1. Introduction
It has become evident in recent years that targeting of proteins to multiple locations is more abundant than was initially assumed [
1,
2,
3,
4,
5,
6,
7]. This phenomenon is termed dual targeting, dual localization, or dual distribution [
3,
8,
9,
10,
11,
12]. In such cases the identical, or nearly identical, forms of the proteins that are localized to different subcellular compartments are termed echoforms or echoproteins (to distinguish them from isoforms/isoproteins) [
12]. Protein dual targeting can be achieved by a variety of molecular mechanisms (for reviews see [
11,
12,
13,
14,
15,
16,
17]). Briefly, dual-targeting mechanisms can either be the result of multiple translation products (for example due to varying transcription/ translation initiation or termination sites) or of a single translation product due to several competing targeting signals or one ambiguous signal. “Reverse translocation” is another example, in which a subpopulation of the molecules moves back into the cytosol during the translocation process [
2,
16,
18,
19].
Since the precise subcellular localization of a protein is critical for its function, one of the challenges facing post-genomic biology is exploring protein subcellular localization to characterize location-specific functions. Indeed, there is a growing number of studies designed to address this challenge by developing global screens of protein localization [
20,
21,
22,
23,
24,
25,
26,
27]. These studies have provided considerable information regarding protein localization. However, these systematic analyses, such as by visualization of fusion proteins with a fluorophore or by mass-spectrometry, often fail to detect one of the echoproteins due to a highly uneven distribution between compartments. This phenomenon is termed
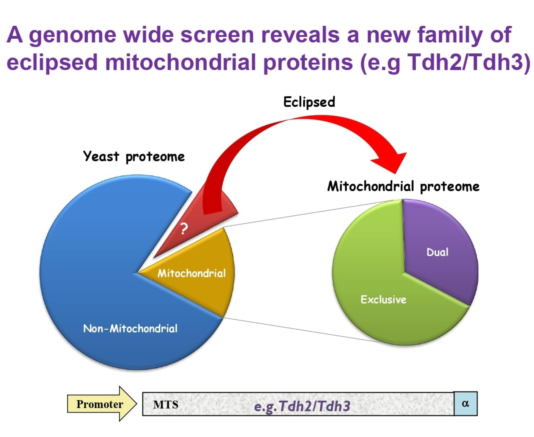
‘eclipsed distribution’ in which the relatively large amount of an echoprotein in one subcellular compartment obscures the detection of the small amount in the other location [
3,
4] (
Figure 1A). Not only the existence, but also the function of one echoprotein may be eclipsed by that of the other [
3,
8,
28,
29]. We maintain that for the above reasons, the eclipsed distribution phenomenon has a wider incidence than currently recorded (
Figure 1B) and will probably be important for understanding mitochondrial disfunction and disease.
Our previous studies in the model organism
Saccharomyces cerevisiae (from here on termed simply yeast) employed an α-complementation technology which uses a split β-galactosidase system to determine protein subcellular localization (
Figure 2A) [
30,
31]. Together with bioinformatics analysis, we estimated that a third of the mitochondrial proteome is dual targeted to other subcellular compartments [
32]. In fact, a major discovery was that many well studied mitochondrial proteins, such as aconitase, were found to be eclipsed in the cytosol [
3,
29]. Furthermore, we found that these proteins are enriched by distinctive properties such as a total net charge of the whole protein and weaker Mitochondrial Targeting Signals (MTS) [
32,
33], and that they are more evolutionary conserved than their exclusive mitochondrial counterparts [
34].
Figure 1.
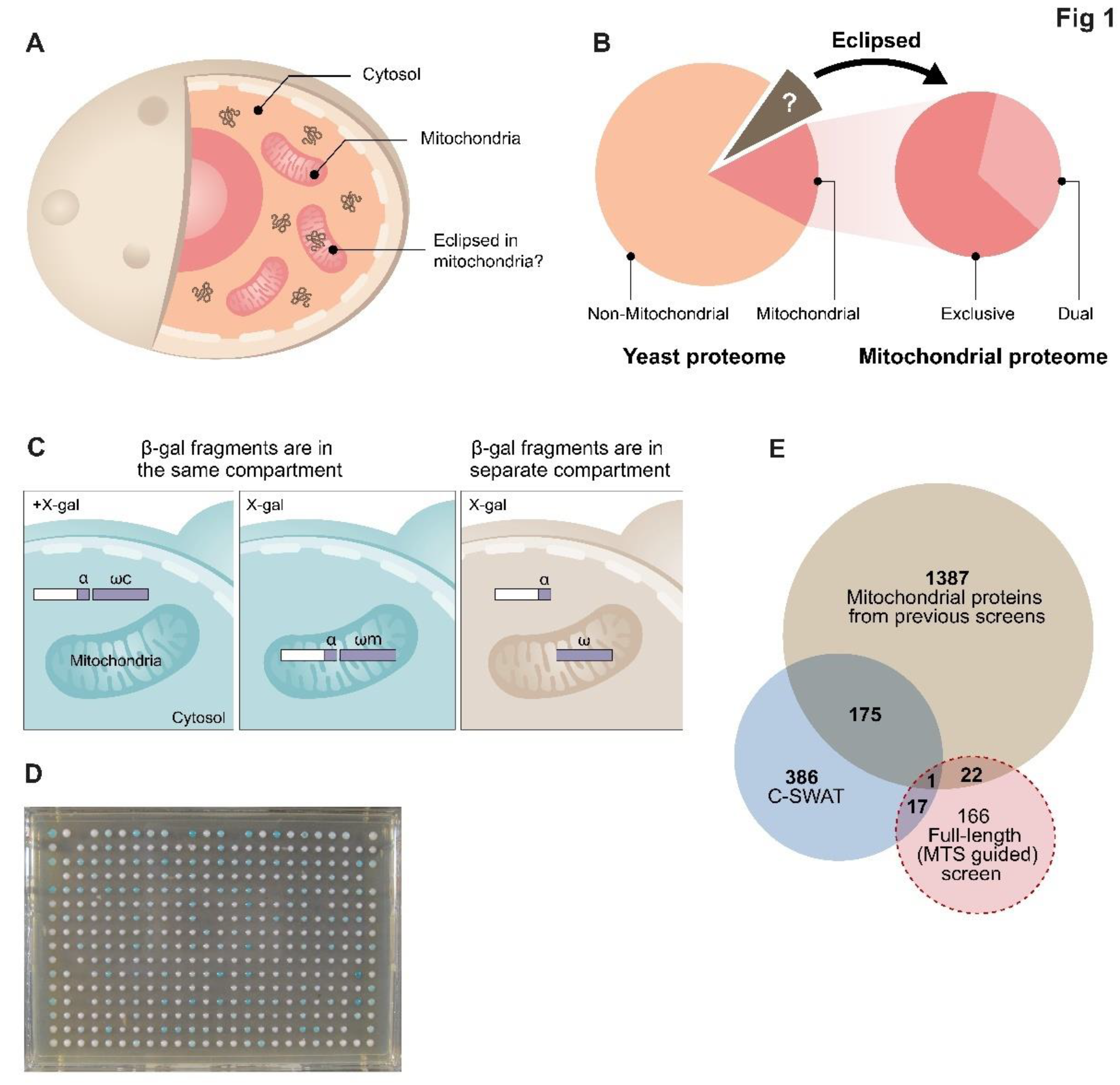
Eclipsed distribution in mitochondria. (
A). Schematic illustration of eclipsed distributed proteins in mitochondria. The relatively large amount of one echoform in one subcellular compartment (cytosol) obscures the detection of the small amount of the other echoform in the second location (mitochondria). (
B). A subset of the yeast proteome (left) is localized to mitochondria. A third of the mitochondrial proteome is dual targeted [
32]. The non-mitochondrial proteome is hypothesized to contain a subset of proteins which are eclipsed distributed in mitochondria (brown section and arrow). (
C). Schematic illustration of the ɑ-complementation assay that is based on co-localization of β-galactosidase enzyme (β-gal) fragments, ɑ and ω, within the same compartment. β-gal activity can be detected by formation of blue colonies on X-gal plates (left). In the case that the two fragments are in separate compartments the colonies appear white (right). (
D). A representative image of a plate from the C-SWAT library plated on X-gal showing colonies in 384-well format submitted to the ɑ-complementation assay (the library contains 30 plates). Genes of the library were tagged with ɑ at their C-terminus and mated with a strain expressing either cytosolic or mitochondrial ω. (
E). Venn diagram of previous studies and both screens from this manuscript. Illustration of the mitochondrial candidate proteins suggested by the C-SWAT (blue sphere, 386 proteins), Full-length (MTS guided) α-complementation screen (red sphere, 166 proteins) and previously alleged mitochondrial proteins (green sphere, 1387 proteins).
Figure 1.
Eclipsed distribution in mitochondria. (
A). Schematic illustration of eclipsed distributed proteins in mitochondria. The relatively large amount of one echoform in one subcellular compartment (cytosol) obscures the detection of the small amount of the other echoform in the second location (mitochondria). (
B). A subset of the yeast proteome (left) is localized to mitochondria. A third of the mitochondrial proteome is dual targeted [
32]. The non-mitochondrial proteome is hypothesized to contain a subset of proteins which are eclipsed distributed in mitochondria (brown section and arrow). (
C). Schematic illustration of the ɑ-complementation assay that is based on co-localization of β-galactosidase enzyme (β-gal) fragments, ɑ and ω, within the same compartment. β-gal activity can be detected by formation of blue colonies on X-gal plates (left). In the case that the two fragments are in separate compartments the colonies appear white (right). (
D). A representative image of a plate from the C-SWAT library plated on X-gal showing colonies in 384-well format submitted to the ɑ-complementation assay (the library contains 30 plates). Genes of the library were tagged with ɑ at their C-terminus and mated with a strain expressing either cytosolic or mitochondrial ω. (
E). Venn diagram of previous studies and both screens from this manuscript. Illustration of the mitochondrial candidate proteins suggested by the C-SWAT (blue sphere, 386 proteins), Full-length (MTS guided) α-complementation screen (red sphere, 166 proteins) and previously alleged mitochondrial proteins (green sphere, 1387 proteins).
In this study we ask the opposite question - whether there are proteins that were known to reside in other cellular areas yet are eclipsed in mitochondria and have failed to be identified by less sensitive methods. We propose that many proteins, which are thought to be non-mitochondrial (e.g. cytosolic,
Figure 1B, orange section) are actually located at very low levels (eclipsed) and may function in mitochondria (
Figure 1B, brown section).
