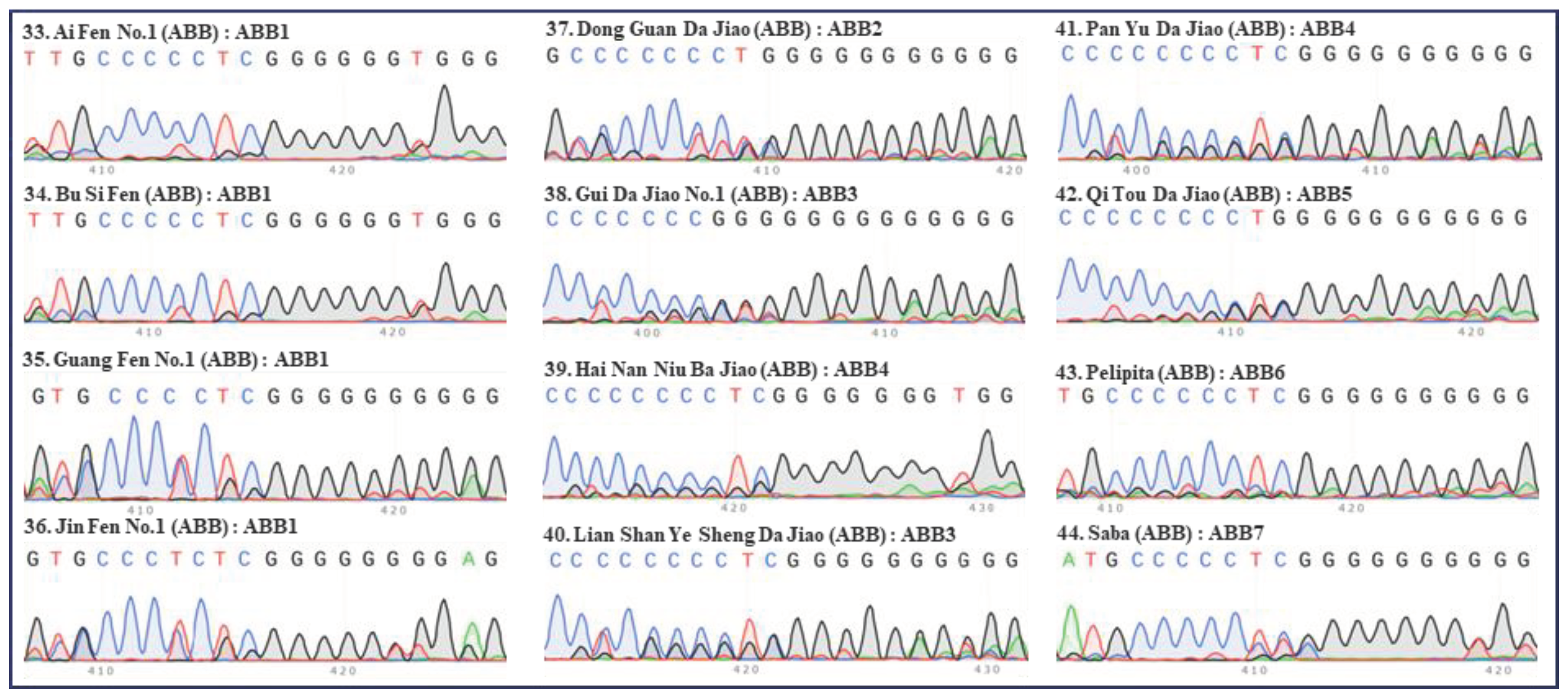
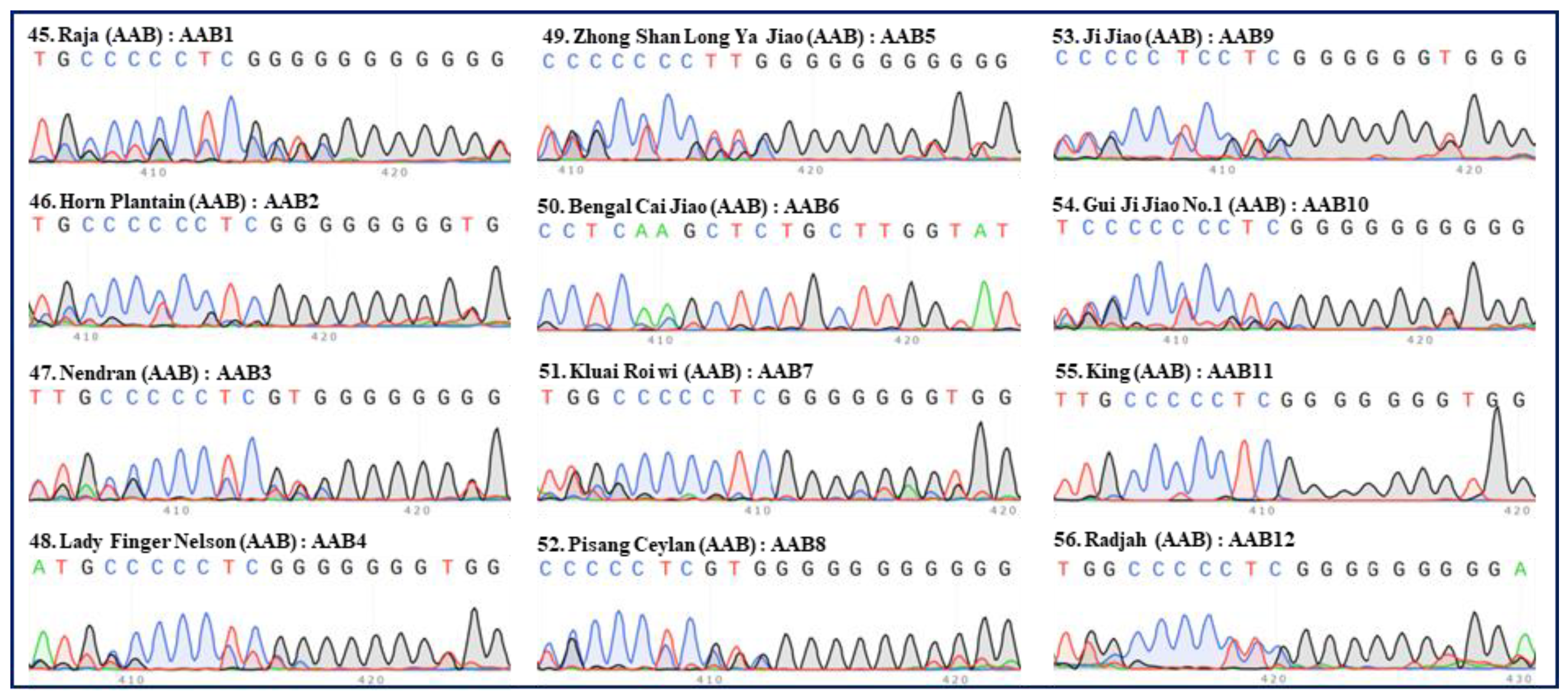
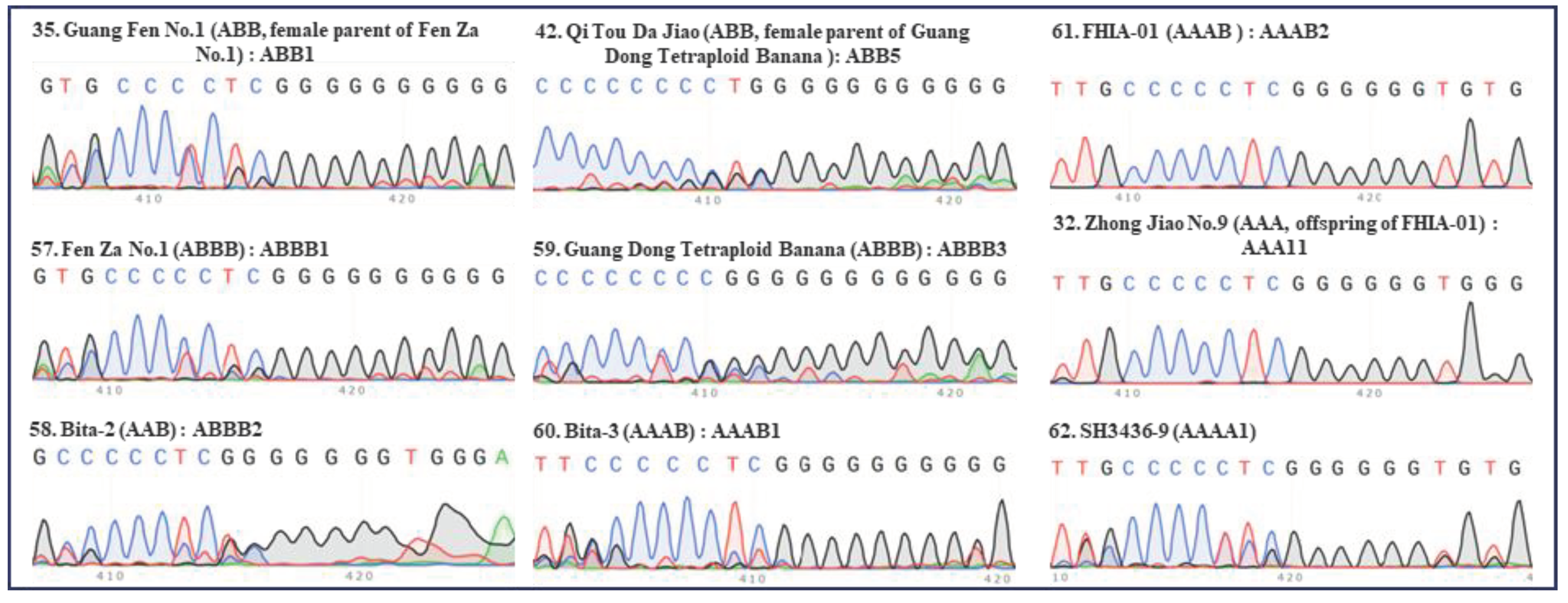
| Serial no. | Names | Groups | Subgroups | ITS types |
|---|---|---|---|---|
| 1 | Inarnibal | AA | Inarnibal | AA1 |
| 2 | Re Gong No.1 | AA | Inarnibal | AA1 |
| 3 | Pisang Mas | AA | Sucrier | AA1 |
| 4 | Pisang Jari Buaya | AA | Pisang Jari Buaya | AA2 |
| 5 | Jia Li (mutant of Kluai lep mu nang) | AA | Unknown | AA3 |
| 6 | Morong Princesa | AA | Unknown | AA4 |
| 7 | Pamotion | AA | Unknown | AA5 |
| 8 | Rose | AA | Unknown | AA6 |
| 9 | Shou Zhi Jiao | AA | Unknown | AA7 |
| 10 | Tudlo Tumbaga | AA | Unknown | AA8 |
| 11 | Veinte cohol | AA | Unknown | AA9 |
| 12 | Ney Pooven | AB | Ney Pooven | AB1 |
| 13 | Brazil | AAA | Cavendish | AAA1 |
| 14 | Costa Rica | AAA | Cavendish | AAA2 |
| 15 | Da Feng No.1 | AAA | Cavendish | AAA1 |
| 16 | Drawf Williams | AAA | Cavendish | AAA1 |
| 17 | Formosana | AAA | Cavendish | AAA1 |
| 18 | Nan Tian Huang | AAA | Cavendish | AAA1 |
| 19 | Nong Ke No.1 | AAA | Cavendish | AAA1 |
