Submitted:
23 May 2023
Posted:
24 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Protein-solid surface interaction in UA
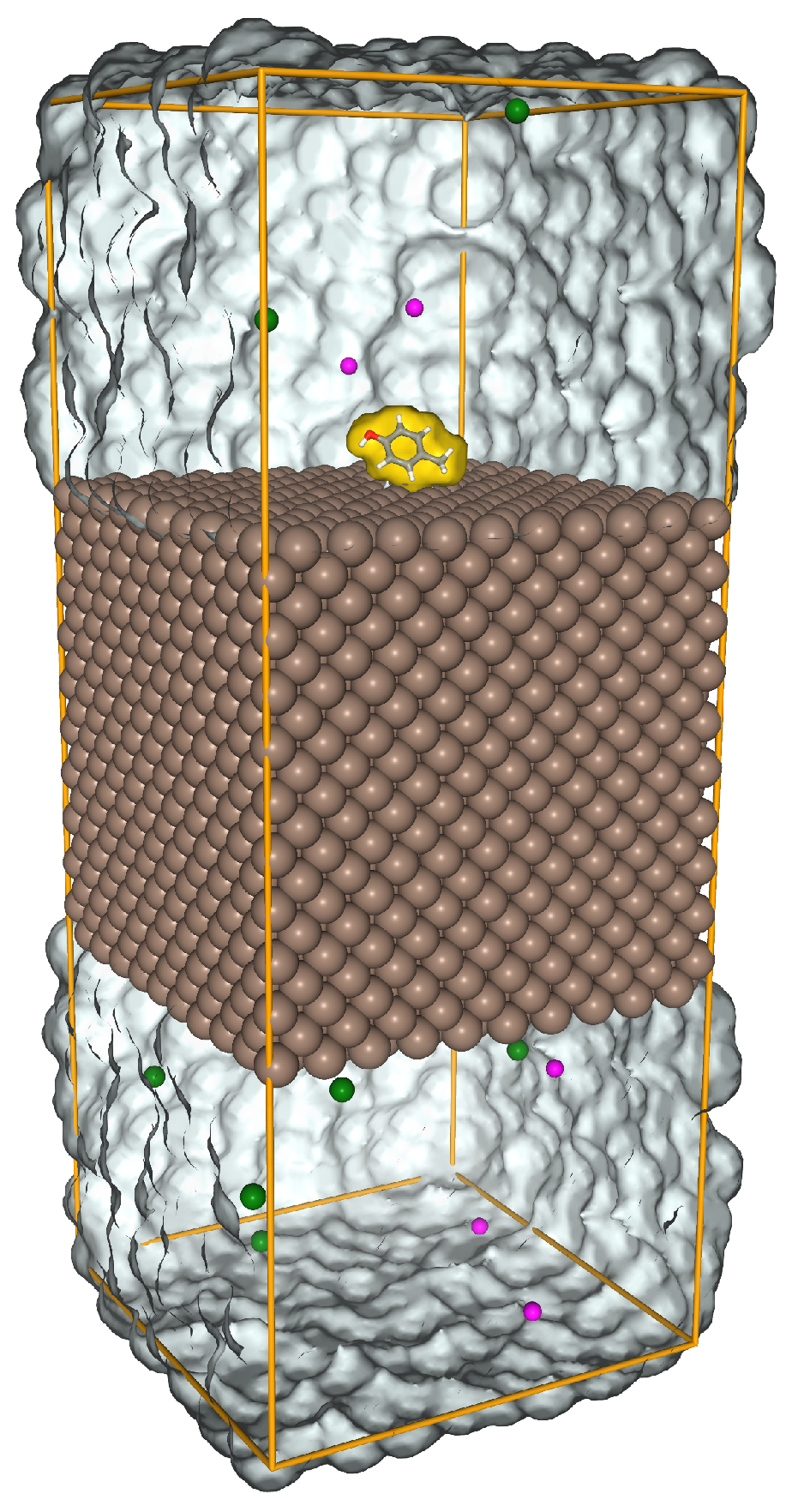
2.2. All-atom model for recovering short-range potentials
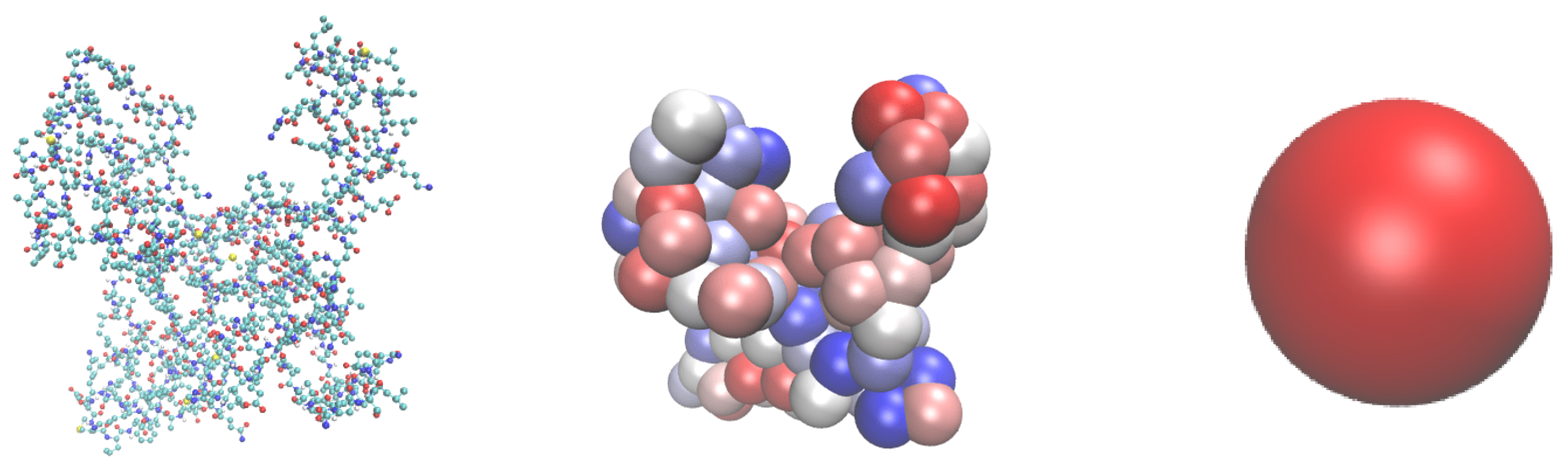
2.3. Preparation of starting coordinates for biomolecules and surfaces
2.4. Competitive adsorption model
3. Results and Discussion
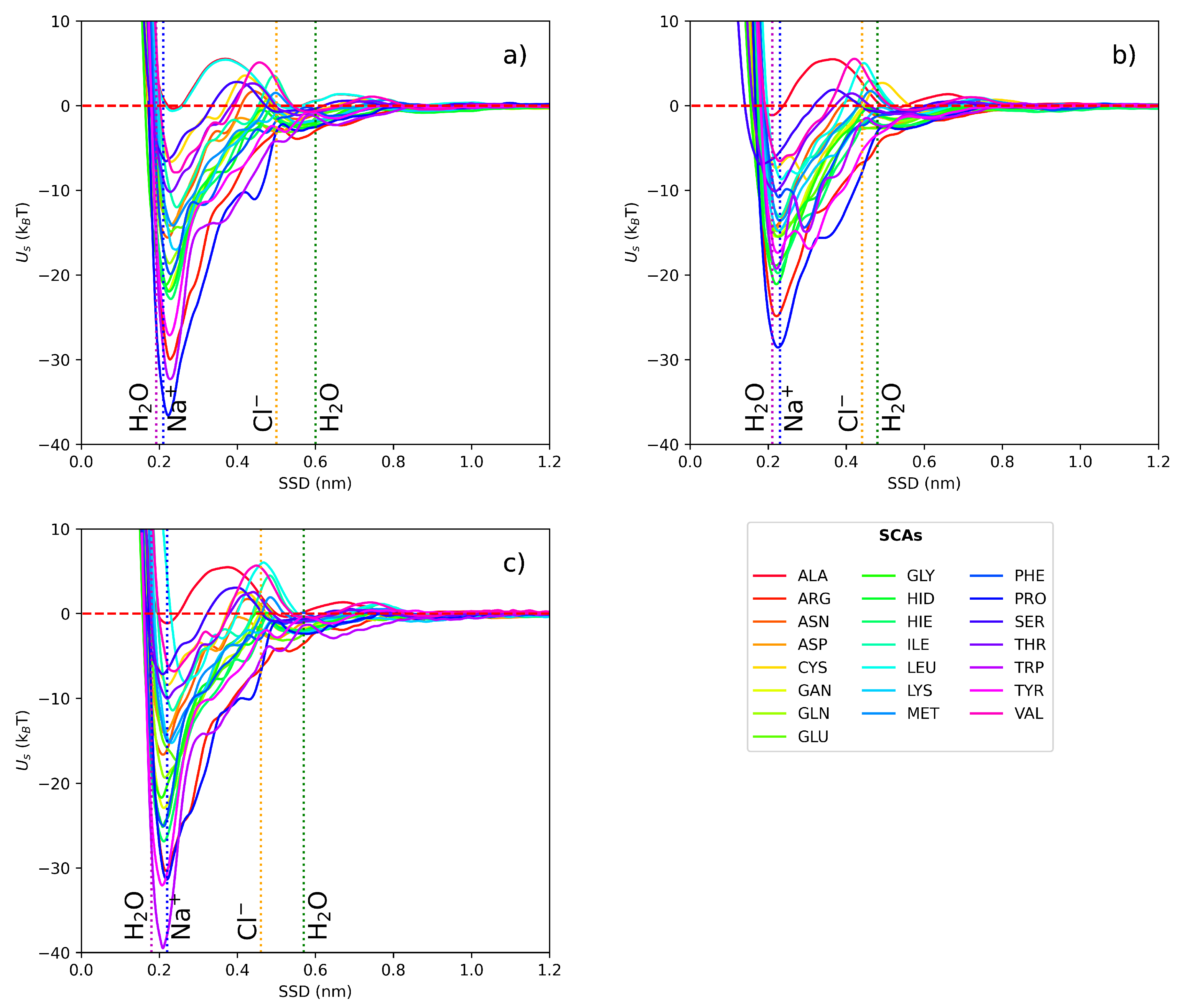
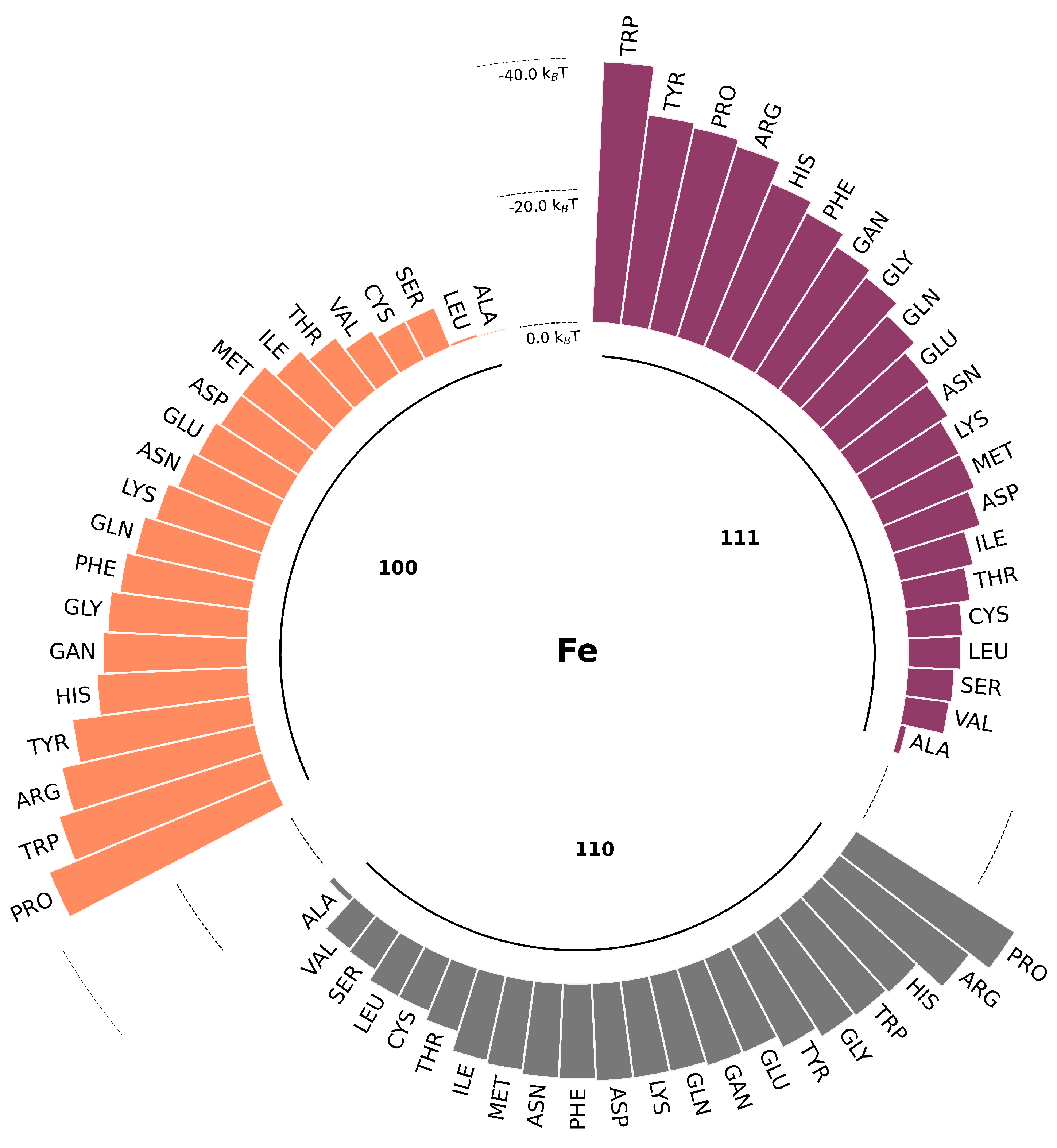
3.1. Short-range potentials for SCAs on iron surfaces
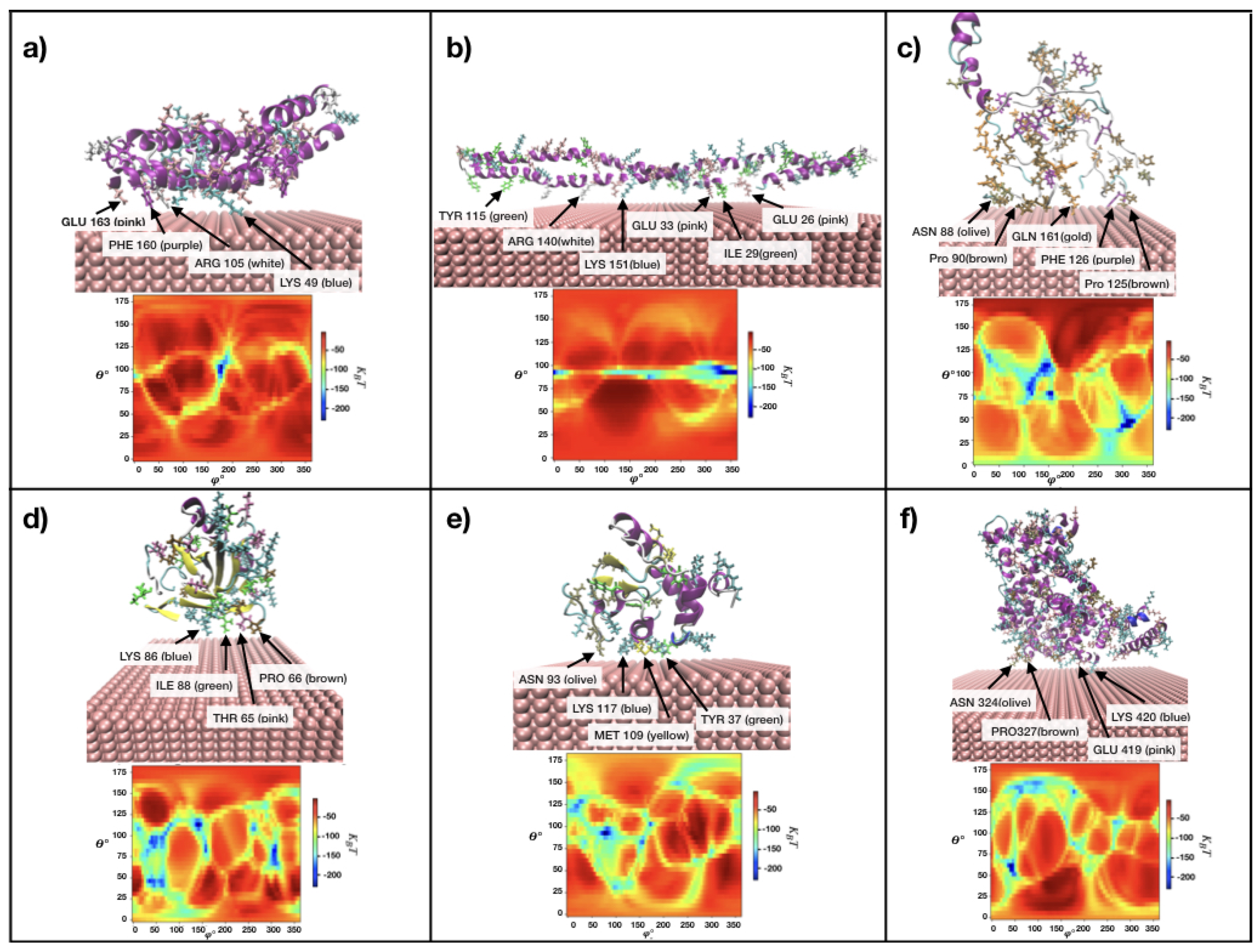
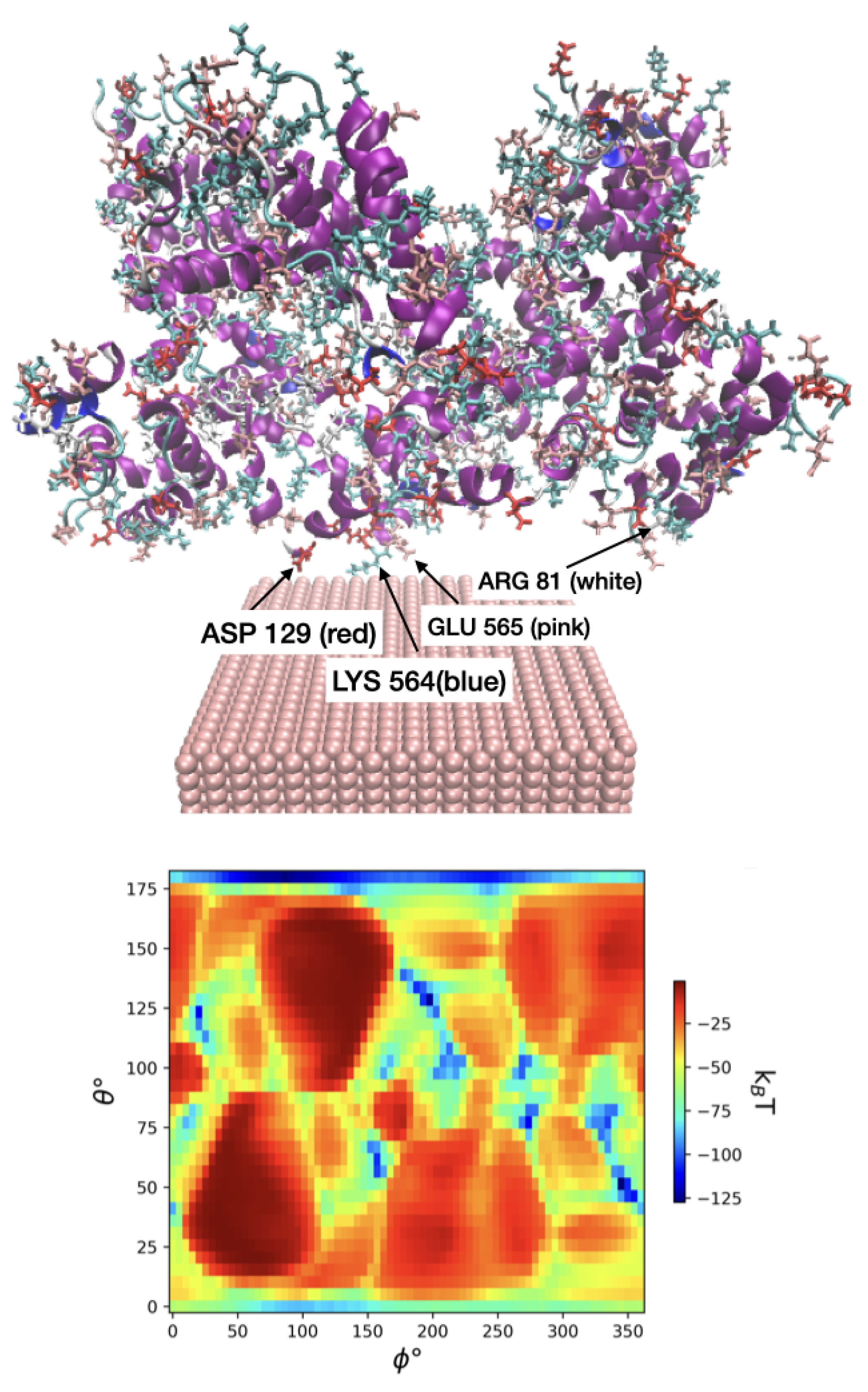
3.2. Protein adsorption energies, preferred orientations and heatmaps
