1. Introduction
Ovarian cancer (OC) is one of the most prevalent malignant tumors in gynecology and the highest mortality in gynecological tumors. The majority of patients present with an insidious onset, and 75% of patients are detected at a late stage due to ineffective diagnosis [
1]. Surgery and chemotherapy remain the cornerstones of OC treatment, and targeted agents appear to have provided little improvement to overall survival while substantially creating clinical adverse effects [
2,
3]. Additionally, OC still has a terrible prognosis, with a 30-50% five-year survival rate [
4]. Existing research has revealed that the five-year overall survival rate is about 92% for early patients and 29% for advanced patients [
5]. Therefore, the discovery of novel prognostic or therapeutic indicators for OC is crucial.
The SET domain containing lysine methyltransferase 7 (SETD7), also called SET7/9, KIAA1717, or KMT7, was first identified as the histone H3 lysine 4-specific (H3K4) methyltransferase [
6]. SETD7 contains one SET domain and three membrane occupation and recognition (MOR) motifs required for enzymatic activity and membrane occupation and recognition of linkages, respectively [
7]. Recently, several basic proteins associated with the development of cancer have been identified as non-histone substrates of SETD7, including P53 [
8], ERα [
9], β-catenin [
10], and NF-κB p65 [
11]. This methylation produces variable results. While methylation of p53 or ERα by SETD7 stabilizes these proteins and promotes transcriptional activity [
8,
9], methylation of β-catenin encourages its proteasomal degradation [
10]. Additionally, methylation of various lysines within a protein can have a variety of effects that offer a complex level of crosstalk [
11].
Given that multiple proteins can prominently regulated by SETD7, SETD7 may play a variety of roles in tumor progression. Existing studies have shown that SETD7 plays a role in tumor progression [
12]. For instance, SETD7 was markedly overexpressed in 80 breast cancer (BC) samples, and upregulation of SETD7 promoted the proliferation, migration, and invasion of BC cells [
13,
14]. Knockdown of SETD7 dramatically diminished the migration ability of hepatoma cells [
15]. Additionally, SETD7 was also discovered to be a target of miR-153. By downregulating SETD7, miR-153 prevented the proliferation and invasive potential of OC cells [
16]. A tumor suppressor role for SETD7, however, is supported by new evidence. SETD7 can inhibit epithelial-mesenchymal transition (EMT) by increasing expression of Cadherin-1 and decreasing Vimentin and EGFR expression in BC cells [
17]. Additionally, methylation of RIOK1 by SETD7 caused its destruction to prevent colorectal cancer (CRC) cell growth and metastasis [
18]. Nevertheless, it is unclear how SETD7 affects OC.
In this study, we found that SETD7 protein was upregulated in human OC tissues and cell lines, and high SETD7 protein levels were associated with M category and FIGO stage. Moreover, the high mRNA levels of SETD7 were correlated to poor progression-free survival (PFS) in all patients and serous patients with OC. Of note, the overexpression of SETD7 significantly facilitated the proliferation and migration of SKOV3 cells, while the knockdown of SETD7 suppressed these effects of A2780 cells. Furthermore, SETD7 might enhance cell migration via accelerating the EMT process in A2780 and SKOV3 cells. Our findings suggest that SETD7 is a key trigger of OC progression, which is essential for the creation of effective treatment options.
2. Materials and Methods
2.1. Antibodies and Reagents
SETD7 (24840-1-AP), E-cadherin (20874-1-AP), and Snail (13099-1-AP) antibodies were purchased from Proteintech (Wuhan, China). N-cadherin (D4R1H) and Vimentin (D21H3) antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA). Tubulin (BS1699) and GAPDH (AP0063) antibodies were obtained from Bioworld Technology (Nanjing, China). Histone H3 antibody (ab1791) was obtained from Abcam (Cambridge, MA, USA). Mouse (31430) and Rabbit (31460) HRP-conjugated secondary antibodies were obtained from ThermoFisher Scientific (MA, USA). Puromycin (A11138-03) was purchased from Invitrogen (Carlsbad, CA, USA). Transwell chamber (TCS020024) was purchased from Jet Bio-Filtration (Guangzhou, China). Cell Counting Kit (CCK)-8 (A311-01) was purchased from Vazyme Biotech (Nanjing, China). The DAB kit, SABC-POD kit, and Mayor’s hematoxylin were obtained from Boster Biological Technology (Wuhan, China).
2.2. Cell Lines and Cell Culture
Human ovarian granulosa cell line KGN and OC cell lines A2780, SKOV3, Hey A8, and ES-2 were obtained from the American Type Culture Collection (ATCC, Shanghai, China) and STR-authenticated by Shanghai Biowing Applied Biotechnology Co. LTD (Shanghai, China). The A2780s cell line originated from undifferentiated ovarian endometrioid adenocarcinoma. The SKOV3, Hey A8, and ES-2 cell lines originated from poorly differentiated ovarian serous cystadenocarcinoma. Every cell line was cultivated in monolayers using the proper media: A2780 cells were grown in RPMI 1640 medium; SKOV3 and ES-2 cells were grown in McCoy's 5A medium; KGN and Hey A8 cells were grown in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum, 100 U/mL penicillin, and 100 μg/mL streptomycin (P/S). Every cell was kept in an atmosphere with 5% CO2 at 37 °C.
2.3. Ethics Statement
The Medical Ethics Committee of Jiangsu University (approval code: JSDX20210303001) approved this study. Human OC tissue specimens for this study were collected after obtaining informed patient consent and the use of the OC specimens was endorsed by The Ethics Committee of Zhuoli Biotechnology (approval code: SHLLS-BA-22101102).
2.4. Human OC Specimen Analysis
Human OC tissue microarray (ZL-OVA961) was created by Shanghai Zhuoli Biotech Company (Shanghai, China) in the form of 4 μm sections of paraffin-embedded tissue on glass slides. The microarray comprised 96 tissue samples, consisting of 49 cases of serous adenocarcinoma, 14 cases of endometrioid carcinoma, 9 cases of clear cell carcinoma, 6 cases of ovarian metastasis of gastric and intestinal cancer, 5 cases of borderline mucinous tumor with intramucosal carcinoma, 4 cases of mucinous adenocarcinoma, 1 case of squamous cell carcinoma, and 8 cases of normal ovary tissues.
To investigate the SETD7 expression profile, the microarray sections were subjected to immunohistochemistry (IHC). The sections were rehydrated in graded ethanol after being deparaffinized with xylene. To retrieve the antigen, the sections were cooked in a citrate buffer (pH 6.0) at 125 °C for 30 sec and 90 °C for 10 sec using a pressure cooker. Protein blocking reagents and peroxidase were incubated for 5 min in each slice. Following that, the sections were treated for 1 h at room temperature with rabbit polyclonal SETD7 antibody (1:100, 24840-1-AP). After incubating with the primary antibody, the sections were treated for 30 minutes at room temperature with HRP-conjugated goat anti-rabbit IgG. The DAB chromogen kit was utilized to develop each section, and hematoxylin was used as a mild counterstain [
19].
The score of the IHC signals was performed using a double-blind reading method by a pathology expert to interpret the results. The expression levels of SETD7 were scored by multiplying staining intensity (0, none; 1, weak; 2, moderate; 3, strong) and the proportion of stained cells (0, ≤5%; 1, 5-25%; 2, 26-50%; 3, 51-75%; 4, ≥76%). SETD7 expression levels were defined as low (score, 0-4) or high (score, 5-12).