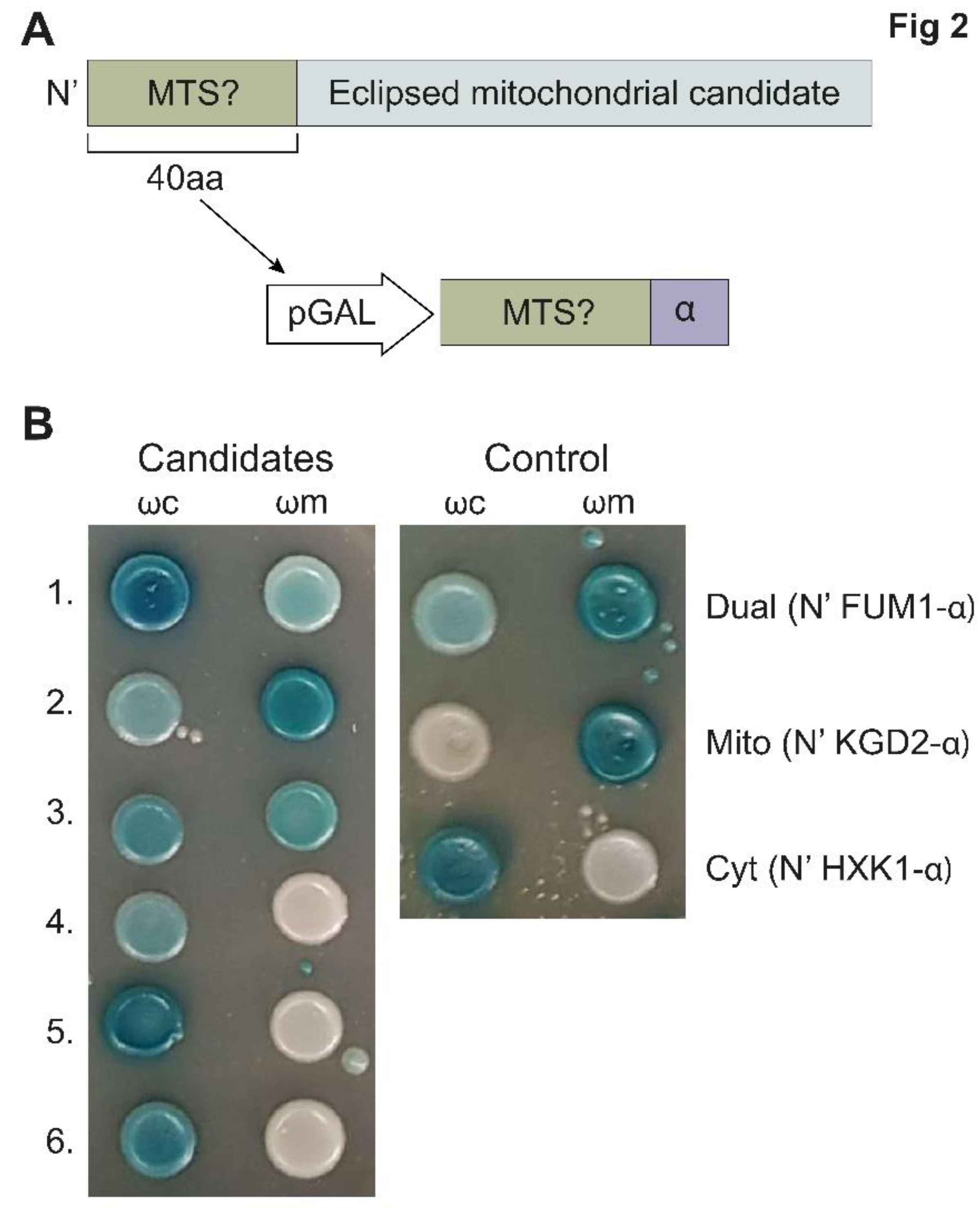
To test this, we again employed a variation of the α-complementation assay (
Figure 2A) [
35] for two complementary screens in the model eukaryote saccharomyces cerevisiae. In the first, we created and screened a genome-wide collection of yeast strains each harboring one small fragment (α) of the β-galactosidase on their C’ terminus of a particular protein and then assayed their complementation capacity with either mitochondrial or cytosolic large complementing β-galactosidase fragments (ω,
Figure 2B) [
35,
36]. The second was a hypothesis driven screen in which we took cytosolic proteins that had a high-scoring predicted MTS-like sequence and assayed them for eclipsed distribution in mitochondria. Using both of these approaches we discovered hundreds of candidates for eclipsed distributed in mitochondria. We chose the Triose-phosphate DeHydrogenases (TDH) protein family as a test case of eclipsed distribution and show that they are indeed eclipsed. Moreover, we demonstrate that their mitochondrial form has a role in mitochondrial activity. This study changes the perception of protein localization and specifically the mitochondrial proteome in eukaryotes.
2. Results
2.1. Genome Wide Analysis of the Yeast Proteome Reveals 137 Candidates of Eclipsed Mitochondrial Proteins
Building on the assumption that there are unidentified eclipsed mitochondrial proteins (
Figure 1B) we employed an α-complementation screen of the whole yeast genome (
Figure 1C,D). A genomic SWAp-Tag (SWAT) library of approximately 6000 yeast genes
, was utilized as a parental library [
36]. Using the SWAT approach, we replaced the endogenous tag with one creating in frame fusions of the α-tag on the C’ of each protein. Since this library contains genes in their native locus, under the regulation of their native promoter, this reduces the risk of false positives due to over- or mis-expression. The agar plates with yeast colonies in 384-well format were mated with yeast that express a mitochondrial-matrix targeted β-galactosidase omega fragment (ωm) (
Figure 1D) and were scored for their ability to create a blue colony phenotype on X-gal plates, indicating mitochondrial localization.
We detected 386 blue colonies (
Supplementary Table S1 and
Supplementary Table S2), out of which, 176 proteins were previously reported to be mitochondrial (
Supplementary Table S1,
Figure 1E, blue sphere). We did not uncover all mitochondrial proteins since our assay only expects to detect matrix proteins and many matrix proteins may not be expressed under the conditions assayed, or may be unstable when fused to the α tag. We found that 116 of our hits are membrane proteins according to the Gene Annotation (GO) database (
Supplementary Table S2). Since complementation demands that their termini face the matrix, these are suggested to be inner membrane proteins, whose C terminus faces the matrix. Moreover, 73 of these were never suggested to be mitochondrial and this presents them as new candidates of inner membrane proteins. Of the remaining soluble proteins, we suggest that 137 are candidates of eclipsed mitochondrial matrix proteins (
Supplementary Table S1).
To verify these results, we randomly chose 20 genes that were positive and 11 that were negative according to the screen (
Supplementary Table S1 and
Figure S1) and cloned them into an overexpression α-complementation vector for validation. When overexpressed, all 20 positive strains exhibited a blue phenotype with both ωm (like the positive control Kgd1) and ω expressed in the cytosol (ωc) indicating dual targeting to both mitochondria and the cytosol, which is in full agreement with the C-SWAT screen (
Figure S1). 7 out of the 11 genes that were negative in the C-SWAT screen were negative in mitochondria (white colonies with ωm) but positive in the cytosol (blue colonies with ωc) which is in full agreement with the C-SWAT screen. Three genes were negative both with ωm and ωc suggesting that these fusion proteins may not be expressed properly from our constructs. Importantly, none of our negative controls gave a mitochondrial signal even when overexpressed attesting to the accuracy of the screen. In summary, the C-SWAT library screen detected a significant number of potential eclipsed proteins targeted to mitochondria.
2.2. A Proteomics Approach to Uncover Eclipsed Mitochondrial Proteins
As an additional avenue to uncover eclipsed proteins we took a proteomics approach using iTRAQ (isobaric tags for relative and absolute quantitation
( [
37] mass spectrometry of mitochondrial versus cytosolic fractions (data not shown, see thesis [
38]). This approach identified mitochondrial, dual targeted and cytosolic proteins with reasonable success (71–98%, 72–93% and 51–73%, coverage of test groups respectively). It appears to be much less precise than the C-SWAT approach, even though some new proteins were detected as eclipsed candidates. Candidate proteins from this approach were over-expressed and examined by the α-complementation assay and 12/17 (70%) were verified as having an eclipsed distribution (
Supplementary Table S3).
2.3. α-Complementation Analysis of Predicted MTSs
To complement our above efforts and create a list of eclipsed proteins candidates, we initiated a hypothesis-based approach in which we bioinformatically examined for traits associated with mitochondrial proteins as a means to directly identify eclipsed candidates. We considered those proteins that were predicted to have a high scoring amino terminal MTS by the MitoProt II algorithm [
33,
34,
39]. Our list contained 542 proteins not known to be mitochondrial but predicted to harbor an MTS with high probability (above 0.55). From this list we removed proteins suggested to be membrane proteins (according to gene ontology (GO) annotation), although they could be new mitochondrial inner membrane proteins, since the downstream determination of their localization is experimentally more complicated. Following this we remained with 313 potential eclipsed mitochondrial proteins (
Supplementary Table S4).
We chose the ~200 highest scoring proteins (MTS score > 0.7,
Supplementary Table S5) and examined only their first 40 amino-acids, which contain the predicted MTS (
Figure 2A) for their capacity to drive the α fragment into the mitochondrial matrix (
Figure 2B,
Supplementary Table S4). α-tagged MTSs whose analysis resulted in blue colonies both with ωc and ωm (BB,
Supplementary Table S5) were considered dual targeted, while those that lead to blue colonies only with ωc and white colonies with ωm (BW,
Supplementary Table S5) were considered not mitochondrial. More than 50 percent (103 out of 201) of the MTSs displayed the capacity to target proteins into the matrix (
Supplementary Table S5). Together, these results suggest the existence of many functional MTSs on proteins previously not identified as mitochondrial.
2.4. Analysis of Full-Length Proteins Harboring an MTS by α-Complementation
The fact that a sequence can act as an MTS in isolation does not necessarily mean that it does act as an MTS when part of an entire polypeptide chain. Hence in parallel we took all 313 soluble proteins with predicted MTS-like sequences (MitoProt II score > 0.55,
Supplementary Table S4) and cloned the full-length protein ORFs fused to the α fragment. Of the 277 that we successfully cloned and sequenced (194 out of which overlap with the previous analysis (MitoProt II score > 0.7,
Supplementary Table S4,
Supplementary Table S5 and
Supplementary Table S6). We found that a significant percentage, 74% (205 out of 277) of these proteins appear to be dual targeted (BB + BLB) to mitochondria according to the α-complementation screen (
Supplementary Table S6). There are a few possible explanations for the rare WW phenotype (7/277,
Supplementary Table S6) as previously discussed [
32] : i) these proteins are not expressed; ii) they are membrane proteins (e.g., OCH1, described below); iii) they are buried in multi subunit protein complexes, such as mitochondrial ribosomes, which in turn may hinder their assembly into active α-ω complexes. Worth mentioning is that we were also capable of detecting predicted MTS (previous section) and full-length protein candidates fused to α, by western blot (
Figure S2).
One negative example was examined in more detail. Och1 harbors an MTS like sequence at its N-terminus (MitoProtII score 0.98,
Supplementary Table S4). Yet it appeared negative in α-complementation with ωm for both the MTS alone (WW;
Figure S2) and the full-length protein fusions (BW;
Figure S2). Further experiments [
40] show that Och1 is associated with mitochondrial membranes and can be released only by detergent (TX-100) but not with Na
2CO
3, and is not sensitive to protease treatment in the absence of Triton. Thus, the N terminus of Och1, even though it has a high MitoProt II score, inserts into the mitochondrial membrane and does not function as an MTS.