3.3. Validation of UA model parameters
3.4. Competitive adsorption and milk protein layer
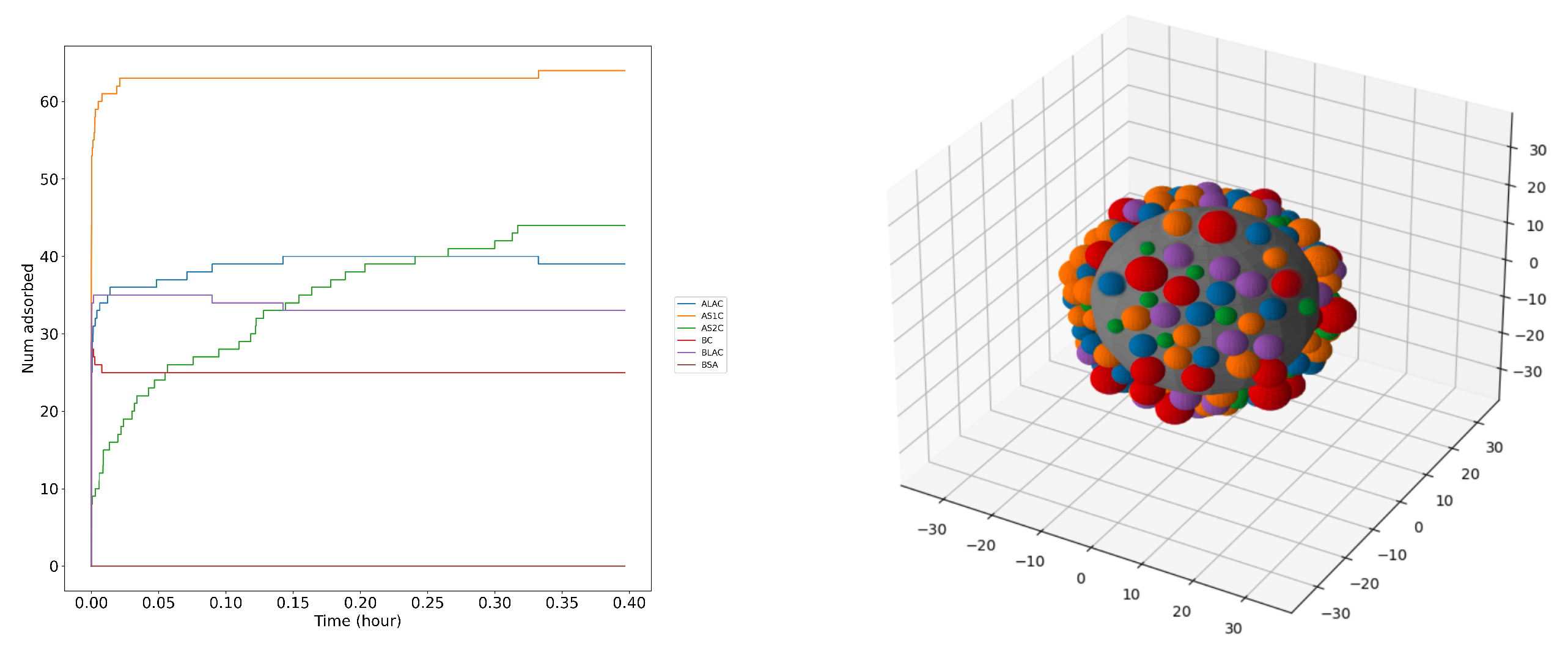
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AA | Amino acid |
| ALAC | -lactalbumins |
| AS1C | -casein |
| AS2C | -casein |
| AWR-MetaD | Adaptive well-tempered metadynamics |
| BC | -caseins |
| BLAC | -lactoglobulins |
| BSA | Bovine Serum Albumin |
| CG | Coarse-grained |
| AA | Amino acid |
| COM | Center of mass |
| EM | Engineered material |
| fcc | Face-centered cubic |
| Fe NP | Fe nanoparticle |
| HS | Hard-sphere |
| HSA | Human serum albumin |
| I-TASSER | Iterative Threading ASSEmbly Refinement |