2.5. The Prognostic Value of SETD7 Analysis
Kaplan-Meier plotter (
https://kmplot.com/analysis) is an online database containing microarray gene expression data and survival information derived from Gene Expression Omnibus, TCGA, and the Cancer Biomedical informatics Grid. In this study, the prognostic value of the mRNA expression of SETD7 was evaluated using the Kaplan-Meier plotter. The overall survival (OS), progression-free survival (PFS), and post-progression survival (PPS) of patients with OC were determined by dividing the patient samples into two groups based on median expression (high vs. low expression) and assessing using a Kaplan-Meier survival plot, with a hazard ratio with 95% confidence intervals and logrank P-value. Subgroup analyses were performed by dividing patients based on pathological and histological subtypes.
2.6. qRT-PCR
Total RNA was extracted using the RNAiso plus (Takara Bio, Shiga, Japan) and processed using the PrimeScript RT Reagent kit (Takara Bio) to create cDNA according to the manufacturer’s instructions. The cDNA was then subjected to qRT-PCR as previously mentioned [
20]. The qRT-PCR was performed on a Bio-Rad CFX96 system (Bio-Rad Laboratories, Inc., Hercules, CA, USA) using the comparative cycle threshold (Ct) method and the SYBR-Green PCR Master Mix (Takara Bio) according to the product specification. The primer sequences were as follows:
SETD7 (GenBank accession no. NM_001306200), 5'-TCACCTACTCCTCCACAGAC-3' (forward) and 5'-TCATCCACATAATACCCCTCCAG-3' (reverse),
GAPDH (GenBank accession no. NM_001256799.3), 5'-CACCAGGGCTGCTTTTAACTC-3' (forward) and 5'-CTTGACGGTGCCATGGAATTTG-3' (reverse). The PCR parameters were as follows: first denaturalization at 95 °C for 30 sec, then denaturalization for 40 sec at 95 °C for 5 sec, annealing at 58 °C for 30 sec, and elongation at 72 °C for 60 sec. With
GAPDH serving as the reference gene, the relative results were examined using the comparative cycle threshold approach (2
-ΔΔCt).
2.7. Western Blotting
Proteins were extracted with the radioimmunoprecipitation assay (RIPA) buffer added with proteinase inhibitors, including aprotinin and leupeptin (Biotopped, Beijing, China). The protein concentration was determined by colorimetric detection with a bicinchoninic acid (BCA) assay (ThermoFisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol. Briefly, protein sample or BSA standard (10 μL) was loaded into a microplate. Working solution (200 μL) was added to each sample and the microplate was incubated for 30 min at 37 °C. The absorbance was measured at 562 nm using a microplate reader (EL808IU-PC, BioTek, Winooski, VT, USA).
For gel electrophoresis, the proteins (30 μg) were detached on 10% sodium dodecyl sulfate-polyacrylamide (SDS) gels. The gels were blotted onto the polyvinylidene fluoride (PVDF; Millipore, MA, USA) membranes. Following a brief wash in tris-buffered saline (TBS), the membrane was blocked with 5% BSA in TBS supplemented with 0.1% Tween-20 (TBS-T). Thereafter, the membrane was incubated with the primary antibodies against SETD7 (1:2000), E-cadherin (1:5000), Snail (1:750), N-cadherin (1:1000), Vimentin (1:1000), Tubulin (1:10,000), GAPDH (1:1000), or Histone H3 (1:1000). Following a brief wash, the membranes were incubated with peroxidase-bound anti-rabbit or anti-mouse IgG (1:10,000) in TBS-T for 2 h at 37 °C, and the ECL western blotting substrate (Thermo Scientific) was used for detection. The proteins were visualized by electrochemiluminescence (Pierce, Rockford, IL, USA) and detected with a bio-imaging system (Fusion Solo, Vilber, France).
2.8. Cell Proliferation Assay
A2780 and SKOV3 cells (3-5 × 104) were inoculated in 12-well plates and cultivated for the allotted time. Using a hemocytometer and a microscope with 100× magnification, the cells in six randomly selected fields were counted after 24 h, 48 h, 72 h, and 96 h. By comparing the increase in cell count since seeding, cell proliferative activity was evaluated. Each specimen was analyzed twice, and three independent experiments were carried out.
2.9. Cell Growth Analysis
Cell viability was evaluated using the CCK-8 assay (Vazyme Biotech, Nanjing, China). In 96-well plates, A2780 and SKOV3 cells (3-5 × 104) were added, and the cells were then pre-cultured in the incubator for 24 h. Following that, 100 µL of CCK-8 (10%) was poured to each hole, and cultured for 2 h at 37°C. A microplate reader measured the absorbance at 450 nm.
2.10. Transwell Assay
OC cell migration was evaluated as previously described (21). Serum-free medium containing A2780 and SKOV3 cells (6 × 103) was planted into the upper Transwell chamber (Jet Bio-Filtration, Guangzhou, China), while medium with 20% FBS served as a chemoattractant in the lower chamber. After 48 h of incubation at 37°C and 5% CO2, the cells in the top chamber were cleaned with a cotton swab, while the cells on the lower layer were stained for 15 min with 1% crystal violet. Microscopical (200×) measurements of the migrated cells were carried out in five random fields, and the results were averaged.
2.11. Wound-Healing Assay
A2780 and SKOV3 cells were sown in 6-well plates and cultivated until they reached 100% confluence. Mitomycin C (10 μg/mL) was administered for 2 h to prevent cell proliferation. A 200-μL pipette tip was used to scratch the single layer of cells to make the artificial linear wounds. The breadth of the scratch gap was observed and photographed at 24 h or 72 h using an inverted microscope (21). ImageJ software was used to measure the distance the cells traveled into the wound areas. The migration rate was calculated as follows: % of scratch closure = [(initial wound length) − (wound length at 24/72 h)] / (initial wound length) × 100.
2.12. Establishment of Stable SETD7 Overexpressing Cell Line
The pCDH-SETD7 expression vector was created by introducing the full-length SETD7 cDNA (GenBank accession number NM_030648) into pCDH-CMV-MCS-EF1-Puro vector (pCDH). In order to generate a cell line that overexpresses either pCDH or SETD7, HEK293T cells were infected with pHR'-CMV-VSVG (0.5 μg), pHR'-CMV-8.2VPR (1.5 μg), pCDH-SETD7 (2 μg) or pCDH (2 μg) using Lipofectamine 8000 reagent. Twenty-four hours after transfection, the virus was separated from the viral supernatant and used to infect SKOV3 cells, which had low SETD7 expression. Following two infection rounds, the SKOV3 cells were selected with 2 μg/mL puromycin for three days. Following this, they were maintained alive for one week using 1 μg/mL puromycin. The stability of the clones was verified by western blotting analysis [
22].
2.13. Generation of Stable SETD7 Knockdown Cell Line
SETD7 short hairpin RNAs (shRNAs) oligos were manufactured by Sangon Biotech Company (Shanghai, China). The specific sequences of SETD7 shRNAs were as follows: shSETD7-1#, GGGAGTTTACACTTACGAAGA; shSETD7-2#, GGACCGCACTTTATGGGAAAT; shSETD7-3#, GCAAACTGGCTACCCTTATGT. A scrambled shRNA sequence (shNC: CCTAAGGTTAAGTCGCCCTCG; Addgene #136035) with no homology to any known human gene was used as negative control. Subsequently, the SETD7 shRNAs or scrambled shRNA sequences were digested using the AgeI and EcoRI restriction enzymes, and cloned into the pLKO.1 vector (Addgene).
As described earlier [
21], a stable cell line expressing shNC of shRNAs targeting SETD7 was generated. Briefly, A2780 cells that expressed high levels of SETD7 were infected with non-target or SETD7-specific shRNA lentiviral particles in 6-well plates in the presence of 6 µg/mL polybrene. The transduced cells were selected with 2 µg/mL puromycin for one week until stable clones were established. For one week, the transduced cells were selected with 2 µg/mL puromycin until stable clones were formed. The levels of SETD7 knockdown were verified by western blotting.
