Submitted:
03 May 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
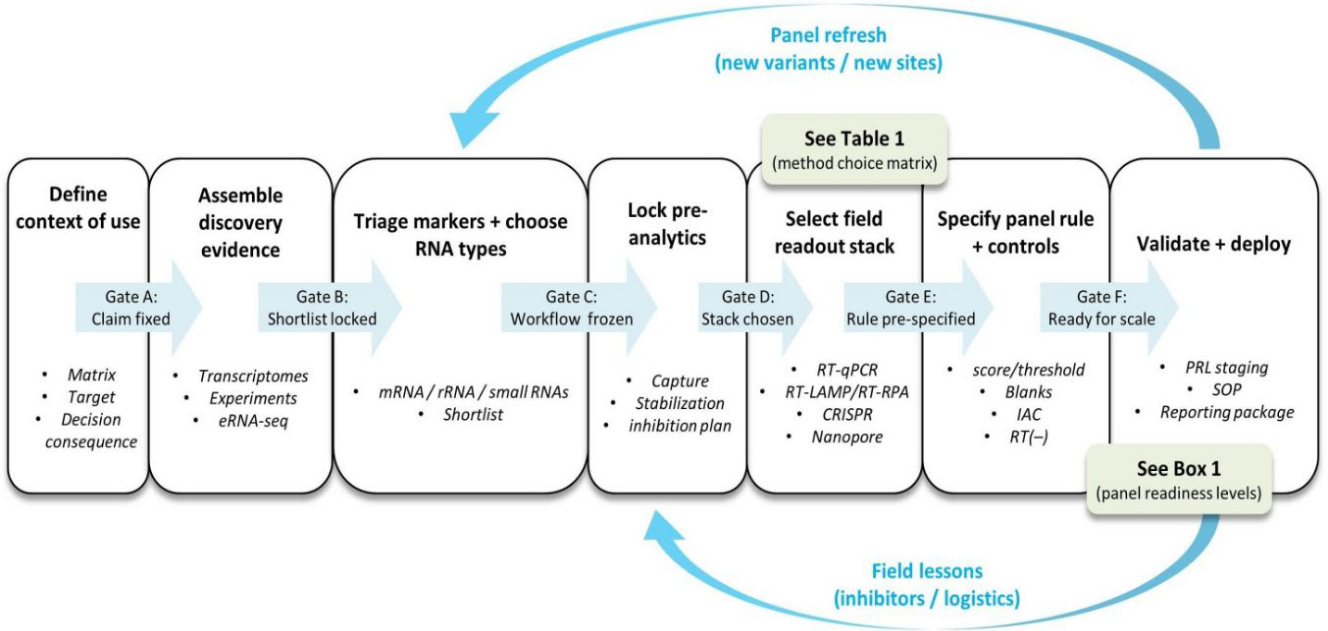
| Box 1 | Panel readiness levels (PRL) for field-ready eRNA panels |
|
PRL0: Concept and context fixed - Context of use, target taxa/community, matrix, and decision consequence are stated. - Claim type is pre-registered (presence; recent activity; phenology; condition). PRL1: Candidate markers and assay blueprint - Candidate targets are justified from discovery data or prior biology. - In-silico specificity is documented against local/non-target references where available. - Assay blueprint is drafted (targets, controls, scoring rule concept). PRL2: Analytical validation on reference material - Assay performance is established on reference templates (LOD/LOQ as relevant, repeatability). - RNA/DNA discrimination is demonstrated where needed (RT(–) and DNase logic). - Core negative/positive controls are defined and shown to behave as intended. PRL3: Matrix resilience and pre-analytics locked - Performance is demonstrated in real extracts or matrix spikes (inhibition frequency, mitigation rule). - Capture, stabilization, and extraction steps are fixed as an SOP (with time-to-stabilization limits). - Contamination-resilience measures are implemented (closed-tube preference; carryover prevention plan). PRL4: Field pilot with independent confirmation - Field detections are replicated across sites/days and compared to an independent line of evidence (survey data, lab confirmatory assay, or orthogonal molecular method). - The scoring rule is finalized (thresholds, replicate concordance rule, “indeterminate” category). PRL5: Transferability and operational QA - Reproducibility is demonstrated across operators (and ideally labs) using the same SOP. - Run-level acceptance criteria are enforced (blank behavior, IAC pass/fail, replicate rules). - Reporting package is complete (metadata + controls + analysis provenance). PRL6: Program integration and lifecycle management - Decision triggers and escalation pathway are codified (screen → confirm → action). - Drift is monitored (seasonal confounders; reagent lots; instrument checks). - “Panel refresh” is scheduled (sequence verification / local variant checks when needed). |
