Submitted:
10 January 2024
Posted:
11 January 2024
You are already at the latest version
Abstract
Keywords:
Introduction
Materials and Methods
Plant materials and field trials
Measurement of Hg content
Genotype and GWAS
Candidate gene identification
Analysis of Expression Level Association of Candidate Genes
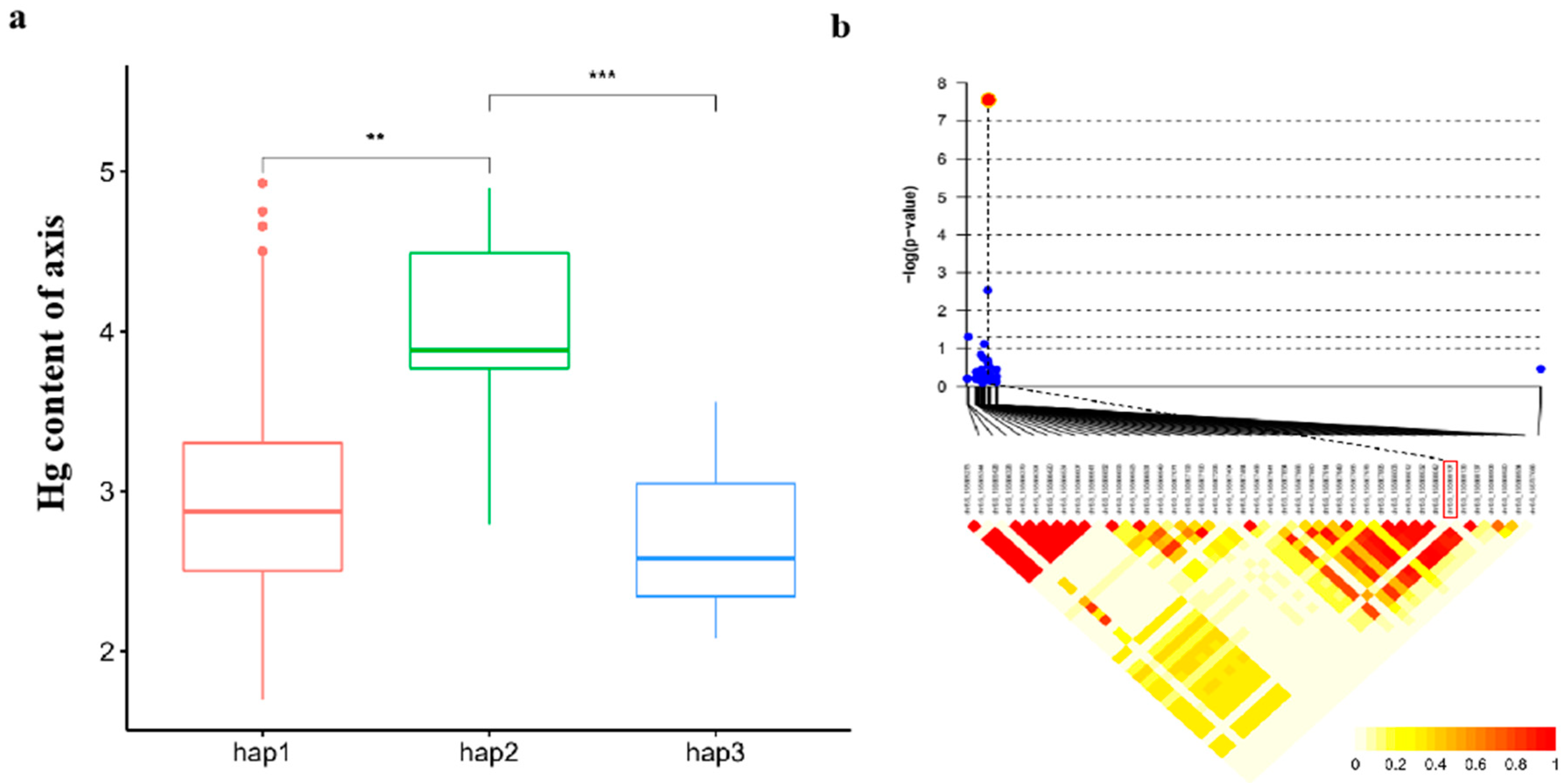
Haplotype analyses of candidate genes
Results
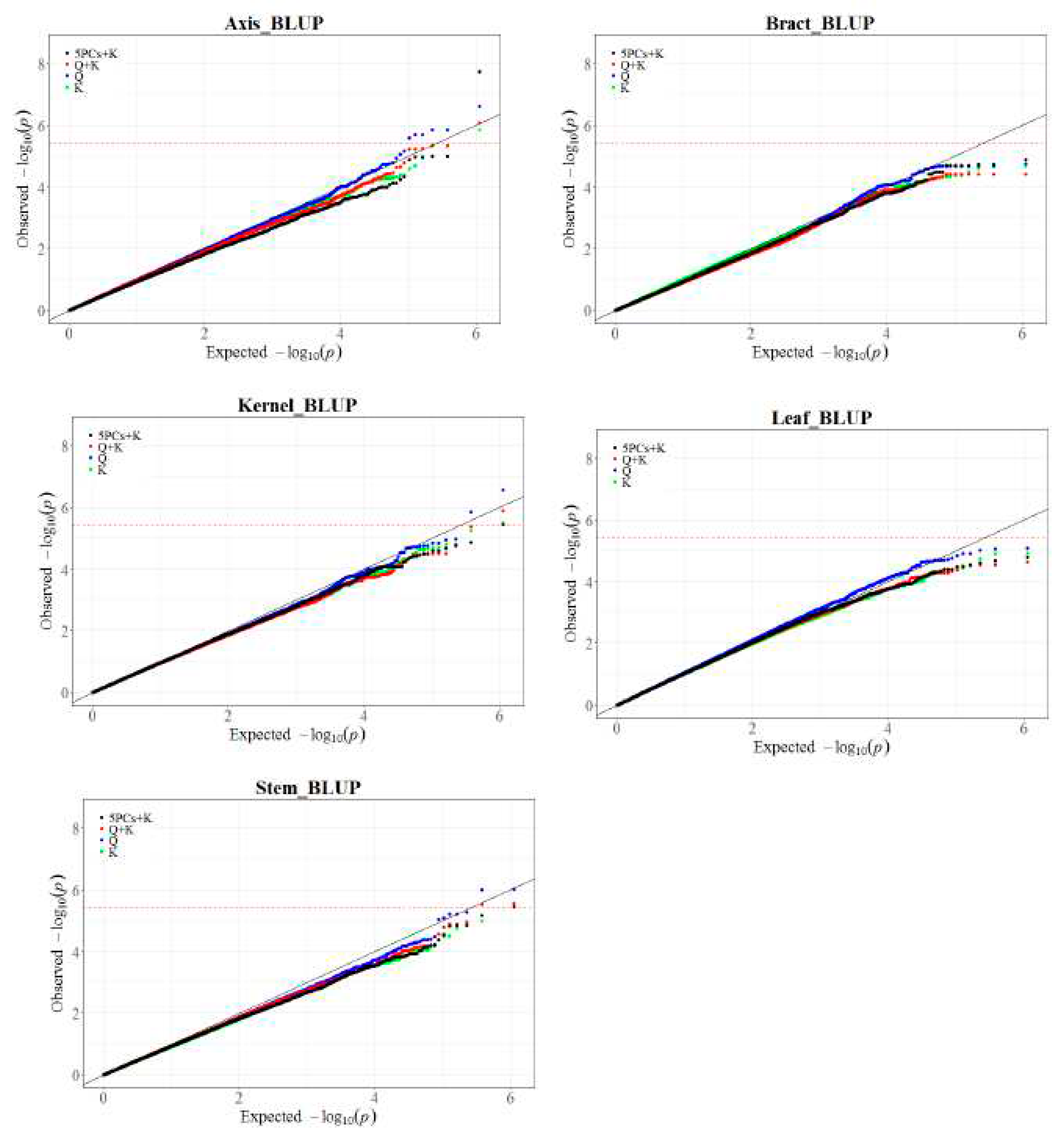
Model comparison and selection
Boosting GWAS Power through Increased Marker Density
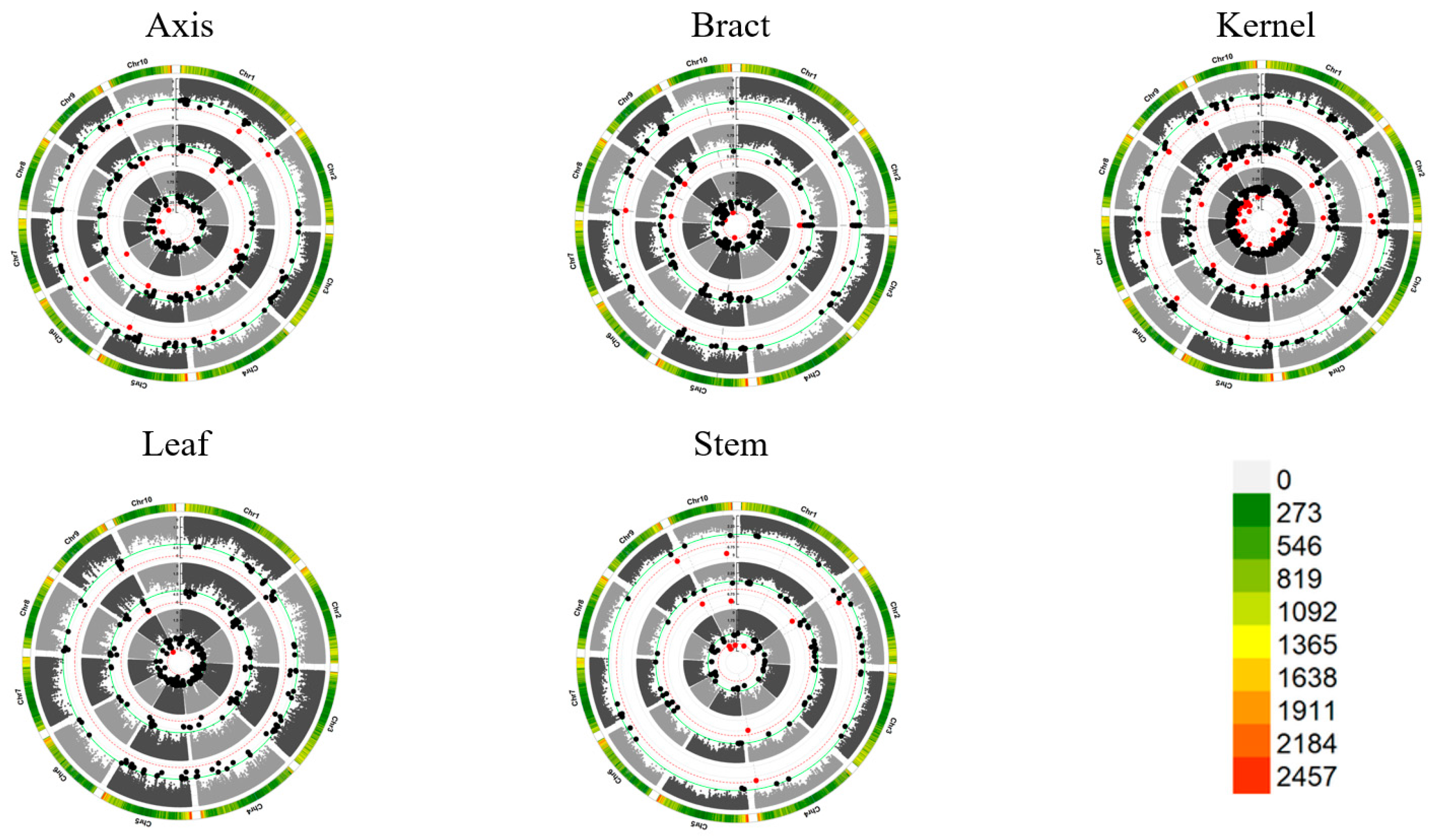
Significant Loci and Tissue-Specific Variability
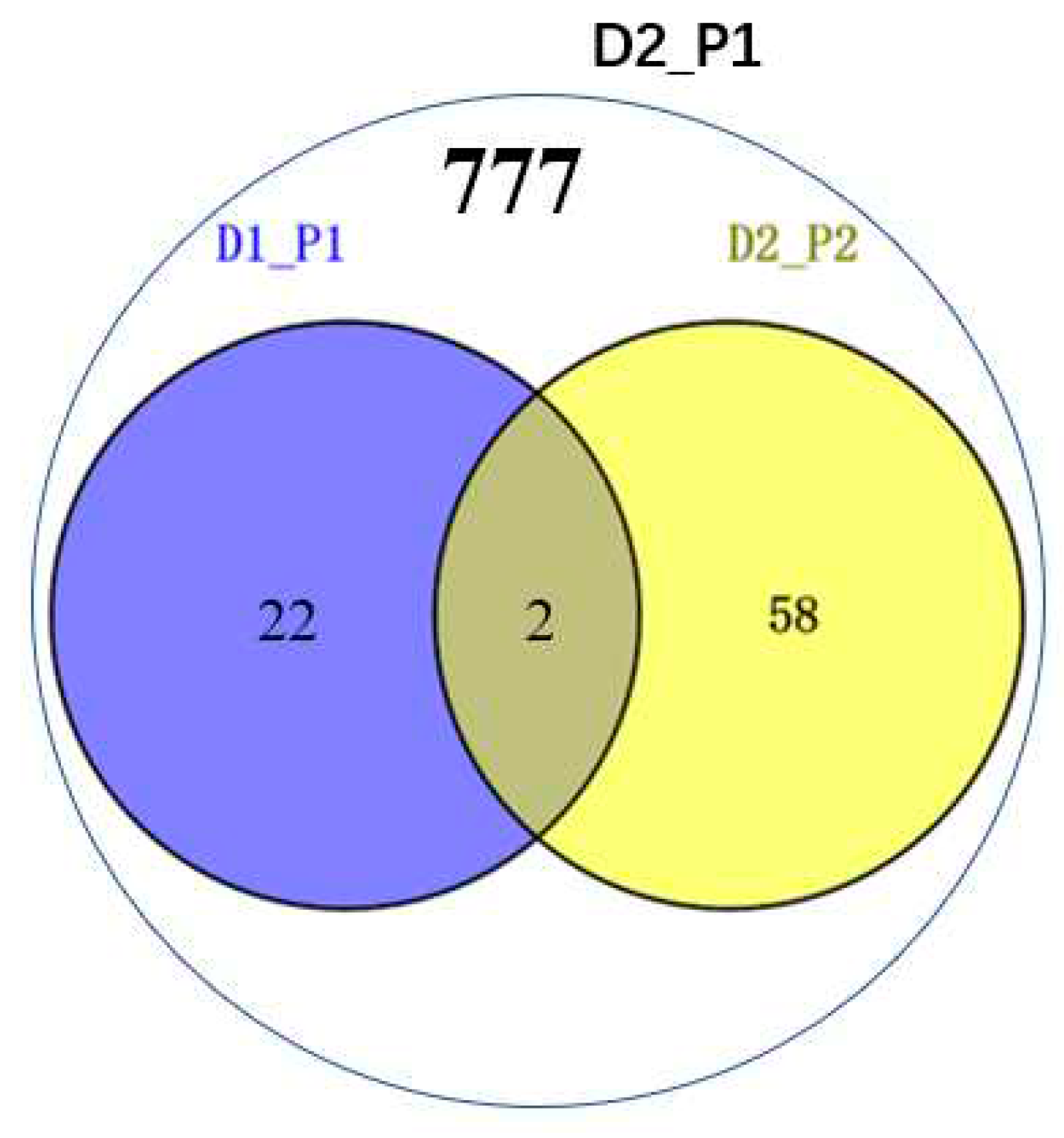
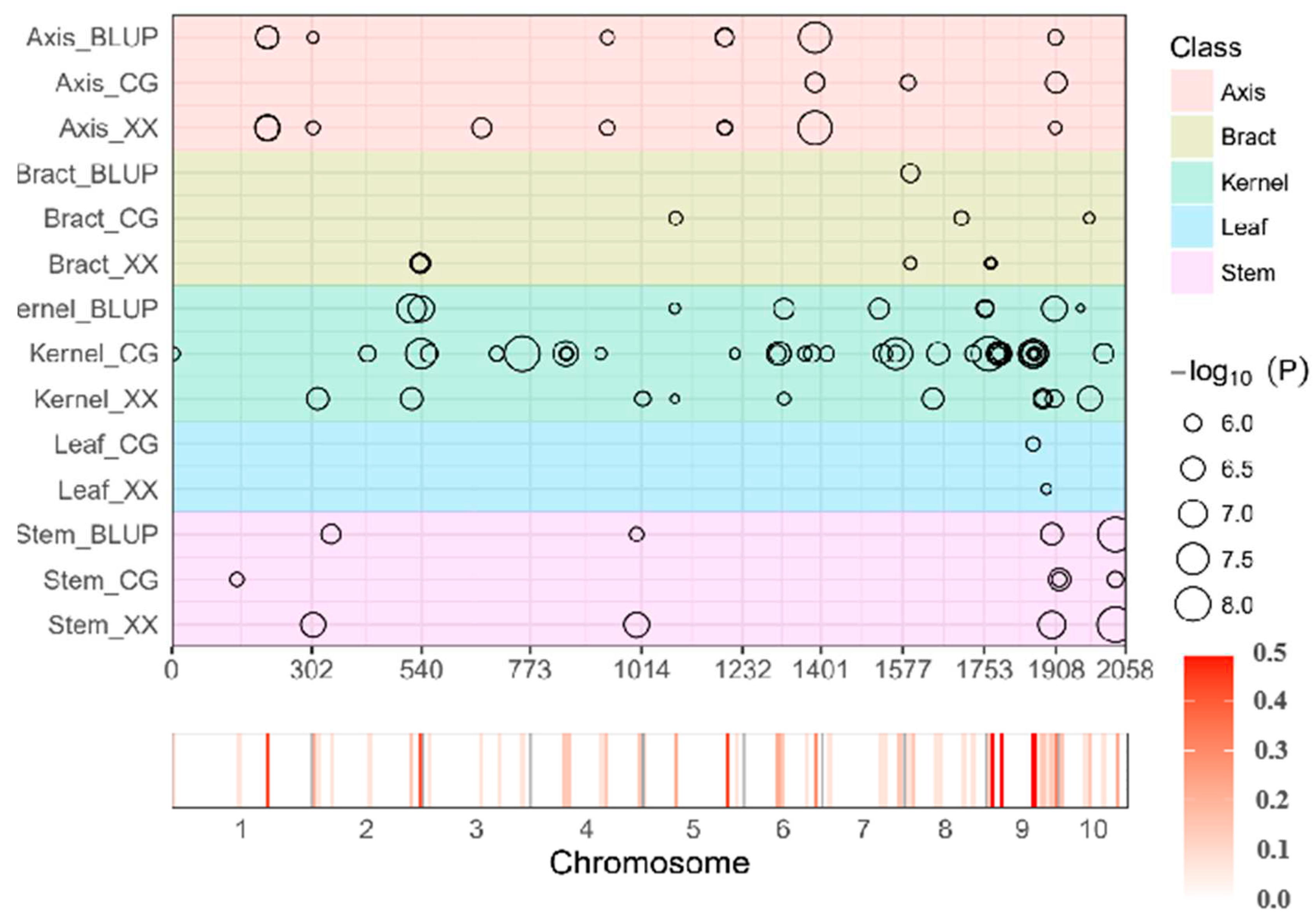
