Submitted:
30 April 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Association of microRNA Expression in the Development and Progression of Cervical Cancer
3. The Regulation of miRNA Expression by High-Risk HPV Viral Oncoproteins
4. MicroRNAs Associated with Signaling Pathways in HPV Infection
5. Clinical Significance of Deregulation of microRNA Expression in Cervical Cancer
6. Computational Strategies for the Identification of Differentially Expressed microRNAs in Cervical Cancer
6.1. Computational Approaches for the Identification of de-miRNAs
6.1.1. Differential Expression Analysis
6.1.2. Integration of Multi-Omics Data
6.1.3. Application of Machine Learning Algorithms
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AKT | Protein kinase B |
| AMER1 | APC membrane recruitment protein 1 |
| ANNs | Artificial Neural Networks |
| AXIN2 | Axis Inhibition Protein 2 |
| CC | Cervical Cancer |
| CDC42 | Cell division control protein 42 homolog |
| CIN | Cervical intraepithelial neoplasia |
| c-MET | C-Mesenchymal-Epithelial Transition Factor |
| CTNNB1 | Catenin beta 1 |
| DE-miRNAs | Differentially expressed miRNAs |
| DGCR8 | DiGeorge Critical Region 8 |
| DGE | Differential gene expression |
| DISC | Death-Inducing Signaling Complex |
| DLL1 | Delta-like notch 1 ligand |
| EGF | Epidermal Growth Factor |
| EGFR | Epidermal Growth Factor Receptor |
| EMT | Epithelial-mesenchymal transition |
| ERK | Extracellular signal-regulated kinase |
| FAK | Focal Adhesion Kinase |
| GEO | Gene Expression Omnibus |
| GLI3 | GLI family zinc finger 3 |
| HIF-1α | Hypoxia-Inducible Factor 1-alpha |
| HPV | Human Papillomavirus |
| hr-HPV | High-risk Human Papillomavirus |
| IL-6 | Interleukin-6 |
| KDM5B | Lysine demethylase 5B |
| lr-HPV | Low-risk Human Papillomavirus |
| MAPK1 | Mitogen-Activated Protein Kinase 1 |
| MHC I | Major Histocompatibility Complex class I |
| miRNAs | MicroRNAs |
| ML | Machine learning |
| MMP2 | Matrix Metalloproteinase-2 |
| MMP9 | Matrix Metalloproteinase-9 |
| MST2 | Mammalian sterile 20-like kinase 2 |
| NRP2 | Neuropilin 2 |
| OncomiRs | Oncogenic miRNAs |
| P18INK4c | p18 inhibitor of cyclin-dependent kinase 4c |
| PD-L1 | Programmed Death-Ligand 1 |
| PI3K | Phosphoinositide-3 kinase |
| PIK3CD | Phosphoinositide-3 kinase delta |
| PLK2 | Polo-like kinase2 |
| pRb | Retinoblastoma protein |
| PTEN | Phosphatase and Tensin Homolog |
| RHOA | Ras homolog gene family member A |
| RNA | Ribonucleic acid |
| RNA-seq | RNA sequencing |
| SP1 | Specificity Protein 1 |
| SVM | Random Forest and Support Vector Machines |
| TCGA | The Cancer Genome Atlas |
| TRIM3 | Tripartite motif-containing 3 |
| ts-miRs | Tumor-suppressor genes |
| UBE2I | Ubiquitin-conjugating enzyme E2I |
| uPA | Urokinase-type plasminogen activator |
| WNT10A | Wnt family member 10a |
| WTX | Wilms tumor gene on the X chromosome |
| ZEB1 | Zinc finger E-box binding homeobox 1 |
| ZNF813 | Zinc finger protein |
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2024, 74, 229–263. [Google Scholar] [CrossRef]
