Submitted:
26 January 2024
Posted:
29 January 2024
Read the latest preprint version here
Abstract
Keywords:
Introduction
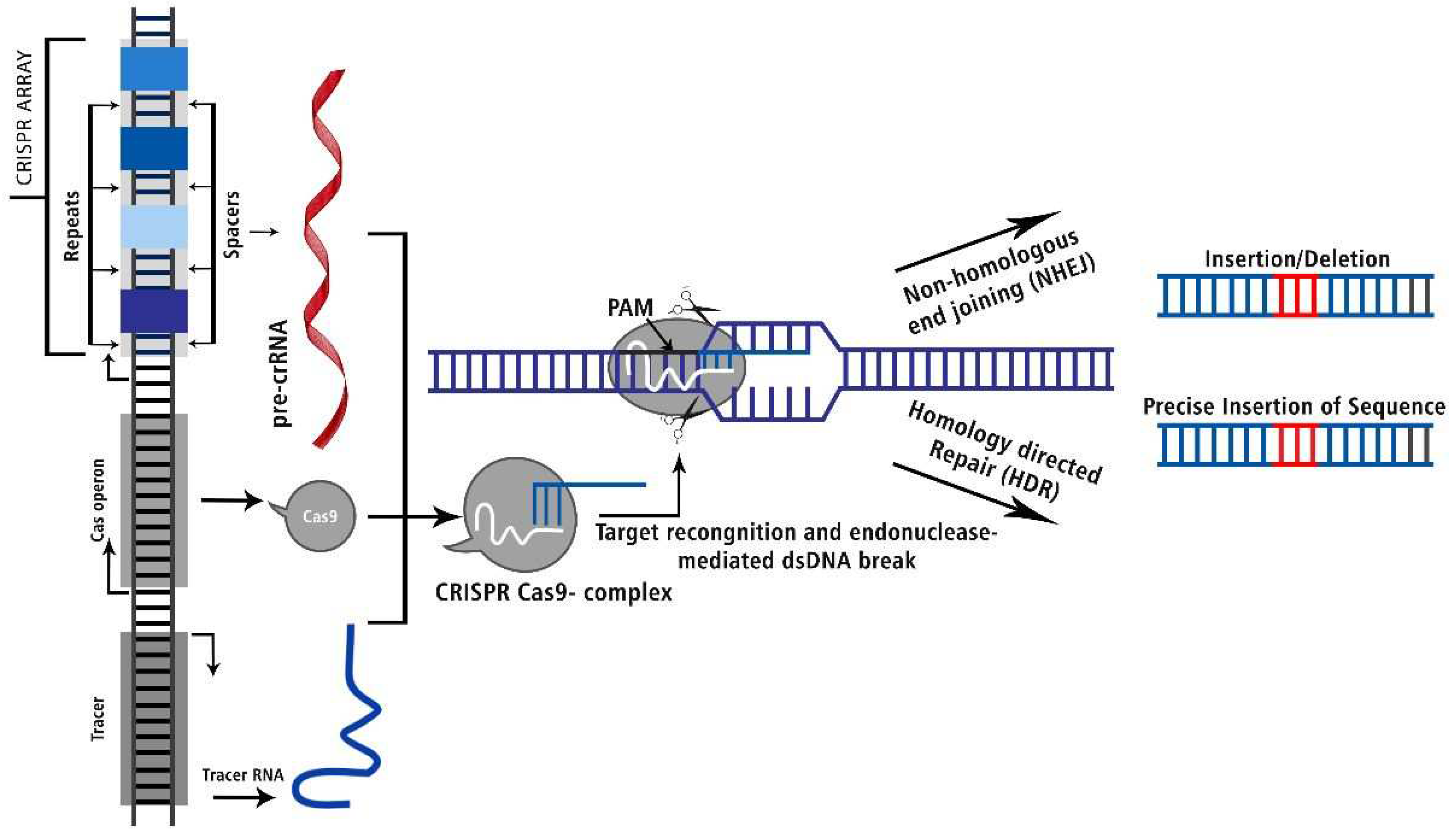
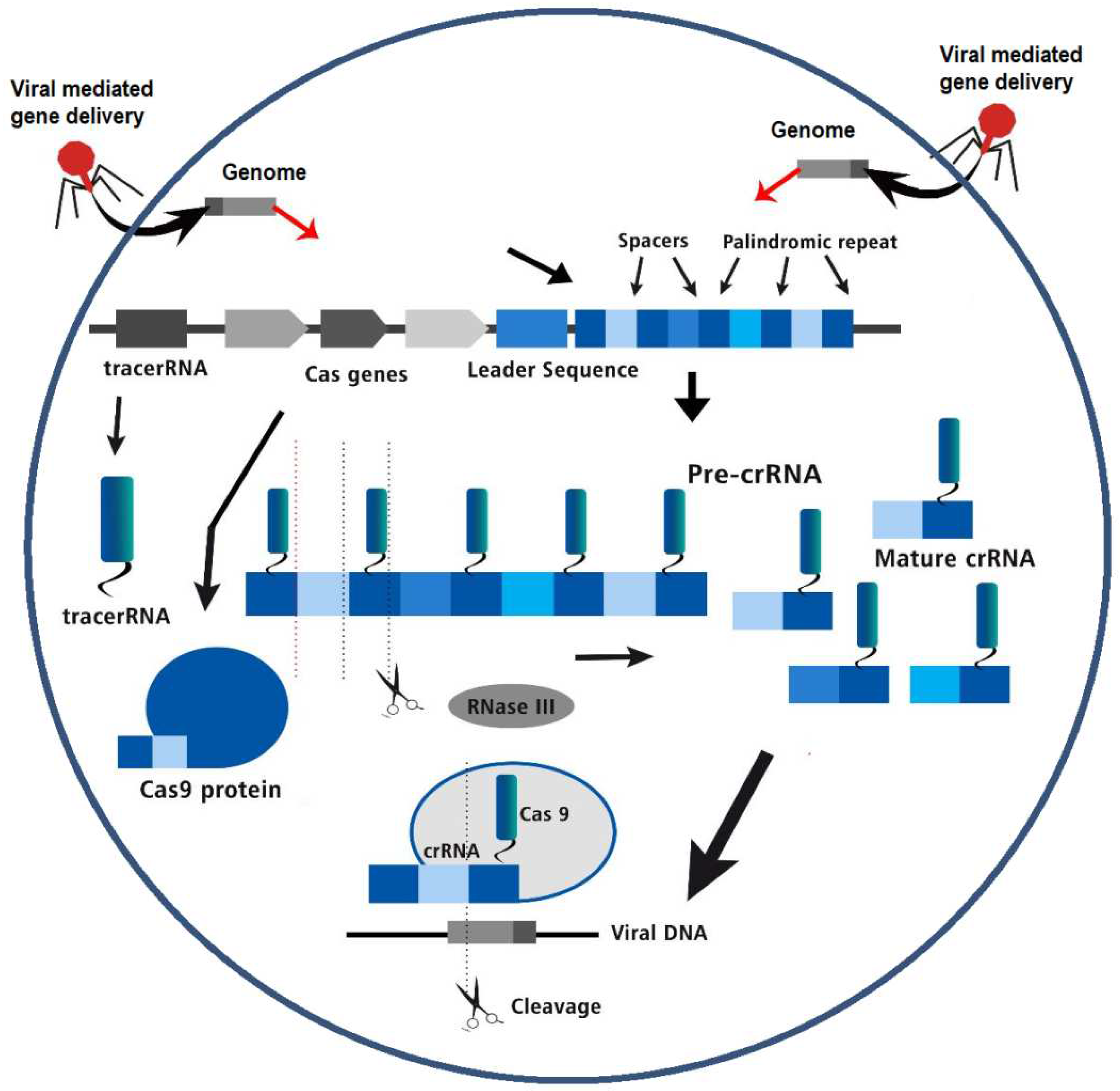
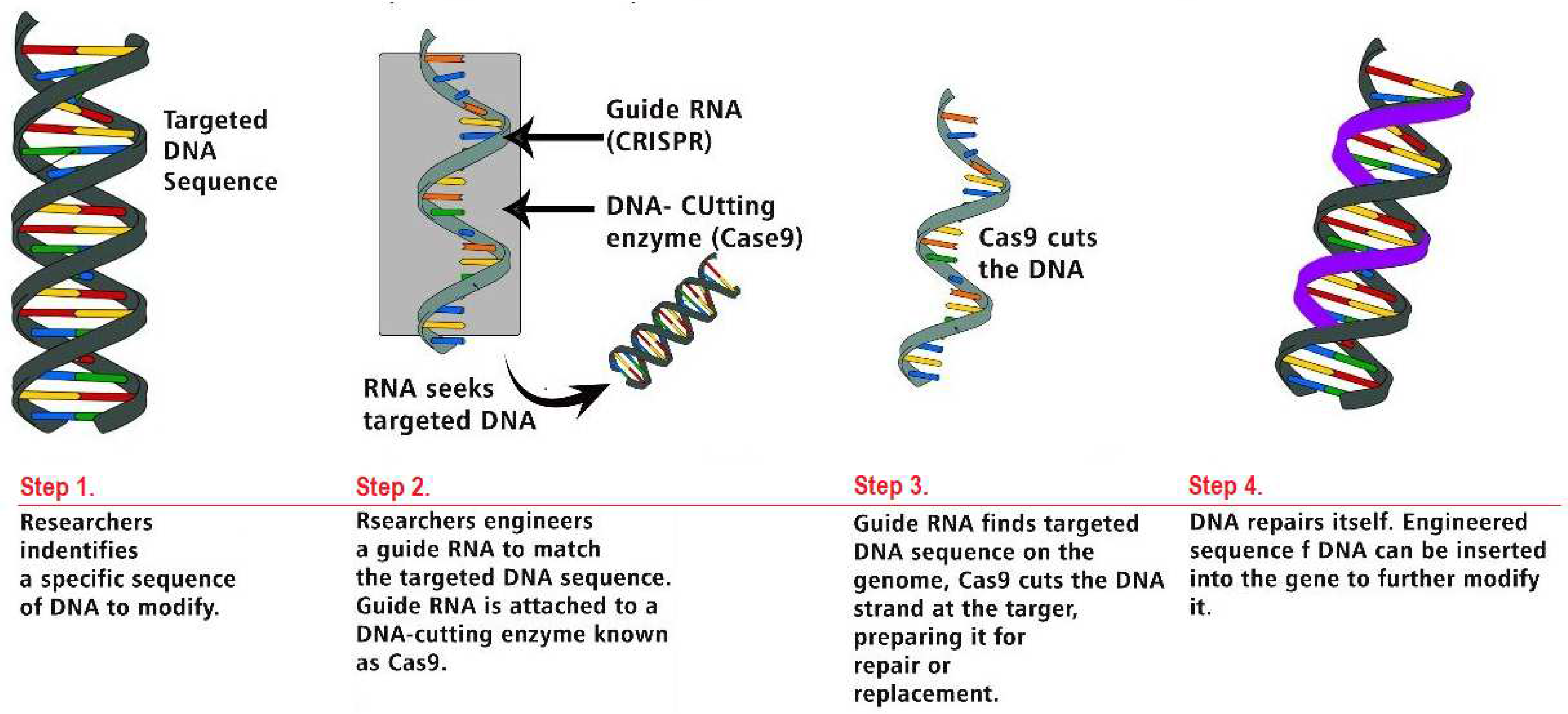
CRISPR-Cas Genome Editing Technology
APPLICATIONS
Genome editing
Transcription regulation
Gene therapy
CHALLENGES
Off-target mutations
PAM dependence
gRNA production
Delivery methods
CONCLUSION
References
- Abudayyeh, O.O.; et al. A cytosine deaminase for programmable single-base RNA editing. Science 2019, 365, 382–386. [Google Scholar] [CrossRef] [PubMed]
- Billon, P.; et al. CRISPR-mediated base editing enables efficient disruption of eukaryotic genes through induction of STOP codons. Mol. Cell 2017, 67, 1068–1079. [Google Scholar] [CrossRef]
- Biotechnol. J. 16, 1918–1927 Okada, A. et al. CRISPR/Cas9-mediated knockout of Ms1 enables the rapid generation of male-sterile hexaploid wheat lines for use in hybrid seed production. Plant Biotechnol. 2019, 17, 1905–1913.