2.14. Statistical Analysis
Statistical analysis was performed using Prism 8 (GraphPad Software). Data were shown as mean ± standard deviation (SD). The relationship between the levels of SETD7 and clinicopathologic characteristics was evaluated by Chi-square test. The data from the migration tests were analyzed with Two-tailed Student’s t test for Figs. 3B, 3F, 4, and 5. The data from the proliferation and viability tests were evaluated by two-way ANOVA for Fig. 3C, 3D, 3G, and 3H. For every analysis, more than two independent tests were conducted. A value of p < 0.05 was considered statistically significant.
3. Results
3.1. SETD7 Expression Is Upregulated in OC Tissues and Cell Lines
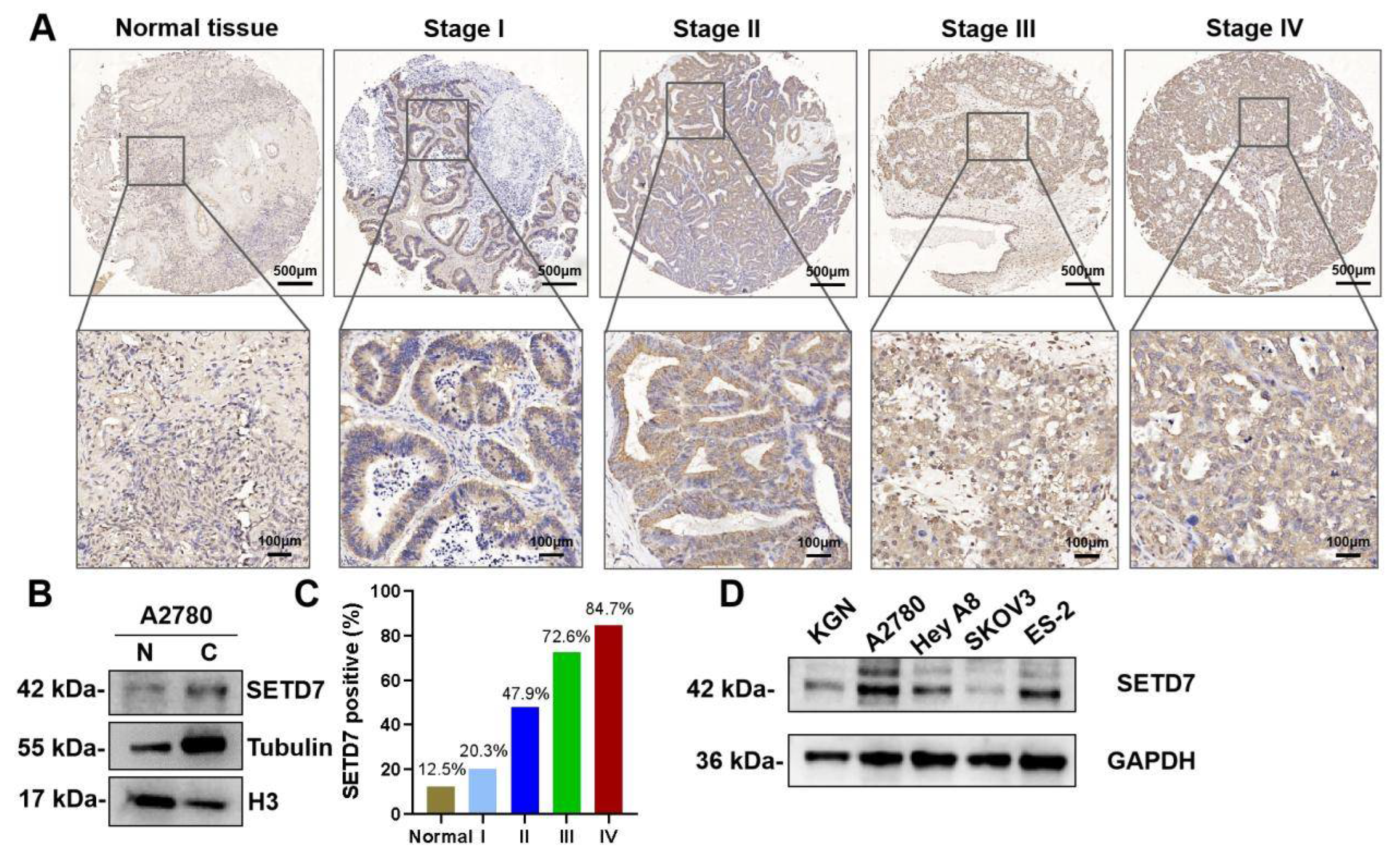
We first used immunohistochemistry (IHC) to assess SETD7 expression in the tissue microarray made up of 88 human OC and 8 normal ovary tissues. IHC showed that SETD7 expression was upregulated in OC tissues and mainly located in the cytoplasm (
Figure 1A). The nuclear and cytoplasmic separation experiment also confirmed the cytoplasmic distribution of SETD7 (
Figure 1B). In addition, the proportion of cells expressing SETD7 varied between 20.3%, 47.9%, 72.6%, and 84.7% in stages I, II, III, and IV of OC, respectively (
Figure 1C), displaying a connection between SETD7 expression and the malignancy of OC. To further confirm the expression of SETD7 in OC tissues, we measured levels of SETD7 protein in various OC cell lines. The higher expression levels of SETD7 were detected in A2780, Hey A8, and ES-2 cell lines than in normal ovarian granulosa cell line KGN (
Figure 1D). Importantly, the microarray analysis showed that 69 out of 88 OC patients (78%) exhibited strong staining for anti-SETD7, while 19 tumor samples (21%) showed weak or no staining (
Table 1). In addition, we studied the relationship between the expression of SETD7 and the pathology of OC. Our analysis revealed a significant correlation between SETD7 levels and the M category and FIGO stage (
p < 0.05;
Table 2). Overall, our data suggest that the expression of SETD7 is elevated in OC tissues and cell lines, and this increase has a positive correlation with malignancy.