2. What is a Field-Ready eRNA Panel?
3. From Discovery to Markers: Choosing Targets and RNA Types
4. Field Sampling for eRNA: Capture, Stabilization, and Inhibitors
5. Field Readout Methods: Amplification, Detection, and Multiplexing
6. Decision-Grade Panels: Controls, Scoring, and Validation
7. Conclusions
Author Contributions
Data Availability Statement
Acknowledgments
References
- Ahi, E. P. (2025). eRNA-Min: Minimum Information Standard for Environmental RNA Reporting. [CrossRef]
- Ahi, E.P.; Khorshid, M. Potentials of RNA biosensors in developmental biology. Dev. Biol. 2025, 526, 173–188. [CrossRef]
- Ahi, E.P.; Schenekar, T. The Promise of Environmental RNA Research Beyond mRNA. Mol. Ecol. 2025, 34, e17787. [CrossRef]
- Alm, E.W.; Zheng, D.; Raskin, L. The Presence of Humic Substances and DNA in RNA Extracts Affects Hybridization Results. Appl. Environ. Microbiol. 2000, 66, 4547–4554. [CrossRef]
- Aminaka, Y.; Wong, M.K.-S.; Yada, T.; Hyodo, S. The use of environmental RNA for inferring fish spawning behavior. Sci. Rep. 2025, 15, 1–14. [CrossRef]
- The External RNA Controls Consortium The External RNA Controls Consortium: a progress report. Nat. Methods 2005, 2, 731–734. [CrossRef]
- Bohmann, K.; Evans, A.; Gilbert, M.T.P.; Carvalho, G.R.; Creer, S.; Knapp, M.; Yu, D.W.; de Bruyn, M. Environmental DNA for wildlife biology and biodiversity monitoring. Trends Ecol. Evol. 2014, 29, 358–367. [CrossRef]
- Bowers, H.A.; Pochon, X.; von Ammon, U.; Gemmell, N.; Stanton, J.-A.L.; Jeunen, G.-J.; Sherman, C.D.H.; Zaiko, A. Towards the Optimization of eDNA/eRNA Sampling Technologies for Marine Biosecurity Surveillance. Water 2021, 13, 1113. [CrossRef]
- Brandão-Dias, P.F.; Shaffer, M.; Guri, G.; Parsons, K.M.; Kelly, R.P.; Allan, E.A. Differential decay of multiple environmental nucleic acid components. Sci. Rep. 2025, 15, 1–13. [CrossRef]
- A Bustin, S.; Ruijter, J.M.; Hoff, M.J.B.v.D.; Kubista, M.; Pfaffl, M.W.; Shipley, G.L.; Tran, N.; Rödiger, S.; Untergasser, A.; Mueller, R.; et al. MIQE 2.0: Revision of the Minimum Information for Publication of Quantitative Real-Time PCR Experiments Guidelines. Clin. Chem. 2025, 71, 634–651. [CrossRef]
- Buxton, A.; Matechou, E.; Griffin, J.; Diana, A.; Griffiths, R.A. Optimising sampling and analysis protocols in environmental DNA studies. Sci. Rep. 2021, 11, 1–10. [CrossRef]
- Carrascosa, L.G.; Huertas, C.S.; Lechuga, L.M. Prospects of optical biosensors for emerging label-free RNA analysis. TrAC Trends Anal. Chem. 2016, 80, 177–189. [CrossRef]
- Çevik, T.; Çevik, N. Environmental DNA (eDNA): A review of ecosystem biodiversity detection and applications. Biodivers. Conserv. 2025, 34, 2999–3035. [CrossRef]
- Cristescu, M.E. Can Environmental RNA Revolutionize Biodiversity Science?. Trends Ecol. Evol. 2019, 34, 694–697. [CrossRef]
- Delica, A.; Nazarov, M.A.; De La Franier, B.; Thompson, M. Antifouling Modification of Gold Surfaces for Acoustic Wave Sensor Applications. Biosensors 2025, 15, 343. [CrossRef]
- Diana, A.; Matechou, E.; Griffin, J.E.; Buxton, A.S.; Griffiths, R.A. An RShiny app for modelling environmental DNA data: accounting for false positive and false negative observation error. Ecography 2021, 44, 1838–1844. [CrossRef]
- Doi, H.; Watanabe, T.; Nishizawa, N.; Saito, T.; Nagata, H.; Kameda, Y.; Maki, N.; Ikeda, K.; Fukuzawa, T. On-site environmental DNA detection of species using ultrarapid mobile PCR. Mol. Ecol. Resour. 2021, 21, 2364–2368. [CrossRef]
