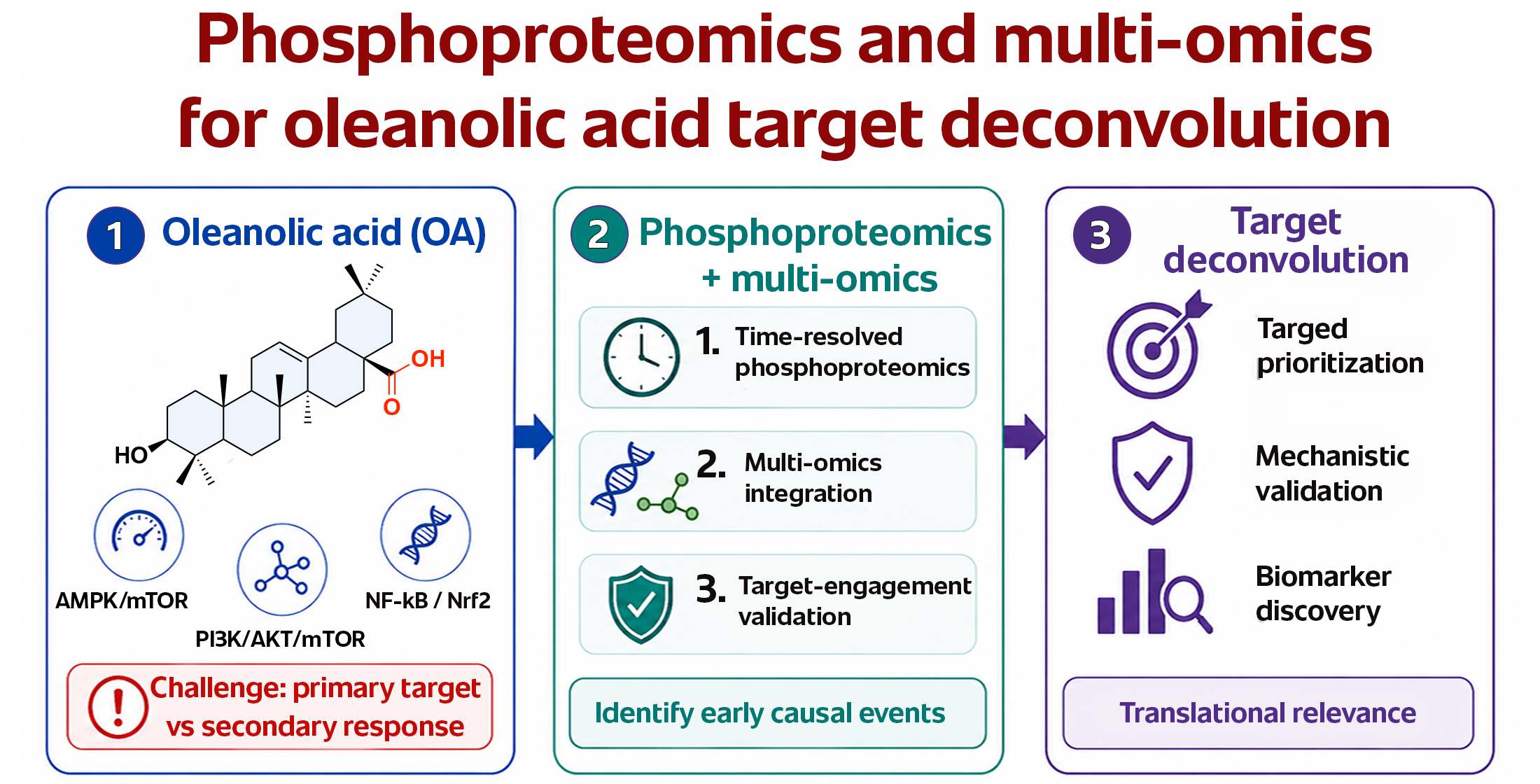
Oleanolic acid (OA) is a pentacyclic triterpenoid with broad biological activity, but its primary molecular points of engagement remain incompletely resolved. Most available studies describe OA through selected pathway markers, particularly within PI3K/AKT/mTOR, AMPK/mTOR, MAPK, NF-κB, and Nrf2 signaling, without clearly distinguishing direct target engagement from downstream adaptive responses. This limits mechanistic interpretation and weakens translational prioritization.
This review examines how phosphoproteomics and integrated multi-omics can support OA target deconvolution. We discuss why phosphoproteomics is particularly informative for capturing early signaling events, how it can be combined with proteomics, tran-scriptomics, metabolomics, and chemoproteomic approaches, and why orthogonal tar-get-engagement methods remain essential for stronger causal inference. We also organize the current signaling evidence for OA and its derivatives, highlighting the strongest support for AMPK/mTOR-linked regulation of autophagy and apoptosis while identi-fying major gaps in systems-level validation across other reported pathways.
Finally, we propose a stepwise workflow for OA target deconvolution based on time-resolved phosphoproteomics, analysis of informative phosphosite subsets, mul-ti-omics integration, kinase/phosphatase activity inference, and experimental target validation. This framework may help move OA research from descriptive pathway pharmacology toward mechanism-based target prioritization and more rational deriva-tive development.