To summarize, we detected 208 (
Supplementary Table S6) candidates of eclipsed mitochondrial proteins, most of which 166 (
Supplementary Table S6), are clearly targeted to the organelle. Out of these, 23 proteins were reported only in the last few years as mitochondrial candidates (
Supplementary Table S6,
Supplementary Table S4 and
Supplementary Table S2). Therefore, our work brings forward 143 (166 minus 23) new mitochondrial matrix candidates. Together our two independent screens, the C-SWAT (spanning 95% of the entire yeast proteome), which is based on the native endogenous expression levels of proteins, and the MTS guided screen, which employed over-expression, identify 137 (
Supplementary Table S1) and 143 (
Supplementary Table S6) novel mitochondrial candidates, respectively.
2.5. Bioinformatics Analysis of the Candidate Eclipsed Mitochondrial Proteins
Previously we have reported that dual-targeted mitochondrial proteins are enriched for specific traits such as lower whole-protein net charge, weaker MTS strength, higher transcript levels and higher evolutionary conservation, when compared to exclusively targeted mitochondrial proteins or non-mitochondrial proteins [
32,
33,
34]. Conservation parameters assayed were low Dn/Ds (ratio of nonsynonymous to synonymous mutations), high CAI (Codon Adaptation Index), a high number of homologs, and a low PGL (propensity for gene loss). We previously hypothesized that such proteins are more evolutionary conserved than exclusive mitochondrial proteins due to separate selective pressures on the proteins’ presence in the different compartments [
34].
To test whether our newly found eclipsed proteins also follow these rules we analyzed the 137 genes found in the C-SWAT screen. Indeed, we found them to be significantly more conserved versus known exclusive mitochondrial and non-mitochondrial proteins. This is true for conservation parameters: CAI; Dn-Ds; Homologs; and PGL (
p Value ≤ 0.001,
Supplementary Table S7). Another parameter that tends to be high for dual targeted and dual functional proteins [
34] is higher expression level which is correlated with higher evolutionary conservation. In fact, when analyzing a previously performed quantification analysis using a different C’ tagged fusion with Green Fluorescent Protein (GFP) [
41], we see that indeed the expression levels of the C-SWAT eclipsed candidates in mitochondria, are significantly higher, when compared to exclusive, dual and non-mitochondrial proteins suggesting higher conservation (
p Value ≤ 0.001,
Supplementary Table S7) [
34,
42,
43].
With regard to net charge on the whole protein sequence, the new C-SWAT candidates appear to be most similar to non-mitochondrial proteins and significantly different from the dual and exclusive mitochondrial proteins (
p Value < 0.001,
Supplementary Table S7). The reason for this is not clear but could be at the basis of why these proteins are distributed in an eclipsed fashion, and were not identified earlier. Using the same rational, it may not be surprising that parameters referring to the strength of a possible N-terminal MTS (hydrophobic moment (µHd) and basic residues in the N-terminus), are low for these proteins (
Supplementary Table S7).
We next repeated these analyses for the 166 mitochondrial eclipsed candidates from the MTS guided screen. Since this screen is based on the candidates containing an MTS like sequence at their N termini, it is not surprising that these proteins have the highest average µHD and positive charges within the N-terminus, and in this regard, are most similar to the dual targeted proteins. Of importance, is that the net charge on the whole protein, is most similar to known exclusive and dual targeted mitochondrial proteins and most significantly different from non-mitochondrial proteins (
p Value < 0.001,
Supplementary Table S7). We found insignificant changes in transcript level. The conservation parameters (number of homologs and PGL) were similar to known exclusive mitochondrial and dual-targeted proteins, which supports our hypothesis that they are eclipsed dual-targeted proteins.
2.6. TDH Genes are Eclipsed Mitochondrial Proteins
Our screens brought forward an extensive array of proteins, many with known cellular activities, that could be functional inside mitochondria. We chose to focus on one example to verify our approach and demonstrate the importance of understanding eclipsed distribution for proteins. To this end we chose the evolutionary conserved
S. cerevisiae Triose-phosphate DeHydrogenase (TDH) enzymes. TDH enzymes constitute a protein family of three, unlinked, highly homologous genes,
TDH1, TDH2 and
TDH3 [
44,
45,
46,
47] that encode 332 amino acid polypeptides.
TDH1 is 84% identical to
TDH2 or
TDH3, whereas
TDH2 and
TDH3 are 95% identical by protein sequence [
46,
47]. None of the three genes on its own is essential for cell viability, however, the presence of at least one functional
TDH2 or
TDH3 gene has been reported to be essential for cell viability [
46]. TDH proteins are amongst the most abundant soluble enzymes in the cytosol [
46,
47,
48], displaying diverse activities in different subcellular locations [
49,
50,
51,
52,
53,
54]. Their most well-characterized roles are in glycolysis, converting glyceraldehyde-3-phosphate (GAP) to 1, 3 bisphosphoglycerate (BPG) in the presence of the cofactor nicotinamide adenine dinucleotide (NAD+) to yield NADH [
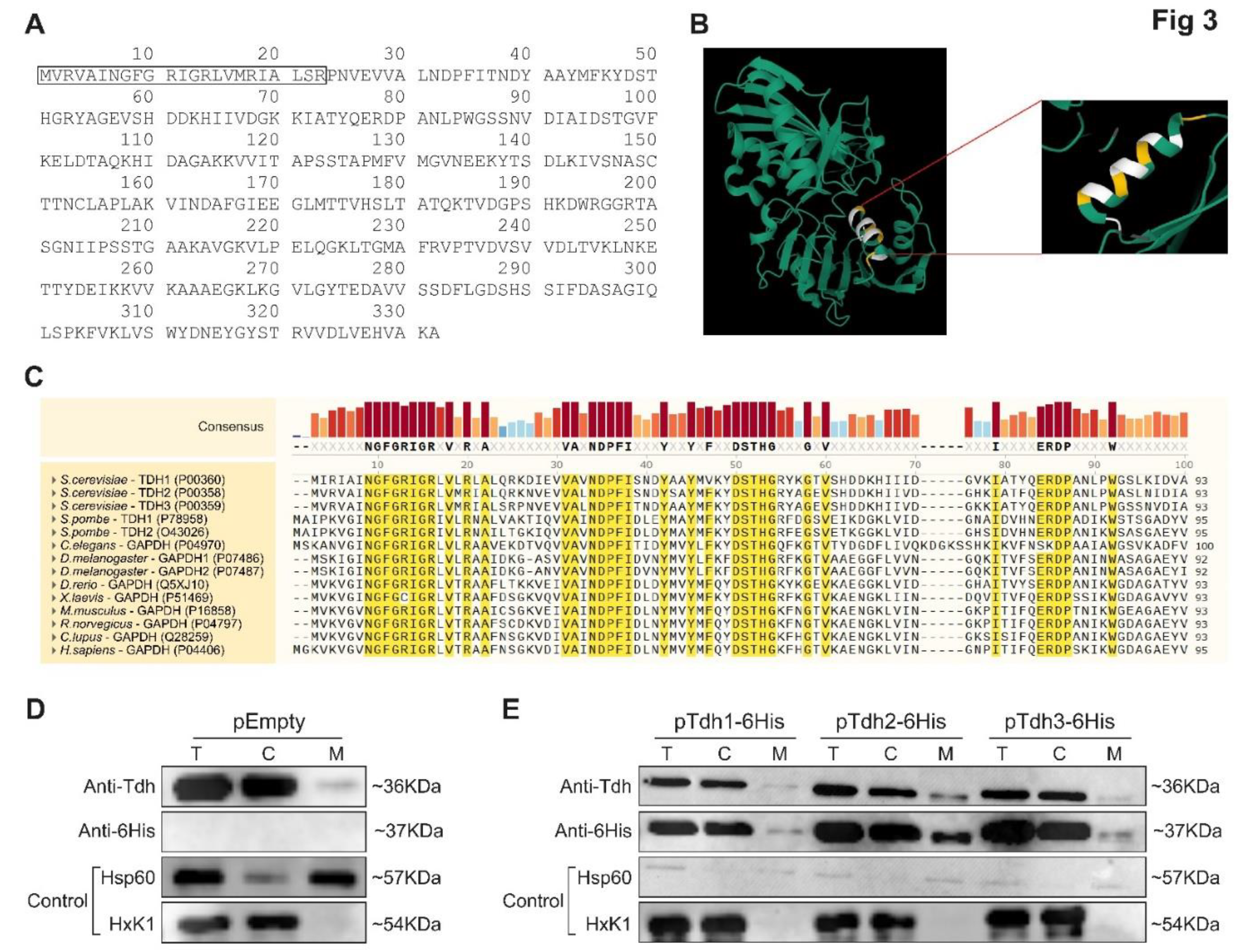
55]. The N-terminal protein sequence of Tdh3 analysis by Uniprot database, based on predicted structures [
56] and MitoProtII [
39], suggest the existence of an MTS at the amino terminus (
Figure 3A,B and
Supplementary Table S8). Moreover, sequence analysis of the first 30 amino acids of the TDH proteins reveals distinct conserved domains across the paralogs throughout eukarya, with a total of 43% amino acid conserved in over 90% of species (
Figure 3C). We chose this family since Tdh2 and Tdh3 were positive in both the MTS guided and C-SWAT screens and they are well studied and have known functions. While several other systematic manuscripts have found them to be associated with mitochondria [
12,
23,
57], they have never been shown to reside in the mitochondrial matrix.
We examined the distribution pattern of Tdh1, Tdh2 and Tdh3 by subcellular fractionation followed by western blot with a polyclonal antibody which recognizes all three wild-type Tdh proteins (
Figure 3D). Using this approach, we found that they exhibit a classic eclipsed distribution with minute amounts in mitochondria (
Figure 3D, M versus C). We also examined the distribution of the full length Tdh proteins when overexpressed (
Figure 3E) using fusion alleles to a 6His-tag, which allows to specifically detect each Tdh protein separately. All tagged Tdh proteins as well as the endogenous (
Figure 3D) proteins, exhibited an eclipsed distribution pattern, which means that the majority of Tdh proteins are found in the cytosolic fraction (C), while minute amounts of the proteins are detected in mitochondria (M) (
Figure 3E).