- Donia, A.; Shahid, M.F.; Hassan, S.-U.; Shahid, R.; Ahmad, A.; Javed, A.; Nawaz, M.; Yaqub, T.; Bokhari, H. Integration of RT-LAMP and Microfluidic Technology for Detection of SARS-CoV-2 in Wastewater as an Advanced Point-of-Care Platform. Food Environ. Virol. 2022, 14, 364–373. [CrossRef]
- Durán-Vinet, B.; Stanton, J.-A.L.; Jeunen, G.-J.; Pochon, X.; Zaiko, A.; Gemmell, N.J. CRISPR as a next-generation environmental biosurveillance tool for air, land, and water. Trends Biotechnol. 2025. [CrossRef]
- A Farrell, J.; Whitmore, L.; Duffy, D.J. The Promise and Pitfalls of Environmental DNA and RNA Approaches for the Monitoring of Human and Animal Pathogens from Aquatic Sources. BioScience 2021, 71, 609–625. [CrossRef]
- Feng, W.; Peng, H.; Xu, J.; Liu, Y.; Pabbaraju, K.; Tipples, G.; Joyce, M.A.; Saffran, H.A.; Tyrrell, D.L.; Babiuk, S.; et al. Integrating Reverse Transcription Recombinase Polymerase Amplification with CRISPR Technology for the One-Tube Assay of RNA. Anal. Chem. 2021, 93, 12808–12816. [CrossRef]
- Giroux, M.S.; Reichman, J.R.; Langknecht, T.; Burgess, R.M.; Ho, K.T. Environmental RNA as a Tool for Marine Community Biodiversity Assessments. Sci. Rep. 2022, 12, 1–13. [CrossRef]
- Goldberg, C.S.; Turner, C.R.; Deiner, K.; Klymus, K.E.; Thomsen, P.F.; Murphy, M.A.; Spear, S.F.; McKee, A.; Oyler-McCance, S.J.; Cornman, R.S.; et al. Critical considerations for the application of environmental DNA methods to detect aquatic species. Methods Ecol. Evol. 2016, 7, 1299–1307. [CrossRef]
- Gou, X.; Liu, X.; Su, X.; Ji, H.; Wang, Q.; Zhang, X. Water Environmental RNA Reveals Dose-Dependent Toxicant Responses in Fish. Environ. Sci. Technol. 2025, 59, 24717–24727. [CrossRef]
- Gygax, D.; Ramirez, S.; Chibesa, M.; Simpamba, T.; Riffel, M.; Riffel, T.; Srivathsan, A.; Nijland, R.; Urban, L. Evaluation of nanopore sequencing for increasing accessibility of eDNA studies in biodiverse countries. PLOS ONE 2025, 20, e0333994. [CrossRef]
- Hartle-Mougiou, K.; Gubili, C.; Xanthopoulou, P.; Kasapidis, P.; Valiadi, M.; Gizeli, E. Development of a quantitative colorimetric LAMP assay for fast and targeted molecular detection of the invasive lionfish Pterois miles from environmental DNA. Front. Mar. Sci. 2024, 11, 1358793. [CrossRef]
- Hayes, E.K.; Gouthro, M.T.; Gagnon, G.A. Isothermal amplification as a water safety tool: rapid detection of viruses in surface water and wastewater. Environ. Sci. Water Res. Technol. 2025, 11, 2141–2151. [CrossRef]
- He, X.; Maruki, T.; Morgado-Gamero, W.B.; Barrett, R.D.H.; Fugère, V.; Fussmann, G.F.; Gonzalez, A.; Shapiro, B.J.; Cristescu, M.E. Environmental RNA -Based Metatranscriptomics as a Novel Biomonitoring Tool: A Case Study of Glyphosate-Based Herbicide Effects on Freshwater Eukaryotic Communities. Mol. Ecol. 2025, 34, e70164. [CrossRef]
- Hechler, R.M.; Yates, M.C.; Chain, F.J.J.; Cristescu, M.E. Environmental transcriptomics under heat stress: Can environmental RNA reveal changes in gene expression of aquatic organisms?. Mol. Ecol. 2023, 34. [CrossRef]
- Hiki, K.; Yamagishi, T.; Yamamoto, H. Environmental RNA as a Noninvasive Tool for Assessing Toxic Effects in Fish: A Proof-of-concept Study Using Japanese Medaka Exposed to Pyrene. Environ. Sci. Technol. 2023, 57, 12654–12662. [CrossRef]
- Hsieh, K.; Mage, P.L.; Csordas, A.T.; Eisenstein, M.; Soh, H.T. Simultaneous elimination of carryover contamination and detection of DNA with uracil-DNA-glycosylase-supplemented loop-mediated isothermal amplification (UDG-LAMP). Chem. Commun. 2014, 50, 3747–3749. [CrossRef]