3.2. SETD7 Overexpression Is Associated with Poor Prognosis in OC
Given that SETD7 is significantly overexpressed in OC tissues and cell lines, we speculate that SETD7 expression may be related to the prognosis of OC patients. KM plotter analysis (
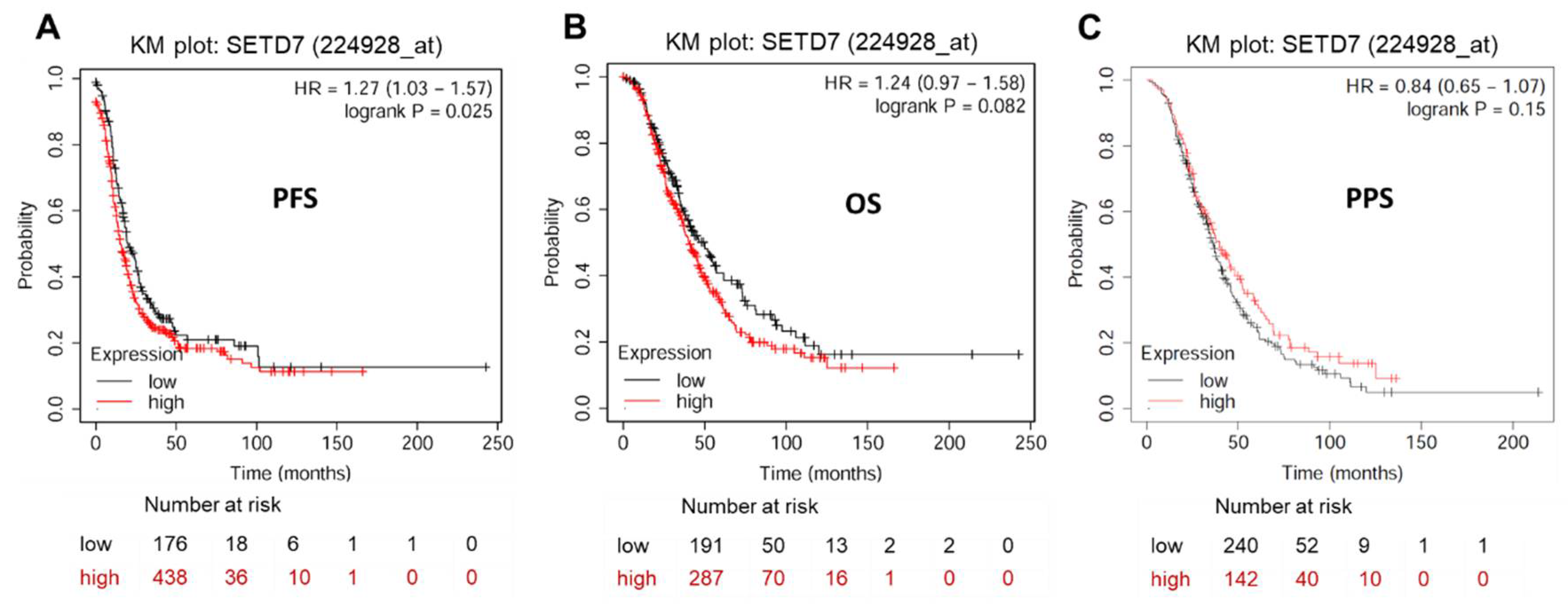
https://kmplot.com/analysis/index.php?p=service&cancer=ovar) exhibited a significant correlation between elevated SETD7 expression and worsened PFS (HR = 1.27, 95% CI: 1.03-1.57,
P < 0.05), but not OS (HR = 1.24, 95% CI: 0.97-1.58,
p > 0.05) and PPS (HR = 0.84, 95% CI: 0.65-1.07,
p > 0.05) in OC patients (
Figure 2 and
Table 3). To yield a further investigation, we assessed the correlation of SETD7 with different histological subtypes of OC, including serous and endometrioid. The high mRNA expression levels of SETD7 were correlated to poor PFS (HR = 1.22, 95% CI: 0.93-1.47,
p < 0.05) in serous OC patients (
Table 3). Data to calculate PPS in patients with endometrial OC were not sufficient. Collectively, our results suggest that SETD7 might function as a prognostic biomarker and contribute to the progression of OC.
3.3. SETD7 Promotes the Proliferation of OC Cells
Based on the association of SETD7 expression with inferior survival in OC patients, we set out to characterize the effects of SETD7 on OC progression. When compared to the normal ovarian granulosa cells KGN, the levels of endogenous SETD7 were clearly greater in the A2780 cells and lower in the SKOV3 cells (
Figure 1E). Therefore, we performed lentivirus-mediated knockdown of SETD7 (SETD7 KD) in the A2780 cells using SETD7-specific shRNAs or overexpression of SETD7 cDNA (SETD7 OE) in the SKOV3 cells. Afterwards, three SETD7-specific shRNAs (shSETD7) were transfected into the A2780 cells to decrease the expression of SETD7. Based on the results of western blotting and qRT-PCR, shSETD7-1 and shSETD7-2, which exhibited higher inhibition rates, were chosen for subsequent functional experiments (
Figure 3A,B). The CCK8 and cell counting assays were used to probe cell proliferation. Interestingly, SETD7 KD significantly blocked the proliferation of A2780 cells (
Figure 3C,D;
p < 0.05). In contrast, SETD7 OE resulted in opposite effects on the proliferation of SKOV3 cells (
Figure 3E-H). Collectively, these results suggest that SETD7 may act as an oncogene in OC.
3.4. SETD7 Promotes the Migration of OC Cells
Afterwards, we insight into the impact of SETD7 on OC cell migration. First, cell migration in Vector and SETD7 KD A2780 cells was investigated using the Transwell assay. SETD7 KD dramatically reduced cell migration at 48 h (
Figure 4A;
p < 0.001), in line with its oncogenic activity (
Figure 3). Subsequently, the wound-healing assay was utilized to confirm the aforementioned findings. As seen in
Figure 4B, SETD7 KD cells traveled noticeably less toward the wound center than did shNC cells (
p < 0.001). Conversely, SETD7 OE SKOV3 cells had opposite effects on cell migration (
Figure 4C,D;
p < 0.001). Together, these findings indicate that SETD7 enhances the migration of A2780 and SKOV3 cells.
3.5. SETD7 Positively Regulates Cell Migration via EMT in OC Cells
EMT, characterized by loss of epithelial markers such as E-cadherin and an increase in mesenchymal markers such as Snail, is a crucial stage in the spread of cancer [
23]. So, we screened EMT-associated proteins to see if the EMT was involved in SETD7-induced cell migration using a western blotting approach. In line with aforementioned biological experiments, SETD7 knockdown in A2780 cells decreased Snail and N-cadherin expression, but increased E-cadherin expression (
Figure 5A). In SETD7 OE SKOV3 cells, Snail and N-cadherin were noticeably elevated, while E-cadherin was decreased (
Figure 5B). Overall, these results show that SETD7 facilitates cell migration by promoting the EMT process in A2780 and SKOV3 cells.
4. Discussion
According to reports, SETD7 has been shown to be either up- or down-regulated in a variety of disease states and particular cell types. The SETD7 function as an oncogene or tumor suppressor depends on the methylation of different substrates [
7]. The importance of SETD7 in the proliferation, invasion, and metastatic processes is becoming increasingly clear [
17,
24]. However, the role of SETD7 in OC is poorly understood. In the present study, the protein expression of SETD7 was significantly up-regulated in OC tissues and cell lines. The survival analysis demonstrated a significant correlation between elevated SETD7 mRNA levels and shorter PFS in all patients and serous patients with OC, thus suggesting its potential as a novel prognostic marker in OC. Importantly, the overexpression of SETD7 facilitated the proliferation and migration of SKOV3 cells, while the knockdown of SETD7 suppressed these effects of A2780 cells. Additionally, SETD7 might promote cell migration via accelerating the EMT process in A2780 and SKOV3 cells.
The IHC results showed that the protein expression of SETD7 was elevated in OC tissues compared to neighboring normal tissues (
Figure 1 and
Table 1). The elevated protein expression of SETD7 was positively correlated with the M category and FIGO stage (
Table 2). Furthermore, the high mRNA levels of SETD7 were correlated to poor PFS in all patients and serous patients with OC (
Figure 2 and
Table 3). These findings imply that SETD7 can be used as a potent independent prognostic factor and may promote OC progression. Recent research has revealed that SETD7 was a poor predictor of disease-free survival (DFS) and OS and that it was greater in 80 breast cancer samples than in non-tumor tissues [
13]. Furthermore, SETD7 has considerably less adjacent tissue area than CRC tissues or colorectal polyps in 176 CRC patients, 20 colon polyps, and paracancerous tissues in 20 colon cancer patients [
25]. These investigations support the potential for SETD7 to be a relevant molecule for the prognosis and treatment of OC.