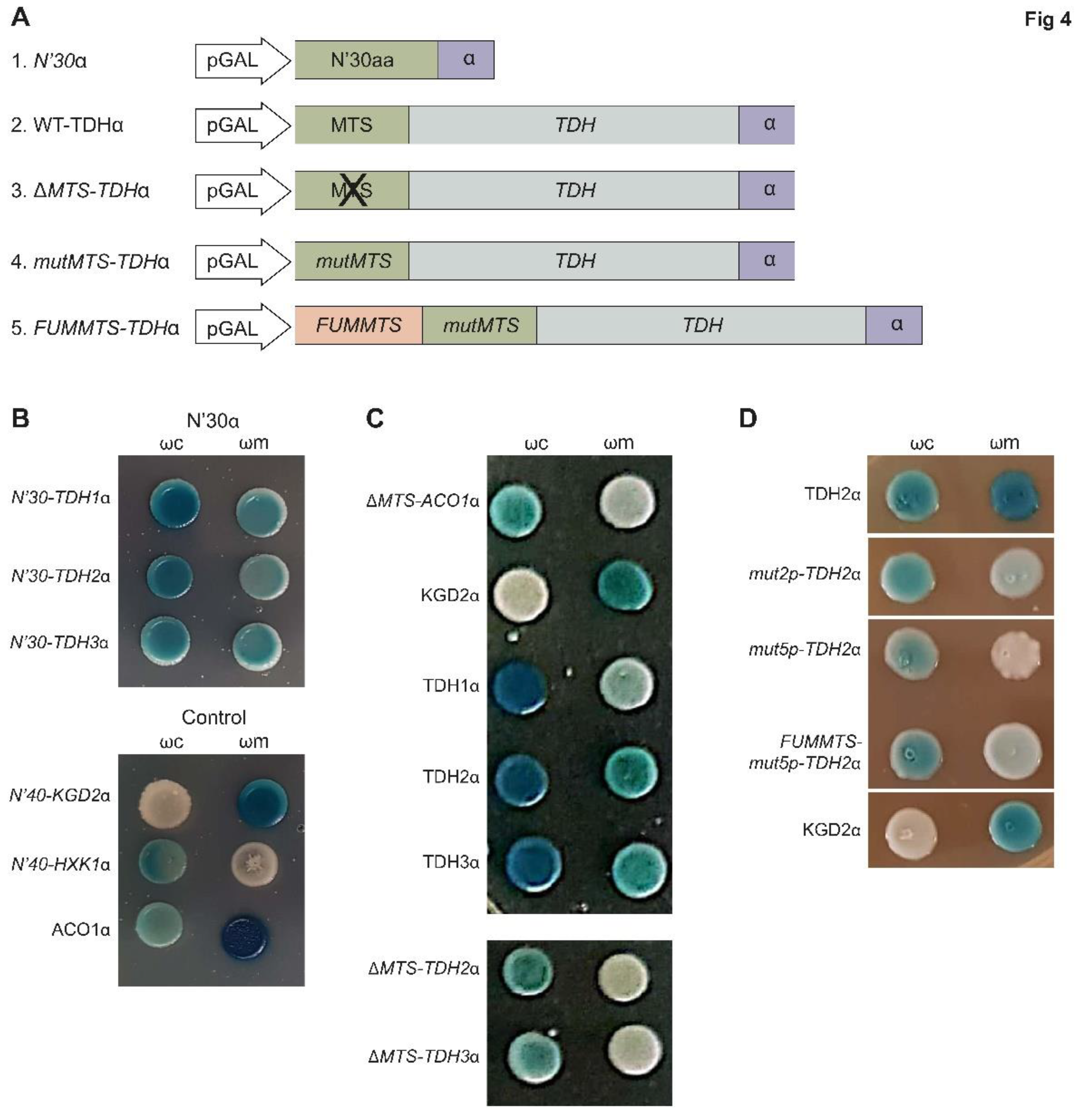
2.7. TDH Proteins Rely on MTS like Signals for Their Eclipsed Distribution
To determine the reliance on the MTS-like sequence for mitochondrial targeting, we fused either their first
30 amino acids or the full-length Tdh proteins to the α fragment at their C-terminus (
Figure 4A). We then followed their localization by α-complementation. Indeed, the MTS alone was sufficient to enable matrix targeting (
Figure 4B) similarly to the full-length proteins. To test if the MTS is also necessary for matrix targeting, we examined Tdh2 and Tdh3 lacking an MTS (
ΔMTS-TDHα) and found that they were absent from mitochondria according to the α-complementation assay (
Figure 4C and
Figure S3). Since these
ΔMTS constructs appear to lack enzymatic activity, we also created a version that lacks a predicted MTS by amino acid substitutions (mutxp,
Figure 4A, Construct 4 illustration). While
mut2p-TDH2α with only two substitutions of arginines to glutamates (Arg11, Glu14), still had residual presence in mitochondria, the mutant with five substitutions (
mut5p-TDH2α in Arg3,11,14,18,23Glu) is already absent from the organelle (
Figure 4D and
Figure S3). These two mutants,
mut2p-TDH2α and
mut5p-TDH2α also appear to lack enzymatic activity hence our experiments cannot clearly distinguish these two traits. Taken together these results indicate that both Tdh2 and Tdh3 harbor a functional MTS [
56].
2.8. Processing of Tdh2/3 Proteins
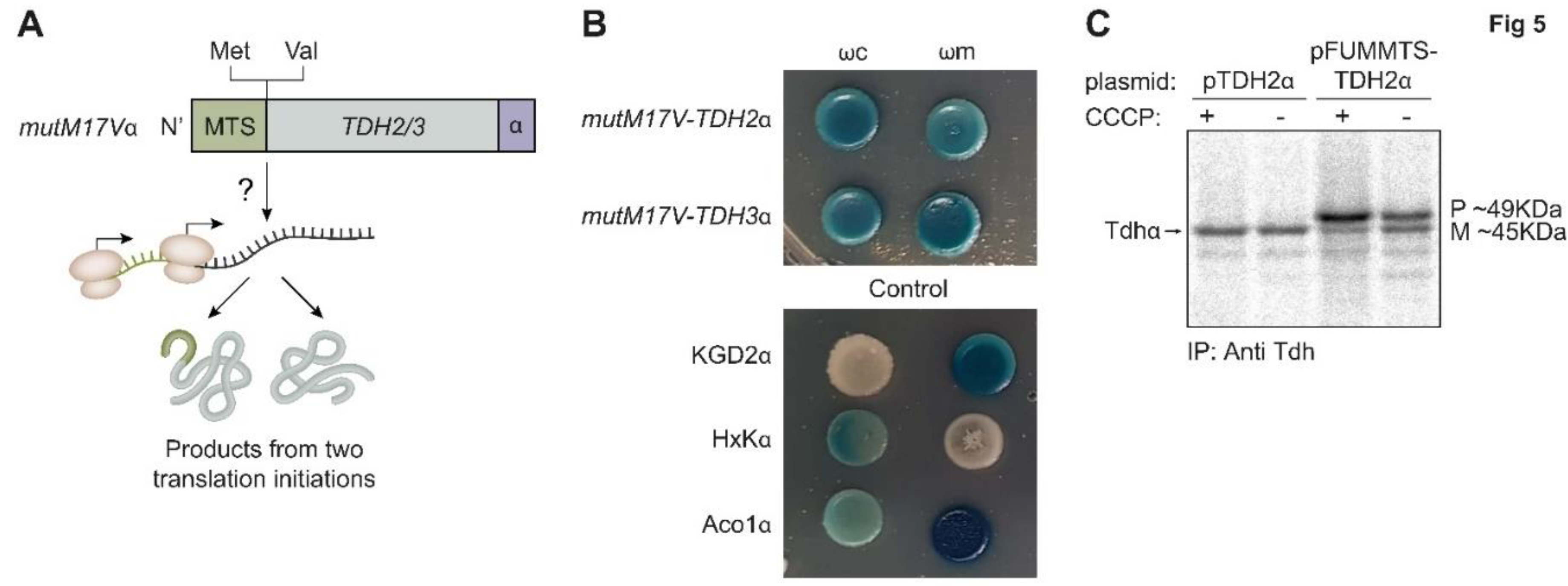
Tdh proteins contain two potential translation initiation methionine codons; one is the first codon that leads to translation of the MTS while the other, from codon 17, will lack the MTS. Hence to determine if the dual targeting of Tdh proteins is a result of two translation products we created a mutant
mutM17Vα, in which the 17th amino acid methionine was replaced by valine, removing the potential second start codon (
Figure 5A, illustration). This mutant was fused to the α tag and followed by the α-complementation assay to probe localization. We found no visible effect of the mutation on the distribution of Tdh2 and Tdh3 proteins in the cell, suggesting that dual localization of Tdh2/3 is not due to two translation products (
Figure 5B). In support of this, no translation from the second methionine was observed by ribosome profiling [
58].
To further examine this question, we fused the whole Tdh2 protein sequence to the strong MTS of fumarase which is expected to be processed by MPP (
FUMMTS-TDH2α, Figure 4A). As expected, this construct is processed by MPP, which removes the
FUM-MTS from some of the molecules, generating two products: A mature (M) species, which displays the same molecular weight as the wild type TDH2α (
Figure 5C, compare lane 4 to lanes 1, 2) and a precursor (P) species which is larger. These two species can be observed more clearly in the presence of the ionophore carbonyl cyanide 3-chlorophenylhydrazone (CCCP) which dissipates the mitochondrial membrane potential and blocks import, resulting in accumulation of unprocessed proteins in the cytosol (
Figure 5C, lane 3). It is important to emphasize that besides these two products, TDH is not further processed indicating that the original Tdh2 is not processed. Accordingly, we show by α-complementation that
mut5p-TDH2α is absent from mitochondria (
Figure 4D and
Figure S3) but its targeting to mitochondria can be restored by the upstream fusion of
FUM-MTS (
FUMMTS-mut5p-TDH2α,
Figure 4A construct 5 illustration and
Figure 4D,
α-complementation). Thus, we conclude that TDH proteins are not processed, suggesting that the MTS amphiphilic α helix is part of the first functional domain of the protein as predicted by the Uniprot database (
Figure 3,
Supplementary Table S8).