- Hur, Y.; Han, J.; Seon, J.; Pak, Y.E.; Roh, Y. Development of an SH-SAW sensor for the detection of DNA hybridization. Sensors Actuators A: Phys. 2005, 120, 462–467. [CrossRef]
- Hwang, M.T.; Heiranian, M.; Kim, Y.; You, S.; Leem, J.; Taqieddin, A.; Faramarzi, V.; Jing, Y.; Park, I.; van der Zande, A.M.; et al. Ultrasensitive detection of nucleic acids using deformed graphene channel field effect biosensors. Nat. Commun. 2020, 11, 1–11. [CrossRef]
- Ikert, H.; Lynch, M.D.J.; Doxey, A.C.; Giesy, J.P.; Servos, M.R.; Katzenback, B.A.; Craig, P.M. High Throughput Sequencing of MicroRNA in Rainbow Trout Plasma, Mucus, and Surrounding Water Following Acute Stress. Front. Physiol. 2021, 11. [CrossRef]
- Jain, U.; Chauhan, N.; Saxena, K. (2023). Fundamentals of sensors and biosensors: An overview. Multifaceted Bio-Sensing Technology: Bioelectrochemical Systems: The Way Forward: Volume IV, 4, 31–44. [CrossRef]
- Jiang, W.; Zhu, T.; Zhou, S.; Pan, L.; Qiao, Z.; Wang, M.; Yang, D. Recent advances in electrochemical-based CRISPR/Cas biosensing for nucleic acid and non-nucleic acid pathogenic microorganism detection. Food Res. Int. 2025, 221, 117213. [CrossRef]
- Jo, T.S. Methodological considerations for aqueous environmental RNA collection, preservation, and extraction. Anal. Sci. 2023, 39, 1711–1718. [CrossRef]
- Jo, T.; Tsuri, K.; Hirohara, T.; Yamanaka, H. Warm temperature and alkaline conditions accelerate environmental RNA degradation. Environ. DNA 2022, 5, 836–848. [CrossRef]
- Kageyama, S.A.; Hoogland, M.R.; Tajjioui, T.; Schreier, T.M.; Erickson, R.A.; Merkes, C.M. Validation of a Portable eDNA Detection Kit for Invasive Carps. Fishes 2022, 7, 363. [CrossRef]
- Kagzi, K.; Hechler, R.M.; Fussmann, G.F.; Cristescu, M.E. Environmental RNA degrades more rapidly than environmental DNA across a broad range of pH conditions. Mol. Ecol. Resour. 2022, 22, 2640–2650. [CrossRef]
- Keneally, C.; Gaget, V.; Kidd, S.P.; Brookes, J.D. Sample Preservation Solution Increases Nucleic Acid Yield and Environmental RNA Quality in Sediments Across an Estuarine Salinity Gradient. Environ. DNA 2024, 6. [CrossRef]
- Khan, M.U.; Rahman, M.; Zahan, N.; Masud, M.K.; Sarker, S.; Haque, H. Electrochemical Biosensing for Antibiotic-Resistant Bacteria: Advances, Challenges, and Future Directions. Micromachines 2025, 16, 986. [CrossRef]
- Khorshid, M.; Ahi, E.P. RNA Aptamers and Epitranscriptomics: Charting Unexplored Territories in RNA Biology. Mol. Diagn. Ther. 2026, 1–20. [CrossRef]
- Kim, K.; Kachiprath, B.; Yeo, I.; Shim, K.; Kwak, I.; Jeong, C. Environmental DNA -based RPA - CRISPR /Cas12a assay for on-site detection of chironomid larvae in aquatic environments. Pest Manag. Sci. 2025, 82, 2458–2467. [CrossRef]
- King, C.E.; Debruyne, R.; Kuch, M.; Schwarz, C.; Poinar, H.N. A Quantitative Approach to Detect and Overcome PCR Inhibition in Ancient DNA Extracts. BioTechniques 2009, 47, 941–949. [CrossRef]
- Kinyua, D.M.; Memeu, D.M.; Mwenda, C.N.M.; Della Ventura, B.; Velotta, R. Advancements and Applications of Lateral Flow Assays (LFAs): A Comprehensive Review. Sensors 2025, 25, 5414. [CrossRef]
- Klymus, K.E.; Baker, J.D.; Abbott, C.L.; Brown, R.J.; Craine, J.M.; Gold, Z.; Hunter, M.E.; Johnson, M.D.; Jones, D.N.; Jungbluth, M.J.; et al. The MIEM guidelines: Minimum information for reporting of environmental metabarcoding data. Metabarcoding Metagenomics 2024, 8, e128689–518. [CrossRef]