- Lima, F.; Oliveira, J.; Costa, L.; Martins, L.; Santos, M.; de Camargo Cancela, M.; Migowski, A.; Schilithz, A.; Corrêa, F.; Oliveira, I.; et al. Estimativa 2023 : Incidência de Câncer No Brasil; 2023; ISBN 978-65-88517-09-3. [Google Scholar]
- Pulliero, A.; Cassatella, G.; Astuni, P.; Khalid, Z.; Fiordoro, S.; Izzotti, A. The Role of microRNA Expression and DNA Methylation in HPV-Related Cervical Cancer: A Systematic Review. Int. J. Mol. Sci. 2024, 25, 12714. [Google Scholar] [CrossRef] [PubMed]
- Burd, E.M. Human Papillomavirus and Cervical Cancer. Clin. Microbiol. Rev. 2003, 16, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Mlynarczyk-Bonikowska, B.; Rudnicka, L. HPV Infections—Classification, Pathogenesis, and Potential New Therapies. Int. J. Mol. Sci. 2024, 25, 7616. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Cao, C.; Wu, P.; Huang, X.; Ma, D. Advances in Cervical Cancer: Current Insights and Future Directions. Cancer Commun. 2025, 45, 77–109. [Google Scholar] [CrossRef] [PubMed]
- Shang, R.; Lee, S.; Senavirathne, G.; Lai, E.C. microRNAs in Action: Biogenesis, Function and Regulation. Nat. Rev. Genet 2023, 24, 816–833. [Google Scholar] [CrossRef]
- Pietrykowska, H.; Sierocka, I.; Zielezinski, A.; Alisha, A.; Carrasco-Sanchez, J.C.; Jarmolowski, A.; Karlowski, W.M.; Szweykowska-Kulinska, Z. Biogenesis, Conservation, and Function of miRNA in Liverworts. J. Exp. Bot. 2022, 73, 4528–4545. [Google Scholar] [CrossRef]
- Vishnoi, A.; Rani, S. MiRNA Biogenesis and Regulation of Diseases: An Overview. Methods Mol. Biol. 2017, 1509, 1–10. [Google Scholar] [CrossRef]
- Banno, K.; Iida, M.; Yanokura, M.; Kisu, I.; Iwata, T.; Tominaga, E.; Tanaka, K.; Aoki, D. MicroRNA in Cervical Cancer: OncomiRs and Tumor Suppressor miRs in Diagnosis and Treatment. ScientificWorldJournal 2014, 2014, 178075. [Google Scholar] [CrossRef]
- Esquela-Kerscher, A.; Slack, F.J. Oncomirs - microRNAs with a Role in Cancer. Nat. Rev. Cancer 2006, 6, 259–269. [Google Scholar] [CrossRef]
- Ratti, M.; Lampis, A.; Ghidini, M.; Salati, M.; Mirchev, M.B.; Valeri, N.; Hahne, J.C. MicroRNAs (miRNAs) and Long Non-Coding RNAs (lncRNAs) as New Tools for Cancer Therapy: First Steps from Bench to Bedside. Targ. Oncol. 2020, 15, 261–278. [Google Scholar] [CrossRef] [PubMed]
- Ali Syeda, Z.; Langden, S.S.S.; Munkhzul, C.; Lee, M.; Song, S.J. Regulatory Mechanism of MicroRNA Expression in Cancer. Int. J. Mol. Sci. 2020, 21, 1723. [Google Scholar] [CrossRef]
- Samad, A.F.A.; Sajad, M.; Nazaruddin, N.; Fauzi, I.A.; Murad, A.M.A.; Zainal, Z.; Ismail, I. MicroRNA and Transcription Factor: Key Players in Plant Regulatory Network. Front Plant Sci. 2017, 8, 565. [Google Scholar] [CrossRef] [PubMed]
- Pajares, M.J.; Alemany-Cosme, E.; Goñi, S.; Bandres, E.; Palanca-Ballester, C.; Sandoval, J. Epigenetic Regulation of microRNAs in Cancer: Shortening the Distance from Bench to Bedside. Int. J. Mol. Sci. 2021, 22, 7350. [Google Scholar] [CrossRef]
- Arif, K.M.T.; Elliott, E.K.; Haupt, L.M.; Griffiths, L.R. Regulatory Mechanisms of Epigenetic miRNA Relationships in Human Cancer and Potential as Therapeutic Targets. Cancers 2020, 12, 2922. [Google Scholar] [CrossRef]