Simultaneous Identification of Significantly Mapped Loci Across Diverse Tissues and Environmental Conditions
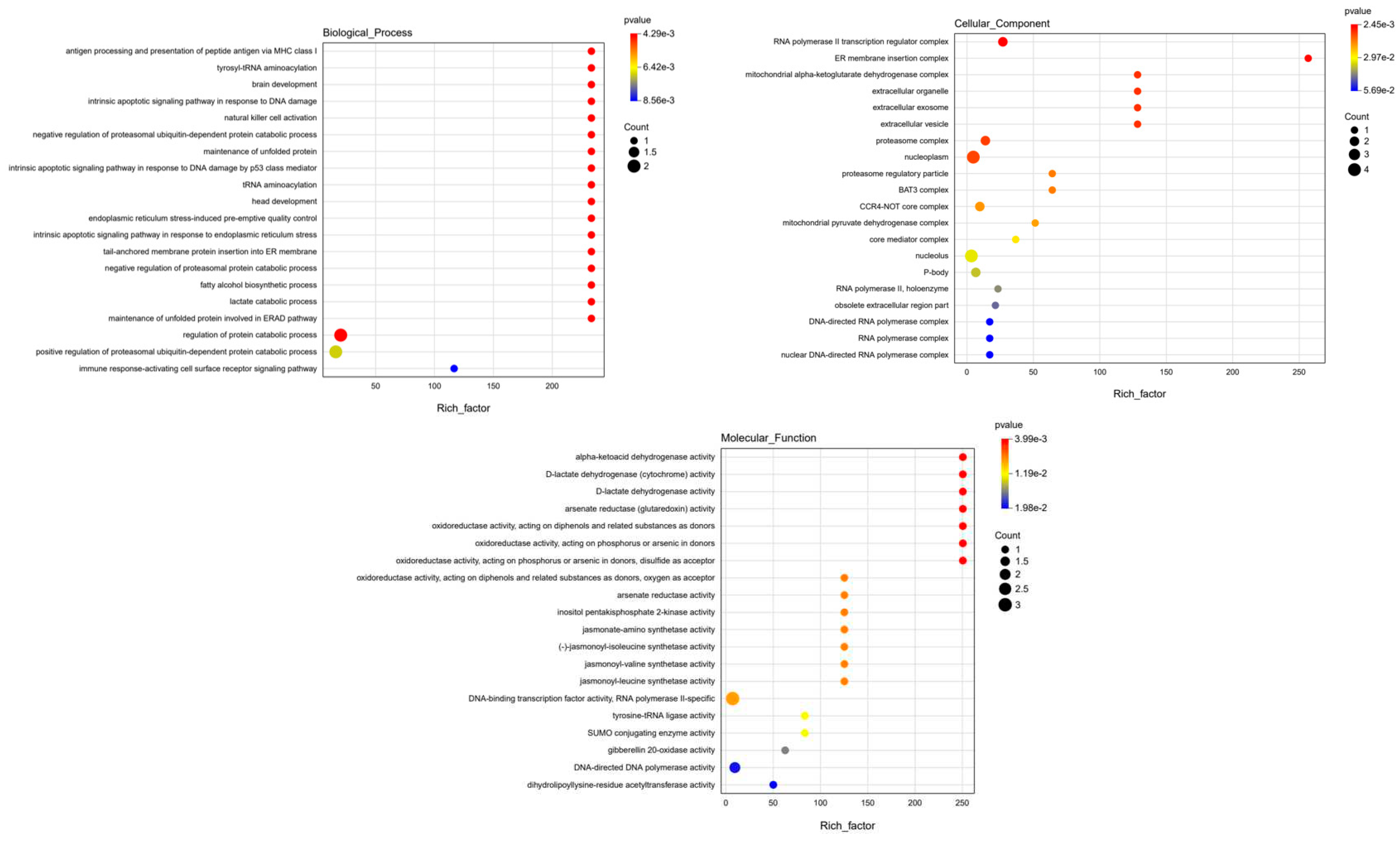
Functional Analysis of Identified Genes
Candidate gene analysis
Discussion
Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Rahman, Z.; Singh, V.P. The relative impact of toxic heavy metals (THMs) (arsenic (As); cadmium (Cd); chromium (Cr)(VI); mercury (Hg); and lead (Pb)) on the total environment: an overview. Environ Monit Assess. 2019, 191, 419. [CrossRef] [PubMed]
- Shao R, Zhang J, Shi W, Wang Y, Tang Y, Liu Z, Sun W, Wang H, Guo J, Meng Y, Kang G, Jagadish KS, Yang Q. Mercury stress tolerance in wheat and maize is achieved by lignin accumulation controlled by nitric oxide. Environ Pollut. 