- Koczula, K.M.; Gallotta, A. Lateral flow assays. Essays Biochem. 2016, 60, 111–120. [CrossRef]
- Leugger, F.; Lüthi, M.; Schmidlin, M.; Kontarakis, Z.; Pellissier, L. Rapid field-based detection of a threatened and elusive species with environmental DNA and CRISPR-Dx. Glob. Ecol. Conserv. 2025, 59. [CrossRef]
- Lim, N.Y.N.; Roco, C.A.; Frostegård, Å. Transparent DNA/RNA Co-extraction Workflow Protocol Suitable for Inhibitor-Rich Environmental Samples That Focuses on Complete DNA Removal for Transcriptomic Analyses. Front. Microbiol. 2016, 7, 1588. [CrossRef]
- Linzner, N.; Bartel, A.; Schumacher, V.; Grau, J.H.; Wyler, E.; Preuß, H.; Garske, S.; Bitzegeio, J.; Kirst, E.B.; Liere, K.; et al. Effective Inhibitor Removal from Wastewater Samples Increases Sensitivity of RT-dPCR and Sequencing Analyses and Enhances the Stability of Wastewater-Based Surveillance. Microorganisms 2024, 12, 2475. [CrossRef]
- Lobato, I.M.; O'SUllivan, C.K. Recombinase polymerase amplification: Basics, applications and recent advances. TrAC Trends Anal. Chem. 2018, 98, 19–35. [CrossRef]
- Longo, M.C.; Berninger, M.S.; Hartley, J.L. Use of uracil DNA glycosylase to control carry-over contamination in polymerase chain reactions. Gene 1990, 93, 125–128. [CrossRef]
- Macher, T.-H.; Arle, J.; Beermann, A.J.; Frank, L.; Hupało, K.; Koschorreck, J.; Schütz, R.; Leese, F. Is it worth the extra mile? Comparing environmental DNA and RNA metabarcoding for vertebrate and invertebrate biodiversity surveys in a lowland stream. PeerJ 2024, 12, e18016. [CrossRef]
- Maggini, S.; Jacobsen, M.W.; Urban, P.; Hansen, B.K.; Kielgast, J.; Bekkevold, D.; Jardim, E.; Martinsohn, J.T.; Carvalho, G.R.; Nielsen, E.E.; et al. Nanopore environmental DNA sequencing of catch water for estimating species composition in demersal bottom trawl fisheries. Environ. DNA 2024, 6. [CrossRef]
- Mannelli, I.; Minunni, M.; Tombelli, S.; Mascini, M. Quartz crystal microbalance (QCM) affinity biosensor for genetically modified organisms (GMOs) detection. Biosens. Bioelectron. 2003, 18, 129–140. [CrossRef]
- Marshall, N.T.; Vanderploeg, H.A.; Chaganti, S.R. Environmental (e)RNA advances the reliability of eDNA by predicting its age. Sci. Rep. 2021, 11, 1–11. [CrossRef]
- Miyata, K.; Inoue, Y.; Yamane, M.; Honda, H. Fish environmental RNA sequencing sensitively captures accumulative stress responses through short-term aquarium sampling. Sci. Total. Environ. 2024, 959, 178182. [CrossRef]
- Nair, M.P.; Teo, A.J.T.; Li, K.H.H. Acoustic Biosensors and Microfluidic Devices in the Decennium: Principles and Applications. Micromachines 2021, 13, 24. [CrossRef]
- Parsley, M.B.; Goldberg, C.S. Environmental RNA can distinguish life stages in amphibian populations. Mol. Ecol. Resour. 2023, 24. [CrossRef]
- Pine, P.S.; Munro, S.A.; Parsons, J.R.; McDaniel, J.; Lucas, A.B.; Lozach, J.; Myers, T.G.; Su, Q.; Jacobs-Helber, S.M.; Salit, M. Evaluation of the External RNA Controls Consortium (ERCC) reference material using a modified Latin square design. BMC Biotechnol. 2016, 16, 1–15. [CrossRef]
- Pochon, X.; Bowers, H.A.; Zaiko, A.; Wood, S.A. Advancing the environmental DNA and RNA toolkit for aquatic ecosystem monitoring and management. PeerJ 2025, 13, e19119. [CrossRef]
