Submitted:
08 December 2023
Posted:
08 December 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
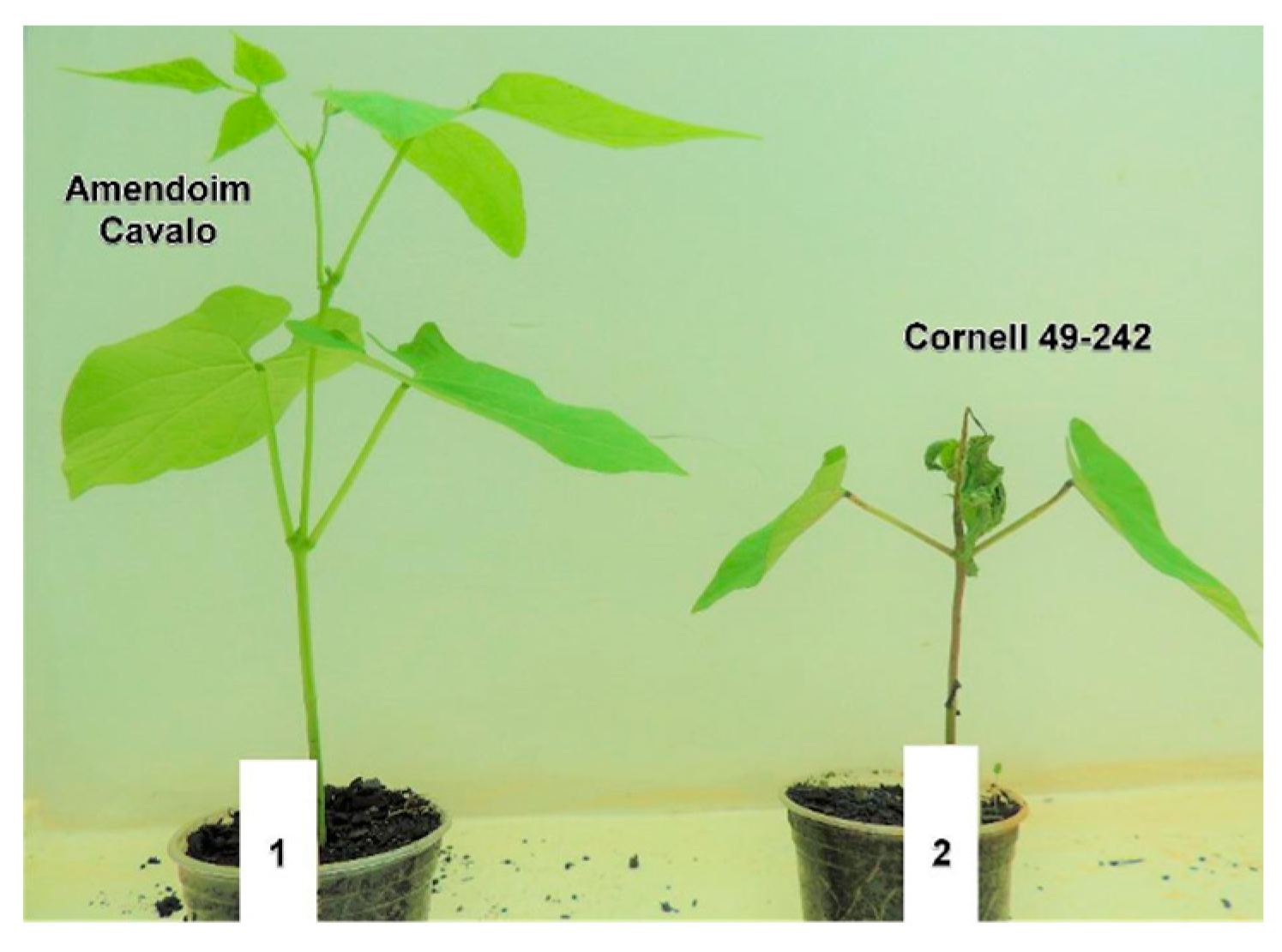
2.1. Phenotypic Evaluation of the Cultivars
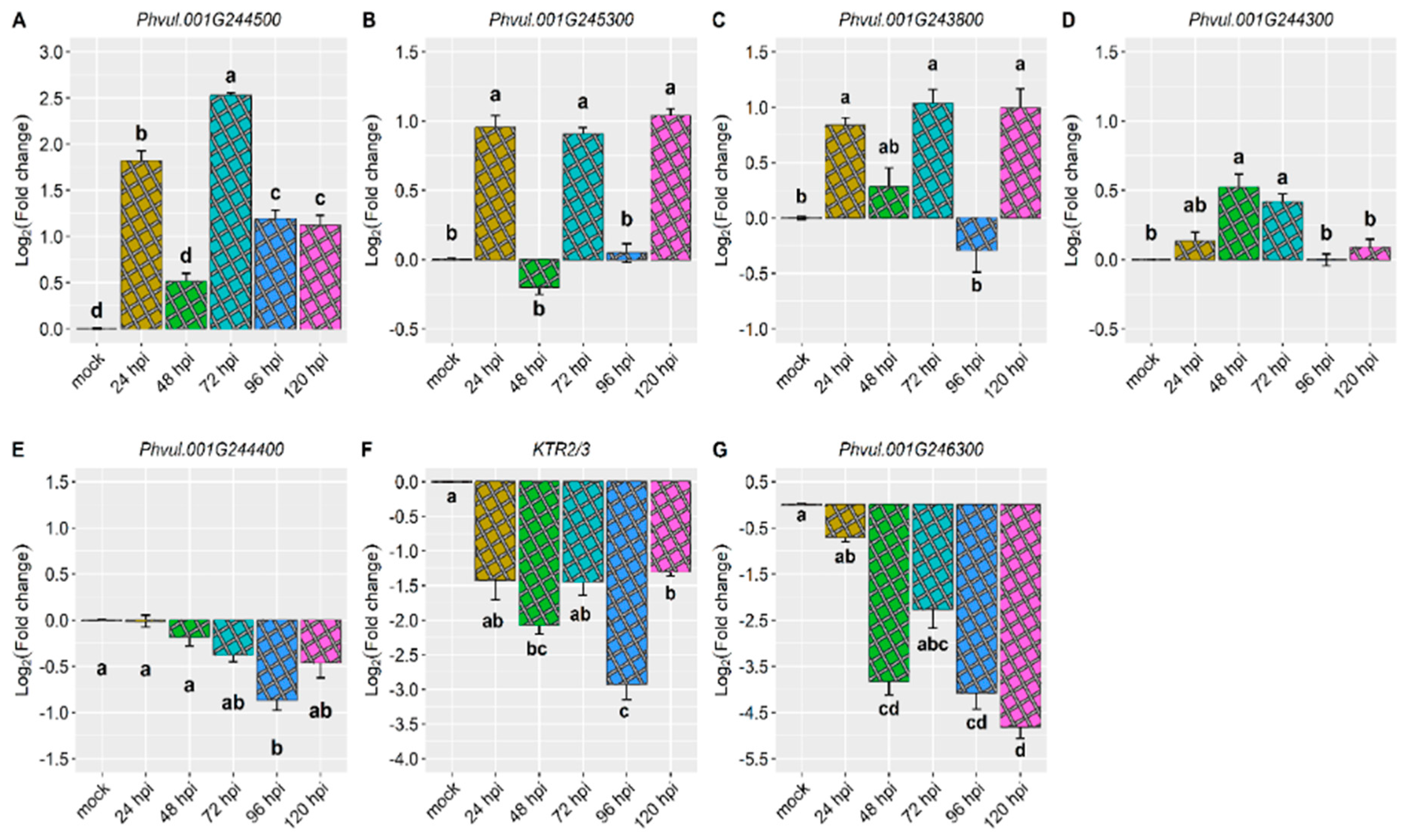
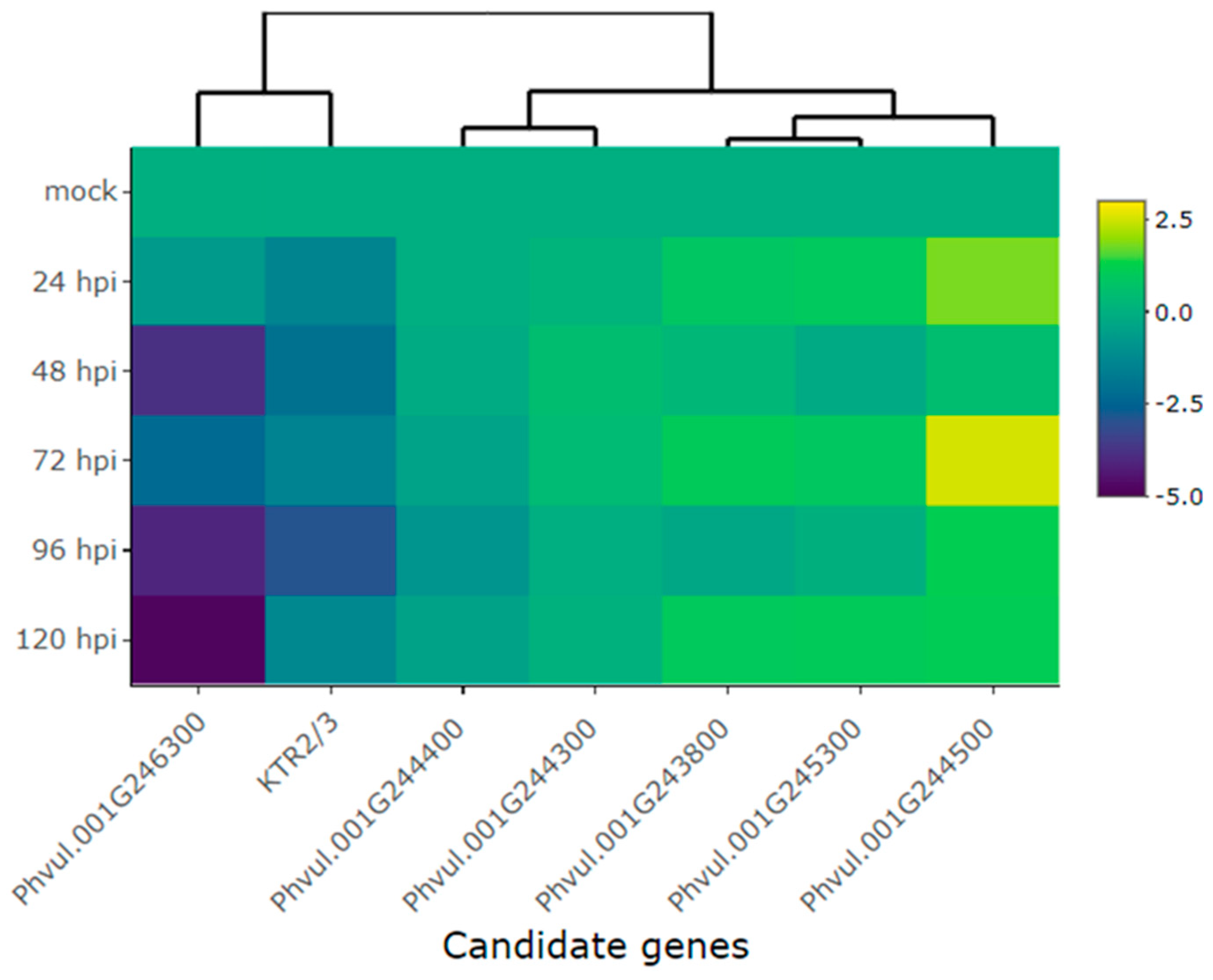
2.2. Differential Expression of Candidate in the Amendoim Cavalo Cultivar Inoculated with Race 73 of C. lindemuthianum
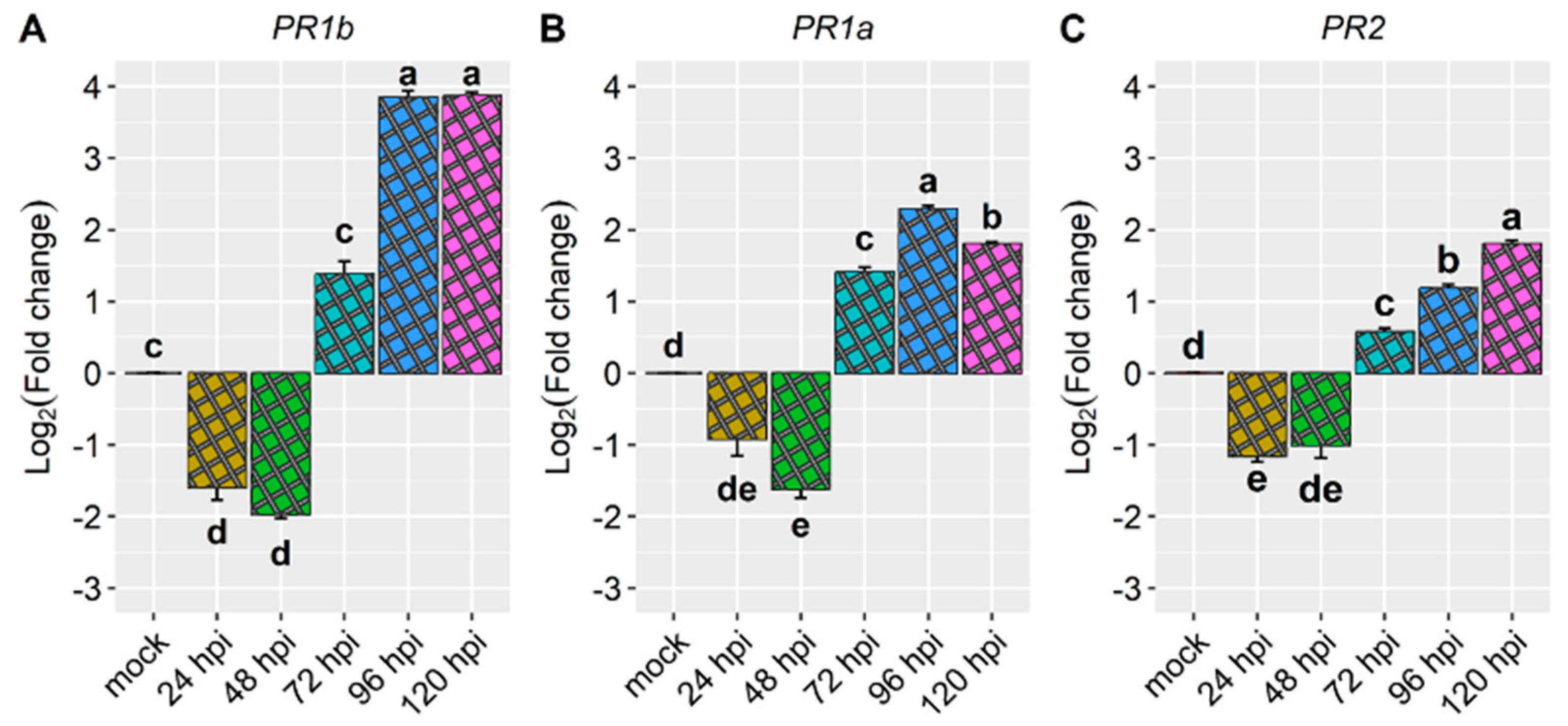
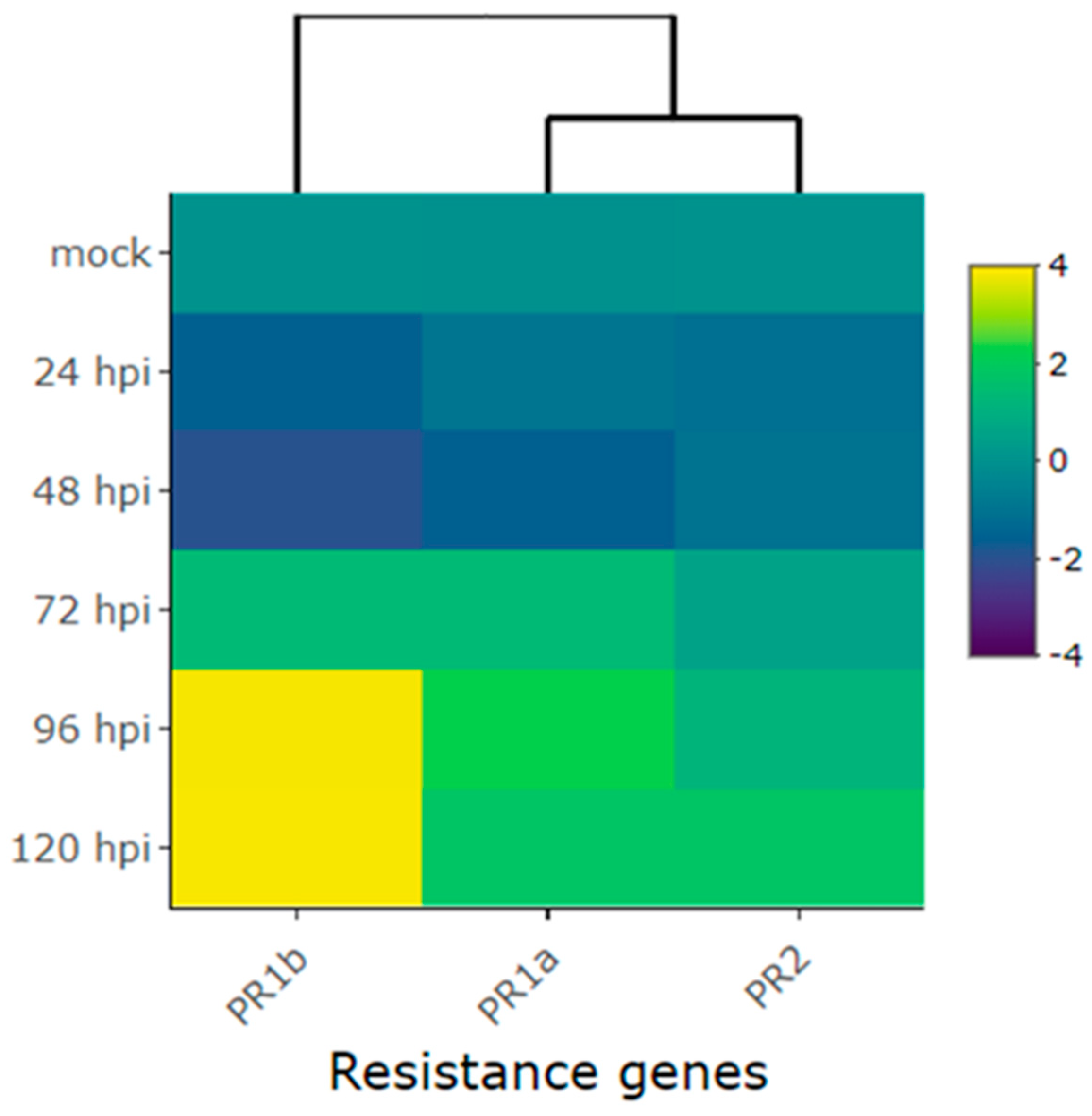
2.3. Expression Profile of Defense Genes in the Amendoim Cavalo Cultivar Inoculated with Race 73 of C. lindemuthianum
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Plant Material and Growth Conditions
5.2. Pathogenesis Assay
5.3. RNA Extraction
5.4. Reverse Transcription (cDNA Synthesis)
5.5. Target Genes and Primer Design
5.6. Quantitative PCR (qPCR) and Data Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bitocchi, E.; Rau, D.; Bellucci, E.; Rodriguez, M.; Murgia, M.L.; Gioia, T.; Santo, D.; Nanni, L.; Attene, G.; Papa, R. Beans (Phaseolus ssp.) as a model for understanding crop evolution. Frontiers in Plant Science 2017, 8, 722. [Google Scholar] [CrossRef] [PubMed]
- Vaz Patto, M.C.; Amarowicz, R.; Aryee, A.N.A.; Boye, J.I.; Chung, H.-J.; Martín-Cabrejas, M.A.; Domoney, C. Achievements and Challenges in Improving the Nutritional Quality of Food Legumes. Critical Reviews in Plant Sciences 2015, 34, 105–143. [Google Scholar] [CrossRef]
- Vidigal Filho, P.S.; Gonçalves-Vidigal, M.C.; Vaz Bisneta, M.; Souza, V.B.; Gilio, T.A.S.; Calvi, A.C.; Lima, L.R.L.; Pastor-Corrales, M.A.; Melotto, M. Genome-wide association study of resistance to anthracnose and angular leaf spot in Brazilian Mesoamerican and Andean common bean cultivars. Crop Science 2020, 60, 2931–2950. [Google Scholar] [CrossRef]
- Padder, B.A.; Sharma, P.N.; Awale, H.E.; Kelly, J.D. Colletotrichum lindemuthianum, the causal agent of bean anthracnose. J. Plant Pathol. 2017, 99, 317–330. [Google Scholar] [CrossRef]