2022, 307, 119488. [CrossRef] [PubMed]
- Z. Boerleider R; Roeleveld N; T.J. Scheepers P. Human biological monitoring of mercury for exposure assessment. AIMS Environmental Science. 2017, 4, 251–276. [CrossRef]
- Wang C; Wang Z; Gao Y; Zhang X. Planular-vertical distribution and pollution characteristics of cropland soil Hg and the estimated soil-air exchange fluxes of gaseous Hg over croplands in northern China. Environ Res. 2021, 195, 110810.
- Zhang L; Wong MH. Environmental mercury contamination in China: sources and impacts. Environ Int. 2017, 33, 108-121.
- Sun T, Wang Z, Zhang X, Niu Z, Chen J. Influences of high-level atmospheric gaseous elemental mercury on methylmercury accumulation in maize (Zea mays L.). Environ Pollut. 2020;265(Pt B):114890.
- Liu S; Wang X; Guo G; Yan Z. Status and environmental management of soil mercury pollution in China: A review. J Environ Manage. 2021, 277, 111442.
- Rocha J; Aschner M; Dorea JG; Ceccatelli S; Farina M; Silveira L. Mercury toxicity. J Biomed Biotechnol. 2012, 831890.
- Wang C; Wang T; Mu P; Li Z; Yang L. Quantitative Trait Loci for Mercury Tolerance in Rice Seedlings. Rice Science. 2013, 20, 238–242.
- Fu Z; Li W; Zhang Q; Wang L; Zhang X; Song G; Fu Z; Ding D; Liu Z; Tang J. Quantitative trait loci for mercury accumulation in maize (Zea mays L.) identified using a RIL population. PLoS ONE 2014, 9, e107243.
- Hu T; Liu Y; Zhu S; Qin J; Li W; Zhou N. Overexpression of OsLea14-A improves the tolerance of rice and increases Hg accumulation under diverse stresses. Environ Sci Pollut Res Int. 2019, 26, 10537-10551.
- Sun L; Ma Y; Wang H; Huang W; Wang X; Han L; Sun W; Han E; Wang B. Overexpression of PtABCC1 contributes to mercury tolerance and accumulation in Arabidopsis and poplar. Biochem Biophys Res Commun. 2018, 497, 997-1002.
- Xu S; Sun B; Wang R; He J; Xia B; Xue Y; Wang R. Overexpression of a bacterial mercury transporter MerT in Arabidopsis enhances mercury tolerance. Biochem Biophys Res Commun. 2017, 490, 528-534.