- Ramirez-Priego, P.; Mauriz, E.; Giarola, J.F.; Lechuga, L.M. Overcoming challenges in plasmonic biosensors deployment for clinical and biomedical applications: A systematic review and meta-analysis. Sens. Bio-Sensing Res. 2024, 46. [CrossRef]
- Rishan, S.T.; Kline, R.J.; Rahman, S. New prospects of environmental RNA metabarcoding research in biological diversity, ecotoxicological monitoring, and detection of COVID-19: a critical review. Environ. Sci. Pollut. Res. 2024, 31, 11406–11427. [CrossRef]
- Rohatensky, M.G.; Livingstone, D.M.; Mintchev, P.; Barnes, H.K.; Nakoneshny, S.C.; Demetrick, D.J.; Dort, J.C.; van Marle, G. Assessing the performance of a Loop Mediated Isothermal Amplification (LAMP) assay for the detection and subtyping of high-risk suptypes of Human Papilloma Virus (HPV) for Oropharyngeal Squamous Cell Carcinoma (OPSCC) without DNA purification. BMC Cancer 2018, 18, 166. [CrossRef]
- Sayad, A.A.; Ibrahim, F.; Uddin, S.M.; Pei, K.X.; Mohktar, M.S.; Madou, M.; Thong, K.L. A microfluidic lab-on-a-disc integrated loop mediated isothermal amplification for foodborne pathogen detection. Sensors Actuators B: Chem. 2016, 227, 600–609. [CrossRef]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors - occurrence, properties and removal. J. Appl. Microbiol. 2012, 113, 1014–1026. [CrossRef]
- Seimon, T.A.; Van Long, N.; Le, M.; McCormack, T.E.M.; Nguyen, T.T.; Ngo, H.; Thang, N.T.; Hoang, T.; Platt, S.G.; Van Ha, H.; et al. Development and Application of a Portable Environmental DNA Test for the Detection of Rafetus swinhoei in Viet Nam. Environ. DNA 2024, 6. [CrossRef]
- Sepulveda, A.J.; Hutchins, P.R.; Forstchen, M.; Mckeefry, M.N.; Swigris, A.M. The Elephant in the Lab (and Field): Contamination in Aquatic Environmental DNA Studies. Front. Ecol. Evol. 2020, 8. [CrossRef]
- Shi, C.; Yu, Z.; Tan, H.; Li, W.; Wang, Y.; Wang, Y.; Zhang, Q.; Man, Y. One-Pot CRISPR-Based Isothermal Amplification for Nucleic Acid Detection: A Comparative Review of Different Strategies. ACS Sensors 2025, 10, 9108–9134. [CrossRef]
- Shi, Y.; Xu, M.; Duan, X.; Li, S.; Ding, J.-W.; Chen, L. WarmStart colorimetric loop-mediated isothermal amplification for the one-tube, contamination-free and visualization detection of Shigella flexneri. Int. J. Infect. Dis. 2021, 112, 55–62. [CrossRef]
- Shu, L.; Ludwig, A.; Peng, Z. Standards for Methods Utilizing Environmental DNA for Detection of Fish Species. Genes 2020, 11, 296. [CrossRef]
- Stevens, J.D.; Parsley, M.B. Environmental RNA applications and their associated gene targets for management and conservation. Environ. DNA 2022, 5, 227–239. [CrossRef]
- Suhag, D. (2025). Electrochemical Biosensors: Principles and Mechanisms. In Biomimetic Sensor Technologies (pp. 79–103). Springer, Singapore. [CrossRef]
- Thalinger, B.; Deiner, K.; Harper, L.R.; Rees, H.C.; Blackman, R.C.; Sint, D.; Traugott, M.; Goldberg, C.S.; Bruce, K. A validation scale to determine the readiness of environmental DNA assays for routine species monitoring. Environ. DNA 2021, 3, 823–836. [CrossRef]
- Tian, M.; Qiao, M.; Shen, C.; Meng, F.; Frank, L.A.; Krasitskaya, V.V.; Wang, T.; Zhang, X.; Song, R.; Li, Y.; et al. Highly-sensitive graphene field effect transistor biosensor using PNA and DNA probes for RNA detection. Appl. Surf. Sci. 2020, 527. [CrossRef]
- Tingley, R.; Coleman, R.; Gecse, N.; van Rooyen, A.; Weeks, A.R. Accounting for false positive detections in occupancy studies based on environmental DNA: A case study of a threatened freshwater fish (Galaxiella pusilla). Environ. DNA 2020, 3, 388–397. [CrossRef]
