Submitted:
15 May 2023
Posted:
16 May 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
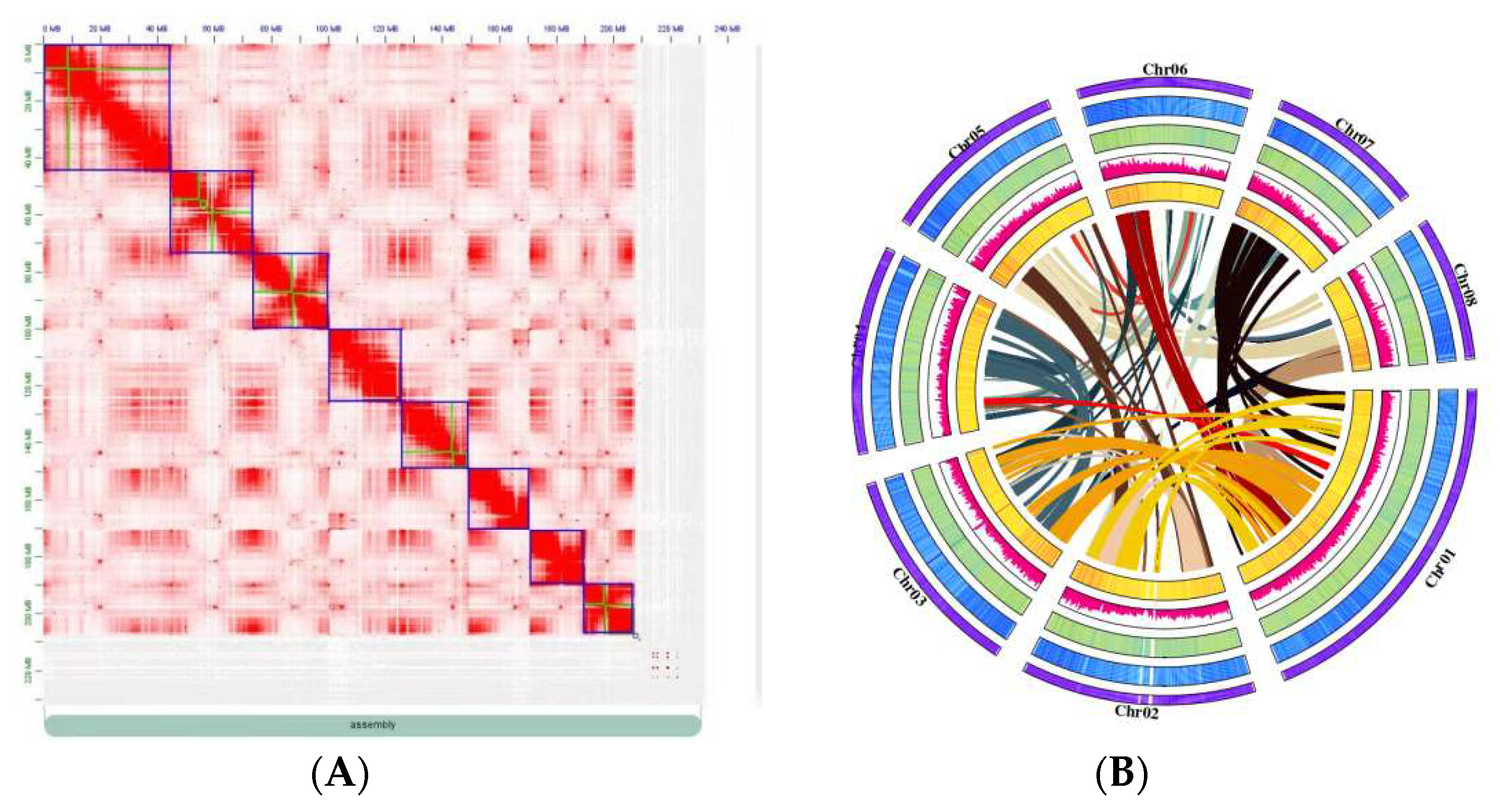
2.1. De novo assembly of the wild almond genome
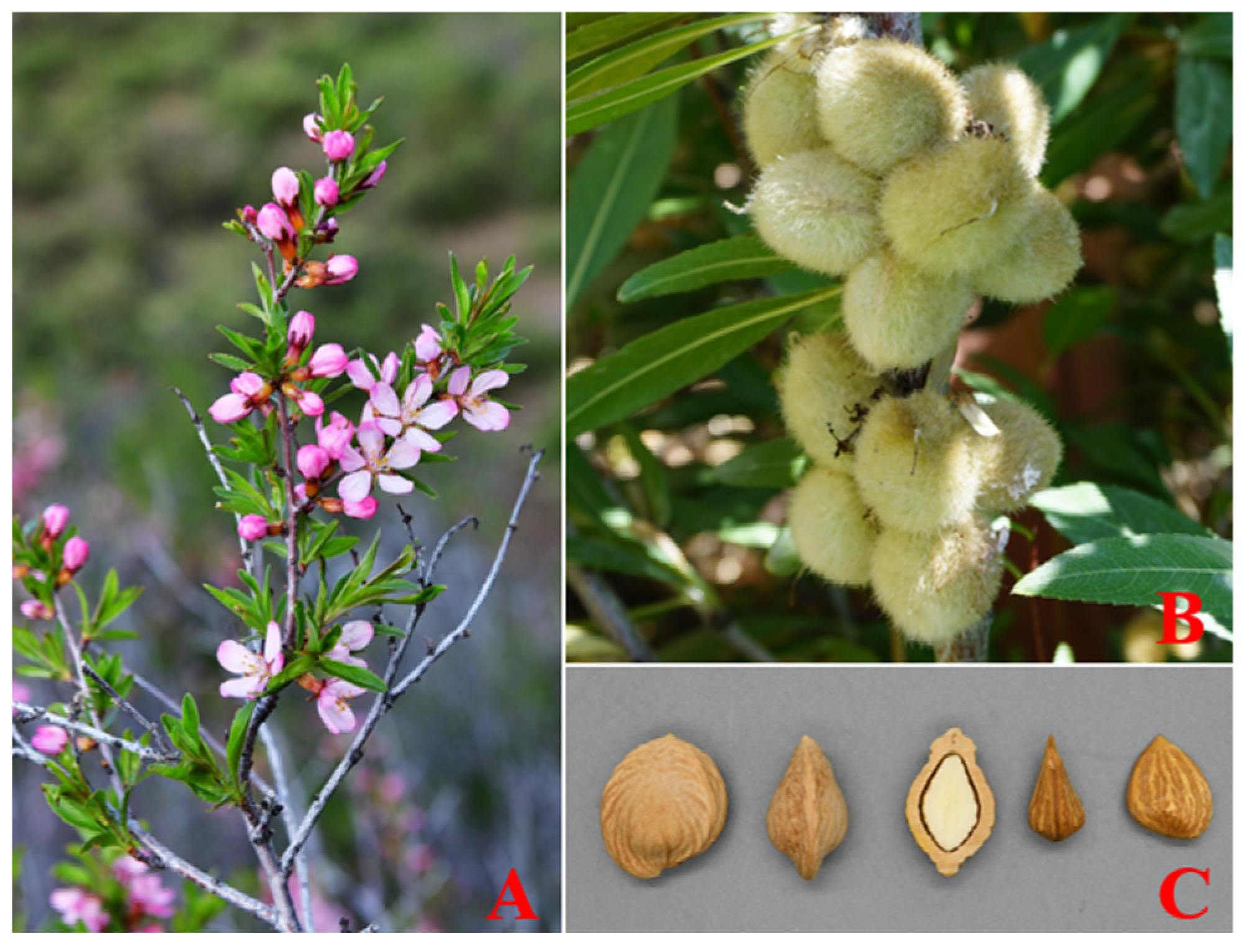

2.2. Functional annonations, gene prediction, and repetitive sequences
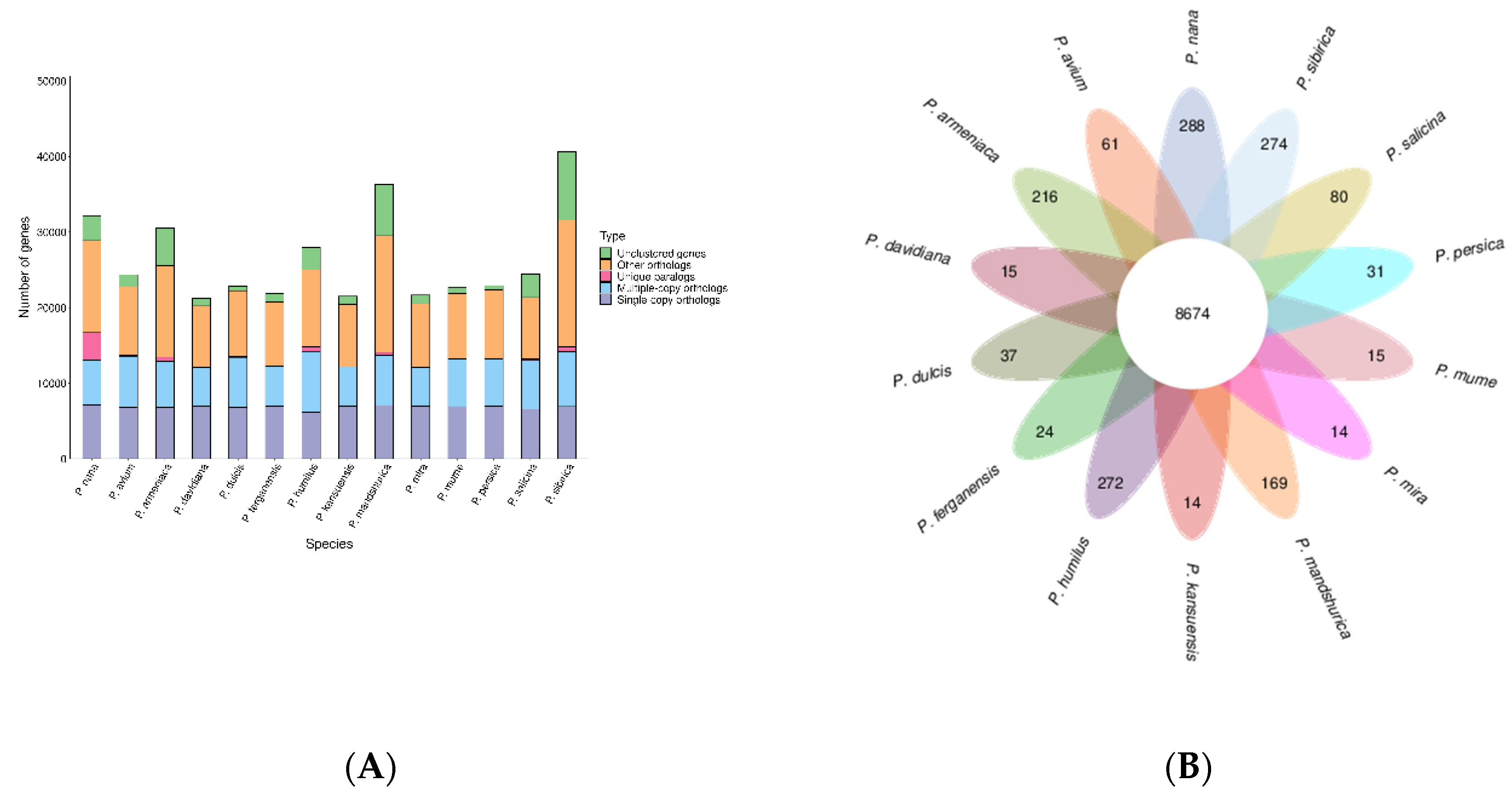
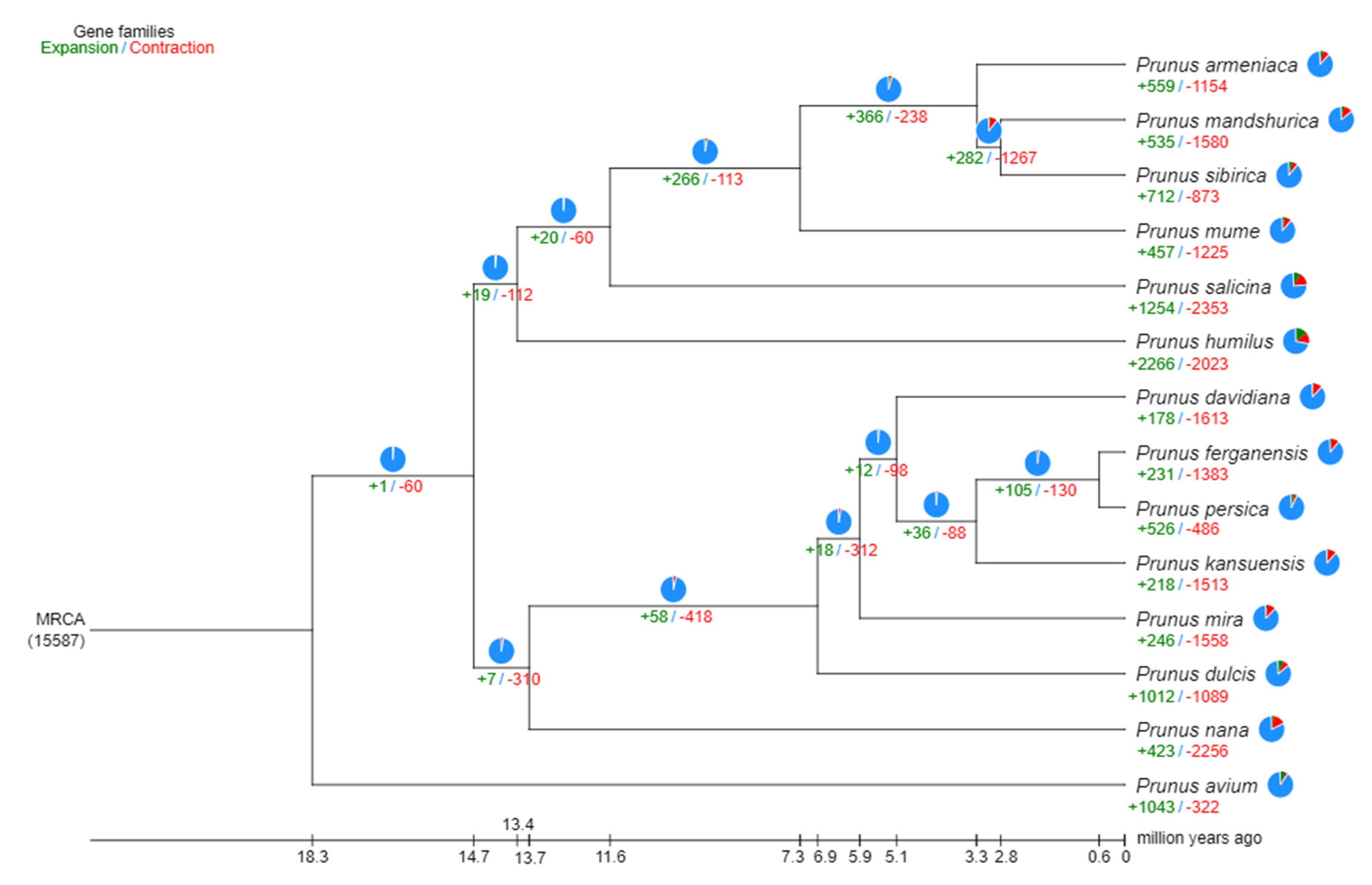
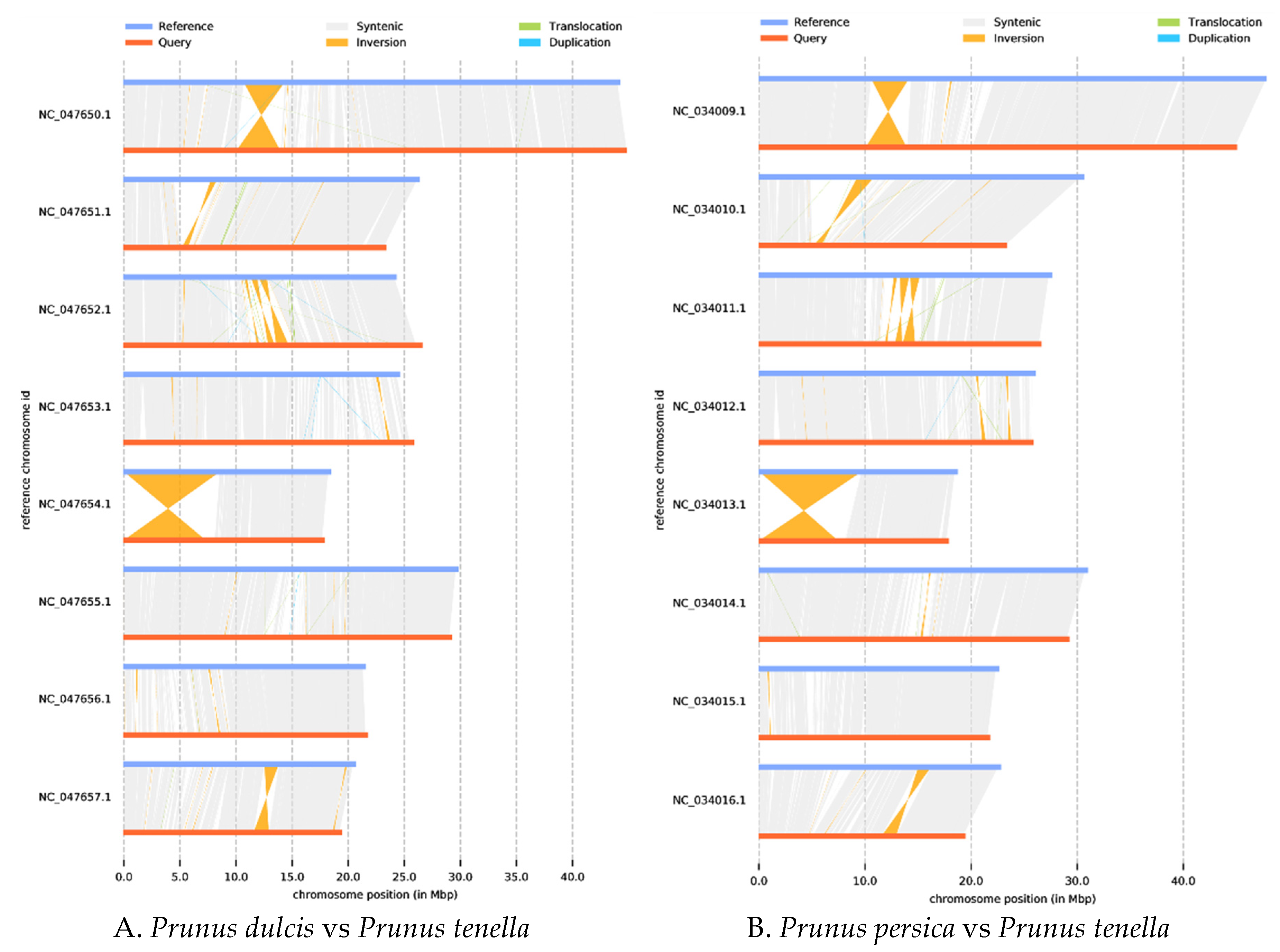
2.3. Synteny analysis, and genome evolution and phylogeny
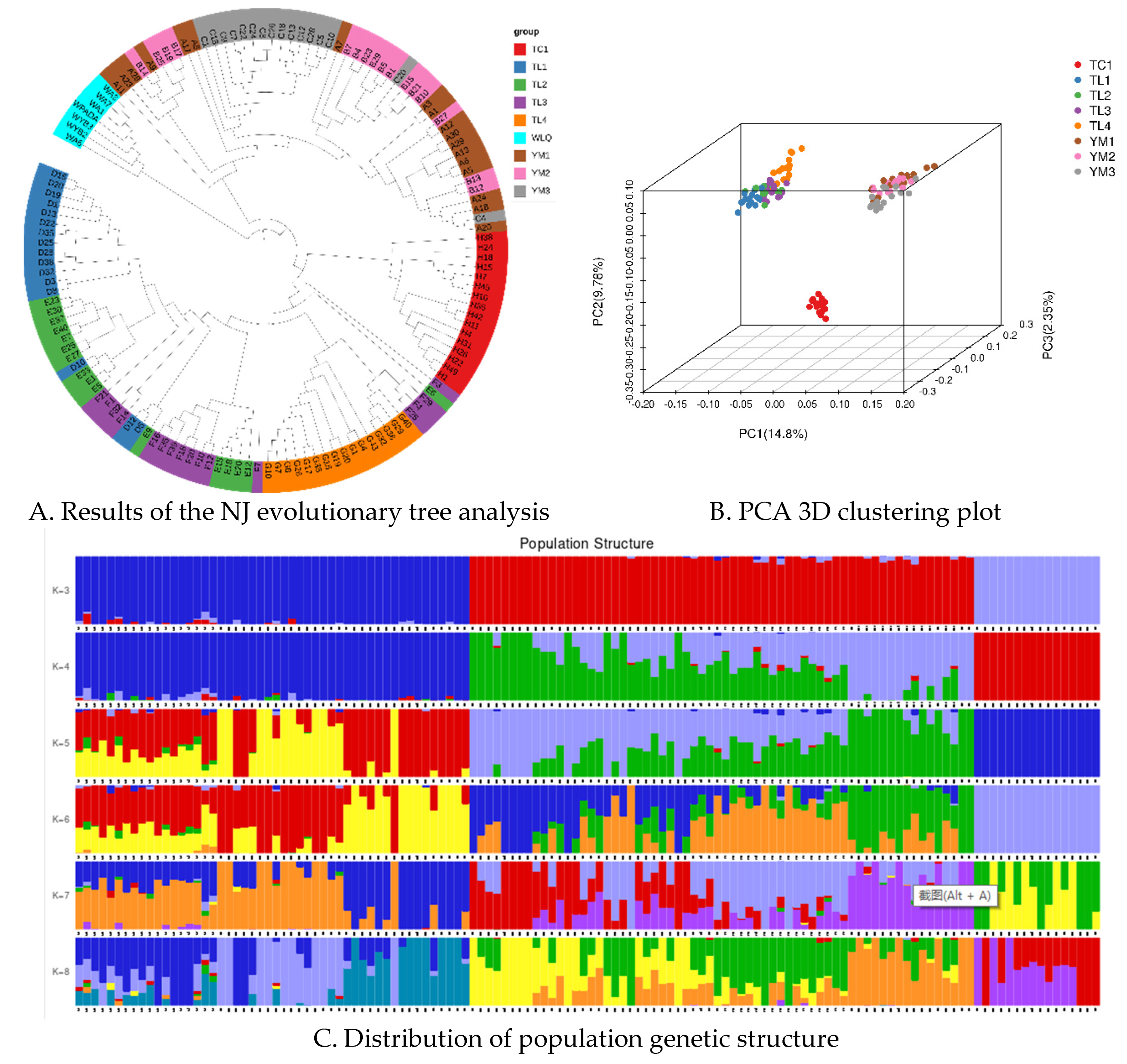
2.4. Population genetic structure and genetic diversity analysis
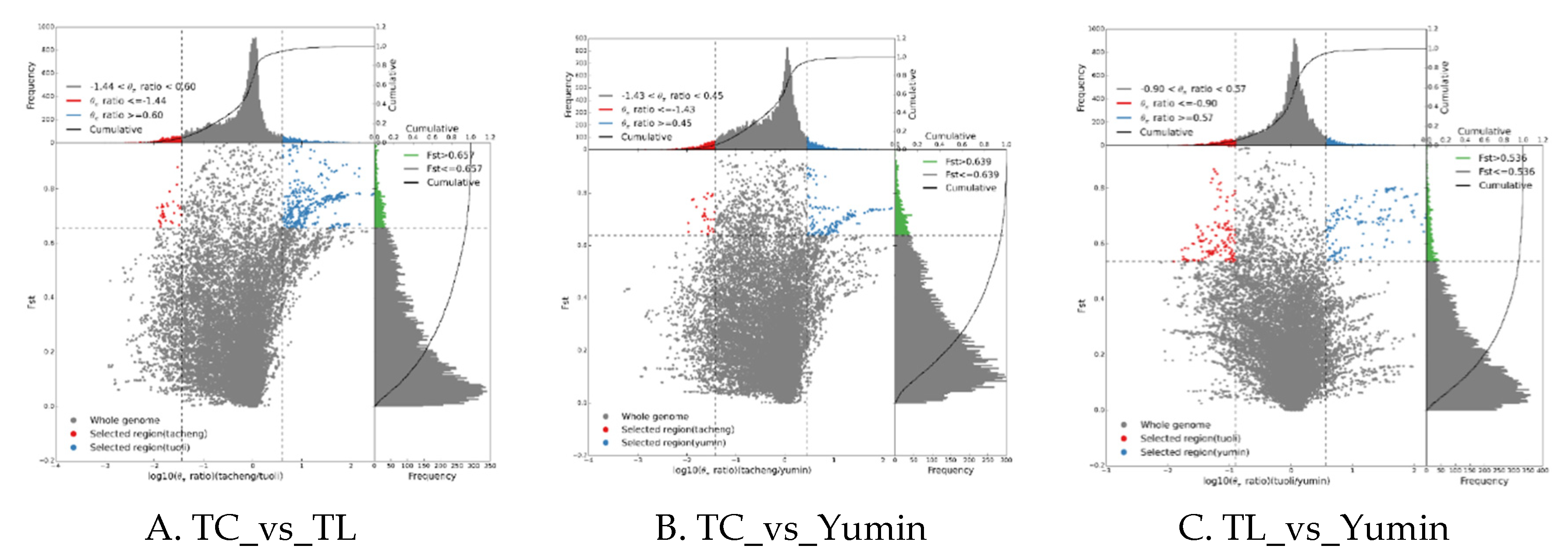
2.5. Genome-wide selection signatures analysis of differentiation
3. Discussion
4. Materials and Methods
4.1. Utilized materials
4.2. Genome sequencing and transcriptome sequencing
4.3. Assurance of Sequencing Data Quality
4.4. Heterozygosity and genome size estimation
4.5. Genome assembly
4.6. Repetitive element annotations