Recent studies have reported that miR-153 prevents the OVCAR-3 cells from proliferating, but SETD7 silencing reverses this effect [
16]. By controlling EMT procedures, SETD7 promotes or inhibits cell migration in a variety of malignancies [
17,
26]. According to the results of our investigation, targeted knockdown of SETD7 suppresses cell proliferation and migration in A2780 cells, whereas upregulation of SETD7 exerts the opposite effect in SKOV3 cells (
Figure 3 and
Figure 4). Furthermore, our research provides the first evidence that SETD7 overexpression in SKOV3 cells results in an increase in Snail and N-cadherin expression and a decrease in E-cadherin expression (
Figure 5). Increased levels of Snail expression can cause EMT and chemotherapy resistance [
27]. Snail and E-cadherin are both implicated in the EMT process [
28]. This perfectly matches the outcomes we found. According to the aforementioned research, SETD7 may play a role in controlling EMT procedures. The changes in the development of OC should not be attributed to the alteration of a small number of genes since it is unknown whether SETD7 has other targets relevant to OC cell migration. Therefore, further research into the potential involvement of SETD7 to OC is required. In the future, the SETD7-Snail or SETD7-E-cadherin pathway could be used in a therapeutic strategy for the treatment of OC.
5. Conclusion
Currently, this study authenticated that SETD7 protein is highly expressed in OC tissues and cells, which is positively related to with M category and FIGO stage. High mRNA levels of SETD7 were correlated to poor PFS. Furthermore, SETD7 is essential to promote OC cell migration and proliferation. Notably, SETD7 accelerates EMT processes by upregulating Snail and N-cadherin and downregulating E-cadherin, which may increase OC cell migration. Further research is necessary, nevertheless, to understand the precise functions of SETD7.
Author Contributions
Conceptualization, G.S. and Q.L.; data curation, Z.Z.; formal analysis, Z.Z., Y.H., T.H., and B.Z.; investigation, Z.Z., Y.H.; methodology, Q.L.; resources, G.S.; supervision, G.S.; validation, Z.Z., visualization, M.L.; writing-original draft, Z.Z., writing-review and editing, G.S. and Q.L. All authors have read and agreed to the published version of the manuscript.
Funding
This study is supported by Jiangsu Provincial Key Research and Development Program (BE2020678).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data needed to evaluate this work are all included in the manuscript and are available upon request.
Acknowledgements
The authors would like to thank Dr Xiaoli Ju (Department of Basic Medicine, School of Medicine, Jiangsu University, Zhenjiang), for reviewing the clinicopathological data in this manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
Ethics Approval and Consent to Participate
Human OC tissue microarray (ZL-OVA961) was purchased from Zhuoli Biotechnology (Shanghai, China). All the tissue specimens for this study were collected after obtaining informed patient consent and the use of the OC specimens was approved by the Ethics Committee of Zhuoli Biotechnology. The present study was endorsed by the Medical Ethics Committee of Jiangsu University.
References
- Lheureux, S.; Gourley, C.; Vergote, I.; Oza, A. M. Epithelial Ovarian Cancer. The Lancet 2019, 393 (10177), 1240–1253. [CrossRef]
- Bartoletti, M.; Musacchio, L.; Giannone, G.; Tuninetti, V.; Bergamini, A.; Scambia, G.; Lorusso, D.; Valabrega, G.; Mangili, G.; Puglisi, F.; Pignata, S. Emerging Molecular Alterations Leading to Histology-Specific Targeted Therapies in Ovarian Cancer beyond PARP Inhibitors. Cancer Treat. Rev. 2021, 101, 102298. [CrossRef]
- Ledermann, J. A.; Kristeleit, R. S. Optimal treatment for relapsing ovarian cancer. Ann. Oncol. 2010, 21 Suppl 7, vii218-22. [CrossRef]
- Xiao, Y.; Bi, M.; Guo, H.; Li, M. Multi-Omics Approaches for Biomarker Discovery in Early Ovarian Cancer Diagnosis. eBioMedicine 2022, 79, 104001. [CrossRef]
- Zhang, R.; Siu, M. K. Y.; Ngan, H. Y. S.; Chan, K. K. L. Molecular Biomarkers for the Early Detection of Ovarian Cancer. Int. J. Mol. Sci. 2022, 23 (19), 12041. [CrossRef]
- Wang, H.; Cao, R.; Xia, L.; Erdjument-Bromage, H.; Borchers, C.; Tempst, P.; Zhang, Y. Purification and Functional Characterization of a Histone H3-Lysine 4-Specific Methyltransferase. Mol. Cell 2001, 8 (6), 1207–1217. [CrossRef]
- Chiang, C.; Yang, H.; Zhu, L.; Chen, C.; Chen, C.; Zuo, Y.; Zheng, D. The Epigenetic Regulation of Nonhistone Proteins by SETD7: New Targets in Cancer. Front. Genet. 2022, 13, 918509. [CrossRef]
- Chuikov, S.; Kurash, J. K.; Wilson, J. R.; Xiao, B.; Justin, N.; Ivanov, G. S.; McKinney, K.; Tempst, P.; Prives, C.; Gamblin, S. J.; Barlev, N. A.; Reinberg, D. Regulation of P53 Activity through Lysine Methylation. Nature 2004, 432 (7015), 353–360. [CrossRef]
- Subramanian, K.; Jia, D.; Kapoor-Vazirani, P.; Powell, D. R.; Collins, R. E.; Sharma, D.; Peng, J.; Cheng, X.; Vertino, P. M. Regulation of Estrogen Receptor α by the SET7 Lysine Methyltransferase. Mol. Cell 2008, 30 (3), 336–347. [CrossRef]
- Shen, C.; Wang, D.; Liu, X.; Gu, B.; Du, Y.; Wei, F.; Cao, L.; Song, B.; Lu, X.; Yang, Q.; Zhu, Q.; Hou, T.; Li, M.; Wang, L.; Wang, H.; Zhao, Y.; Yang, Y.; Zhu, W. SET7/9 Regulates Cancer Cell Proliferation by Influencing Β-catenin Stability. FASEB J. 2015, 29 (10), 4313–4323. [CrossRef]
- Ea, C.-K.; Baltimore, D. Regulation of NF-κB Activity through Lysine Monomethylation of P65. Proc. Natl. Acad. Sci. 2009, 106 (45), 18972–18977. [CrossRef]
- Monteiro, F. L.; Williams, C.; Helguero, L. A. A Systematic Review to Define the Multi-Faceted Role of Lysine Methyltransferase SETD7 in Cancer. Cancers 2022, 14 (6), 1414. [CrossRef]
- Zhang, Y.; Liu, J.; Lin, J.; Zhou, L.; Song, Y.; Wei, B.; Luo, X.; Chen, Z.; Chen, Y.; Xiong, J.; Xu, X.; Ding, L.; Ye, Q. The Transcription Factor GATA1 and the Histone Methyltransferase SET7 Interact to Promote VEGF-Mediated Angiogenesis and Tumor Growth and Predict Clinical Outcome of Breast Cancer. Oncotarget 2016, 7 (9), 9859–9875. [CrossRef]
- Si, W.; Zhou, J.; Zhao, Y.; Zheng, J.; Cui, L. SET7/9 Promotes Multiple Malignant Processes in Breast Cancer Development via RUNX2 Activation and Is Negatively Regulated by TRIM21. Cell Death Dis. 2020, 11 (2), 151. [CrossRef]
- Gu, Y.; Wang, X.; Liu, H.; Li, G.; Yu, W.; Ma, Q. SET7/9 Promotes Hepatocellular Carcinoma Progression through Regulation of E2F1. Oncol. Rep. 2018. [CrossRef]