2.9. Tdh Proteins are Important for Growth on Carbon Sources Requiring Mitochondria
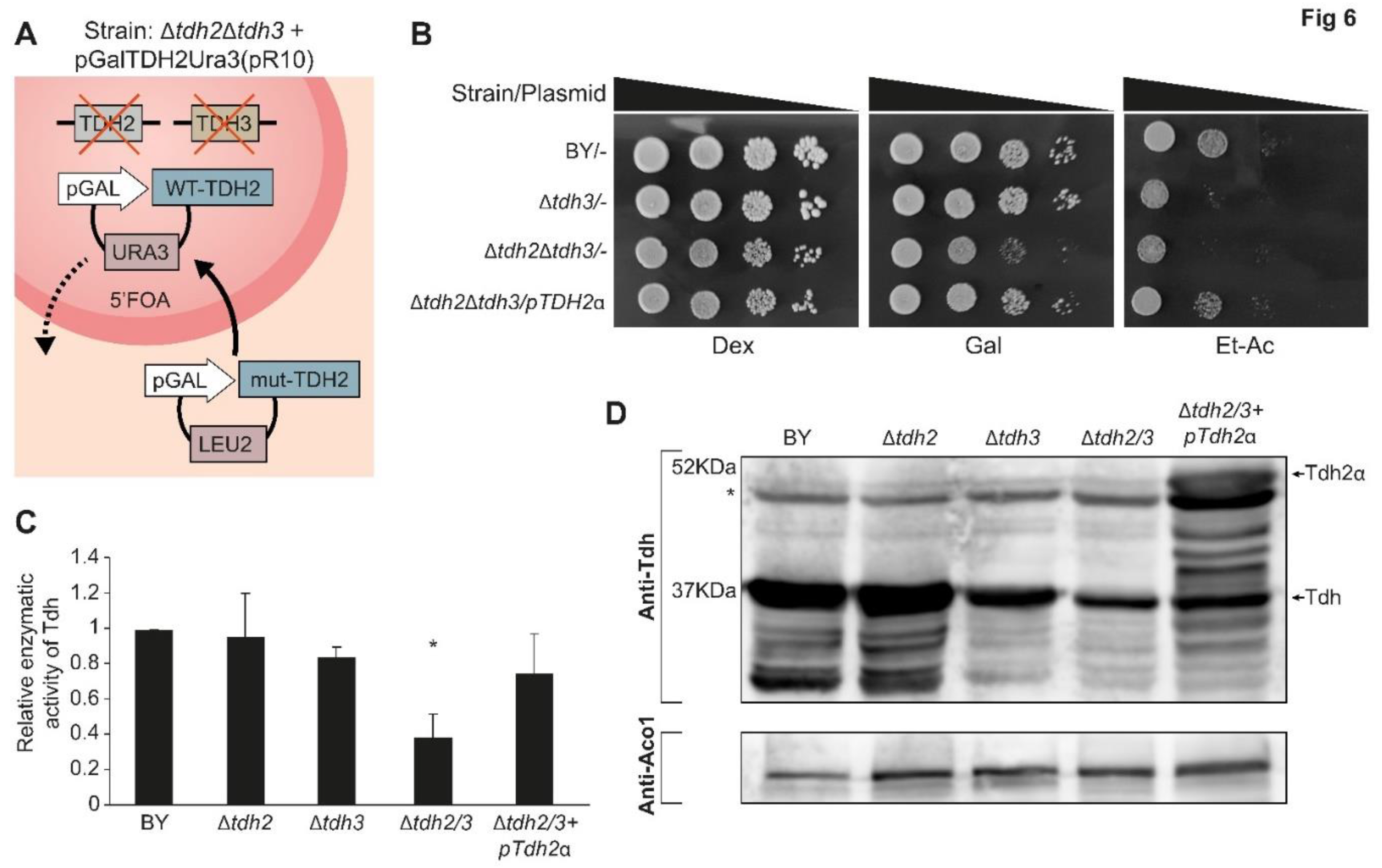
Since we verified that Tdh proteins indeed have an eclipsed distribution in mitochondria we wanted to explore their functional role in the organelle. To do this we constructed a double deletion strain of both
TDH2 and
TDH3 kept viable by a rescue plasmid expressing WT-Tdh2 under an inducible
GAL10 promoter with a counter-selectable URA3 marker [
47] (
Figure 6A). Use of 5FOA (5 Fluoroorotic acid) can therefore be used to replace plasmids (shuffle) with a plasmid enabling biosynthesis of leucine (LEU) expressing specific variants (
Figure 6A).
To examine the different shuffled strains, growth was examined on agar medium supplemented with either fermentable (glucose), oxygen requiring (galactose) or respiration requiring (ethanol-acetate) carbon/energy sources (
Figure 6B). All the strains appear to readily grow on glucose medium which indicates that the double deletion is viable in our strains and growth conditions (
Figure 6B, Row 3). Nevertheless, the double deletion mutant exhibited poor growth on galactose medium when compared to the wild type or to a single knockout strain,
Δtdh3 (
Figure 6B, Row 3). On ethanol-acetate medium, which requires functional mitochondria, both the double deletion mutants and the single
Δtdh3 exhibit poor growth (
Figure 6B, Rows 2, 3). These galactose and ethanol-acetate phenotypes can be restored by a complementing expression plasmid of
TDH2ɑ (
Figure 6B, Row 4).
We assayed the enzymatic activity of the Tdh protein-shuffled-strains (
Figure 6C). The Tdh enzymatic activity of the single mutants
Δtdh2 or
Δtdh3 show similarity to that of the wild type. As expected, the double mutant
Δtdh2Δtdh3 displays significantly lower activity (
Figure 6C, bar 4,
p<0.05) that can be complemented by Tdh2 expression from a plasmid (Figure 6C bar 5). The background Tdh activity can be attributed to the
TDH1 gene which exists in all our strains. The
Δtdh3-strain activity is slightly lower corresponding to the lower protein expression of Tdh in this strain (
Figure 6D lane 3).
2.10. Tdh Proteins affect Mitochondrial Activity when in the Mitochondrial Matrix
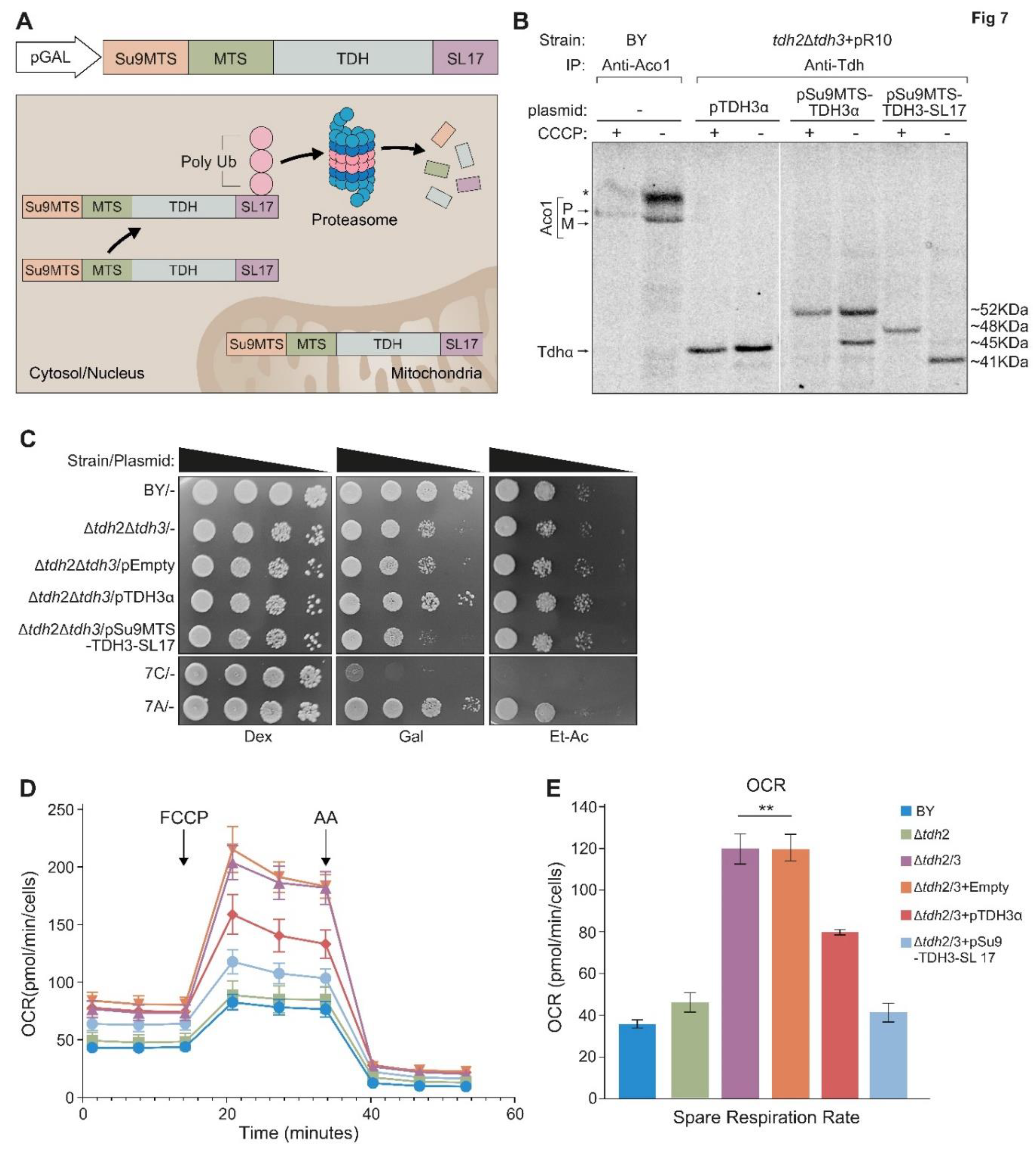
Our attempts to isolate mutant variants in which Tdh2/3 are absent from mitochondria but are both present and active outside mitochondria were unsuccessful. As an alternative approach, we expressed the Tdh2/3 proteins solely in mitochondria. To do this we fused upstream to Tdh3, the sequence of the MTS of subunit 9 of F0-ATPase of
Neurospora crassa (
Su9-MTS (1-69 aa)
Figure 7A in grey). Su9-MTS is the strongest known MTS and is processed twice by MPP. In addition, we fused the degron tag SL17 (colored blue) to the C terminus of Tdh, to target the cytosolic pool of the protein for proteasomal degradation. Expressing
Su9MTS-Tdh3-SL17 creates a hybrid protein that is efficiently targeted to mitochondria and there it is protected from degradation while the cytosolic form of the protein is degraded.
When expressing
Su9MTS-Tdh3-SL17 in the absence of CCCP, only the mature processed mitochondrial form of the protein can be detected (~41KDa), indicating that the cytosolic form has, as expected, been eliminated (
Figure 7B). In the presence of CCCP only the precursor form (~48KDa) can be detected (
Figure 7B, lanes 7, 8). Tdh3α (~45KDa) which lacks an external MTS, as expected, does not exhibit processing (
Figure 7B, lanes 3, 4).
Su9MTS-Tdh3α (~52KDa) which harbors an
Su9MTS displays two molecular bands in the absence of CCCP (precursor ~52KDa and mature ~45KDa) indicating either partial import into mitochondria or partial
Su9-MTS cleavage (
Figure 7B, lane 6). In the presence of CCCP only the precursor is detected (
Figure 7B, lane 5).
The mutant variant
Su9MTS-Tdh3-SL17 cannot rescue the growth phenotype of
Δtdh2Δtdh3 on galactose media (
Figure 7C) demonstrating an essential role under these conditions for Tdh proteins in the cytosol. Growth on ethanol-acetate media, however, which requires mitochondrial respiration showed a slight, but reproducible, increase in the growth of
Δtdh2Δtdh3 expressing
Su9MTS-Tdh3-SL17 which is localized only to the organelle relative to its controls (
Figure 7C rows 2,3 and 5). This supports the notion that under these conditions there is a functional role for the eclipsed Tdh proteins in mitochondria.
To examine this in more depth we analyzed the quantitative respiratory capacity of yeast cultures grown in ethanol-acetate medium by determining the oxygen consumption rate (OCR). Basal respiration was measured, which was followed by the addition of FCCP, a mitochondrial oxidative phosphorylation uncoupler, which leads to maximal respiration. Then antimycin A (AA), which inhibits complex 3 in the respiratory chain and blocks respiration, was added (
Figure 7D). The spare respiration capacity (Maximal minus basal respiration) was calculated (
Figure 7E and
Figure S4). We found that, surprisingly, the respiratory capacity of
Δtdh2Δtdh3 (or
Δtdh2Δtdh3 containing an empty vector) was 3-fold higher compared to the control or a single
Δtdh2 deletion. We hypothesize that this may be a result of a compensatory mechanism of the cells. Importantly,
pSu9MTS-Tdh3-SL17 which is efficiently expressed only in mitochondria, completely restored the normal respiration capacity phenotype of the double mutant. A plasmid expressing full length pTdh3α only partially complemented the
Δtdh2Δtdh3 strain as expected since it has only a minor fraction targeted to mitochondria and cannot replace the loss of both homologues.