- Nunes, M.P.B.A.; Gonçalves-Vidigal, M.C.; Martins, V.S.R.; Xavier, L.F.S.; Valentini, G.; Vaz Bisneta, M.; Vidigal Filho, P.S. Relationship of Colletotrichum lindemuthianum races and resistance loci in the Phaseolus vulgaris L. genome. Crop Science 2021, 61, 3877–3893. [Google Scholar] [CrossRef]
- Singh, S.P.; Schwartz, H.F. Breeding common bean for resistance to diseases: A review. Crop Sci. 2010, 50, 2199–2223. [Google Scholar] [CrossRef]
- Kelly, J.D.; Afanador, L.; Cameron, L.S. New races of Colletotrichum lindemuthianum in Michigan and implications in dry bean resistance breeding. Plant Disease 1994, 78, 892–894. [Google Scholar] [CrossRef]
- Pastor-Corrales, M.A.; Otoya, M.M.; Molina, A.; Singh, S.P. Resistance to Colletotrichum lindemuthianum isolates from middle America and Andean South America in different common bean races. Plant Dis. 1995, 79, 63–67. [Google Scholar] [CrossRef]
- Pacheco, L.M.; Berrouet, K.V.; Yepes, M.S.; Sánchez, P.G.; Montoya, M.M. Detección por PCR de Colletotrichum lindemuthianum en cultivos y semillas de frijol en Antioquia, Colombia. Acta Agrómica 2014, 63, 377–387. [Google Scholar] [CrossRef]
- Vaz Bisneta, M.; Gonçalves-Vidigal, M.C. Integration of anthracnose resistance loci and RLK and NBS-LRR-encoding genes in the Phaseolus vulgaris L. genome. Crop Sci. 2020, 60, 2901–2918. [Google Scholar] [CrossRef]
- Gonçalves-Vidigal, M.C.; Cruz, A.S.; Garcia, A.; Kami, J.; Vidigal Filho, P.S.; Sousa, L.L.; McClean, P.; Gepts, P.; Pastor-Corrales, M.A. Linkage mapping of the Phg-1 and Co-14 genes for resistance to angular leaf spot and anthracnose in the common bean cultivar AND 277. Theor. Appl. Genet. 2011, 122, 893–903. [Google Scholar] [CrossRef] [PubMed]
- Zuiderveen, G.H.; Padder, B.A.; Kamfwa, K.; Song, Q.; Kelly, J.D. Genome-wide association study of anthracnose resistance in Andean beans (Phaseolus vulgaris). PLoS One 2016, 11, e0156391. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wu, J.; Wang, L.; Mantri, N.; Zhang, X.; Zhu, Z.; Wang, S. Mapping and genetic structure analysis of the anthracnose resistance locus Co-1HY in the common bean (Phaseolus vulgaris L.). PLoS One, 2017; 12, e0169954. [Google Scholar] [CrossRef]