- Yu J; Buckler ES. Genetic association mapping and genome organization of maize. Curr Opin Biotechnol.2006, 17, 155–160.
- Xiao Y; Liu H; Wu L; Warburton M; Yan J. Genome-wide Association Studies in Maize: Praise and Stargaze. Mol Plant. 2017, 10:359-374.
- Li H; Peng Z; Yang X; Wang W; Fu J; Wang J; Han Y; Chai Y; Guo T; Yang N; Liu J; Warburton ML; Cheng Y; Hao X; Zhang P; Zhao J; Liu Y; Wang G; Li J; Yan J. Genome-wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat Genet. 2013, 45:43-50.
- Beló A; Zheng P; Luck S; Shen B; Meyer DJ; Li B; Tingey S; Rafalski A. Whole genome scan detects an allelic variant of fad2 associated with increased oleic acid levels in maize. Mol Genet Genomics. 2008, 279, 1–10.
- Zhao Z; Zhang H; Fu Z; Chen H; Lin Y; Yan P; Li W; Xie H; Guo Z; Zhang X; Tang J. Genetic-based dissection of arsenic accumulation in maize using a genome-wide association analysis method. Plant Biotechnol J. 2018, 16, 1085-1093.
- Korte A, Farlow A. The advantages and limitations of trait analysis with GWAS: a review. Plant Methods 2013, 9, 29.
- Huang X, Zhao Y, Wei X, Li C, Wang A, Zhao Q, Li W, Guo Y, Deng L, Zhu C, Fan D, Lu Y, Weng Q, Liu K, Zhou T, Jing Y, Si L, Dong G, Huang T, Lu T, Feng Q, Qian Q, Li J, Han B. Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet. 2011, 44, 32-9.
- Wang SB, Feng JY, Ren WL, Huang B, Zhou L, Wen YJ, Zhang J, Dunwell JM, Xu S, Zhang YM. Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci Rep. 2016, 6, 19444.
- Wang Q, Tian F, Pan Y, Buckler ES, Zhang Z. A SUPER powerful method for genome wide association study. PLoS ONE 2014, 9, e107684.
- Li X, Zhu C, Yeh CT, Wu W, Takacs EM, Petsch KA, Tian F, Bai G, Buckler ES, Muehlbauer GJ, Timmermans MC, Scanlon MJ, Schnable PS, Yu J. Genic and nongenic contributions to natural variation of quantitative traits in maize. Genome Res. 2012, 22, 2436-44.
- Yang N; Lu Y; Yang X; Huang J; Zhou Y; Ali F; Wen W; Liu J; Li J; Yan J. Genome wide association studies using a new nonparametric model reveal the genetic architecture of 17 agronomic traits in an enlarged maize association panel. PLoS Genet. 2014, 10, e1004573. [CrossRef]
- Tian F; Bradbury P; Brown P; Hung H; Sun Q; Flint-Garcia S; Rocheford TR; McMullen MD; Holland JB; Buckler ES. Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat Genet. 2011, 43, 159-162.
- Zhao K; Tung CW; Eizenga GC; Wright MH; Ali ML; Price AH; Norton GJ; Islam MR; Reynolds A; Mezey J; McClung AM; Bustamante CD; McCouch SR. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun. 2011, 2, 467.
- Zhao Z; Fu Z; Lin Y; Chen H; Liu K; Xing X; Liu Z; Li W; Tang J. Genome-wide association analysis identifies loci governing mercury accumulation in maize. Sci Rep. 2017, 7, 247.
- Wang H; Xu S; Fan Y; Liu N; Zhan W; Liu H; Xiao Y; Li K; Pan Q; Li W; Deng M; Liu J; Jin M; Yang X; Li J; Li Q; Yan J. Beyond pathways: genetic dissection of tocopherol content in maize kernels by combining linkage and association analyses. Plant Biotechnol J. 2018, 16, 1464-1475. 1475.
- Liu H; Luo X; Niu L; Xiao Y; Chen L; Liu J; Wang X; Jin M; Li W; Zhang Q; Yan J. Distant eQTLs and Non-coding Sequences Play Critical Roles in Regulating Gene Expression and Quantitative Trait Variation in Maize. Mol Plant. 2017, 10, 414-426.
- Li Q; Yang X; Xu S; Cai Y; Zhang D; Han Y; Li L; Zhang Z; Gao S; Li J; Yan J. Genome-wide association studies identified three independent polymorphisms associated with alpha-tocopherol content in maize kernels. PLoS One 2012, 7, e36807.
- Fu J; Cheng Y; Linghu J; Yang X; Kang L; Zhang Z; Zhang J; He C; Du X; Peng Z; Wang B; Zhai L; Dai C; Xu J; Wang W; Li X; Zheng J; Chen L; Luo L; Liu J; Qian X; Yan J; Wang J; Wang G. RNA sequencing reveals the complex regulatory network in the maize kernel. Nat Commun. 2013, 4, 2832.
- Elshire R; Glaubitz J; Sun Q; Poland J; Kawamoto K; Buckler E; Mitchell S. A robust; simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 2011, 6, e19379.
- Deng M; Li D; Luo J; Xiao Y; Liu H; Pan Q; Zhang X; Jin M; Zhao M; Yan J. The genetic architecture of amino acids dissection by association and linkage analysis in maize. Plant Biotechnol J. 2017, 15, 1250-1263.