| KMC | Kinetic Monte Carlo |
| MD | Molecular dynamics |
| MW | Molecular weight |
| NP | Nanoparticle |
| PMF | Potential of mean force |
| PDB | Protein data bank |
| SCA | Side chain analogues |
| SSD | Surface separation distance |
| UA | United Atom |
| vdW | van der Waals |
References
- Hesterberg, T.W.; Long, C.M.; Lapin, C.A.; Hamade, A.K.; Valberg, P.A. Diesel exhaust particulate (DEP) and nanoparticle exposures: what do DEP human clinical studies tell us about potential human health hazards of nanoparticles. Inhalation toxicology 2010, 22, 679–694. [CrossRef]
- Gopal, N.; Hill, C.; Ross, P.R.; Beresford, T.P.; Fenelon, M.A.; Cotter, P.D. The prevalence and control of Bacillus and related spore-forming bacteria in the dairy industry. Frontiers in microbiology 2015, 6, 1418. [CrossRef]
- Marchand, S.; De Block, J.; De Jonghe, V.; Coorevits, A.; Heyndrickx, M.; Herman, L. Biofilm formation in milk production and processing environments; influence on milk quality and safety. Comprehensive Reviews in Food Science and Food Safety 2012, 11, 133–147. [CrossRef]
- Flint, S.; Bremer, P.; Brooks, J. Biofilms in dairy manufacturing plant-description, current concerns and methods of control. Biofouling 1997, 11, 81–97. [CrossRef]
- Bansal, B.; Chen, X.D. A critical review of milk fouling in heat exchangers. Comprehensive reviews in food science and food safety 2006, 5, 27–33. [CrossRef]
- Rosmaninho, R.; Santos, O.; Nylander, T.; Paulsson, M.; Beuf, M.; Benezech, T.; Yiantsios, S.; Andritsos, N.; Karabelas, A.; Rizzo, G.; others. Modified stainless steel surfaces targeted to reduce fouling–Evaluation of fouling by milk components. Journal of Food Engineering 2007, 80, 1176–1187. [CrossRef]
- Rahman, M.; Laurent, S.; Tawil, N.; Yahia, L.; Mahmoudi, M.; Rahman, M.; Laurent, S.; Tawil, N.; Yahia, L.; Mahmoudi, M. Nanoparticle and protein corona. Protein-Nanoparticle Interactions: The Bio-Nano Interface 2013, pp. 21–44.