- Lima, L.R.L.; Gonçalves-Vidigal, M.C.; Vaz Bisneta, M.; Valentini, G.; Vidigal Filho, P.S.; Martins, V.S.R.; Souza, T.L.P.O. Genetic fine-mapping of anthracnose disease-resistance allele Co-14 present in the Andean common bean cultivar AND 277. Crop Sci. 2023; 1–14. [Google Scholar] [CrossRef]
- Richard, M.M.S.; Pflieger, S.; Sevignac, M.; Thareau, V.; Blanchet, S.; Li, Y.; Jackson, S.A.; Jackson, S.A.; Geffroy, V. Fine mapping of Co-x, an anthracnose resistance gene to a highly virulent strain of Colletotrichum lindemuthianum in common bean. Theor. Appl. Genet. 2014, 127, 1653–1666. [Google Scholar] [CrossRef] [PubMed]
- Richard, M.M.S.; Gratias, A.; Diaz, J.C.A.; Thareau, V.; Pflieger, S.; Meziadi, C.; Blanchet, S.; Marande, W.; Bitocchi, E.; Papa, R.; Miklas, P.N.; Geffroy, V. A common bean truncated CRINKLY4 kinase controls gene-for-gene resistance to the fungus Colletotrichum lindemuthianum. J. Exp. Bot. 2021, 72, 3569–3581. [Google Scholar] [CrossRef]
- Lima-Castro, S. A., Gonçalves-Vidigal, M. C., Gilio, T. A. S., Lacanallo, G. F., Valentini, G., Martins, V. S. R. et al. (2017). Genetics and mapping of a new anthracnose resistance locus in Andean common bean Paloma. BMC Genom. 18, 306. [CrossRef]
- Gonçalves-Vidigal, M.C.; Gilio, T.A.S.; Valentini, G.; Vaz Bisneta, M.; Vidigal Filho, P.S.; Song, Q.; Oblessuc, P.R.; Melotto, M. (2020). New Andean source of resistance to anthracnose and angular leaf spot: Fine-mapping of disease-resistance genes in California Dark Red Kidney common bean cultivar. PLoS One. 15, e0235215. [CrossRef]