- Braun, C.J.; et al. Versatile in vivo regulation of tumor phenotypes by dCas9-mediated transcriptional perturbation. Proc. Natl. Acad. Sci. U. S. A. 2016, 113, E3892–E3900. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; et al. CRISPR/Cas9-mediated targeted mutagenesis of GmFT2a delays flowering time in soya bean. Plant Biotechnol. J. 2018, 16, 176–185. [Google Scholar] [CrossRef]
- Chandrasekaran, J.; et al. Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol. Plant Pathol. 2016, 17, 1140–1153. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; et al. Whole genome sequencing reveals rare off-target mutations and considerable inherent genetic or/and somaclonal variations in CRISPR/ Cas9-edited cotton plants. Plant Biotechnol. J. 2018, 17, 858–868. [Google Scholar] [CrossRef]
- Li, J.F.; et al. Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 2013, 31, 688–691. [Google Scholar] [CrossRef]
- Li, X.; et al. Base editing with a Cpf1-cytidine deaminase fusion. Nat. Biotechnol. 2018, 36, 324–327. [Google Scholar] [CrossRef]
- Li, Z.; et al. Cas9-guide RNA directed genome editing in soybean. Plant Physiol. 2015, 169, 960–970. [Google Scholar] [CrossRef]
- Li, Z.; et al. A potent Cas9-derived gene activator for plant and mammalian cells. Nat. Plants 2017, 3, 930–936. [Google Scholar] [CrossRef]
- Liang, Z.; et al. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 2017, 8, 14261. [Google Scholar] [CrossRef]
- Liu, L.; et al. C2c1-sgRNA complex structure reveals RNA-guided DNA cleavage mechanism. Mol. Cell 2017, 65, 310–322. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; et al. Hi-TOM: a platform for high[1]throughput tracking of mutations induced by CRISPR/Cas systems. Sci. China Life Sci. 2018, 62, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; et al. Application of CRISPR/Cas9 in plant biology. Acta Pharm. Sin. B 2017, 7, 292–302. [Google Scholar] [CrossRef] [PubMed]
- Odipio, J.; et al. Efficient CRISPR/Cas9 genome editing of phytoene desaturase in cassava. Front. Plant Sci. 2017, 8, 1780. [Google Scholar] [CrossRef] [PubMed]
- Ordon, J.; et al. Generation of chromosomal deletions in dicotyledonous plants employing a user-friendly genome editing toolkit. Plant J. 2017, 89, 155–168. [Google Scholar] [CrossRef]
- Ortigosa, A.; et al. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol. J. 2019, 17, 665–673. [Google Scholar] [CrossRef] [PubMed]
- Pan, C.; et al. CRISPR/Cas9-mediated efficient and heritable targeted mutagenesis in tomato plants in the first and later generations. Sci. Rep. 2016, 6, 24765. [Google Scholar] [CrossRef]
- Peng, A.; et al. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 2017, 15, 1509–1519. [Google Scholar] [CrossRef]
- Ren, B.; et al. Cas9-NG greatly expands the targeting scope of the genome-editing toolkit by recognizing NG and other atypical PAMs in rice. Mol. Plant 2019, 12, 1015–1026. [Google Scholar] [CrossRef]
- Safari, F.; et al. CRISPR Cpf1 proteins: structure, function and implications for genome editing. Cell Biosci 2019, 9, 36. [Google Scholar] [CrossRef]
- Schaeffer, S.M.; Nakata, P.A. CRISPR/ Cas9-mediated genome editing and gene replacement in plants: transitioning from lab to field. Plant Sci. 2015, 240, 130–142. [Google Scholar]
- Scheben, A.; et al. Towards CRISPR/Cas crops–bringing together genomics and genome editing. New Phytol. 2017, 216, 682–698. [Google Scholar] [CrossRef]
- Schiml, S.; et al. Repair of adjacent single[1]strand breaks is often accompanied by the formation of tandem sequence duplications in plant genomes. Proc. Natl. Acad. Sci. U. S. A. 2016, 113, 7266–7271. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; et al. Efficient CRISPR/Cas9-based gene knockout in watermelon. Plant Cell Rep. 2017, 36, 399–406. [Google Scholar] [CrossRef] [PubMed]
- Ueta, R.; et al. Rapid breeding of parthenocarpic tomato plants using CRISPR/Cas9. Sci. Rep. 2017, 7, 507. [Google Scholar] [CrossRef] [PubMed]
- Veres, A.; et al. Low incidence of off-target mutations in individual CRISPR-Cas9 and TALEN targeted human stem cell clones detected by whole-genome sequencing. Cell Stem Cell 2014, 15, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; et al. Clonal seeds from hybrid rice by simultaneous genome engineering of meiosis and fertilization genes. Nat Biotechnol. 2019, 37, 283–286. [Google Scholar] [CrossRef]
- Wang, J.; et al. xCas9 expands the scope of genome editing with reduced efficiency in rice. Plant Biotechnol. J. 2018, 17, 709–711. [Google Scholar] [CrossRef]
- Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346, 1258096. [Google Scholar]
- Yang, H.; et al. CRISPR/Cas9-mediated genome editing efficiently creates specific mutations at multiple loci using one sgRNA in Brassica napus. Sci. Rep. 2017, 7, 7489. [Google Scholar] [CrossRef] [PubMed]
- Yin, K.; et al. Progress and prospects in plant genome editing. Nat. Plants 2017, 3, 17107. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, S.S.; et al. CRISPR-Cpf1: a new tool for plant genome editing. Trends Plant Sci. 2017, 22, 550–553. [Google Scholar] [CrossRef] [PubMed]
- Zischewski, J.; et al. Detection of on-target and off-target mutations generated by CRISPR/ Cas9 and other sequence-specific nucleases. Biotechnol. Adv. 2017, 35, 95–104. [Google Scholar] [CrossRef]
- Zong, Y.; et al. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 438–440. [Google Scholar] [CrossRef]
- Zong, Y.; et al. Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat. Biotechnol. 2018, 36, 950–953. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