- van Doorn, R.; Klerks, M.M.; van Gent-Pelzer, M.P.E.; Speksnijder, A.G.C.L.; Kowalchuk, G.A.; Schoen, C.D. Accurate Quantification of Microorganisms in PCR-Inhibiting Environmental DNA Extracts by a Novel Internal Amplification Control Approach Using Biotrove OpenArrays. Appl. Environ. Microbiol. 2009, 75, 7253–7260. [CrossRef]
- Varzandi, A.R.; Reska, T.; Urban, L.; Zanet, S.; Ferroglio, E. Environmental screening through nanopore native sequencing leads to the detection of Batrachochytrium dendrobatidis in La Mandria Regional Park, Italy. Glob. Ecol. Conserv. 2025, 59. [CrossRef]
- Wang, F.; Xiong, W.; Huang, X.; Zhan, A. Influence of short-term water sample storage on environmental RNA metabarcoding-based biodiversity assessment. J. Environ. Sci. 2025. [CrossRef]
- The dMIQE Group; Huggett, J.F. The Digital MIQE Guidelines Update: Minimum Information for Publication of Quantitative Digital PCR Experiments for 2020. Clin. Chem. 2020, 66, 1012–1029. [CrossRef]
- Williams, M.A.; Briciu-Burghina, C.; Power, S.; O’gRady, J.; de Eyto, E.; Kent, N.; Parle-McDermott, A.; Regan, F. SensEDNA: An innovative optical CRISPR platform for rapid environmental DNA monitoring. Talanta Open 2025, 12. [CrossRef]
- Wu, J.; Liu, H.; Chen, W.; Ma, B.; Ju, H. Device integration of electrochemical biosensors. Nat. Rev. Bioeng. 2023, 1, 346–360. [CrossRef]
- Wu, Y.; Wang, G.; Yu, X.; Fan, Y.; Chen, X.; Liu, S. Label-Free DNA Hybridization Detection Using a Highly Sensitive Fiber Microcavity Biosensor. Sensors 2024, 24, 278. [CrossRef]
- Xu, X.; Li, G.; Xue, L.; Dong, S.; Luo, J.; Cao, Z. Microfluidic devices integrated with plasmonic nanostructures for sensitive fluorescent immunoassays. Biomicrofluidics 2024, 18, 011303. [CrossRef]
- Xu, Z.; Asakawa, S. Release and degradation of dissolved environmental RNAs from zebrafish cells. RNA Biol. 2025, 22, 1–12. [CrossRef]
- Xue, M.; Gonzalez, D.H.; Osikpa, E.; Gao, X.; Lillehoj, P.B. Rapid and automated interpretation of CRISPR-Cas13-based lateral flow assay test results using machine learning. Sensors Diagn. 2024, 4, 171–181. [CrossRef]
- Xue, Z.; Tian, W.; Han, Y.; Li, S.; Guo, J.; He, H.; Yu, P.; Zhang, W. Environmental RNA metabarcoding for ballast water microbial diversity: Minimizing false positives. Sci. Total. Environ. 2024, 955, 176902. [CrossRef]
- Yang, J.; Matsushita, S.; Xia, F.; Yoshizawa, S.; Iwasaki, W. Rapid, easy, sensitive, low-cost and on-site detection of environmental DNA and RNA using CRISPR-Cas13. Methods Ecol. Evol. 2024, 15, 1408–1421. [CrossRef]
- Yang, Y.; Wang, F.; Xue, B.; Zhou, X. Field-deployable assay based on CRISPR-Cas13a coupled with RT-RPA in one tube for the detection of SARS-CoV-2 in wastewater. J. Hazard. Mater. 2023, 459, 132077. [CrossRef]
- Yates, M.C.; Derry, A.M.; Cristescu, M.E. Environmental RNA: A Revolution in Ecological Resolution?. Trends Ecol. Evol. 2021, 36, 601–609. [CrossRef]
- Zakiyyah, S.N.; Irkham; Sima, K.S.P.; Kharismasari, C.Y.; Xi, M.; Gaffar, S.; Ozsoz, M.; Paolucci, F.; Valenti, G.; Hartati, Y.W. CRISPR-Cas13a-powered electrochemical biosensors for RNA-based disease diagnostic and monitoring. Sensors Actuators Rep. 2025, 10. [CrossRef]
- Zarrabian, M.; Sherif, S.M. Overcoming soil matrix interference for reliable environmental monitoring of dsRNA. Environ. Technol. Innov. 2025, 40. [CrossRef]