3. Discussion
This study changes the perception of the mitochondrial proteome. We put forward hundreds of eclipsed mitochondrial protein candidates due to the complementarity of our approaches. First, the C-SWAT library enabled us an unbiased screen of genes under their natural promoter providing limited sensitivity but a wide coverage. From this approach the 137 candidates are more abundant proteins that are, on average, significantly more conserved than exclusive mitochondrial and non-mitochondrial proteins supporting their identification as eclipsed proteins.
Our complementary approach was more limited in scope but with much higher sensitivity. Choosing proteins based on the presence of an MTS-like sequence at their amino terminus uncovered 143 candidate eclipsed mitochondrial proteins. These exhibit an enrichment of mitochondrial protein properties such as their whole-protein net charge that is similar to known exclusive and dual targeted mitochondrial proteins. With regard to conservation parameters, such as the number of homologs and PGL, they are similar to known exclusive mitochondrial and dual localized proteins which follows our interpretation that they are eclipsed dual-targeted proteins.
Our approaches were complementary and allowed us to capture a plethora of hits as can be seen by the low (~10%) overlap between the candidates raised by each approach (
Figure 1E). In other words, we are looking at the tip of the iceberg. This suggests that additional guided screens, based on metabolic/enzymatic activity, protein traits (such as net charge on the whole protein), internal MTSs [
59,
60] or evolutionary conservation, may identify additional eclipsed mitochondrial proteins.
To prove that the eclipsed proteins are indeed deliberately targeted to mitochondria and have a function there, it is important to go into in-depth, mechanistic, exploration of each example. We chose the
TDH1/2/3 gene family since it appeared in a number of our screens, and also in recent studies [
21,
26,
61]. Focusing on Tdh2/3 that came up in both screens, we found that the natural and overexpressed subcellular pattern of
Tdh2/3 is of eclipsed distribution (highly abundant in the cytosol with minute amounts in mitochondria). Tdh2/3 harbor an N terminal “MTS like” sequence which can function independently of the whole protein but is not cleaved since its removal or site-specific mutagenesis results in loss of enzymatic activity. We could show that the dual targeting is not a result of two translation products and that the N terminus of these proteins is both necessary and sufficient for targeting. Most importantly, while we do not yet understand the exact role of Tdh proteins in specific mitochondrial metabolic pathways, we could show that the matrix form of the proteins affects mitochondrial related phenotypes (the capacity to grow on respiration requiring medium and the overall respiration rate). Since, the rescue by
Su9MTS-TDH3-SL17 is stronger than the rescue by WT-TDH3α, our results suggest that the abundance of matrix echoforms is of importance. Put together our results provide evidence that Tdh proteins are involved in mitochondrial respiration and confirms echoform dual function. While we have chosen here to present the Tdh1/2/3 family as an example of eclipsed distribution of proteins in mitochondria, this is not the only case that we are pursuing. Recently we have found that ubiquitination occurs in the mitochondrial matrix by eclipsed targeted components of the ubiquitin machinery [
38]. Future single gene studies should reveal many more examples of functional eclipsed distribution of mitochondrial proteins.
Generally, we have come a long way since the “One gene one enzyme (polypeptide) hypothesis” [
62]. Today, not only do we understand that a single gene can give rise to multiple protein products, but we also realize that these protein products are not necessarily targeted to a single organelle or subcellular location. Such cases of dual targeting or localization, which we have shown are highly abundant, provide cellular flexibility and tunability. Our work therefore not only expands our understanding of mitochondrial function but also changes the way we perceive protein localization in eukaryotic cells. This study provides a cue for the investigation of eclipsed mitochondrial proteins’ functions in higher eukaryotes and involvement in human disorders. Our work provides a paradigm of deliberate eclipsed mitochondrial localization, targeting and function, and should expand our understanding of mitochondrial function in health and disease.
4. Materials and Methods
4.1. Strains
Saccharomyces cerevisiae Yeast strains were provided by Euroscarf; BY4741 (MATa; his3Δ1; leu2Δ0; met15Δ0; ura3Δ0), Δtdh2 (BY4741; MATa; his3Δ1; leu2Δ0; met15Δ0; ura3Δ0; YJR009c::kanMX4), and Δtdh3 (BY4741; MATa; his3Δ1; leu2Δ0; met15Δ0; ura3Δ0; YGR192c::kanMX4). S. Cerevisiae wild type 7A (YPH499, Sc-7A) and TCA cycle (tricarboxylic acid cycle) mutant 7C (YPH500, Sc-7C) were provided from our lab stock.
Additional null and composite null mutants were generated for this study using homologous recombination technique. The Δtdh2Δtdh3 double null mutant strain was constructed using transformants Δtdh3 carrying pGalTDH2Ura(R10) plasmid. Essentially, the cells were made competent using the Frozen-EZ Yeast Transformation II™ (ZYMO RESEARCH), according to the manufacturer’s instructions. A cloNAT cassette with flanking upstream and downstream sequences of TDH2 gene was cloned and amplified by PCR, using pAV10.KN (AddGene) plasmid as a template using the following primers: 5’-CCAAGAACTTAGTTTCAAATTAAATTCATCACACAAACAAACAAAACAAAgacatggaggcccagaatac-3’ and 5’-AATTATTAATAATAAAAACTAAATCATTAAAGTAACTTAAGGAGTTAAATcagtatagcgaccagcattc-3’.
The amplified cassette was transformed into the Δtdh3+pGalTDH2Ura(R10) strain and yeast were plated on 2% galactose lacking uracil and containing 200µg/ml cloNAT as a selective medium. After primary selection, mutant colonies were purified by PCR to confirm complete parental gene knockout, using the following primers: 5’-tcggagacctgcaatttt-3’ (TDH2 chromosomal flanking upstream sequence); 5’-ggttgtttatgttcggatgt-3’ (cloNAT TEF promoter sequence, positive control); and 5’-ccagcgtcaatgtgcttt-3’ (TDH2 mid chromosomal sequence, negative control).
4.2. Plasmids
pωc (pYES/M15) was kindly provided by Picard [
30] ,pFumα, pHxk1α, pKgd1α and BS-Su9ω (pωm) were described elsewhere [
35]. All the screened proteins used in this study were created by synthesis and amplification of the corresponding fragments by Bio-Basic Singapore. The resulting products were cloned into pFumα using an Orientation Enrichment Reaction (OER). All plasmids described above were introduced into strain BY4741.
Su9-MTS - MASTRVLASRLASQMAASAKVARPAVRVAQVSKRTIQTGSPLQTLKRTQMTSIVNATTRQ
AFQKRAYSS
SL17 – SISFVIRSHASIRMGASNDFFHKLYFTKCLTSVILSKFLIHLLLRSTPRV
FUM-MTS - MLRFTNCSCKTFVKSSYKLNIRRMNSS
pTDH(1-3)-6His–TDH(1-3) were tagged with 6His at their C-terminus and cloned into pRS423Gal. These plasmids were created by amplifying the respective sequences from genomic DNA using the indicated primers below with BamHI and EcoRI at the '5 and '3 respectively. The amplified PCR fragments were cut with BamHI and EcoRI and cloned into pRS423Gal cut with BamHI and EcoRI.
Primers: For TDH1–5’-aaatttggatccATGATCAGAATTGCTATTAACG-3’ and 5’-tttaaagaattcGTGATGGTGATGGTGATGTTAAGCCTTGGCAACATATTC-3’.
For TDH(2-3)-5'-aaatttggatccATGGTTAGAGTTGCTATTAACG-3' and 5’-aaatttgaattcGTGATGGTGATGGTGATGTTAAGCCTTGGCAACGTGTTC-3’
4.3. Growth Conditions
Strains were grown at 30°C or as indicated in synthetic depleted medium containing 0.67% (w/v) yeast nitrogen base without amino acids (Difco Laboratories), 2% glucose or galactose (w/v), CSM dropout mix (Qbiogene) supplemented with the appropriate amino acids (50 μg/ml). For agar plates, 2% agar was added. X-gal plates were prepared as follows; a stock of 2% galactose, 1% raffinose, 0.008% X-gal (dissolved in 100% N,Ndimethylformamide) and 1× BU salts (25 mM sodium phosphate buffer titrated to pH 7.0) was added to autoclaved medium at a temperature of 50°C. For 5’FOA selection, strains containing pGalTDH2Ura3 (pR10) plasmid, were streaked on 2% galactose plates supplemented with 0.1% 5-fluoroorotic acid for selection of the loss of pGalTDH2Ura3. Colonies were picked and streaked on galactose media lacking URA3 to confirm loss of pR10.
4.4. β-Galactosidase α-Complementation Assay
Yeast cells were transformed with plasmids encoding various α fusion proteins and either ωc or ωm. Colonies were plated on X-gal plates and incubated at 30°C for 72 h.
4.5. C-SWAT Library
The C-terminus SWAp Tag (SWAT) library of the yeast Saccharomyces cerevisiae [
63] was used to generate a library with C-terminus tag as previously published (same reference). In short, a SWAT donor strain (yMS2085) was transformed with a donor plasmid (pMS859) containing the alpha tag and then SWATted as described. The final library genotype is
his3∆1 leu2∆0 met15∆0 ura3∆0, can1∆::GAL1pr-SceI-NLS::STE2pr-SpHIS5 lyp1∆::STE3pr-LEU2 (XXX::L3-alpha-ADH1ter-Hygro-ALG9term-L4. Once generated the swatting efficiency and integrity were evaluated (figure S5). For α-complementation assay, library was mated with BY4742 (MATα;
his3Δ1;leu2Δ0;lys2Δ0; ura3Δ0), contain ωm expression plasmid (described above), and plated on X-gal. The plates were incubated at 30°C for 72 h.
4.6. Metabolic Labeling
Yeast cultures were grown in 5ml SD–glucose or SD–galactose at 30°C or as indicated to OD600nm=1.5. Cells were collected by centrifugation and resuspended in 400 μl fresh media lacking methionine. Cultures were labeled with 40 μCi/ml [35S] methionine and further incubated for 30 min at 30°C. When required, carbonyl cyanide chlorophenylhydrazone (CCCP) was added to a final concentration of 20 μM at the start of labeling. Labeling was stopped by addition of 10mM sodium azide. The labeled cells were collected by centrifugation and resuspended in TE buffer (Tris10 mM, EDTA 1 mM, pH 8.0) containing 1 mM phenylmethylsulfonylfluoride (PMSF), broken with glass beads, and centrifuged to obtain the supernatant fraction. Supernatants were denatured by boiling in 1% SDS, immunoprecipitated with the indicated anti-serum and protein A–Sepharose (Amersham Biosciences) or magnetic Dynabeads protein A (Dynal Biotech ASA). Samples were then analyzed by SDS-PAGE followed by visualization with the BAS2000 image analyzing system (Fuji Corp.) and autoradiography.