- Remenyi, J.; Bajan, S.; Fuller-Pace, F.V.; Arthur, J.S.C.; Hutvagner, G. The Loop Structure and the RNA Helicase P72/DDX17 Influence the Processing Efficiency of the Mice miR-132. Sci. Rep. 2016, 6, 22848. [Google Scholar] [CrossRef]
- Hashemi, M.; Khosroshahi, E.M.; Daneii, P.; Hassanpoor, A.; Eslami, M.; Koohpar, Z.K.; Asadi, S.; Zabihi, A.; Jamali, B.; Ghorbani, A.; et al. Emerging Roles of CircRNA-miRNA Networks in Cancer Development and Therapeutic Response. Non-Coding RNA Res. 2025, 10, 98–115. [Google Scholar] [CrossRef]
- Ross, S.A.; Davis, C.D. MicroRNA, Nutrition, and Cancer Prevention1. Adv. Nutr. 2011, 2, 472–485. [Google Scholar] [CrossRef]
- Tsang, J.; Zhu, J.; van Oudenaarden, A. MicroRNA-Mediated Feedback and Feedforward Loops Are Recurrent Network Motifs in Mammals. Mol. Cell 2007, 26, 753–767. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef]
- Causin, R.L.; de Freitas, A.J.A.; Trovo Hidalgo Filho, C.M.; dos Reis, R.; Reis, R.M.; Marques, M.M.C. A Systematic Review of MicroRNAs Involved in Cervical Cancer Progression. Cells 2021, 10, 668. [Google Scholar] [CrossRef]
- Otmani, K.; Rouas, R.; Berehab, M.; Lewalle, P. The Regulatory Mechanisms of oncomiRs in Cancer. Biomed. Pharmacother. 2024, 171, 116165. [Google Scholar] [CrossRef] [PubMed]
- Amin, F.A.S.; Un Naher, Z.; Ali, P.S.S. Molecular Markers Predicting the Progression and Prognosis of Human Papillomavirus-Induced Cervical Lesions to Cervical Cancer. J. Cancer Res. Clin. Oncol. 2023, 149, 8077–8086. [Google Scholar] [CrossRef] [PubMed]
- Pedroza-Torres, A.; López-Urrutia, E.; García-Castillo, V.; Jacobo-Herrera, N.; Herrera, L.A.; Peralta-Zaragoza, O.; López-Camarillo, C.; De Leon, D.C.; Fernández-Retana, J.; Cerna-Cortés, J.F.; et al. MicroRNAs in Cervical Cancer: Evidences for a miRNA Profile Deregulated by HPV and Its Impact on Radio-Resistance. Molecules 2014, 19, 6263–6281. [Google Scholar] [CrossRef] [PubMed]
- Abolfathi, H.; Arabi, M.; Sheikhpour, M. A Literature Review of microRNA and Gene Signaling Pathways Involved in the Apoptosis Pathway of Lung Cancer. Respir. Res. 2023, 24, 55. [Google Scholar] [CrossRef]
- Wang, J.; Chen, L. The Role of miRNAs in the Invasion and Metastasis of Cervical Cancer. Biosci. Rep. 2019, 39, BSR20181377. [Google Scholar] [CrossRef]
- Gu, Y.; Feng, X.; Jin, Y.; Liu, Y.; Zeng, L.; Zhou, D.; Feng, Y. Upregulation of miRNA-10a-5p Promotes Tumor Progression in Cervical Cancer by Suppressing UBE2I Signaling. J. Obstet. Gynaecol. 2023, 43, 2171283. [Google Scholar] [CrossRef]
- Passaro, A.; Al Bakir, M.; Hamilton, E.G.; Diehn, M.; André, F.; Roy-Chowdhuri, S.; Mountzios, G.; Wistuba, I.I.; Swanton, C.; Peters, S. Cancer Biomarkers: Emerging Trends and Clinical Implications for Personalized Treatment. Cell 2024, 187, 1617–1635. [Google Scholar] [CrossRef]
- Tommasino, M. The Human Papillomavirus Family and Its Role in Carcinogenesis. Semin. Cancer Biol. 2014, 26, 13–21. [Google Scholar] [CrossRef]