- Gilio, T.A.S.; Hurtado-Gonzales, O.P.; Gonçalves-Vidigal, M.C.; Valentini, G.; Elias, J.C.F.; Song, Q.; Pastor-Corrales, M.A. Fine mapping of an anthracnose-resistance locus in Andean common bean cultivar Amendoim Cavalo. PLoS One 2020, 15, e0239763. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.L.; Puttick, M.N.; Clark, J.W.; Edwards, D.; Kenrick, P.; Pressel, S.; Wellman, C.H.; Yang, Z.; Schneider, H.; Donoghue, P.C.J. The timescale of early land plant evolution. Proceedings of the National Academy of Sciences 2018, 115, E2274–E2283. [Google Scholar] [CrossRef]
- Gu, J.; Sun, J.; Liu, N.; Sun, X.; Liu, C.; Wu, L.; Liu, G.; Zeng, F.; Hou, C.; Han, S.; Zhen, W.; Wang, D. A novel cysteine-rich receptor-like kinase gene, TaCRK2, contributes to leaf rust resistance in wheat. Molecular Plant Pathology 2020, 21, 732–746. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Ngou, B.P.M.; Ding, P.; Xin, X.-F. PTI-ETI crosstalk: an integrative view of plant immunity. Current Opinion in Plant Biology 2021, 62, 102030. [Google Scholar] [CrossRef] [PubMed]
- Ngou, B.P.M.; Ding, P.; Jones, J.D.G. Thirty years of resistance: Zig-zag through the plant immune system. The Plant Cell 2022, 34, 1447–1478. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Deng, Y.; Ning, Y.; He, Z.; Wang, G.-L. Exploiting broad-spectrum disease resistance in crops: from molecular dissection to breeding. Annual Review of Plant Biology 2020, 71, 575–603. [Google Scholar] [CrossRef] [PubMed]
- Kankanala, P.; Nandety, R.S.; Mysore, K.S. Genomics of plant disease resistance in legumes. Frontiers in Plant Science 2019, 10, 1345. [Google Scholar] [CrossRef] [PubMed]
- Borges, A.; Melotto, M.; Tsai, S.M.; Caldas, D.G.G. Changes in spatial and temporal gene expression during incompatible interaction between common bean and anthracnose pathogen. J. Plant Physiol. 2012, 169, 1216–1220. [Google Scholar] [CrossRef] [PubMed]
- Mahiya-Farooq; Padder, B.A.; Bhat, N.N.; Shah, M.D.; Shikari, A.B.; Awale, H.E.; Kelly, J.D. Temporal expression of candidate genes at the Co-1 locus and their interaction with other defense related genes in common bean. Physiol. Mol. Plant Pathol. 2019, 108, 101424. [Google Scholar] [CrossRef]