- Zhou, J.; Xie, M.; Shi, Y.; Luo, B.; Gong, G.; Li, J.; Wang, J.; Zhao, W.; Zi, Y.; Wu, X.; Wen, J. MicroRNA-153 Functions as a Tumor Suppressor by Targeting SET7 and ZEB2 in Ovarian Cancer Cells. Oncol. Rep. 2015, 34 (1), 111–120. [CrossRef]
- Montenegro, M. F.; Sánchez-del-Campo, L.; González-Guerrero, R.; Martínez-Barba, E.; Piñero-Madrona, A.; Cabezas-Herrera, J.; Rodríguez-López, J. N. Tumor Suppressor SET9 Guides the Epigenetic Plasticity of Breast Cancer Cells and Serves as an Early-Stage Biomarker for Predicting Metastasis. Oncogene 2016, 35 (47), 6143–6152. [CrossRef]
- Hong, X.; Huang, H.; Qiu, X.; Ding, Z.; Feng, X.; Zhu, Y.; Zhuo, H.; Hou, J.; Zhao, J.; Cai, W.; Sha, R.; Hong, X.; Li, Y.; Song, H.; Zhang, Z. Targeting Posttranslational Modifications of RIOK1 Inhibits the Progression of Colorectal and Gastric Cancers. eLife 2018, 7, e29511. [CrossRef]
- Zhao, K.; Li, Y.; Han, T.; Sun, A.; Bian, W.; Li, M.; Lin, Q.; Shao, G. Comparison of the Expression Levels of Lysine-specific Demethylase 1 and Survival Outcomes between Triple-negative and Non-triple-negative Breast Cancer. Oncol. Lett. 2020, 21 (2), 102. [CrossRef]
- Wei, Y.; Han, T.; Wang, R.; Wei, J.; Peng, K.; Lin, Q.; Shao, G. LSD1 Negatively Regulates Autophagy through the mTOR Signaling Pathway in Ovarian Cancer Cells. Oncol. Rep. 2018. [CrossRef]
- Shao, G.; Wang, R.; Sun, A.; Wei, J.; Peng, K.; Dai, Q.; Yang, W.; Lin, Q. The E3 Ubiquitin Ligase NEDD4 Mediates Cell Migration Signaling of EGFR in Lung Cancer Cells. Mol. Cancer 2018, 17 (1), 24. [CrossRef]
- Shao, G.; Wang, J.; Li, Y.; Liu, X.; Xie, X.; Wan, X.; Yan, M.; Jin, J.; Lin, Q.; Zhu, H.; Zhang, L.; Gong, A.; Shao, Q.; Wu, C. Lysine-Specific Demethylase 1 Mediates Epidermal Growth Factor Signaling to Promote Cell Migration in Ovarian Cancer Cells. Sci. Rep. 2015, 5 (1), 15344. [CrossRef]
- Yang, X.; Bai, Q.; Chen, W.; Liang, J.; Wang, F.; Gu, W.; Liu, L.; Li, Q.; Chen, Z.; Zhou, A.; Long, J.; Tian, H.; Wu, J.; Ding, X.; Zhou, N.; Li, M.; Yang, Y.; Cai, J. M 6 A-Dependent Modulation via IGF2BP3/MCM5/Notch Axis Promotes Partial EMT and LUAD Metastasis. Adv. Sci. 2023, 10 (20), 2206744. [CrossRef]
- Song, Y.; Zhang, J.; Tian, T.; Fu, X.; Wang, W.; Li, S.; Shi, T.; Suo, A.; Ruan, Z.; Guo, H.; Yao, Y. SET7/9 Inhibits Oncogenic Activities through Regulation of Gli-1 Expression in Breast Cancer. Tumor Biol. 2016, 37 (7), 9311–9322. [CrossRef]
- Duan, B.; Bai, J.; Qiu, J.; Wang, J.; Tong, C.; Wang, X.; Miao, J.; Li, Z.; Li, W.; Yang, J.; Huang, C. Histone-Lysine N-Methyltransferase SETD7 Is a Potential Serum Biomarker for Colorectal Cancer Patients. EBioMedicine 2018, 37, 134–143. [CrossRef]
- Xie, H.; Li, J.; Ying, Y.; Yan, H.; Jin, K.; Ma, X.; He, L.; Xu, X.; Liu, B.; Wang, X.; Zheng, X.; Xie, L. METTL3/YTHDF2 m 6 A Axis Promotes Tumorigenesis by Degrading SETD7 and KLF4 mRNAs in Bladder Cancer. J. Cell. Mol. Med. 2020, 24 (7), 4092–4104. [CrossRef]
- Kaufhold, S.; Bonavida, B. Central Role of Snail1 in the Regulation of EMT and Resistance in Cancer: A Target for Therapeutic Intervention. J. Exp. Clin. Cancer Res. 2014, 33 (1), 62. [CrossRef]
- Elloul, S.; Bukholt Elstrand, M.; Nesland, J. M.; Tropé, C. G.; Kvalheim, G.; Goldberg, I.; Reich, R.; Davidson, B. Snail, Slug, and Smad-Interacting Protein 1 as Novel Parameters of Disease Aggressiveness in Metastatic Ovarian and Breast Carcinoma: Cadherin Regulation in Ovarian and Breast CA. Cancer 2005, 103 (8), 1631–1643. [CrossRef]
Figure 1.
SETD7 is strikingly upregulated in OC tissues and cell lines. (A) IHC analysis of SETD7 on a tissue array of OC patients (n=88) and healthy adjacent tissues (n=8). (B) Western blot analysis of SETD7 in the nuclear (N) and cytosolic (C) fractions from A2780 cells. Histone H3 and α-tubulin were used as markers for the nuclear and cytoplasmic fractions, respectively. (C) The proportion of OC tissues expressing SETD7 in various stages of malignancy. (D) The expression levels of SETD5 protein in four OC cell lines and one normal ovarian granulosa cell line (KGN) were analyzed by western blot.
Figure 1.
SETD7 is strikingly upregulated in OC tissues and cell lines. (A) IHC analysis of SETD7 on a tissue array of OC patients (n=88) and healthy adjacent tissues (n=8). (B) Western blot analysis of SETD7 in the nuclear (N) and cytosolic (C) fractions from A2780 cells. Histone H3 and α-tubulin were used as markers for the nuclear and cytoplasmic fractions, respectively. (C) The proportion of OC tissues expressing SETD7 in various stages of malignancy. (D) The expression levels of SETD5 protein in four OC cell lines and one normal ovarian granulosa cell line (KGN) were analyzed by western blot.
Figure 2.
Prognostic impact of SETD7 in OC. (A) Progression-free survival (PFS) rate for 614 patients with OC. (B) Overall survival (OS) rate for 478 patients with OC. (C) Post-progression survival (PPS) rate for 382 patients with OC.
Figure 2.
Prognostic impact of SETD7 in OC. (A) Progression-free survival (PFS) rate for 614 patients with OC. (B) Overall survival (OS) rate for 478 patients with OC. (C) Post-progression survival (PPS) rate for 382 patients with OC.
Figure 3.
SETD7 stimulates cell proliferation in OC. (A) Western blot for SETD7 in A2780 cells infected with shSETD7-1, shSETD7-2, shSETD7-3, or negative control shRNA (shNC). (B) The relative mRNA levels of SETD7 in A2780 cells infected with shSETD7-1, shSETD7-2, shSETD7-3, or shNC were determined by qRT-PCR. (C) SETD7 knockdown inhibited the proliferation of A2780 cells determined by CCK-8 assay. (D) SETD7 knockdown inhibited the proliferation of A2780 cells determined by cell counting assay. Left panels provide a graphical representation of the accumulated number of proliferated cells at 0, 24, 48, 72, and 96 h, and the right panels show representative images (200× magnification) at 48 h. (E) Western blot for SETD7 in SKOV3 cells infected with pCDH-SETD7 or pCDH vector. (F) The relative mRNA levels of SETD7 in SKOV3 cells infected with pCDH-SETD7 or pCDH vector were determined by qRT-PCR. (G) Ectopic expression of SETD7 promoted the proliferation of SKOV3 cells determined by CCK-8 assay. (H) Ectopic expression of SETD7 promoted the proliferation of SKOV3 cells determined by cell counting assay. Left panels provide a graphical representation of the accumulated number of proliferated cells at 0, 24, 48, 72, and 96 h, and the right panels show representative images (200× magnification) at 48 h. The results are presented as are means ± SD (n=3). The data were analyzed with Two-tailed Student’s t test for (B) and (F), or two-way ANOVA for (C), (D), (G), and (H). Statistical significance was concluded at ***p < 0.001.