- Kabsch, K.; Alonso, A. The Human Papillomavirus Type 16 E5 Protein Impairs TRAIL- and FasL-Mediated Apoptosis in HaCaT Cells by Different Mechanisms. J. Virol. 2002, 76, 12162–12172. [Google Scholar] [CrossRef]
- Cortese, M.S.; Ashrafi, G.H.; Campo, M.S. All 4 Di-Leucine Motifs in the First Hydrophobic Domain of the E5 Oncoprotein of Human Papillomavirus Type 16 Are Essential for Surface MHC Class I Downregulation Activity and E5 Endomembrane Localization. Int. J. Cancer 2010, 126, 1675–1682. [Google Scholar] [CrossRef]
- DiMaio, D.; Petti, L.M. The E5 Proteins. Virology 2013, 445, 99–114. [Google Scholar] [CrossRef]
- Mantovani, F.; Banks, L. The Human Papillomavirus E6 Protein and Its Contribution to Malignant Progression. Oncogene 2001, 20, 7874–7887. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, S.L.; Stremlau, M.; He, X.; Basile, J.R.; Münger, K. Degradation of the Retinoblastoma Tumor Suppressor by the Human Papillomavirus Type 16 E7 Oncoprotein Is Important for Functional Inactivation and Is Separable from Proteasomal Degradation of E7. J. Virol. 2001, 75, 7583–7591. [Google Scholar] [CrossRef]
- Zheng, Z.-M.; Wang, X. Regulation of Cellular miRNA Expression by Human Papillomaviruses. Biochim Biophys. Acta 2011, 1809, 668–677. [Google Scholar] [CrossRef]
- Greco, D.; Kivi, N.; Qian, K.; Leivonen, S.-K.; Auvinen, P.; Auvinen, E. Human Papillomavirus 16 E5 Modulates the Expression of Host microRNAs. PLoS ONE 2011, 6, e21646. [Google Scholar] [CrossRef]
- Sommerova, L.; Anton, M.; Bouchalova, P.; Jasickova, H.; Rak, V.; Jandakova, E.; Selingerova, I.; Bartosik, M.; Vojtesek, B.; Hrstka, R. The Role of miR-409-3p in Regulation of HPV16/18-E6 mRNA in Human Cervical High-Grade Squamous Intraepithelial Lesions. Antivir. Res. 2019, 163, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Konstantopoulos, G.; Leventakou, D.; Saltiel, D.-R.; Zervoudi, E.; Logotheti, E.; Pettas, S.; Karagianni, K.; Daiou, A.; Hatzistergos, K.E.; Dafou, D.; et al. HPV16 E6 Oncogene Contributes to Cancer Immune Evasion by Regulating PD-L1 Expression through a miR-143/HIF-1a Pathway. Viruses 2024, 16, 113. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Zhang, S.; Zhao, Z.; Mao, X.; Huang, J.; Wu, Z.; Zheng, L.; Wang, Q. MicroRNA-27b up-Regulated by Human Papillomavirus 16 E7 Promotes Proliferation and Suppresses Apoptosis by Targeting Polo-like Kinase2 in Cervical Cancer. Oncotarget 2016, 7, 19666–19679. [Google Scholar] [CrossRef]
- Kong, Q.; Wang, W.; Li, P. Regulator Role of HPV E7 Protein on miR-21 Expression in Cervical Carcinoma Cells and Its Functional Implication. Int. J. Clin. Exp. Pathol. 2015, 8, 15808–15813. [Google Scholar]
- Honegger, A.; Schilling, D.; Bastian, S.; Sponagel, J.; Kuryshev, V.; Sültmann, H.; Scheffner, M.; Hoppe-Seyler, K.; Hoppe-Seyler, F. Dependence of Intracellular and Exosomal microRNAs on Viral E6/E7 Oncogene Expression in HPV-Positive Tumor Cells. PLoS Pathog. 2015, 11, e1004712. [Google Scholar] [CrossRef]
- Ling, J.; Sun, Q.; Tian, Q.; Shi, H.; Yang, H.; Ren, J. Human Papillomavirus 16 E6/E7 Contributes to Immune Escape and Progression of Cervical Cancer by Regulating miR-142-5p/PD-L1 Axis. Arch. Biochem Biophys. 2022, 731, 109449. [Google Scholar] [CrossRef]