- Oblessuc, P.R.; Baroni, R.M.; Garcia, A.A.F.; Chioratto, A.F.; Carbonell, S.A.M.; Camargo, L.E.A.; Benchimol, L.L. Mapping of angular leaf spot resistance QTL in common bean (Phaseolus vulgaris L.) under different environments. BMC Genet. 2012, 13, 50. [Google Scholar] [CrossRef] [PubMed]
- Shams, E.; Javan-Nikkhah, M.; Mirzadi Gohari, A. Dissecting molecular events and gene expression signatures involved in Colletotrichum lindemuthianum-Phaseolus vulgaris pathosystem in compatible and incompatible interactions. Eur. J. Plant Pathol. 2020, 156, 925–937. [Google Scholar] [CrossRef]
- Lovatto, M.; Gonçalves-Vidigal, M.C.; Calvi, A.C.; Vaz Bisneta, M.; Vidigal Filho, P.S. Gene expression analysis in California Dark Red Kidney common bean cultivar during incompatible interaction with Colletotrichum lindemuthianum. Annual Report of Bean Improvement Cooperative 2022, 65, 71–72. [Google Scholar] [CrossRef]
- Nanami, D.S.Y.; Gonçalves-Vidigal, M.C.; Castro, S.A.L.; Frias, A.A.T.; Vidigal Filho, P.S.; Elias, H.T. Characterization of genetic resistance in Andean common bean cultivar Amendoim Cavalo to Colletotrichum lindemuthianum. Agronomy Science and Biotechnology 2017, 3, 43–52. [Google Scholar] [CrossRef]
- Friesen, T.L.; Faris, J.D. Characterization of Effector–Target Interactions in Necrotrophic Pathosystems Reveals Trends and Variation in Host Manipulation. Annual Review of Phytopathology 2021, 59, 77–98. [Google Scholar] [CrossRef] [PubMed]
- Riviere, M.-P.; Marais, A.; Ponchet, M.; Willats, W.; Galiana, E. Silencing of acidic pathogenesis-related PR-1 genes increases extracellular beta-(1->3)-glucanase activity at the onset of tobacco defence reactions. Journal of Experimental Botany 2008, 59, 1225–1239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Guo, N.; Zhang, Y.; Yu, Y.; Liu, S. Genome-wide characterization and expression analysis of pathogenesis-related 1 (PR-1) gene family in tea plant (Camellia sinensis (L.) O. Kuntze) in response to blister-blight disease stress. International Journal of Molecular Sciences 2022, 23, 1292. [Google Scholar] [CrossRef]