4.7. Functional annonations and gene prediction
4.8. Phylogenetic and gene family analysis
4.9. Whole-genome synteny analysis
4.10. Single-nucleotide polymorphism (SNP) calling
4.11. Phylogenetic analysis
4.12. Population genetic structure and genetic diversity analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Li, J.; Zeng, B.; Luo, S. P.; Li,H.L.; and Madaniyati,W. Protection and propagation ofAmygdalus ledebouriana Schleche in China. Xinjiang AgriculturalSciences. 2006, 43, 61-62.
- Yin, L. K.; Tan, L. X.;Wang, B. Rare endangered endemic higherplants in Xinjiang of China. Urumqi: Xinjiang Science & TechnologyPublishing House. 2006.
- Zhong, H. X.; Lu, C. S.; Luo, S. P.;Li, J. The study of cold resistancetest of dormancy branches and buds of Amygdalus ledebourianaSchleche in Xinjiang. Xinjiang Agricultural Sciences. 2016, 53, 120-125.
- Perazzolli, M., Malacarne, G., Baldo, A., Righetti, L., Bailey, A., Fontana, P., Velasco, R., Malnoy, M., Characterization of resistance gene analogues (RGAs) in apple (Malus × domestica Borkh.) and their evolutionary history of the Rosaceae family. PLoS One. 2014, Feb 5;9(2):e83844. PMID: 24505246; PMCID: PMC3914791. [CrossRef]
- Vinceti, B., Elias, M., Azimov, R., Turdieva, M., Aaliev, S., Bobokalonov, F., Butkov, E., Kaparova, E., Mukhsimov, N., Shamuradova, S., Turgunbaev, K., Azizova, N., & Loo, J. Home gardens of Central Asia: Reservoirs of diversity of fruit and nut tree species. PloS one. 2022, 17(7), e0271398. [CrossRef]
- Singh, R. K., Singh, C., Ambika, Chandana, B. S., Mahto, R. K., Patial, R., Gupta, A., Gahlaut, V., Gayacharan, Hamwieh, A., Upadhyaya, H. D., & Kumar, R. Exploring Chickpea Germplasm Diversity for Broadening the Genetic Base Utilizing Genomic Resourses. Frontiers in genetics. 2022, 13, 905771. [CrossRef]