- Fujii, T.; Shimada, K.; Asano, A.; Tatsumi, Y.; Yamaguchi, N.; Yamazaki, M.; Konishi, N. MicroRNA-331-3p Suppresses Cervical Cancer Cell Proliferation and E6/E7 Expression by Targeting NRP2. Int. J. Mol. Sci. 2016, 17, 1351. [Google Scholar] [CrossRef]
- Han, M.-S.; Lee, J.M.; Kim, S.-N.; Kim, J.-H.; Kim, H.-S. Human Papillomavirus 16 Oncoproteins Downregulate the Expression of miR-148a-3p, miR-190a-5p, and miR-199b-5p in Cervical Cancer. BioMed Res. Int. 2018, 2018, 1942867. [Google Scholar] [CrossRef]
- Wang, F.; Li, B.; Xie, X. The Roles and Clinical Significance of microRNAs in Cervical Cancer. Histol. Histopathol. 2016, 131–139. [Google Scholar] [CrossRef]
- Gupta, S.; Kumar, P.; Das, B.C. HPV: Molecular Pathways and Targets. Curr. Probl. Cancer 2018, 42, 161–174. [Google Scholar] [CrossRef] [PubMed]
- Doghish, A.S.; Ali, M.A.; Elyan, S.S.; Elrebehy, M.A.; Mohamed, H.H.; Mansour, R.M.; Elgohary, A.; Ghanem, A.; Faraag, A.H.I.; Abdelmaksoud, N.M.; et al. miRNAs Role in Cervical Cancer Pathogenesis and Targeted Therapy: Signaling Pathways Interplay. Pathol.-Res. Pract. 2023, 244, 154386. [Google Scholar] [CrossRef]
- Hazazi, A.; Khan, F.R.; Albloui, F.; Arif, S.; Abdulaziz, O.; Alhomrani, M.; Sindi, A.A.A.; Abu-Alghayth, M.H.; Abalkhail, A.; Nassar, S.A.; et al. Signaling Pathways in HPV-Induced Cervical Cancer: Exploring the Therapeutic Promise of RNA Modulation. Pathol.-Res. Pract. 2024, 263, 155612. [Google Scholar] [CrossRef]
- Pal, A.; Kundu, R. Human Papillomavirus E6 and E7: The Cervical Cancer Hallmarks and Targets for Therapy. Front Microbiol. 2019, 10, 3116. [Google Scholar] [CrossRef]
- Wang, X.; Meyers, C.; Guo, M.; Zheng, Z.-M. Up-Regulation of p18Ink4c Expression by Oncogenic HPV E6 via P53-miR-34a Pathway. Int. J. Cancer 2011, 129, 1362–1372. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, H.-K.; McCoy, J.P.; Banerjee, N.S.; Rader, J.S.; Broker, T.R.; Meyers, C.; Chow, L.T.; Zheng, Z.-M. Oncogenic HPV Infection Interrupts the Expression of Tumor-Suppressive miR-34a through Viral Oncoprotein E6. RNA 2009, 15, 637–647. [Google Scholar] [CrossRef]
- Au Yeung, C.L.; Tsang, T.Y.; Yau, P.L.; Kwok, T.T. Human Papillomavirus Type 16 E6 Induces Cervical Cancer Cell Migration through the P53/microRNA-23b/Urokinase-Type Plasminogen Activator Pathway. Oncogene 2011, 30, 2401–2410. [Google Scholar] [CrossRef]
- Jung, H.M.; Phillips, B.L.; Chan, E.K. miR-375 Activates P21 and Suppresses Telomerase Activity by Coordinately Regulating HPV E6/E7, E6AP, CIP2A, and 14-3-3ζ. Mol. Cancer 2014, 13, 80. [Google Scholar] [CrossRef]
- Song, L.; Liu, S.; Zeng, S.; Zhang, L.; Li, X. miR-375 Modulates Radiosensitivity of HR-HPV-Positive Cervical Cancer Cells by Targeting UBE3A through the P53 Pathway. Med. Sci. Monit. 2015, 21, 2210–2217. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Li, S.; Yan, Q.; Chen, X.; Yang, Y.; Liu, X.; Wan, X. Interferon-β Induced microRNA-129-5p Down-Regulates HPV-18 E6 and E7 Viral Gene Expression by Targeting SP1 in Cervical Cancer Cells. PLoS ONE 2013, 8, e81366. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Zhang, Y.; Zhang, S. MicroRNA-92 Regulates Cervical Tumorigenesis and Its Expression Is Upregulated by Human Papillomavirus-16 E6 in Cervical Cancer Cells. Oncol. Lett. 2013, 6, 468–474. [Google Scholar] [CrossRef] [PubMed]