- Liu H; Wang X; Warburton ML; Wen W; Jin M; Deng M; Liu J; Tong H; Pan Q; Yang X; Yan J. Genomic; Transcriptomic; and Phenomic Variation Reveals the Complex Adaptation of Modern Maize Breeding. Mol Plant. 2015, 8, 871-884.
- iu B; Zhang B; Yang Z; Liu Y; Yang S; Shi Y; Jiang C; Qin F. Manipulating ZmEXPA4 expression ameliorates the drought-induced prolonged anthesis and silking interval in maize. Plant Cell. 2021, 33, 2058-2071. 2071.
- Huang X; Wei X; Sang T; Zhao Q; Feng Q; Zhao Y; Li C; Zhu C; Lu T; Zhang Z; Li M; Fan D; Guo Y; Wang A; Wang L; Deng L; Li W; Lu Y; Weng Q; Liu K; Huang T; Zhou T; Jing Y; Li W; Lin Z; Buckler ES; Qian Q; Zhang QF; Li J; Han B. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet. 2010, 42, 961–967.
- Khorchid A; Ikura M. Bacterial histidine kinase as signal sensor and transducer. International. Int J Biochem Cell Biol. 2006, 38, 307-312.
- Yu X; Meng X; Xu M; Zhang X; Zhang Y; Ding G; Huang S; Zhang A; Jia Z. Celastrol ameliorates cisplatin nephrotoxicity by inhibiting NF-kappaB and improving mitochondrial function. EBioMedicine 2018, 36, 266-280.
- Stratton A, Ericksen M, Harris TV, Symmonds N, Silverstein TP. Mercury(II) binds to both of chymotrypsin's histidines, causing inhibition followed by irreversible denaturation/aggregation. Protein Sci. 2017;26(2):292-305.
- Cao Y; Ma C; Chen H; Chen G; White JC; Xing B. Copper stress in flooded soil: Impact on enzyme activities; microbial community composition and diversity in the rhizosphere of Salix integra. Sci Total Environ. 2020, 704, 135350.
- Liu YN; Liu BY; Ma YC; Yang HL; Liu GQ. Analysis of reference genes stability and histidine kinase expression under cold stress in Cordyceps militaris. PLoS One 2020, 15, e0236898.
- Kumar M; Verslues P. Stress physiology functions of the Arabidopsis histidine kinase cytokinin receptors. Physiol Plant. 2015, 154, 369-380.
- Abbehausen C. Zinc finger domains as therapeutic targets for metal-based compounds - an update. Metallomics. 2019, 11, 15-28.
- Banerjee M; Ferragut Cardoso AP; Lykoudi A; Wilkey DW; Pan J; Watson WH; Garbett NC; Rai SN; Merchant ML; States JC. Arsenite Exposure Displaces Zinc from ZRANB2 Leading to Altered Splicing. Chem Res Toxicol. 2020, 33, 1403-1417.
- Razmiafshari M; Kao J; d'Avignon A; Zawia NH. NMR identification of heavy metal-binding sites in a synthetic zinc finger peptide: toxicological implications for the interactions of xenobiotic metals with zinc finger proteins. Toxicol Appl Pharmacol. 2001, 172, 1-10.
- Sivo V; D'Abrosca G; Baglivo I; Iacovino R; Pedone PV; Fattorusso R; Russo L; Malgieri G; Isernia C. Ni(II); Hg(II); and Pb(II) Coordination in the Prokaryotic Zinc-Finger Ros87. Inorg Chem. 2019, 58, 1067-1080.
- Zhang X; Warburton ML; Setter T; Liu H; Xue Y; Yang N; Yan J; Xiao Y. Genome-wide association studies of drought-related metabolic changes in maize using an enlarged SNP panel. Theor Appl Genet. 2016, 129, 1449-63.
- Tibbs Cortes L; Zhang Z; Yu J. Status and prospects of genome-wide association studies in plants. Plant Genome. 2021, 14, e20077.
- Liao X; Sun J; Li Q; Ding W; Zhao B; Wang B; Zhou S; Wang H. ZmSIZ1a and ZmSIZ1b play an indispensable role in resistance against Fusarium ear rot in maize. Mol Plant Pathol. 2023, 24, 711-724.
- Kumari N; Jagadevan S. Genetic identification of arsenate reductase and arsenite oxidase in redox transformations carried out by arsenic metabolising prokaryotes - A comprehensive review. Chemosphere 2016, 163, 400-412.
- An B; Lan J; Deng X; Chen S; Ouyang C; Shi H; Yang J; Li Y. Silencing of D-Lactate Dehydrogenase Impedes Glyoxalase System and Leads to Methylglyoxal Accumulation and Growth Inhibition in Rice. Front Plant Sci. 2017, 8, 2071.
- Sun Y; Xu W; Wu L; Wang R; He Z; Ma M. An Arabidopsis mutant of inositol pentakisphosphate 2-kinase AtIPK1 displays reduced arsenate tolerance. Plant Cell Environ. 2016, 39, 416-426.
- Maruyama K; Yorifuji T; Tsuda T; Sekikawa T; Nakadaira H; Saito H. Methyl mercury exposure at Niigata; Japan: results of neurological examinations of 103 adults. J Biomed Biotechnol. 2012, 635075.
- Huang C; Liu S; Lin-Shiau S. Neurotoxicological effects of cinnabar (a Chinese mineral medicine; HgS) in mice. Toxicol Appl Pharmacol. 2007, 224, 192-201.