- Dixon, D.C.; Cutt, J.R.; Klessig, D.F. Differential targeting of the tobacco PR-1 pathogenesis-related proteins to the extracellular space and vacuoles of crystal idioblasts. The EMBO Journal 1991, 10, 1317–1324. [Google Scholar] [CrossRef]
- Santamaria, M.; Thomson, C.J.; Read, N.D.; Loake, G.J. The promoter of a basic PR1-like gene, AtPRB1, from Arabidopsis establishes an organ-specific expression pattern and responsiveness to ethylene and methyl jasmonate. Plant Molecular Biology 2001, 47, 641–652. [Google Scholar] [CrossRef]
- Walter, M.H.; Liu, J.-W.; Grand, C.; Lamb, C.J.; Hess, D. Bean pathogenesis-related (PR) proteins deduced from elicitor-induced transcripts are members of a ubiquitous new class of conserved PR proteins including pollen allergens. Molecular and Genetics and Genomics 1990, 222, 353–360. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Diaz, J.C.; Laugé, R.; Delannoy, E.; Huguet, S.; Roux, C.P.-L.; Gratias, A.; Geffroy, V. Genome-Wide Transcriptomic Analysis of the Effects of Infection with the Hemibiotrophic Fungus Colletotrichum lindemuthianum on Common Bean. Plants 2022, 11, 1995. [Google Scholar] [CrossRef] [PubMed]
- Edington, B.V.; Lamb, C.J.; Dixon, R.A. cDNA cloning and characterization of a putative 1,3-b-Dglucanase transcript induced by fungal elicitor in bean cell suspension cultures. Plant Molecular Biology 1991, 16, 81–94. [Google Scholar] [CrossRef] [PubMed]
- de Jonge, R.; van Esse H.P.; Kombrink, A.; Shinya, T.; Desaki, Y.; Bours, R.; ... Thomma, B.P.H.J. Conserved fungal lysM effector ecp6 prevents chitin-triggered immunity in Plants. Science, 2010, 329, 953–955. [CrossRef] [PubMed]
- Barreto-Bergter, E.; Figueiredo, R.T. Fungal glycans and the innate immune recognition. Frontiers in Cellular and Infection Microbiology 2014, 4, 145. [Google Scholar] [CrossRef] [PubMed]
- Mathur, R.S.; Barnett, H.l.; Lilly, V.G. Sporulation of Colletotrichum lindemuthianum in culture. Phytopathol. 1950, 40, 104–114. [Google Scholar] [CrossRef]