Figure 3.
SETD7 stimulates cell proliferation in OC. (A) Western blot for SETD7 in A2780 cells infected with shSETD7-1, shSETD7-2, shSETD7-3, or negative control shRNA (shNC). (B) The relative mRNA levels of SETD7 in A2780 cells infected with shSETD7-1, shSETD7-2, shSETD7-3, or shNC were determined by qRT-PCR. (C) SETD7 knockdown inhibited the proliferation of A2780 cells determined by CCK-8 assay. (D) SETD7 knockdown inhibited the proliferation of A2780 cells determined by cell counting assay. Left panels provide a graphical representation of the accumulated number of proliferated cells at 0, 24, 48, 72, and 96 h, and the right panels show representative images (200× magnification) at 48 h. (E) Western blot for SETD7 in SKOV3 cells infected with pCDH-SETD7 or pCDH vector. (F) The relative mRNA levels of SETD7 in SKOV3 cells infected with pCDH-SETD7 or pCDH vector were determined by qRT-PCR. (G) Ectopic expression of SETD7 promoted the proliferation of SKOV3 cells determined by CCK-8 assay. (H) Ectopic expression of SETD7 promoted the proliferation of SKOV3 cells determined by cell counting assay. Left panels provide a graphical representation of the accumulated number of proliferated cells at 0, 24, 48, 72, and 96 h, and the right panels show representative images (200× magnification) at 48 h. The results are presented as are means ± SD (n=3). The data were analyzed with Two-tailed Student’s t test for (B) and (F), or two-way ANOVA for (C), (D), (G), and (H). Statistical significance was concluded at ***p < 0.001.
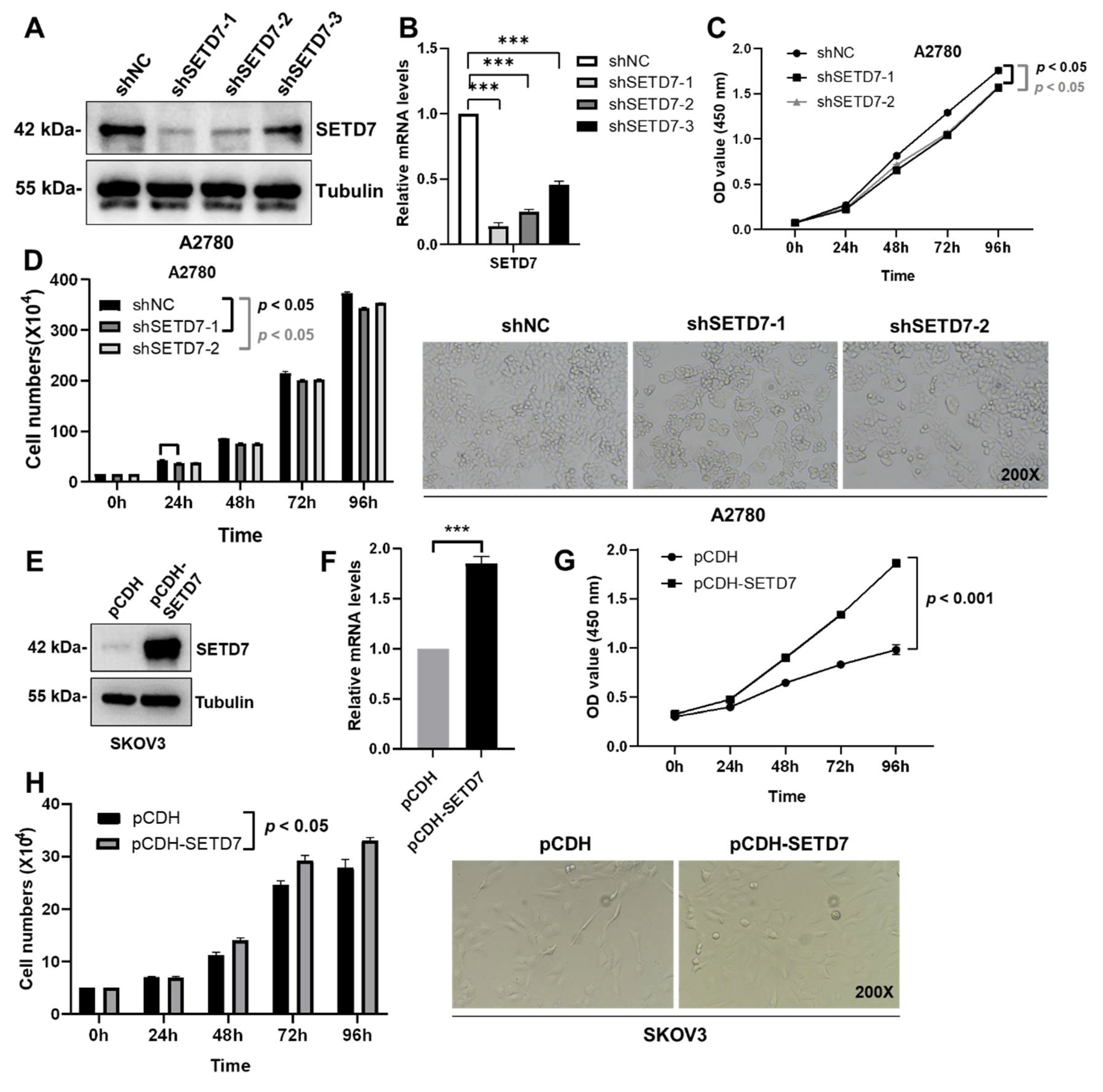
Figure 4.
SETD7 promotes OC cell migration. (A) The effects of knockdown of SETD7 or negative control shRNA (shNC) on the migration of A2780 cells were determined by Transwell assay. Left panels show representative images (200× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 48 h. (B) The effects of knockdown of SETD7 or negative control shRNA (shNC) on the migration of A2780 cells were determined by wound-healing assay. Left panels show representative images (40× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 72 h. (C) The effects of overexpression of pCDH-SETD7 or pCDH on the migration of SKOV3 cells were measured by Transwell assay. Left panels show representative images (200× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 24 h. (D) The effects of overexpression of pCDH-SETD7 or pCDH on the migration of SKOV3 cells were measured by wound-healing assay. Left panels show representative images (40× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 24 h. The results are presented as means ± SD (n=3). The data were analyzed with Two-tailed Student’s t test. Statistical significance was concluded at ***p < 0.001.
Figure 4.
SETD7 promotes OC cell migration. (A) The effects of knockdown of SETD7 or negative control shRNA (shNC) on the migration of A2780 cells were determined by Transwell assay. Left panels show representative images (200× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 48 h. (B) The effects of knockdown of SETD7 or negative control shRNA (shNC) on the migration of A2780 cells were determined by wound-healing assay. Left panels show representative images (40× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 72 h. (C) The effects of overexpression of pCDH-SETD7 or pCDH on the migration of SKOV3 cells were measured by Transwell assay. Left panels show representative images (200× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 24 h. (D) The effects of overexpression of pCDH-SETD7 or pCDH on the migration of SKOV3 cells were measured by wound-healing assay. Left panels show representative images (40× magnification), and the right panels provide a graphical representation of the accumulated number of migrated cells at 24 h. The results are presented as means ± SD (n=3). The data were analyzed with Two-tailed Student’s t test. Statistical significance was concluded at ***p < 0.001.