- Barroso da Silva, F.L.; Carloni, P.; Cheung, D.; Cottone, G.; Donnini, S.; Foegeding, E.A.; Gulzar, M.; Jacquier, J.C.; Lobaskin, V.; MacKernan, D.; others. Understanding and controlling food protein structure and function in foods: perspectives from experiments and computer simulations. Annual review of food science and technology 2020, 11, 365–387. [CrossRef]
- Changani, S.; Belmar-Beiny, M.; Fryer, P. Engineering and chemical factors associated with fouling and cleaning in milk processing. Experimental Thermal and Fluid Science 1997, 14, 392–406. [CrossRef]
- Förster, M.; Bohnet, M. Influence of the interfacial free energy crystal/heat transfer surface on the induction period during fouling. International journal of thermal sciences 1999, 38, 944–954. [CrossRef]
- Tang, L.; Thevenot, P.; Hu, W. Surface chemistry influences implant biocompatibility. Current topics in medicinal chemistry 2008, 8, 270–280. [CrossRef]
- Wilson, C.J.; Clegg, R.E.; Leavesley, D.I.; Pearcy, M.J. Mediation of biomaterial–cell interactions by adsorbed proteins: a review. Tissue engineering 2005, 11, 1–18. [CrossRef]
- Tang, L.; Hu, W. Molecular determinants of biocompatibility. Expert review of medical devices 2005, 2, 493–500. [CrossRef]
- Landsiedel, R.; Ma-Hock, L.; Kroll, A.; Hahn, D.; Schnekenburger, J.; Wiench, K.; Wohlleben, W. Testing metal-oxide nanomaterials for human safety. Advanced Materials 2010, 22, 2601–2627. [CrossRef]
- Winkler, D.A.; Burden, F.R.; Yan, B.; Weissleder, R.; Tassa, C.; Shaw, S.; Epa, V.C. Modelling and predicting the biological effects of nanomaterials. SAR and QSAR in Environmental Research 2014, 25, 161–172. [CrossRef]
- Dell’Orco, D.; Lundqvist, M.; Oslakovic, C.; Cedervall, T.; Linse, S. Modeling the time evolution of the nanoparticle-protein corona in a body fluid. PloS one 2010, 5, e10949. [CrossRef]
- Darabi Sahneh, F.; Scoglio, C.; Riviere, J. Dynamics of nanoparticle-protein corona complex formation: analytical results from population balance equations. PloS one 2013, 8, e64690. [CrossRef]
- Shao, Q.; Hall, C.K. Protein adsorption on nanoparticles: model development using computer simulation. Journal of Physics: Condensed Matter 2016, 28, 414019. [CrossRef]
- Vilaseca, P.; Dawson, K.A.; Franzese, G. Understanding and modulating the competitive surface-adsorption of proteins through coarse-grained molecular dynamics simulations. Soft Matter 2013, 9, 6978–6985. [CrossRef]
- Angioletti-Uberti, S.; Ballauff, M.; Dzubiella, J. Competitive adsorption of multiple proteins to nanoparticles: the Vroman effect revisited. Molecular Physics 2018, 116, 3154–3163. [CrossRef]
- Rouse, I.; Power, D.; Brandt, E.G.; Schneemilch, M.; Kotsis, K.; Quirke, N.; Lyubartsev, A.P.; Lobaskin, V. Advanced in silico characterization of nanomaterials for nanoparticle toxicology. arXiv preprint arXiv:2007.04017 2020. [CrossRef]
- Rouse, I.; Lobaskin, V. A hard-sphere model of protein corona formation on spherical and cylindrical nanoparticles. Biophysical Journal 2021, 120, 4457–4471. [CrossRef]
- Monopoli, M.P.; Walczyk, D.; Campbell, A.; Elia, G.; Lynch, I.; Baldelli Bombelli, F.; Dawson, K.A. Physical- chemical aspects of protein corona: relevance to in vitro and in vivo biological impacts of nanoparticles. Journal of the American Chemical Society 2011, 133, 2525–2534. [CrossRef]
- Vilanova, O.; Mittag, J.J.; Kelly, P.M.; Milani, S.; Dawson, K.A.; Rädler, J.O.; Franzese, G. Understanding the kinetics of protein–nanoparticle corona formation. ACS nano 2016, 10, 10842–10850. [CrossRef]
- Hasenkopf, I.; Mills-Goodlet, R.; Johnson, L.; Rouse, I.; Geppert, M.; Duschl, A.; Maier, D.; Lobaskin, V.; Lynch, I.; Himly, M. Computational prediction and experimental analysis of the nanoparticle-protein corona: Showcasing an in vitro-in silico workflow providing FAIR data. Nano Today 2022, 46, 101561. [CrossRef]
- Lopez, H.; Lobaskin, V. Coarse-grained model of adsorption of blood plasma proteins onto nanoparticles. The Journal of chemical physics 2015, 143, 12B620_1. [CrossRef]
- Walkey, C.D.; Olsen, J.B.; Song, F.; Liu, R.; Guo, H.; Olsen, D.W.H.; Cohen, Y.; Emili, A.; Chan, W.C. Protein corona fingerprinting predicts the cellular interaction of gold and silver nanoparticles. ACS nano 2014, 8, 2439–2455. [CrossRef]
- Eskin, N.A.M.; Goff, H.D. Chapter 4 - Milk. In Biochemistry of Foods (Third Edition), Third Edition ed.; Eskin, N.M.; Shahidi, F., Eds.; Academic Press: San Diego, 2013; pp. 187–214. doi:https://doi.org/10.1016/B978-0-08-091809-9.00004-2. [CrossRef]
- Brandt, E.G.; Lyubartsev, A.P. Molecular dynamics simulations of adsorption of amino acid side chain analogues and a titanium binding peptide on the TiO2 (100) surface. The Journal of Physical Chemistry C 2015, 119, 18126–18139. [CrossRef]
- Subbotina, J.; Lobaskin, V. Multiscale Modeling of Bio-Nano Interactions of Zero-Valent Silver Nanoparticles. The Journal of Physical Chemistry B 2022, 126, 1301–1314. [CrossRef]
- Power, D.; Rouse, I.; Poggio, S.; Brandt, E.; Lopez, H.; Lyubartsev, A.; Lobaskin, V. A multiscale model of protein adsorption on a nanoparticle surface. Modelling and Simulation in Materials Science and Engineering 2019, 27, 084003. [CrossRef]
- Alsharif, S.A.; Power, D.; Rouse, I.; Lobaskin, V. In silico prediction of protein adsorption energy on titanium dioxide and gold nanoparticles. Nanomaterials 2020, 10, 1967. [CrossRef]
- del Giudice, G.; Serra, A.; Saarimäki, L.; Kotsis, K.; Rouse, I.; Colibaba, A.; Jagiello, K.; Mikolajczyk, A.; Papadiamantis, A.; Sanabria, N.; others. A gene regulation model reveals an ancestral adaptation response to particulate exposure triggered by nanomaterials 2022. [CrossRef]
- Repositories of United Atom code and tools. https://bitbucket. org/softmattergroup/ (accessed Dec 11, 2020).