- Farrell, R.E. RNA Methodologies: Laboratory guide for isolation and characterization, 5th ed.; Elsevier: Amsterdam, Netherlands, 2017; pp. 1–855. [Google Scholar]
- Borges, A.; Tsai, S.M.; Caldas, D.G.G. Validation of reference genes for RT-qPCR normalization in common bean during biotic and abiotic stresses. Plant Cell Rep. 2012, 31, 827–838. [Google Scholar] [CrossRef] [PubMed]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; Rokhsar, D.S. Phytozome: a comparative platform for green plant genomics. Nucleic. Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T.L. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012, 13, 134. [Google Scholar] [CrossRef] [PubMed]
- Stothard, P. The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. BioTechniques 2000, 28, 1102–1104. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M. The MIQE Guidelines: minimum information for publication of quantitative real-time PCR Experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.; Huggett, J. qPCR primer design revisited. Biomol. Detect. Quantif. 2017, 14, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Svec, D.; Tichopad, A.; Novosadova, V.; Pfaffl, M.W.; Kubista, M. How good is a PCR efficiency estimate: Recommendations for precise and robust qPCR efficiency assessments. Biomol. Detect. Quantif. 2015, 3, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, R. Quantification on the LightCycler. In Rapid Cycle Real-Time PCR; Meuer, S., Wittwer, C., Nakagawara, K-I., Eds.; Springer: Heidelberg, Berlin, 2001; pp. 21–34. [Google Scholar] [CrossRef]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Dag, O.; Dolgun, A.; Konar, N.M. onewaytests: An R Package for One-Way Tests in Independent Groups Designs. The R Journal 2018, 10, 175–199. [Google Scholar] [CrossRef]
- R Core Team. R: A language and environment for statistical computing. R Foundation for Statistics Computing: Vienna, Austria. 2020. Available online: http://www.R-project.org/.