- Kumar, S., Jacob, S. R., Mir, R. R., Vikas, V. K., Kulwal, P., Chandra, T., Kaur, S., Kumar, U., Kumar, S., Sharma, S., Singh, R., Prasad, S., Singh, A. M., Singh, A. K., Kumari, J., Saharan, M. S., Bhardwaj, S. C., Prasad, M., Kalia, S., & Singh, K. Indian Wheat Genomics Initiative for Harnessing the Potential of Wheat Germplasm Resources for Breeding Disease-Resistant, Nutrient-Dense, and Climate-Resilient Cultivars. Frontiers in genetics. 2022, 13, 834366. [CrossRef]
- Kefale, H., & Wang, L. Discovering favorable genes, QTLs, and genotypes as a genetic resource for sesame (Sesamum indicum L.) improvement. Frontiers in genetics. 2022,13, 1002182. [CrossRef]
- García-Gómez, B. E., Salazar, J. A., Nicolás-Almansa, M., Razi, M., Rubio, M., Ruiz, D., & Martínez-Gómez, P. Molecular Bases of Fruit Quality in Prunus Species: An Integrated Genomic, Transcriptomic, and Metabolic Review with a Breeding Perspective. International journal of molecular sciences. 2020, 22(1), 333. [CrossRef]
- Filip, E., Woronko, K., Stępień, E., & Czarniecka, N. An Overview of Factors Affecting the Functional Quality of Common Wheat (Triticum aestivum L.). International journal of molecular sciences. 2023, 24(8), 7524. [CrossRef]
- Li, Z., Xue, Y., Zhou, H., Li, Y., Usman, B., Jiao, X., Wang, X., Liu, F., Qin, B., Li, R., & Qiu, Y. High-resolution mapping and breeding application of a novel brown planthopper resistance gene derived from wild rice (Oryza. rufipogon Griff). Rice (New York, N.Y.). 2019,12(1), 41. [CrossRef]
- Mamidi, S., Healey, A., Huang, P., Grimwood, J., Jenkins, J., Barry, K., Sreedasyam, A., Lovell, J. T., Feldman, M., Wu, J., Yu, Y., Chen, C., Johnson, J., Sakakibara, H., Kiba, T., Sakurai, T., Tavares, R., Nusinow, D. A., Baxter, I., Schmutz, J., Brutnell, T. P., Kellogg, E. A. A genome resource for green millet Setaria viridis enables discovery of agronomically valuable loci. Nat Biotechnol, 2020, 38,1203-1210.
