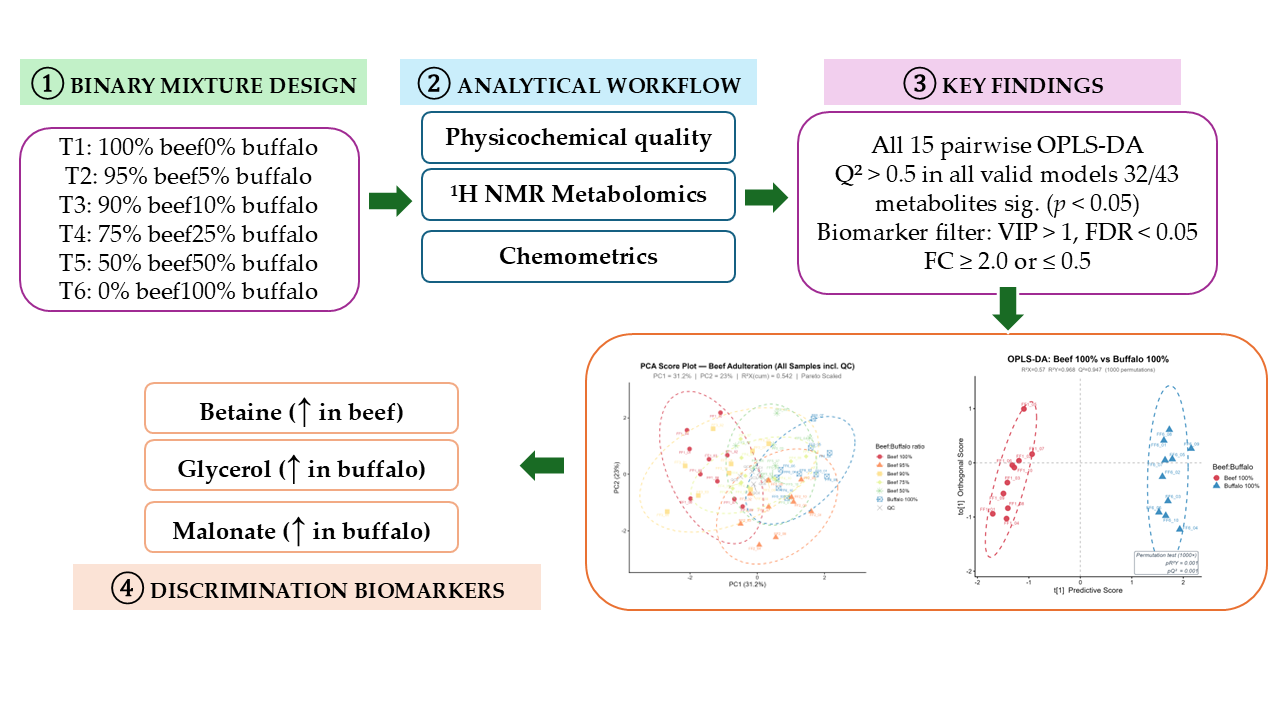
Adulteration of beef (Bos indicus) with buffalo meat (Bubalus bubalis) is a common form of food fraud with economic and religious implications, but quantitatively detecting its presence in ground beef products is difficult. Ten replicates of each of six binary mixtures (100:0 to 0:100 % w/w) of ground beef and buffalo meat were characterized using untargeted 1H NMR metabolomics (43 metabolites after QC filtering), physicochemical measurements (pH, CIE L*a*b* color, water activity, and electronic nose), and proximate composition. Fifteen pairwise OPLS-DA models and a 1000-fold permutation test were performed for discrimination and biomarker identification. PCA explained 54.2% of the total variance, and the adulteration groups separated along the PC1 axis. All OPLS-DA models were statistically valid (R2Y = 0.738–0.981; Q2 = 0.532–0.961; pQ2 < 0.001), with no evidence of overfitting. Three metabolites met all three criteria (VIP > 1.0, FDR < 0.05, < FC > 2 or < 0.5) and had AUC = 1.00 in the internal data set: betaine (−82.6% in buffalo vs. beef), glycerol (+154.7%), and malonate (+656%). No individual biomarker exceeded the multi-criterion threshold at buffalo substitution levels less than10%. The selection of external discovery-phase candidates for beef authentication using NMR includes betaine, glycerol, and malonate.