4.7. Subcellular Fractionation
Induced yeast (in galactose) cultures were grown to OD
600nm=1.5. Mitochondria were isolated as described previously [
19]. Spheroplasts were prepared in the presence of Zymolyase-20T. Subcellular fractionation experiments were assayed for cross contamination of fractions using Hsp60 as a mitochondrial marker and Hexokinase1 as a cytosolic marker in Western blots.
4.8. Western Blot Analysis
S. cerevisiae cells were harvested using lysis buffer containing the following: 10 mM Tris pH 8, 1 mM EDTA, and 100 mM PMSF. Protein concentrations were determined using the Bradford method [
64]. Samples were separated on 10% or 12% SDS-PAGE gels and transferred onto PVDF membranes (Millipore). The indicated primary antibodies were used. All blots were incubated with the appropriate IgG-HRP–conjugated secondary antibody. Protein bands were visualized and developed using Enhanced Chemiluminescent immunoblotting detection system (ImageQuant LAS 4000 mini, GE Healthcare) and Gel Documentation System (Bio-Rad, Hercules, CA, USA).
4.9. Enzymatic Activity
S. cerevisiae cultures grown in galactose medium for translation induction were harvested and resuspended in TE buffer (pH 8.0) containing 1 mM PMSF, broken with glass beads, and centrifuged to obtain the supernatant fraction. Glyceraldehyde-3-phosphate dehydrogenase activity was assayed in 1.0-ml reactions containing 0.1 M potassium phosphate (pH 7.4), 1.0 mM NAD+, 10 mM EDTA, 0.1 mM dithiothreitol, and 4.0 mM glyceraldehyde 3-phosphate (29). NADH formation was monitored spectrophotometrically at 340nm. Glyceraldehyde-3-phosphate dehydrogenase activity units are expressed as µmol of NADH formed/min at 25°C.
4.10. Oxygen Consumption Evaluation by Seahorse
Mitochondrial function was assessed using the XF96 extracellular flux analyzer (Agilent, Saint Clara, CA, USA) as previously described [
65], with minor modifications. Yeast cultures were grown overnight at 30°C in galactose or galactose selected medium. Cells were diluted to OD
600nm = 0.1 in ethanol-acetate medium and 180µl were seeded in Seahorse XF96 microplate coated in poly-L-lysine (22µl of 0.1mg/ml per well). The loaded plate was centrifuged at 500 X g for 3 min at room temperature following incubation for 30 minutes at 30°C. Basal Oxygen Consumption Rate (OCR) was measured following an injection of 20 µM of FCCP and 2.5 µM of antimycin A according to the manufactures’ manual Seahorse XF Cell Mito Stress Test (Agilent). OCR were normalized to cells at OD
600nm = 0.1 and Basal respiration, Maximal respiration and spare respiration capacity were calculated according to Agilent protocol.
4.11. Microscopic Analysis of Mitochondria Phenotype
Glass-bottom, 384-well microscopy plates (Matrical Bioscience, Spokane, WA, USA) coated with Concanavalin A (Sigma-Aldrich, St. Louis, MO, USA) were used for imaging. Cells in mid-logarithmic phase were adhered to the plates by incubating at RT for 15 min. Upon adherence to the plate, media was replaced with media containing 50 nM MitoTracker (MitoTracker Orange CMTMRos; Invitrogen, Waltham, MA, USA), and cells were incubated at RT for 10 min, washed once, and imaged using automated inverted fluorescence microscope system (Olympus) harboring a spinning disk high-resolution module (Yokogawa CSU-W1 SoRa confocal scanner with double micro lenses and 50-µm pinholes). Images of cells in the 384-well plates were using a 60× oil lens (NA 1.42) and with a Hamamatsu ORCA-Flash 4.0 camera. Mitottracker was excited by a 561 laser and the signal detected through a 617/73 nm emission filter. For each well, 9 positions were imaged and over 6000 yeast cells were segmented for mitochondria, according to mitotracker signal, and analyzed using ScanR software (Olympus).
4.12. Statistical Analysis
All statistical analysis in
Supplementary Table S3 were carried by GraphPed Prism, version 9.3.1.
p value was calculated for unpaired and non-parametric tests, using the Mann-Whitney test (to compare ranks) or the Kolmogorov-Smirnov (to compare cumulative distribution).
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org,
Figure S1. C-SWAT library controls and validations. (A). C-SWAT yeast library strains expressing either cytosolic or mitochondrial ω were submitted to the α-complementation assay as described in
Figure 2. Control genes: KGD2α, mitochondrial; TDH3α, dual targeted; ERG1α, cytosolic. Many library genes show white colonies both with cytosolic and mitochondrial ω (WW), such as CDC3α. Yeast strains expressing cytosolic ω (ωc) or mitochondrial ω (ωm) were co-expressed with either α-fused genes from the C-SWAT library (B), or with over expressed genes (C) (7/31 proteins from the “validation screen”) and submitted to the α-complementation assay (first and second columns of the two panels). Strain extracts were analyzed by western blotting using antibodies against either α or β-Gal (third columns of the two panels).
Figure S2. Examples of assays for predicted MTSs and corresponding full-length proteins. Yeast cultures expressing cytosolic ω (ωc) or mitochondrial ω (ωm) were co-expressed with various α-fused N-terminal peptides (first 40 aa, N’40-α) or with the corresponding full-length α-fused proteins (WT). Yeast cultures were grown on galactose medium plates containing X-gal. Blue colonies detect fragments that are associated with the indicated ω fragments as described in
Figure 2A. Controls: TDH2α (dual targeted), KGD2α (mitochondrial) and mut5p-TDH2α (cytosolic, as described in
Figure 6D). Equivalent portions from each strain extract were analyzed by western blotting using antibodies against α and Aco1 as a control.
Figure S3. Tdh2/3 protein expression. Yeast strains harboring the indicated plasmids were grown on galactose media. Equivalent portions from each strain extract were analyzed by western blotting using antibodies against β-Gal (A) or α (B). Tdhα (A) and
mut5p-Tdh2α (B) are indicated by arrows. * Nonspecific bands.
Figure S4. Mitochondrial effects of TDH proteins. Yeast strains harboring plasmids were grown and diluted to A600=0.1 in ethanol-acetate seahorse buffer as described in Figure 9D/E. Quantification of basal, maximal (maximal electron transport activity induced by the uncoupler FCCP) and the spare respiration capacity (maximal minus basal) OCR are presented. The results were obtained from three independent experiments (mean ± SEM [n = 3]) two-tailed Student’s t test **P < 0.001).
Figure S5. Evaluation of C-tagged alpha library SWATting efficiency and integrity. (A). Pie chart of the percent of yeast strains that survived the entire SWATting procedure. (B). Pie chart of SWATting efficiency. To test if there were any residual parental strains remaining in the library, the library was plated on -Ura agar plates, the resistance of the mother C-SWAT library. No growth was observed after 4 days in 37
0c, indicating that the new alpha library does not contain unswatted strains. C. Pie chart of the integration efficiency of the alpha tag. 33 random strains were tested by PCR for the proper C-tagging of their respective genes with the alpha tag. Only 2 colonies failed to produce the desired PCR product, indicating that the rate of successful swatting is 93.94%.
Author Contributions
The author contributions are as follows: OK, MM, KD, AE and ZY performed the experiments; MM, ZY and OK analyzed the data in the supplementary Tables; MS and OP mentored the students performing the work, raised funds for the research and generated ideas. MM, OP, OK and MS wrote the manuscript. All authors reviewed the final manuscript, contributed to the final version and approved it. All authors have read and agreed to the published version of the manuscript.
Funding
The study was supported by the German Israeli Project Cooperation (Grant P17516) DIP to MS and OP. This work was supported by grants to OP from the Israel Science Foundation (Grant 1455/17) and The CREATE Project of the National Research Foundation of Singapore (Singapore-Hebrew University of Jerusalem MMID2). Work in the Schuldiner lab was additionally supported by the ERC CoG OnTarget (864068). The funders had no role in study design, data collection, and interpretation or in the decision to submit the work for publication. MS is an Incumbent of the Dr. Gilbert Omenn and Martha Darling Professorial Chair in Molecular Genetics and OP is an Incumbent of the Harry and Helen L. Brenner Chair in Molecular Biology.
Acknowledgments
We wish to thank Michael Knop for graciously sharing the C’ SWAT library with us. We thank Amir Fadel for help with the analysis of the SWAT complementation. We wish to thank Tali Wiesel from the Weizmann Graphics Unit for help with Figure design. We would also like to thank Zohar Gazi and Dr. Tsviya Olender for help with bioinformatic analysis.
Conflicts of Interest
Declare conflicts of interest or state “The authors declare no conflict of interest.” Authors must identify and declare any personal circumstances or interest that may be perceived as inappropriately influencing the representation or interpretation of reported research results. Any role of the funders in the design of the study; in the collection, analyses or interpretation of data; in the writing of the manuscript, or in the decision to publish the results must be declared in this section. If there is no role, please state “The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results”.
References
- Carrie, C., E. Giraud, and J. Whelan, Protein transport in organelles: Dual targeting of proteins to mitochondria and chloroplasts. FEBS J, 2009. 276(5): p. 1187-95. [CrossRef]
- Kalderon, B. and O. Pines, Protein folding as a driving force for dual protein targeting in eukaryotes. Front Mol Biosci, 2014. 1: p. 23. [CrossRef]
- Regev-Rudzki, N., et al., Yeast aconitase in two locations and two metabolic pathways: seeing small amounts is believing. Mol Biol Cell, 2005. 16(9): p. 4163-71. [CrossRef]
- Regev-Rudzki, N. and O. Pines, Eclipsed distribution: a phenomenon of dual targeting of protein and its significance. Bioessays, 2007. 29(8): p. 772-82. [CrossRef]
- Silva-Filho, M.C., One ticket for multiple destinations: dual targeting of proteins to distinct subcellular locations. Curr Opin Plant Biol, 2003. 6(6): p. 589-95. [CrossRef]
- Weill, U., et al., Genome-wide SWAp-Tag yeast libraries for proteome exploration. Nat Methods, 2018. 15(8): p. 617-622. [CrossRef]
- Yofe, I., et al., One library to make them all: streamlining the creation of yeast libraries via a SWAp-Tag strategy. Nat Methods, 2016. 13(4): p. 371-378. [CrossRef]
- Naamati, A., et al., Dual targeting of Nfs1 and discovery of its novel processing enzyme, Icp55. J Biol Chem, 2009. 284(44): p. 30200-8. [CrossRef]
- Regev-Rudzki, N., et al., Dual localization of fumarase is dependent on the integrity of the glyoxylate shunt. Mol Microbiol, 2009. 72(2): p. 297-306. [CrossRef]
- Yogev, O., A. Naamati, and O. Pines, Fumarase: a paradigm of dual targeting and dual localized functions. FEBS J, 2011. 278(22): p. 4230-42. [CrossRef]
- Yogev, O. and O. Pines, Dual targeting of mitochondrial proteins: mechanism, regulation and function. Biochim Biophys Acta. 1808(3): p. 1012-20. [CrossRef]
- Yogev, O. and O. Pines, Dual targeting of mitochondrial proteins: mechanism, regulation and function. Biochim Biophys Acta, 2011. 1808(3): p. 1012-20. [CrossRef]
- Avadhani, N.G., Targeting of the same proteins to multiple subcellular destinations: mechanisms and physiological implications. FEBS J, 2011. 278(22): p. 4217. [CrossRef]
- Carrie, C. and J. Whelan, Widespread dual targeting of proteins in land plants: when, where, how and why. Plant Signal Behav, 2013. 8(8). [CrossRef]
- Duchene, A.M. and P. Giege, Dual localized mitochondrial and nuclear proteins as gene expression regulators in plants? Front Plant Sci, 2012. 3: p. 221.