- Canabady-Rochelle LL; Harscoat-Schiavo C; Kessler V; Aymes A; Fournier F; Girardet JM. Determination of reducing power and metal chelating ability of antioxidant peptides: revisited methods. Food Chem. 2015, 183, 129-135.
- Torres-Fuentes C; Alaiz M; Vioque J. Iron-chelating activity of chickpea protein hydrolysate peptides. Food Chem. 2012, 134, 1585-1588.
- Chang J; Zhou Y; Wang Q; Aschner M; Lu R. Plant components can reduce methylmercury toxication: A mini-review. Biochimica et biophysica acta. Biochim Biophys Acta Gen Subj. 2019, 1863, 129290.
- Li Y; Takada M; Mizuno N. Premotor neurons projecting simultaneously to two orofacial motor nuclei by sending their branched axons. A study with a fluorescent retrograde double-labeling technique in the rat. Neurosci Lett. 1993, 152, 29–32.
| ID | Chr. | Position | Trait | Location | SNP | P value | R2 | Candidate gene | Annotation | Expressed or not | Verified by expression GWAS |
| 1 | 4 | 167403585 | Axis | BLUP | chr4.S_167403585 | 1.54E-06 | 0.176900204 | GRMZM2G125943 | histidine kinase | express | Ture |
| GRMZM2G125991 | endoglucanase 7-like | express | Ture | ||||||||
| GRMZM2G029859 | pentatricopeptide repeat-containing protein At3g16610 | express | Ture | ||||||||
| 2 | 4 | 167403585 | Axis | XX | chr4.S_167403585 | 1.33E-06 | 0.177900282 | GRMZM2G125943 | histidine kinase | express | Ture |
| GRMZM2G125991 | endoglucanase 7-like | express | Ture | ||||||||
| GRMZM2G029859 | pentatricopeptide repeat-containing protein At3g16610 | express | Ture | ||||||||
| 3 | 5 | 2356674 | Kernel | XX | chr5.S_2356674 | 1.31E-06 | 0.096393321 | GRMZM2G002825 | actin-depolymerizing factor 3 | express | Ture |
| GRMZM2G002805 | zinc finger protein ZAT5 | express | Ture | ||||||||
| GRMZM2G002815 | NA | express | Ture | ||||||||
| GRMZM2G144188 | dof zinc finger protein DOF2.4-like | not | |||||||||
| GRMZM2G144172 | dof zinc finger protein DOF2.4-like | express | Ture | ||||||||
| GRMZM2G003068 | NA | express | Ture | ||||||||
| GRMZM2G003108 | CRAL/TRIO domain containing protein | express | Ture | ||||||||
| 4 | 6 | 155668107 | Axis | BLUP | chr6.S_155668107 | 2.82E-08 | 0.127072513 | GRMZM2G566873 | NA | not | |
| GRMZM2G140805 | NA | not | |||||||||
| GRMZM2G440949 | dr1-associated corepressor | express | Ture | ||||||||
| GRMZM2G440968 | cystatin 3 | express | Ture | ||||||||
| GRMZM2G140817 | putative cytochrome P450 superfamily protein | express | Ture | ||||||||
| 5 | 6 | 155668107 | Axis | CG | chr6.S_155668107 | 6.77E-07 | 0.103439562 | GRMZM2G566873 | NA | not | |
| GRMZM2G140805 | NA | not | |||||||||
| GRMZM2G440949 | dr1-associated corepressor | express | Ture | ||||||||
| GRMZM2G440968 | cystatin 3 | express | Ture | ||||||||
| GRMZM2G140817 | putative cytochrome P450 superfamily protein | express | Ture | ||||||||
| 6 | 6 | 155668107 | Axis | XX | chr6.S_155668107 | 1.46E-08 | 0.131892015 | GRMZM2G566873 | NA | not | |
| GRMZM2G140805 | NA | not | |||||||||
| GRMZM2G440949 | dr1-associated corepressor | express | Ture | ||||||||
| GRMZM2G440968 | cystatin 3 | express | Ture | ||||||||
| GRMZM2G140817 | putative cytochrome P450 superfamily protein | express | Ture | ||||||||
| 7 | 10 | 127359876 | Stem | BLUP | chr10.S_127359876 | 8.01E-09 | 0.232693323 | GRMZM2G005633 | Endochitinase B | express | Ture |
| GRMZM2G006428 | NA | express | Ture | ||||||||
| GRMZM2G006216 | S-adenosyl-L-methionine-dependent methyltransferase superfamily protein | express | Ture | ||||||||
| GRMZM2G005939 | NA | express | Ture | ||||||||
| 8 | 10 | 127359876 | Stem | CG | chr10.S_127359876 | 1.11E-06 | 0.171467848 | GRMZM2G005633 | Endochitinase B | express | Ture |
| GRMZM2G006428 | NA | express | Ture | ||||||||
| GRMZM2G006216 | S-adenosyl-L-methionine-dependent methyltransferase superfamily protein | express | Ture | ||||||||
| GRMZM2G005939 | NA | express | Ture | ||||||||
| 9 | 10 | 127359876 | Stem | XX | chr10.S_127359876 | 5.81E-09 | 0.237084701 | GRMZM2G005633 | Endochitinase B | express | Ture |
| GRMZM2G006428 | NA | express | Ture | ||||||||
| GRMZM2G006216 | S-adenosyl-L-methionine-dependent methyltransferase superfamily protein | express | Ture | ||||||||
| GRMZM2G005939 | NA | express | Ture |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