- Prahalada, G. D., Shivakumar, N., Lohithaswa, H. C., Sidde Gowda, D. K., Ramkumar, G., Kim, S. R., Ramachandra, C., Hittalmani, S., Mohapatra, T., & Jena, K. K. Identification and fine mapping of a new gene, BPH31 conferring resistance to brown planthopper biotype 4 of India to improve rice, Oryza sativa L. Rice (New York, N.Y.). 2017, 10(1), 41. [CrossRef]
- Laugerotte, J., Baumann, U., & Sourdille, P. Genetic control of compatibility in crosses between wheat and its wild or cultivated relatives. Plant biotechnology journal. 2022, 20(5), 812-832. [CrossRef]
- Sharma, S., Schulthess, A. W., Bassi, F. M., Badaeva, E. D., Neumann, K., Graner, A., Özkan, H., Werner, P., Knüpffer, H., & Kilian, B. Introducing Beneficial Alleles from Plant Genetic Resources into the Wheat Germplasm. Biology. 2021, 10(10), 982. [CrossRef]
- Aleem, M., Aleem, S., Sharif, I., Aleem, M., Shahzad, R., Khan, M. I., Batool, A., Sarwar, G., Farooq, J., Iqbal, A., Jan, B. L., Kaushik, P., Feng, X., Bhat, J. A., & Ahmad, P. Whole-Genome Identification of APX and CAT Gene Families in Cultivated and Wild Soybeans and Their Regulatory Function in Plant Development and Stress Response. Antioxidants (Basel, Switzerland). 2022, 11(8), 1626. [CrossRef]
- Mk, A.; Shw, B.; and Ss, A. Wheat wild germplasm: a hidden treasure. Wild Germplasm for Genetic Improvement in Crop Plants.2011, 2021, 55-63.
- Yumurtaci, A. Utilization of wild relatives of wheat, barley, maize and oat in developing abiotic and biotic stress tolerant new varieties. Emirates Journal of Food & Agriculture. 2015, 27.
- Haus, M. J.; Pierz, L. D.; Jacobs, J. L.; Wiersma, A. T.; and Cichy, K. E. Preliminary evaluation of wild bean (phaseolus spp.) germplasm for resistance to fusarium cuneirostrum and fusarium oxysporum. Crop Science. 2021, 3.
- Rostad, H. E.; Reen, R. A.; Mumford, M. H.; Zwart, R.; and Thompson, J. P. Resistance to root-lesion nematode pratylenchus neglectus identified in a new collection of two wild chickpea species (cicer reticulatum and c. echinospermum) from turkey. Plant Pathology. 2022, 5, 71.
- Jeff, E.; Olumide, S.T.; Bruce,D.; Andre, H.; Julianne, A.; Olufemi, A. Resistance in wild macadamia germplasm to phytophthora cinnamomi and phytophthora multivora. Annals of Applied Biology. 2021, 178.
- Wang, B.; Yu, Z.F.; Zeng, B.; Xia, J.H.;Ma,X.X. Self-incompatibility Gene Cullin1 Cloning and Bioinformatics Analysis of Wild Almond in Xinjiang. Chinese Agricultural Science Bulletin. 2017, 33, 63-68.
- Zeng, B.; Liu, N.N.; Xia, J.H.; Liu, M.W.; Wang,J.Y.; Wang, B. Molecular Cloning and Bioinformatics Analysis of SFB Genes Controlling Self-incompatibility in Xinjiang Wild Almond (Prunus tenella Batsch.). Chinese Agricultural Science Bulletin. 2017, 33, 22-30.
- Yu, Z. F.; Wang, B.; Zeng, B.; Wang, J.Y. Cloning and sequence analysisof self - incompatibility gene SBPI of wild almond in Xinjiang.Molecular Plant Breeding. 2018, 16, 6955-6960.
- Zeng, B.; Li,J.; Luo, S. P.; Cheng, Y. J. Identification of Genetic Relationship of Amygdalus Plants by SSR. Xinjiang Agricultural Sciences. 2009, 46, 18-22.
- Lu, Z.J.; Li, J.; Omir, S.T.; Zeng.B.; Luo, S.P. ISSR analysis for genetic diversity of Amygdalus ledebouriana germplasmfrom Xinjiang,China. Journal of Fruit Science. 2010, 27, 918-923.
- Chen, D. X.; Pan, Y.; Wang, Y.; Cui, Y. Z.; & Li, L. Y. The chromosome-level reference genome of coptis chinensis provides insights into genomic evolution and berberine biosynthesis. Horticultural Research. 2021, 8, 11.