| 20 | Pei Chiao | AAA | Cavendish | AAA1 |
| 21 | Philippines | AAA | Cavendish | AAA1 |
| 22 | Qi Wei | AAA | Cavendish | AAA1 |
| 23 | Williams | AAA | Cavendish | AAA1 |
| 24 | Xian Ba | AAA | Cavendish | AAA1 |
| 25 | Green Banana | AAA | Red | AAA3 |
| 26 | Gui Hong Jiao No.1 | AAA | Red | AAA3 |
| 27 | Hong Jiao Wang | AAA | Red | AAA4 |
| 28 | Red Banana | AAA | Red | AAA5 |
| 29 | Gros Michel | AAA | Gros Michel | AAA6 |
| 30 | Berangan | AAA | Lakatan | AAA7 |
| 31 | Yangambi KM5 | AAA | Ibota Bota | AAA8 |
| 32 | Zhong Jiao No.9 | AAA | FHIA-01×SH-3142 | AAA9 |
| 33 | Ai Fen No.1 | ABB | Pisang Awak | ABB1 |
| 34 | Bu Si Fen | ABB | Pisang Awak | ABB1 |
| 35 | Guang Fen No.1 | ABB | Pisang Awak | ABB1 |
| 36 | Jin Fen No.1 | ABB | Pisang Awak | ABB1 |
| 37 | Dong Guan Da Jiao | ABB | Da Jiao | ABB2 |
| 38 | Gui Da Jiao No.1 | ABB | Da Jiao | ABB3 |
| 39 | Hai Nan Niu Ba Jiao | ABB | Da Jiao | ABB4 |
| 40 | Lian Shan Ye Sheng Da Jiao | ABB | Da Jiao | ABB3 |
| 41 | Pan Yu Da Jiao | ABB | Da Jiao | ABB4 |
| 42 | Qi Tou Da Jiao | ABB | Da Jiao | ABB5 |