- NanoCommons. Nano-Knowledge Community. https:// www.nanocommons.eu/ (accessed Jan 15, 2021).
- Bellion, M.; Santen, L.; Mantz, H.; Hähl, H.; Quinn, A.; Nagel, A.; Gilow, C.; Weitenberg, C.; Schmitt, Y.; Jacobs, K. Protein adsorption on tailored substrates: long-range forces and conformational changes. Journal of Physics: Condensed Matter 2008, 20, 404226. [CrossRef]
- Darve, E.; Pohorille, A. Calculating free energies using average force. The Journal of chemical physics 2001, 115, 9169–9183. [CrossRef]
- Wang, F.; Landau, D.P. Efficient, multiple-range random walk algorithm to calculate the density of states. Physical review letters 2001, 86, 2050. [CrossRef]
- Marsili, S.; Barducci, A.; Chelli, R.; Procacci, P.; Schettino, V. Self-healing umbrella sampling: a non-equilibrium approach for quantitative free energy calculations. The Journal of Physical Chemistry B 2006, 110, 14011–14013. [CrossRef]
- Maragliano, L.; Vanden-Eijnden, E. A temperature accelerated method for sampling free energy and determining reaction pathways in rare events simulations. Chemical physics letters 2006, 426, 168–175. [CrossRef]
- Dickson, B.M.; Legoll, F.; Lelievre, T.; Stoltz, G.; Fleurat-Lessard, P. Free energy calculations: An efficient adaptive biasing potential method. The Journal of Physical Chemistry B 2010, 114, 5823–5830. [CrossRef]
- Laio, A.; Parrinello, M. Escaping free-energy minima. Proceedings of the national academy of sciences 2002, 99, 12562–12566. [CrossRef]
- Laio, A.; Gervasio, F.L. Metadynamics: a method to simulate rare events and reconstruct the free energy in biophysics, chemistry and material science. Reports on Progress in Physics 2008, 71, 126601. [CrossRef]
- Hess, B.; Kutzner, C.; Van Der Spoel, D.; Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. Journal of chemical theory and computation 2008, 4, 435–447. [CrossRef]
- Pronk, S.; Páll, S.; Schulz, R.; Larsson, P.; Bjelkmar, P.; Apostolov, R.; Shirts, M.R.; Smith, J.C.; Kasson, P.M.; Van Der Spoel, D.; others. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 2013, 29, 845–854. [CrossRef]
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. Journal of computational chemistry 2008, 29, 1859–1865. [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. The Journal of chemical physics 1983, 79, 926–935. [CrossRef]
- Berendsen, H.J.; Postma, J.v.; Van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. The Journal of chemical physics 1984, 81, 3684–3690. [CrossRef]
- Tribello, G.A.; Bonomi, M.; Branduardi, D.; Camilloni, C.; Bussi, G. PLUMED 2: New feathers for an old bird. Computer physics communications 2014, 185, 604–613. [CrossRef]
- Brooks, B.R.; Brooks III, C.L.; Mackerell Jr, A.D.; Nilsson, L.; Petrella, R.J.; Roux, B.; Won, Y.; Archontis, G.; Bartels, C.; Boresch, S.; others. CHARMM: the biomolecular simulation program. Journal of computational chemistry 2009, 30, 1545–1614. [CrossRef]
- Lee, J.; Cheng, X.; Swails, J.M.; Yeom, M.S.; Eastman, P.K.; Lemkul, J.A.; Wei, S.; Buckner, J.; Jeong, J.C.; Qi, Y.; others. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. Journal of chemical theory and computation 2016, 12, 405–413. [CrossRef]
- Roy, A.; Kucukural, A.; Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nature protocols 2010, 5, 725–738. [CrossRef]
- Meurant, G. Handbook of milk composition; Elsevier, 1995.
- Reinhardt, T.A.; Lippolis, J.D.; Nonnecke, B.J.; Sacco, R.E. Bovine milk exosome proteome. Journal of proteomics 2012, 75, 1486–1492. [CrossRef]
- Olsson, M.H.; Søndergaard, C.R.; Rostkowski, M.; Jensen, J.H. PROPKA3: consistent treatment of internal and surface residues in empirical p K a predictions. Journal of chemical theory and computation 2011, 7, 525–537.
- Søndergaard, C.R.; Olsson, M.H.; Rostkowski, M.; Jensen, J.H. Improved treatment of ligands and coupling effects in empirical calculation and rationalization of p K a values. Journal of chemical theory and computation 2011, 7, 2284–2295. [CrossRef]
- Delboni, L.A.; Da Silva, F.L.B. On the complexation of whey proteins. Food Hydrocolloids 2016, 55, 89–99. [CrossRef]
- Dupont, D.; Croguennec, T.; Pochet, S. Milk Proteins-Analytical Methods. Reference Module in Food Science 2018, p. np.
- Davies, D.T.; Law, A.J. The content and composition of protein in creamery milks in south-west Scotland. Journal of Dairy Research 1980, 47, 83–90. [CrossRef]
- Kalyankar, S.; Khedkar, C.; Patil, A.; Deosarkar, S. Milk: Sources and composition 2016.
- Davoodi, S.H.; Shahbazi, R.; Esmaeili, S.; Sohrabvandi, S.; Mortazavian, A.; Jazayeri, S.; Taslimi, A. Health-related aspects of milk proteins. Iranian journal of pharmaceutical research: IJPR 2016, 15, 573.