- Rush, D. W.; and Epstein, E. Breeding and selection for salt tolerance by the incorporation of wild germplasm into a domestic tomato. Journal American Society for Horticultural Science. 1981, 106, 699-704.
- D'Amico-Willman, K. M.; Ouma, W. Z.; Meulia, T.; Sideli, G. M.; Gradziel, T. M.; and Fresnedo-Ramírez, J. Whole-genome sequence and methylome profiling of the almond ( prunus dulcis [mill.] d.a.webb) cultivar ‘nonpareil’. G3 Genes|Genomes|Genetics. 2022,12(2022)jkac065.
- Liu, J. F.; Wei, H.; Zhang, X.; and Wang, D. Chromosome-level genome assembly and hazelomics database construction provides insights into unsaturated fatty acid synthesis and cold resistance in hazelnut (corylus heterophylla). Frontiers in Plant Science. 2021, 12, 766548.
- Verde, I.; Abbott, A. G.; Scalabrin, S.; Jung, S.; and Rokhsar, D. S. The high-quality draft genome of peach (Prunus persica) identifies unique patterns of genetic diversity, domestication and genome evolution-Supplementary Information.Nature Genetics. 2014, 45, 486-495.
- Suguiyama, V. F., Vasconcelos, L. A. B., Rossi, M. M., Biondo, C., & de Setta, N. The population genetic structure approach adds new insights into the evolution of plant LTR retrotransposon lineages. PloS one. 2019, 14(5), e0214542. [CrossRef]
- Ikeda, H. Decades-long phylogeographic issues: complex historical processes and ecological factors on genetic structure of alpine plants in the Japanese Archipelago. Journal of plant research. 2022, 135(2), 191-201. [CrossRef]
- Mori, G. M., Zucchi, M. I., & Souza, A. P. Multiple-geographic-scale genetic structure of two mangrove tree species: the roles of mating system, hybridization, limited dispersal and extrinsic factors. PloS one.2015, 10(2), e0118710. [CrossRef]
- Nishio, S., Takada, N., Terakami, S., Takeuchi, Y., Kimura, M. K., Isoda, K., Saito, T., & Iketani, H. Genetic structure analysis of cultivated and wild chestnut populations reveals gene flow from cultivars to natural stands. Scientific reports. 2021, 11(1), 240. [CrossRef]
- Deb, S., Della Lucia, M. C., Ravi, S., Bertoldo, G., & Stevanato, P. (2023). Transcriptome-Assisted SNP Marker Discovery for Phytophthora infestans Resistance in Solanum lycopersicum L. International journal of molecular sciences, 24(7), 6798. [CrossRef]
- Bali, S., Robinson, B. R., Sathuvalli, V., Bamberg, J., & Goyer, A. Single Nucleotide Polymorphism (SNP) markers associated with high folate content in wild potato species. PloS one. 2018,13(2), e0193415. [CrossRef]
- Roncallo, P. F., Beaufort, V., Larsen, A. O., Dreisigacker, S., & Echenique, V. Genetic diversity and linkage disequilibrium using SNP (KASP) and AFLP markers in a worldwide durum wheat (Triticum turgidum L. var durum) collection. PloS one. 2019,14(6), e0218562. [CrossRef]
- Castilla, A. R., Méndez-Vigo, B., Marcer, A., Martínez-Minaya, J., Conesa, D., Picó, F. X., & Alonso-Blanco, C. Ecological, genetic and evolutionary drivers of regional genetic differentiation in Arabidopsis thaliana. BMC evolutionary biology. 2020, 20(1), 71. [CrossRef]
- Oh, A., & Oh, B. U. Genetic differentiation that is exceptionally high and unexpectedly sensitive to geographic distance in the absence of gene flow: Insights from the genus Eranthis in East Asian regions. Ecology and evolution. 2022,12(6), e9007. [CrossRef]
- Santangelo, J. S., Johnson, M. T. J., & Ness, R. W. Modern spandrels: the roles of genetic drift, gene flow and natural selection in the evolution of parallel clines. Proceedings. Biological sciences. 2018, 285(1878), 20180230. [CrossRef]
- Marcais,G.; and Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics. 2011, 27, 764-770. [CrossRef]
- Roach,M. J.; Schmidt, S. A.; and Borneman,A. R. Purge Haplotigs:allelic contig reassignment for third-gen diploid genome assemblies. BMC Bioinformatics. 2018, 19, 460. [CrossRef]
- Dudchenko, O.; Batra, S. S.; Omer, A. D.; Nyquist, S. K.; Hoeger, M.; Durand, N.C. De novo assembly of the Aedes aegypti genome using Hi-C yieldschromosome-length scaffolds.Science. 2017, 356, 92-95. [CrossRef]
- Durand, N.; Robinson, J.; Shamim, S.; Aiden, E. L. Juicebox provides a visualization system for Hi-C contactmaps with unlimited zoom. Cell Syst. 2016, 3, 99-101. [CrossRef]
- Simão, F. A.; Waterhouse, R. M.; Loannidis, P.; Kriventseva, E. V.; and Zdobnov,E.M. BUSCO: assessing genome assembly and annotation completenesswith single-copy orthologs. Bioinformatics. 2015, 31, 3210-3212. [CrossRef]
- Kim, D.; Langmead, B.; and Salzberg, S. L. HISAT: a fast spliced alignerwith low memory requirements.Nat. Methods. 2015, 12, 357-360. [CrossRef]
- Ou, S.; Su, W.; Liao, Y.; Chougule, K.; Doreen, W.; Thomas, P.; Ning, J.; Candice, N. H.; Hufford, M. B. Benchmarking transposable element annotation methods for creationofa streamlined,comprehensive pipeline. Cold Spring Harbor Laboratory. 2019, 1.
- Bao, W.; Kojima, K. K.; and Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA. 2015, 6, 11. [CrossRef]
- Tempel, S. Using and Understanding RepeatMasker. Methods Mol. Biol. 2012, 859, 29-51. [CrossRef]
- Grabherr, M. G.; Haas, B. J.; Yassour, M.; Levin, J. Z.; Thompson, D. A.; Amit,I. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644-652. di: 10.1038/nbt1883.