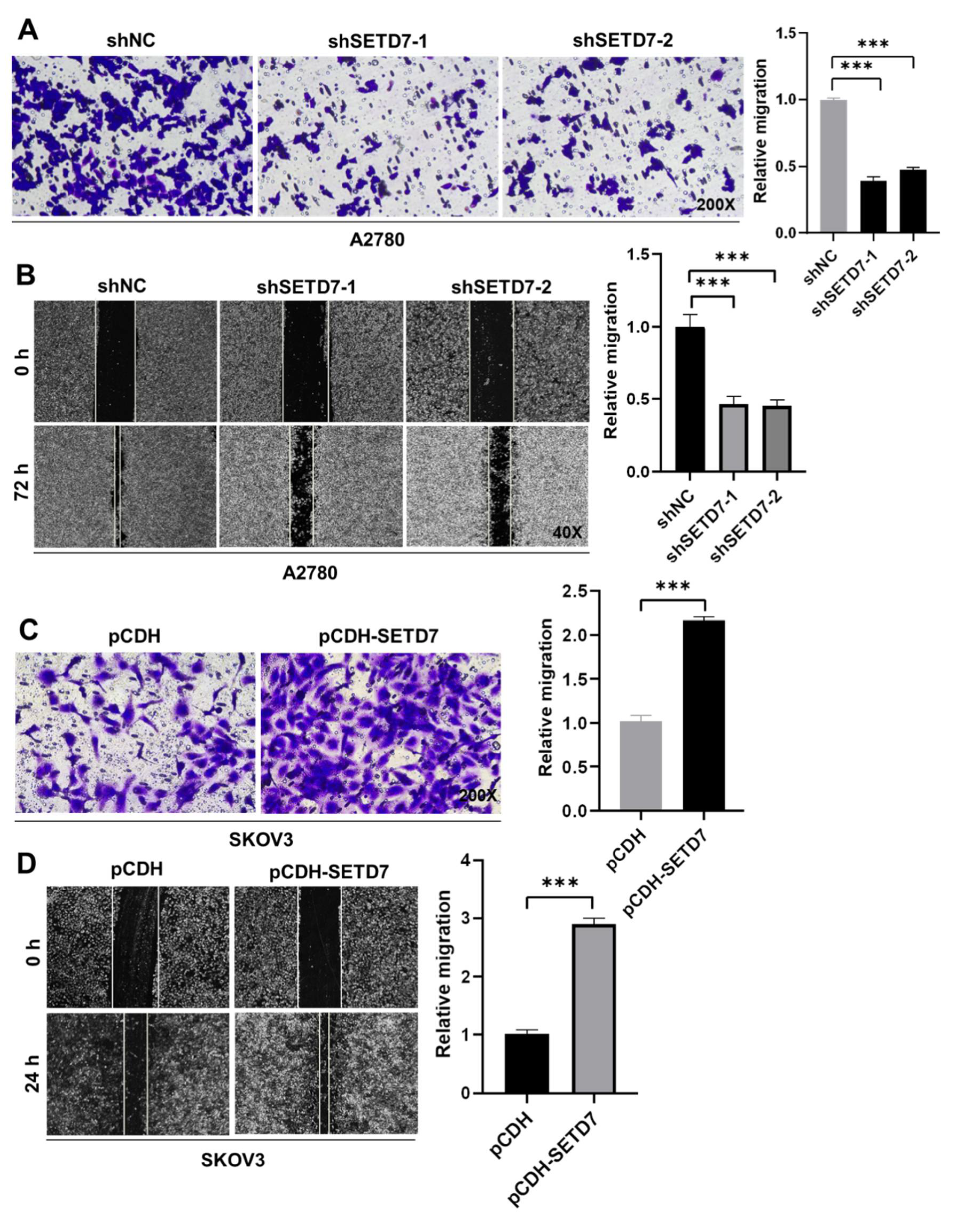
Figure 5.
SETD7 promotes cell migration via EMT. (A) A2780 cells downregulated of SETD7 or negative control shRNA (shNC) were subjected to western blot to test the expression of SETD7, E-cadherin, N-cadherin, Vimentin, and Snail, taking α-tubulin as an internal reference. (B) SKOV3 cells overexpressed of pCDH-SETD7 or pCDH were subjected to western blot to test the expression of SETD7, E-cadherin, N-cadherin, Vimentin, and Snail, taking α-tubulin as an internal reference.
Figure 5.
SETD7 promotes cell migration via EMT. (A) A2780 cells downregulated of SETD7 or negative control shRNA (shNC) were subjected to western blot to test the expression of SETD7, E-cadherin, N-cadherin, Vimentin, and Snail, taking α-tubulin as an internal reference. (B) SKOV3 cells overexpressed of pCDH-SETD7 or pCDH were subjected to western blot to test the expression of SETD7, E-cadherin, N-cadherin, Vimentin, and Snail, taking α-tubulin as an internal reference.
Table 1.
Association of SETD7 expression in 88 OC tissue and 8 adjacent normal tissue samples.
Table 1.
Association of SETD7 expression in 88 OC tissue and 8 adjacent normal tissue samples.
| Tissues |
|
Total |
SETD7 |
p-value |
| |
Positive |
Negative |
| Tumor |
|
88 |
69 (78%) |
19 (21%) |
0.004*
|
| Normal |
|
8 |
2 (25%) |
6 (75%) |
|
Table 2.
Clinical characterization of SETD7 expression in 88 OC patients.
Table 2.
Clinical characterization of SETD7 expression in 88 OC patients.
| Pathological category |
Total |
SETD7 |
p-value |
| Positive |
Negative |
| Age |
≤45 |
11 |
7 |
4 |
0.378 |
| >45 |
77 |
62 |
15 |
|
| Tumor size (cm) |
≤3 |
6 |
4 |
2 |
0.833 |
| >3 |
82 |
65 |
17 |
|
| T category |
T1 |
50 |
38 |
12 |
0.529 |
| T2/T3 |
38 |
31 |
7 |
|
|
& N category |
N0 |
31 |
20 |
11 |
0.466 |
| N1 |
15 |
8 |
7 |
|
| M category |
M0
M1 |
63
25 |
45
24 |
18
1 |
0.012*
|
|
# FIGO stage |
I/II |
30 |
21 |
9 |
0.029*
|
| III/IV |
41 |
37 |
4 |
|
Table 3.
The prognostic values of SETD7 in all and different histological subtypes of OC patients.
Table 3.
The prognostic values of SETD7 in all and different histological subtypes of OC patients.
| Gene |
Histology |
OS |
PFS |
PPS |
| Cases |
HR |
95% CI |
P-value |
Cases |
HR |
95% CI |
P-value |
Cases |
HR |
95% CI |
P-value |
SETD7
224928_at |
Overall |
655 |
1.24 |
0.97-1.58 |
0.082 |
614 |
1.27 |
1.03-1.57 |
0.025 |
382 |
0.84 |
0.65-1.07 |
0.15 |
| Serous |
523 |
1.13 |
0.91-1.26 |
0.11 |
483 |
1.22 |
0.93-1.47 |
0.037 |
346 |
0.78 |
0.6-1.01 |
0.061 |
| Endometrioid |
30 |
4.03 |
042-38.71 |
0.19 |
44 |
0.31 |
0.07-1.4 |
0.11 |
10 |
− |
− |
− |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).