- Zhang, B.; Li, X.; Huang, M.; Jiang, M.; Du, L.; Yin, P.; Fang, X.; Jiang, X.; Qi, F.; Lin, Y.; et al. A Portable Dual-Mode Microfluidic Device Integrating RT-qPCR and RT-LAMP for Rapid Nucleic Acid Detection in Point-of-Care Testing. Biosensors 2026, 16, 51. [CrossRef]
- Zhang, Y.; Qiu, Y.; Liu, K.; Zhong, W.; Yang, J.; Altermatt, F.; Zhang, X. Evaluating eDNA and eRNA metabarcoding for aquatic biodiversity assessment: From bacteria to vertebrates. Environ. Sci. Ecotechnology 2024, 21, 100441. [CrossRef]
- Zhong, K.; Khorshid, M.; Li, J.; Markey, K.; Wagner, P.H.; Song, K.; Van Cleuvenbergen, S.; Clays, K. Fabrication of optomicrofluidics for real-time bioassays based on hollow sphere colloidal photonic crystals with wettability patterns. J. Mater. Chem. C 2016, 4, 7853–7858. [CrossRef]
- Zhu, L.; Gao, Y.; Shen, H.; Yang, Y.; Yuan, L. A Quartz Crystal Microbalance (QCM) Study of Single-Strand DNA Hybridization and Hydrolytic Cleavage. J. Anal. Chem. 2005, 60, 780–783. [CrossRef]
- Zou, N.; Wang, S.; Qiu, W.; Kong, W.; Wang, G.; Wang, S. Environmental RNA as a transformative tool for aquatic ecosystem health assessment: progress and challenges. Ecol. Indic. 2025, 180. [CrossRef]
| Readout option (typical stack) | Typical time | Minimum field setup | Realistic panel scale | Output | Best fit in workflow | Key constraints to plan for | Ref. |
| portable RT-qPCR / POD qPCR (one-step RT-qPCR) | 45–120 min | portable qPCR + pipettes; cold chain or lyophilized reagents | small multiplex (≈4–6) or parallel singleplex | quantitative / semi-quant | confirmatory testing; decision thresholds | inhibition; RT(–)/blank discipline; RNase control; logistics | (Bustin et al., 2025; Doi et al., 2021; Seimon et al., 2024) |
| lab RT-dPCR / ddPCR | same day (near-lab) | benchtop dPCR system | singleplex to modest multiplex | high-confidence quant | adjudicate borderline results; enforcement-grade confirmation | not field-portable; transport/turnaround | (Whale et al., 2020) |
| RT-LAMP (colorimetric/ fluor) |
20–45 min | heat block; closed tubes; optional simple reader | parallel singleplex; very small multiplex | qualitative / semi-quant | rapid screen; low-instrument settings | spurious amp/primer crosstalk; carryover risk; inhibitors; replicate rules | (Hartle-Mougiou et al., 2024; Hayes et al., 2025) |
| RT-RPA / RT-RAA (+ lateral flow/fluor) | 15–30 min | 37–42 °C incubator; strips or small reader | parallel singleplex; small multiplex (fragile) | qualitative / semi-quant | rapid screen; battery-friendly workflows | design-sensitive; nonspecific amp if poorly optimized; reagent ecosystem varies | (Feng et al., 2021; Lobato & O’Sullivan, 2018) |
| isothermal + CRISPR (RT-RPA/LAMP → Cas12/13; one-tube when possible) | 30–75 min | incubator; lateral flow or compact optics | low-plex; staged testing works well | high-specificity qual / semi-quant | specificity upgrade where false positives are costly | integration complexity; inhibition near LOD; contamination control still needed | (Leugger et al., 2025; Williams et al., 2025; J. Yang et al., 2024) |
| portable nanopore sequencing (amplicon or native) | 6–24 h | nanopore + laptop + power; basic wet lab | broad (sequence-based) | sequence confirmation / discovery | on-site verification; panel refresh; unexpected taxa/variants | operator skill; bioinformatics; sensitivity may lag targeted assays at very low template | (Gygax et al., 2025; Maggini et al., 2024; Varzandi et al., 2025) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