- Haas, B. J.; Delcher, A. L.; Mount, S. M.; Wortman, J. R.; Smith, J.; Hannick, L. I.; Rama, M.; Ronning, C. M.; Rusch, D. B.; Town, C. D. Improving the Arabidopsis genome annotation using maximaltranscript alignment assemblies.Nucleic Acids Res. 2003, 31, 5654-5666. [CrossRef]
- Majoros, W.; Pertea, M.; and Salzberg, S. TigrScan and GlimmerHMM: twoopen source ab initio eukaryotic gene-finders. Bioinformatics. 2004, 20, 2878-2879. [CrossRef]
- Stanke, M.; Keller, O.; Gunduz, I.; Hayes, A.; Waack, S.; and Morgenstern, B. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 2006, 34,W435-W439. [CrossRef]
- Johnson, A. D.; Handsaker, R. E.; Pulit, S. L.; Nizzari, M. M.; O’donnell, C.J.; De Bakker, P.I. SNAP: a web-based tool for identification andannotation of proxy SNPs using HapMap. Bioinformatics. 2008, 24, 2938-2939. [CrossRef]
- Haas, B. J.; Salzberg, S. L.; Zhu, W.; Pertea, M.; Allen, J. E.; Orvis, J.; White, O.; Buell, C. R.; Wortman, J. R. Automated eukaryotic gene structure annotation using evidence modeler and the program to assemble spliced alignments. Genome Biol. 2008, 9, R7. [CrossRef]
- Emms, D. M.; Kelly, S. OrthoFinder: phylogenetic orthology inferencefor comparative genomics. Genome Biol. 2019, 20, 238. [CrossRef]
- Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis andpost-analysis of large phylogenies. Bioinformatics. 2014, 30, 1312-1313. [CrossRef]
- Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Bio.Evol. 2007, 24,1586-1591. [CrossRef]
- Han,M. V.; Thomas,G.W.C.; Lugo-Martinez, J.; and Hahn,M. W. Estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol Biol. Evol. 2013, 30, 1987-1997. di: 10.1093/molbev/mst100.
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; and Yu, J. KaKs_Calculator 2.0:a toolkit incorporating gamma-series methods and sliding window strategiesGenomics Proteomics Bioinform. 2010, 8, 77-80. [CrossRef]
- Marais,G.; Delcher, A. L.; Phillippy, A. M.; Coston, R.; and Zimin, A. MUMmer4: a fast and versatile genome alignment system. PLOS Comput. Biol. 2018, 14, e1005944. [CrossRef]
- Sudhir, K.; Glen, S.; Michael, L.; Christina, K.; and Koichiro, T. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Molecular biology and evolution. 2018, 35, 1547-1549.
- Alkes, L. P.; Nick, J. P.; Robert, M.P.; Michael,E.W.; Nanch,A.S.; and David. Principal components analysis corrects for stratification in genome-wide association studies. Nature Genetics. 2006, 38, 904-909.
- Vilella, A.J.; Blanco-Garcia, A.; Hutter, S.; Rozas, J. VariScan: analysis of evolutionary patterns from large-scale DNA sequence polymorphism data.Bioinformatics. 2005, 21, 2791-3.
- Alexander, D. H.; Novembre, J.; Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome research. 2009, 19, 1655-1664.
| Term | Contig number | Contig size (bp) | Scaffold number | Scaffold size (bp) |
| N90 | 5 | 270515 | 9 | 1233122 |
| N80 | 12 | 8938573 | 7 | 19226261 |
| N70 | 11 | 10302541 | 6 | 21538263 |
| N60 | 15 | 14010243 | 5 | 23141640 |
| N50 | 1 | 18100976 | 4 | 25637364 |
| Max length (bp) | 35886393 | 44825466 | ||
| Total size (bp) | 231191648 | 231208648 | ||
| Total number | 513 | 479 | ||
| Average length | 450665.98 | 482690.29 | ||
| Number >= 10kb | 513 | 479 |
| Chr ID | Length (bp) |
| Chr1 | 44825466 |
| Chr2 | 29024987 |
| Chr3 | 26387986 |
| Chr4 | 25637364 |
| Chr5 | 23141640 |
| Chr6 | 21538263 |
| Chr7 | 19226261 |
| Chr8 | 17664456 |
| Total chromosome level contig length | 207446423 |
| Total contig length | 231208648 |
| Chromosome length/Total length | 89.7% |
| Library | eudicotyledons_odb10 |
| Fragmented BUSCOs (F) | 48 |
| Missing BUSCOs (M) | 64 |
| Complete and duplicated BUSCOs (D) | 42 |
| Complete and single-copy BUSCOs (S) | 1967 |
| Complete BUSCOs (C) | 2009 |
| Total BUSCO groups searched | 2121 |
| Summary | 94.7% |
| Class | Length (bp) | Type | Sub-Class | (%) |
| retrotransposons | 8454829 | Ty1/Copia | LTR | 3.66% |
| 18247107 | Ty3/Gypsy | 7.89% | ||
| 15756928 | unknown | 6.82% | ||
| - | LINE | Non-LTR | - | |
| - | unknown | - | ||
| DNA transposons | 1845596 | CACTA | TIR | 0.80% |
| 7818414 | Mutator | 3.38% | ||
| 4181524 | PIF/Harbinger | 1.81% | ||
| 278071 | Tc1/Mariner | 0.12% | ||
| 3345229 | hAT | 1.45% | ||
| 7038992 | helitron | Non-TIR | 3.04% | |
| Total | 66966690 | 28.97% |
| Database | Gene numbers | (%) |
| GO | 10761 | 33.54 |
| KEGG | 11435 | 35.64 |
| KOG | 19251 | 59.99 |
| Swissprot | 19449 | 60.61 |
| Pfam annotation | 22815 | 71.1 |
| Nr annotation | 31202 | 97.24 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