- Cai, N.; Wang, Y.-D.; Zheng, P.-S. The microRNA-302-367 Cluster Suppresses the Proliferation of Cervical Carcinoma Cells through the Novel Target AKT1. RNA 2013, 19, 85–95. [Google Scholar] [CrossRef]
- Hou, T.; Ou, J.; Zhao, X.; Huang, X.; Huang, Y.; Zhang, Y. MicroRNA-196a Promotes Cervical Cancer Proliferation through the Regulation of FOXO1 and p27Kip1. Br. J. Cancer 2014, 110, 1260–1268. [Google Scholar] [CrossRef]
- Cui, F.; Li, X.; Zhu, X.; Huang, L.; Huang, Y.; Mao, C.; Yan, Q.; Zhu, J.; Zhao, W.; Shi, H. MiR-125b Inhibits Tumor Growth and Promotes Apoptosis of Cervical Cancer Cells by Targeting Phosphoinositide 3-Kinase Catalytic Subunit Delta. Cell Physiol. Biochem 2012, 30, 1310–1318. [Google Scholar] [CrossRef]
- Qin, W.; Dong, P.; Ma, C.; Mitchelson, K.; Deng, T.; Zhang, L.; Sun, Y.; Feng, X.; Ding, Y.; Lu, X.; et al. MicroRNA-133b Is a Key Promoter of Cervical Carcinoma Development through the Activation of the ERK and AKT1 Pathways. Oncogene 2012, 31, 4067–4075. [Google Scholar] [CrossRef]
- Li, J.; Ping, Z.; Ning, H. MiR-218 Impairs Tumor Growth and Increases Chemo-Sensitivity to Cisplatin in Cervical Cancer. Int. J. Mol. Sci. 2012, 13, 16053–16064. [Google Scholar] [CrossRef]
- Campos-Viguri, G.E.; Peralta-Zaragoza, O.; Jiménez-Wences, H.; Longinos-González, A.E.; Castañón-Sánchez, C.A.; Ramírez-Carrillo, M.; Camarillo, C.L.; Castañeda-Saucedo, E.; Jiménez-López, M.A.; Martínez-Carrillo, D.N.; et al. MiR-23b-3p Reduces the Proliferation, Migration and Invasion of Cervical Cancer Cell Lines via the Reduction of c-Met Expression. Sci. Rep. 2020, 10, 3256. [Google Scholar] [CrossRef]
- Xu, J.; Zhang, W.; Lv, Q.; Zhu, D. Overexpression of miR-21 Promotes the Proliferation and Migration of Cervical Cancer Cells via the Inhibition of PTEN. Oncol. Rep. 2015, 33, 3108–3116. [Google Scholar] [CrossRef]
- Hao, Z.; Yang, J.; Wang, C.; Li, Y.; Zhang, Y.; Dong, X.; Zhou, L.; Liu, J.; Zhang, Y.; Qian, J. MicroRNA-7 Inhibits Metastasis and Invasion through Targeting Focal Adhesion Kinase in Cervical Cancer. Int. J. Clin. Exp. Med. 2015, 8, 480–487. [Google Scholar] [PubMed]
- Lu, M.; Gao, Q.; Wang, Y.; Ren, J.; Zhang, T. LINC00511 Promotes Cervical Cancer Progression by Regulating the miR-497-5p/MAPK1 Axis. Apoptosis 2022, 27, 800–811. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Guo, Q.; Gao, S.; Hua, K. miR-454-3p Promotes Proliferation and Induces Apoptosis in Human Cervical Cancer Cells by Targeting TRIM3. Biochem. Biophys. Res. Commun. 2019, 516, 872–879. [Google Scholar] [CrossRef]
- Leung, C.O.N.; Deng, W.; Ye, T.-M.; Ngan, H.Y.S.; Tsao, S.W.; Cheung, A.N.Y.; Pang, R.T.K.; Yeung, W.S.B. miR-135a Leads to Cervical Cancer Cell Transformation through Regulation of β-Catenin via a SIAH1-Dependent Ubiquitin Proteosomal Pathway. Carcinogenesis 2014, 35, 1931–1940. [Google Scholar] [CrossRef] [PubMed]