| 43 | Pelipita | ABB | Pelipita | ABB6 |
| 44 | Saba | ABB | Saba | ABB7 |
| 45 | Raja | AAB | Pisang Raja | AAB1 |
| 46 | Horn Plantain | AAB | Plantain | AAB2 |
| 47 | Nendran | AAB | Plantain | AAB3 |
| 48 | Lady Finger Nelson | AAB | Pome | AAB4 |
| 49 | Zhong Shan Long Ya Jiao | AAB | Silk | AAB5 |
| 50 | Bengal Cai Jiao | AAB | Unknown | AAB6 |
| 51 | Kluai Roi wi | AAB | Unknown | AAB7 |
| 52 | Pisang Ceylan | AAB | Unknown | AAB8 |
| 53 | Ji Jiao | AAB | Unknown | AAB9 |
| 54 | Gui Ji Jiao No.1 | AAB | Unknown | AAB10 |
| 55 | King | AAB | Unknown | AAB11 |
| 56 | Radjah | AAB | Unknown | AAB12 |
| 57 | Fen Za No.1 | ABBB | Guang Fen No.1×Musa Balbisiana | ABBB1 |
| 58 | Bita-2 | ABBB | Fougamou(AAB, Pisang Awak) x Musa Balbisiana 1-63 | ABBB2 |
| 59 | Guang Dong Tetraploid Banana | ABBB | Qi Tou Da Jiao×BB | ABBB3 |
| 60 | Bita-3 | AAAB | (AAAB, Laknau (AAB, Laknau) x Tjau Lagada(AA)) | AAAB1 |
| 61 | FHIA-01 | AAAB | Prata Ana×SH-3142 | AAAB2 |
| 62 | SH3436-9 | AAAA | Unknown | AAAA1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