- Wickham, H. ggplot2: elegant graphics for data analysis. 2nd ed. Cham: Springer International Publishing: Imprint: Springer. 2016. [CrossRef]
- Galili, T.; O’Callaghan, A.; Sidi, J.; Sievert, C. Heatmaply: an R package for creating interactive cluster heatmaps for online publishing. Wren, J., editor. Bioinformatics 2018, 34, 1600–1602. [Google Scholar] [CrossRef] [PubMed]

| Gene | Gene model | C. lindemuthianum race 73 | ||||
|---|---|---|---|---|---|---|
| 24 hpi | 48 hpi | 72 hpi | 96 hpi | 120 hpi | ||
| Co-x | KTR2/3 | -1.4 | -2.1 | -1.4 | -2.9 | -1.3 |
| Co-1 | Phvul.001G243800 | 0.8 | 0.3 | 1.0 | -0.3 | 1.0 |
| Co-AC | Phvul.001G244300 | 0.1 | 0.5 | 0.4 | 0.0 | 0.1 |
| Phvul.001G244400 | 0.0 | -0.2 | -0.4 | -0.9 | -0.5 | |
| Phvul.001G244500 | 1.8 | 0.5 | 2.5 | 1.2 | 1.1 | |
| CoPv01CDRK /PhgPv01CDRK | Phvul.001G245300 | 1.0 | -0.2 | 0.9 | 0.0 | 1.0 |
| Phvul.001G246300 | -0.7 | -3.8 | -2.3 | -4.1 | -4.8 | |
| Pathogenesis-related genes | Phvul.003G109100 (PR1a) | -0.9 | -1.6 | 1.4 | 2.3 | 1.8 |
| Phvul.006G196900 (PR1b) | -1.6 | -2.0 | 1.4 | 3.8 | 3.9 | |
| Phvul.009G256400 (PR2) | -1.2 | -1.0 | 0.6 | 1.2 | 1.8 | |
| Genea | Reference | Primers Forward (F) and | Tm | Amplicon | E | R² |
|---|---|---|---|---|---|---|
| Reverse (R) (5`-3`) | (°C) | (bp) | ||||
| Phvul.001G133200* | IDE | F: AAGCAGGTATCTTGGCCATCTC | F: 60.16 | 126 | 0.92 | 0.99 |
| R: AAAGCAAACTCCAAGCTCCAATC | R: 59.99 | |||||
| Phvul.008G011000* | ACT | F: ACAGCCAGGACCAGTTCATC | F: 59.67 | 154 | 0.93 | 0.98 |
| R: TGTATGTGGTCTCGTGAATGC | R: 58.38 | |||||
| Phvul.001G243800 | Co-1 | F: CCTCAAGGTGGGGCTTTTGAG | F: 61.16 | 118 | 1.01 | 0.99 |
| R: TCACCGAGAAACTCCCATTGC | R: 60.61 | |||||
| KTR2/3 | Co-x | F: ATGCACAGGGGAATGGGATG | F: 60.11 | 279 | 1.06 | 0.98 |
| R: GCCATAGCGAGTGAGAGTGCG | R: 63.42 | |||||
| Phvul.001G244300 | Co-AC | F: GAAACGTCTCCGCAGAATAGTG | F: 59.40 | 150 | 0.99 | 0.99 |
| R: GTCTTGTGTTGTTCCTTGGAGTTG | R: 60.44 | |||||
| Phvul.001G244400 | Co-AC | F: TACAGCAAGAGAGCGGTTAAAGG | F: 60.62 | 121 | 1.07 | 0.99 |
| R: CCCTTTGTCACTTTGTTTTGAAGC | R: 59.67 | |||||
| Phvul.001G244500 | Co-AC | F: CAATGCACAGCTCGCAACTC | F: 60.45 | 141 | 1.09 | 0.98 |
| R: GGAACTGTGAAAGCTCTGCTAAC | R: 59.81 | |||||
| Phvul.001G245300 | CoPv01CDRK | F: TCTGCTGGAAGGGTGGTAGTC | F: 61.17 | 93 | 1.04 | 0.99 |
| R: GGACGTTATGTGAACAAGGTTTGC | R: 61.08 | |||||
| Phvul.001G246300 | CoPv01CDRK | F: CTTCTTCCCTTCACTTCGATACC | F: 58.57 | 87 | 0.95 | 0.99 |
| R: GTTGAGAGTGTTTGTGGCAGT | R: 58.98 | |||||
| Phvul.003G109100 | PR1a | F: GTCCTAACGGAGGATCACTCA | F: 58.62 | 148 | 1.01 | 0.98 |
| R: CAGGGATTGGCCAGAAGGTAT | R: 59.50 | |||||
| Phvul.006G196900 | PR1b | F: GGTTTGCCTATGATCCCAATGC | F: 59.96 | 115 | 0.99 | 0.99 |
| R: TGTTGTGAGCGTTGAGGAAGTC | R: 61.06 | |||||
| Phvul.009G256400 | PR2 | F: CAGAGGTTCTCATTTGCTGCTTTC | F: 60.62 | 98 | 1.09 | 0.99 |
| R: ATGCCATAACACACCCCGATTTG | R: 61.75 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