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; others. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [CrossRef]
- Kharazian, B.; Hadipour, N.; Ejtehadi, M. Understanding the nanoparticle–protein corona complexes using computational and experimental methods. The international journal of biochemistry & cell biology 2016, 75, 162–174. [CrossRef]
- Casals, E.; Pfaller, T.; Duschl, A.; Oostingh, G.J.; Puntes, V. Time evolution of the nanoparticle protein corona. ACS nano 2010, 4, 3623–3632. [CrossRef]
- Talbot, J.; Tarjus, G.; Van Tassel, P.; Viot, P. From car parking to protein adsorption: an overview of sequential adsorption processes. Colloids and Surfaces A: Physicochemical and Engineering Aspects 2000, 165, 287–324. [CrossRef]
- Wright, L.B.; Rodger, P.M.; Corni, S.; Walsh, T.R. GolP-CHARMM: first-principles based force fields for the interaction of proteins with Au (111) and Au (100). Journal of chemical theory and computation 2013, 9, 1616–1630. [CrossRef]
- Feng, J.; Pandey, R.B.; Berry, R.J.; Farmer, B.L.; Naik, R.R.; Heinz, H. Adsorption mechanism of single amino acid and surfactant molecules to Au {111} surfaces in aqueous solution: design rules for metal-binding molecules. Soft Matter 2011, 7, 2113–2120.
- Nawrocki, G.; Cieplak, M. Aqueous amino acids and proteins near the surface of gold in hydrophilic and hydrophobic force fields. The Journal of Physical Chemistry C 2014, 118, 12929–12943. [CrossRef]
- Hoefling, M.; Iori, F.; Corni, S.; Gottschalk, K.E. Interaction of amino acids with the Au (111) surface: adsorption free energies from molecular dynamics simulations. Langmuir 2010, 26, 8347–8351. [CrossRef]
- Shao, Q.; Hall, C.K. Binding preferences of amino acids for gold nanoparticles: a molecular simulation study. Langmuir 2016, 32, 7888–7896. [CrossRef]
- Fazlzadeh, M.; Ansarizadeh, M.; Leili, M. Data of furfural adsorption on nano zero valent iron (NZVI) synthesized from Nettle extract. Data in brief 2018, 16, 341–345. [CrossRef]
- Xu, L.; Kirvassilis, D.; Bai, Y.; Mavrikakis, M. Atomic and molecular adsorption on Fe (110). Surface Science 2018, 667, 54–65. [CrossRef]
- Anbouhi, T.S.; Esfidvajani, E.M.; Nemati, F.; Haghighat, S.; Sari, S.; Attar, F.; Pakaghideh, A.; Sohrabi, M.J.; Mousavi, S.E.; Falahati, M. Albumin binding, anticancer and antibacterial properties of synthesized zero valent iron nanoparticles. International journal of nanomedicine 2019, 14, 243. [CrossRef]
- Power, D.; Poggio, S.; Lopez, H.; Lobaskin, V. Bionano interactions: a key to mechanistic understanding of nanoparticle toxicity. In Computational Nanotoxicology; Jenny Stanford Publishing, 2019; pp. 189–215.
| Abbreviation | UniProt ID | Protein Name | , Da | Charge, e | ,mol/l | |
|---|---|---|---|---|---|---|
| AS1C | P02662 | -casein | 24528.00 | -8.5 | 214 | 4 |
| AS2C | P02663 | -casein | 26018.69 | 4.5 | 222 | 1 |
| BC | P02666 | -casein | 25107.33 | -4.5 | 224 | 4 |
| ALAC | P00711 | -lactalbumin | 16246.61 | -5 | 142 | 0.9 |
| BLAC | P02754 | -lactoglobulin | 19883.25 | -6 | 178 | 2 |
| BSA | P02769 | Bovine Serum Albumin | 69293.41 | -4.5 | 607 | 0.1 |
| Individual protein adsorption description on Fe-10 | ||||
| Protein, | ,T | , | , | , nm |
| AS1C | -343.07 | 175 | 100 | 0.14 |
| AS2C | -238.64 | 335 | 90 | 0.08 |
| BC | -211.96 | 315 | 45 | 0.04 |
| BSA | -202.99 | 40 | 60 | 0.08 |
| ALAC | -159.12 | 80 | 90 | 0.18 |
| BLAC | -142.98 | 140 | 110 | 0.19 |
| Individual protein adsorption description on Fe-110 | ||||
| Protein, | ,T | , | , | , nm |
| AS1C | -325.92 | 175 | 100 | 0.14 |
| AS2C | -229.90 | 335 | 90 | 0.07 |
| BSA | -189.87 | 40 | 60 | 0.06 |
| BC | -162.43 | 310 | 45 | 0.04 |
| BLAC | -147.43 | 140 | 110 | 0.18 |
| ALAC | -144.41 | 75 | 90 | 0.16 |
| Individual protein adsorption description on Fe-111 | ||||
| Protein, | ,T | , | , | , nm |
| AS1C | -330.36 | 175 | 100 | 0.14 |
| AS2C | -230.99 | 330 | 90 | 0.10 |
| BC | -186.44 | 310 | 40 | 0.02 |
| BSA | -178.25 | 40 | 60 | 0.06 |
| ALAC | -154.60 | 75 | 90 | 0.15 |
| BLAC | -151.56 | 140 | 110 | 0.18 |
| Fe-100 | Fe-100 | Fe-110 | Fe-110 | Fe-111 | Fe-111 | |
|---|---|---|---|---|---|---|
| Protein | , nm] | ,% | , nm] | , % | , nm] | , % |
| AS1C | 9.8 | 41.22 | 9.0 | 38.77 | 9.6 | 40.33 |
| BLAC | 5.6 | 19.18 | 5.5 | 19.37 | 5.4 | 18.50 |
| BS | 4.1 | 17.40 | 4.2 | 18.80 | 4.3 | 18.29 |
| ALAC | 5.5 | 15.43 | 5.6 | 16.29 | 5.7 | 15.94 |
| AS2C | 1.4 | 6.19 | 1.3 | 6.15 | 1.4 | 6.19 |
| BSA | 0.05 | 0.59 | 0.05 | 0.60 | 0.06 | 0.73 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