- Karniely, S. and O. Pines, Single translation--dual destination: mechanisms of dual protein targeting in eukaryotes. EMBO Rep, 2005. 6(5): p. 420-5. [CrossRef]
- Monteuuis, G., et al., Non-canonical translation initiation in yeast generates a cryptic pool of mitochondrial proteins. Nucleic Acids Res, 2019. 47(11): p. 5777-5791. [CrossRef]
- Knox, C., et al., Import into mitochondria, folding and retrograde movement of fumarase in yeast. J Biol Chem, 1998. 273(40): p. 25587-93. [CrossRef]
- Yogev, O., et al., Fumarase: a mitochondrial metabolic enzyme and a cytosolic/nuclear component of the DNA damage response. PLoS Biol, 2010. 8(3): p. e1000328.
- Danpure, C.J., How can the products of a single gene be localized to more than one intracellular compartment? Trends Cell Biol, 1995. 5(6): p. 230-8.
- Di Bartolomeo, F., et al., Absolute yeast mitochondrial proteome quantification reveals trade-off between biosynthesis and energy generation during diauxic shift. Proc Natl Acad Sci U S A, 2020. 117(13): p. 7524-7535. [CrossRef]
- Prokisch, H., et al., Integrative analysis of the mitochondrial proteome in yeast. PLoS Biol, 2004. 2(6): p. e160. [CrossRef]
- Reinders, J., et al., Toward the complete yeast mitochondrial proteome: multidimensional separation techniques for mitochondrial proteomics. J Proteome Res, 2006. 5(7): p. 1543-54. [CrossRef]
- Sickmann, A., et al., The proteome of Saccharomyces cerevisiae mitochondria. Proc Natl Acad Sci U S A, 2003. 100(23): p. 13207-12. [CrossRef]
- Small, I., et al., Two birds with one stone: genes that encode products targeted to two or more compartments. Plant Mol Biol, 1998. 38(1-2): p. 265-77.
- Vogtle, F.N., et al., Landscape of submitochondrial protein distribution. Nat Commun, 2017. 8(1): p. 290. [CrossRef]
- Breker, M., M. Gymrek, and M. Schuldiner, A novel single-cell screening platform reveals proteome plasticity during yeast stress responses. J Cell Biol, 2013. 200(6): p. 839-50. [CrossRef]
- Nakai, Y., et al., Nuclear localization of yeast Nfs1p is required for cell survival. J Biol Chem, 2001. 276(11): p. 8314-20. [CrossRef]
- Shlevin, L., et al., Location-specific depletion of a dual-localized protein. Traffic, 2007. 8(2): p. 169-76. [CrossRef]
- Ullmann, A., F. Jacob, and J. Monod, Characterization by in vitro complementation of a peptide corresponding to an operator-proximal segment of the beta-galactosidase structural gene of Escherichia coli. J Mol Biol, 1967. 24(2): p. 339-43.
- Abbas-Terki, T. and D. Picard, Alpha-complemented beta-galactosidase. An in vivo model substrate for the molecular chaperone heat-shock protein 90 in yeast. Eur J Biochem, 1999. 266(2): p. 517-23. [CrossRef]
- Ben-Menachem, R., et al., A third of the yeast mitochondrial proteome is dual localized: a question of evolution. Proteomics, 2011. 11(23): p. 4468-76. [CrossRef]
- Dinur-Mills, M., M. Tal, and O. Pines, Dual targeted mitochondrial proteins are characterized by lower MTS parameters and total net charge. PLoS One, 2008. 3(5): p. e2161. [CrossRef]
- Kisslov, I., et al., Dual-targeted proteins tend to be more evolutionarily conserved. Mol Biol Evol, 2014. 31(10): p. 2770-9. [CrossRef]
- Karniely, S., et al., Alpha-complementation as a probe for dual localization of mitochondrial proteins. Exp Cell Res, 2006. 312(19): p. 3835-46.
- Meurer, M., et al., Genome-wide C-SWAT library for high-throughput yeast genome tagging. Nat Methods, 2018. 15(8): p. 598-600. [CrossRef]
- Velez-Bermudez, I.C., et al., Isobaric Tag for Relative and Absolute Quantitation (iTRAQ)-Based Protein Profiling in Plants. Methods Mol Biol, 2016. 1450: p. 213-21.
- Zhang Y, Karmon O, Das K, Wiener R, Lehming N, Pines O. 2022. Ubiquitination occurs in the mitochondrial matrix by eclipsed targeted components of the ubiquitin machinery. Cells. 11(24): 4109. [CrossRef]
- Andreoli, C., et al., MitoP2, an integrated database on mitochondrial proteins in yeast and man. Nucleic Acids Res, 2004. 32(Database issue): p. D459-62. [CrossRef]
- Elbaz, A.M., Study of mitochondrial eclipsed dually targeted proteins in yeast, in Microbiology and Molecular genetics. 2021, The Hebrew university of Jerusalem.
- Breker, M., et al., LoQAtE--Localization and Quantitation ATlas of the yeast proteomE. A new tool for multiparametric dissection of single-protein behavior in response to biological perturbations in yeast. Nucleic Acids Res, 2014. 42(Database issue): p. D726-30. [CrossRef]
- Suzuki, R. and N. Saitou, Exploration for functional nucleotide sequence candidates within coding regions of mammalian genes. DNA Res, 2011. 18(3): p. 177-87. [CrossRef]
- Zhang, F. and R.E. Broughton, Mitochondrial-nuclear interactions: compensatory evolution or variable functional constraint among vertebrate oxidative phosphorylation genes? Genome Biol Evol, 2013. 5(10): p. 1781-91.
- Holland, J.P. and M.J. Holland, The primary structure of a glyceraldehyde-3-phosphate dehydrogenase gene from Saccharomyces cerevisiae. J Biol Chem, 1979. 254(19): p. 9839-45. [CrossRef]
- Holland, J.P. and M.J. Holland, Structural comparison of two nontandemly repeated yeast glyceraldehyde-3-phosphate dehydrogenase genes. J Biol Chem, 1980. 255(6): p. 2596-605. [CrossRef]
- McAlister, L. and M.J. Holland, Isolation and characterization of yeast strains carrying mutations in the glyceraldehyde-3-phosphate dehydrogenase genes. J Biol Chem, 1985. 260(28): p. 15013-8. [CrossRef]
- McAlister, L. and M.J. Holland, Differential expression of the three yeast glyceraldehyde-3-phosphate dehydrogenase genes. J Biol Chem, 1985. 260(28): p. 15019-27. [CrossRef]
- Martin, W.F. and R. Cerff, Physiology, phylogeny, early evolution, and GAPDH. Protoplasma, 2017. 254(5): p. 1823-1834.
- Almeida, B., et al., NO-mediated apoptosis in yeast. J Cell Sci, 2007. 120(Pt 18): p. 3279-88. [CrossRef]
- Boucherie, H., et al., Differential synthesis of glyceraldehyde-3-phosphate dehydrogenase polypeptides in stressed yeast cells. FEMS Microbiol Lett, 1995. 125(2-3): p. 127-33. [CrossRef]
- Delgado, M.L., M.L. Gil, and D. Gozalbo, Starvation and temperature upshift cause an increase in the enzymatically active cell wall-associated glyceraldehyde-3-phosphate dehydrogenase protein in yeast. FEMS Yeast Res, 2003. 4(3): p. 297-303. [CrossRef]
- Delgado, M.L., et al., The glyceraldehyde-3-phosphate dehydrogenase polypeptides encoded by the Saccharomyces cerevisiae TDH1, TDH2 and TDH3 genes are also cell wall proteins. Microbiology (Reading), 2001. 147(Pt 2): p. 411-417. [CrossRef]
- Ringel, A.E., et al., Yeast Tdh3 (glyceraldehyde 3-phosphate dehydrogenase) is a Sir2-interacting factor that regulates transcriptional silencing and rDNA recombination. PLoS Genet, 2013. 9(10): p. e1003871. [CrossRef]
- Silva, A., et al., Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) is a specific substrate of yeast metacaspase. Biochim Biophys Acta, 2011. 1813(12): p. 2044-9. [CrossRef]
- Shen, H., et al., Structural insights into RNA recognition properties of glyceraldehyde-3-phosphate dehydrogenase 3 from Saccharomyces cerevisiae. IUBMB Life, 2014. 66(9): p. 631-8. [CrossRef]
- Liu, Q., Wang, H., Liu, H., Teng, M., Li, X. Preliminary crystallographic analysis of glyceraldehyde-3-phosphate dehydrogenase 3 from Saccharomyces cerevisiae. 2012. Acta Crystallogr Sect F Struct Biol. Cryst Commun 68: 978-980. [CrossRef]
- Brandina, I., et al., Enolase takes part in a macromolecular complex associated to mitochondria in yeast. Biochim Biophys Acta, 2006. 1757(9-10): p. 1217-28. [CrossRef]
- Eisenberg, A.R., et al., Translation Initiation Site Profiling Reveals Widespread Synthesis of Non-AUG-Initiated Protein Isoforms in Yeast. Cell Syst, 2020. 11(2): p. 145-160 e5. [CrossRef]
- Backes, S., et al., Tom70 enhances mitochondrial preprotein import efficiency by binding to internal targeting sequences. J Cell Biol, 2018. 217(4): p. 1369-1382. [CrossRef]
- Schneider, K., et al., iMLP, a predictor for internal matrix targeting-like sequences in mitochondrial proteins. Biol Chem, 2021. 402(8): p. 937-943. [CrossRef]
- Morgenstern, M., et al., Definition of a High-Confidence Mitochondrial Proteome at Quantitative Scale. Cell Rep, 2017. 19(13): p. 2836-2852. [CrossRef]
- Beadle, G.W. and E.L. Tatum, Genetic Control of Biochemical Reactions in Neurospora. Proc Natl Acad Sci U S A, 1941. 27(11): p. 499-506. [CrossRef]
- Meurer, M.; Duan, Y.; Sass, E.; Kats, I.; Herbst, K.; Buchmuller, B.C.; Dederer, V.; Huber, F.; Kirrmaier, D.; Štefl, M.; et al. Genome-wide C-SWAT library for high-throughput yeast genome tagging. Nat. Methods 2018, 15, 598–600,. [CrossRef]
- Kruger, N.J., The Bradford method for protein quantitation. Methods Mol Biol, 1994. 32: p. 9-15.
- Bradley, M.C., et al., COQ11 deletion mitigates respiratory deficiency caused by mutations in the gene encoding the coenzyme Q chaperone protein Coq10. J Biol Chem, 2020. 295(18): p. 6023-6042. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).