- Niu, Z.; Wang, F.; Lv, S.; Lv, Y.; Liu, M.; Fu, L.; Yao, Y.; Wang, L.; Lin, W.; Yuan, F. HNRNPU-AS1 Regulates Cell Proliferation and Apoptosis via the MicroRNA 205-5p/AXIN2 Axis and Wnt/β-Catenin Signaling Pathway in Cervical Cancer. Mol. Cell. Biol. 2021, 41, e00115-21. [Google Scholar] [CrossRef]
- Sathyanarayanan, A.; Chandrasekaran, K.S.; Karunagaran, D. microRNA-146a Inhibits Proliferation, Migration and Invasion of Human Cervical and Colorectal Cancer Cells. Biochem. Biophys. Res. Commun. 2016, 480, 528–533. [Google Scholar] [CrossRef]
- Li, T.; Zhou, W.; Li, Y.; Gan, Y.; Peng, Y.; Xiao, Q.; Ouyang, C.; Wu, A.; Zhang, S.; Liu, J.; et al. MiR-4524b-5p/WTX/β-Catenin Axis Functions as a Regulator of Metastasis in Cervical Cancer. PLoS ONE 2019, 14, e0214822. [Google Scholar] [CrossRef]
- Zhou, Y.; An, Q.; Guo, R.; Qiao, Y.; Li, L.; Zhang, X.; Zhao, X. miR424-5p Functions as an Anti-Oncogene in Cervical Cancer Cell Growth by Targeting KDM5B via the Notch Signaling Pathway. Life Sci. 2017, 171, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Pang, R.T.K.; Leung, C.O.N.; Ye, T.-M.; Liu, W.; Chiu, P.C.N.; Lam, K.K.W.; Lee, K.-F.; Yeung, W.S.B. MicroRNA-34a Suppresses Invasion through Downregulation of Notch1 and Jagged1 in Cervical Carcinoma and Choriocarcinoma Cells. Carcinogenesis 2010, 31, 1037–1044. [Google Scholar] [CrossRef]
- Chen, Y.; Wu, Q.; Lin, J.; Wei, J. DARS-AS1 Accelerates the Proliferation of Cervical Cancer Cells via miR-628-5p/JAG1 Axis to Activate Notch Pathway. Cancer Cell Int. 2020, 20, 535. [Google Scholar] [CrossRef] [PubMed]
- Causin, R.L.; Van Helvoort Lengert, A.; Gomes, I.N.F.; De Freitas, A.J.A.; Rosa, M.N.; Dos Reis, R.; Reis, R.M.; Marques, M.M.C. MicroRNA-130a-3p Inhibition Suppresses Cervical Cancer Cell Progression. Oncol. Rep. 2023, 49, 109. [Google Scholar] [CrossRef]
- Wen, S.-Y.; Lin, Y.; Yu, Y.-Q.; Cao, S.-J.; Zhang, R.; Yang, X.-M.; Li, J.; Zhang, Y.-L.; Wang, Y.-H.; Ma, M.-Z.; et al. miR-506 Acts as a Tumor Suppressor by Directly Targeting the Hedgehog Pathway Transcription Factor Gli3 in Human Cervical Cancer. Oncogene 2015, 34, 717–725. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Jia, J.; Zhao, L.; Li, X.; Xie, Q.; Chen, X.; Wang, J.; Lu, F. Down-Regulation of microRNA-9 Leads to Activation of IL-6/Jak/STAT3 Pathway through Directly Targeting IL-6 in HeLa Cell. Mol. Carcinog. 2016, 55, 732–742. [Google Scholar] [CrossRef]
- Xu, J.; Wang, H.; Wang, H.; Chen, Q.; Zhang, L.; Song, C.; Zhou, Q.; Hong, Y. The Inhibition of miR-126 in Cell Migration and Invasion of Cervical Cancer through Regulating ZEB1. Hereditas 2019, 156, 11. [Google Scholar] [CrossRef]
- Chen, G.; Huang, P.; Xie, J.; Li, R. microRNA-211 Suppresses the Growth and Metastasis of Cervical Cancer by Directly Targeting ZEB1. Mol. Med. Rep. 2018, 17, 1275–1282. [Google Scholar] [CrossRef]
- Liang, H.; Zhao, Y.; Pi, J.; Luo, R. MiR-875-5p Suppresses Cervical Cancer Cell Proliferation and Metastasis via Negative Regulation of EGFR. Trop. J. Pharm. Res. 2021, 20, 939–946. [Google Scholar] [CrossRef]
- Lei, C.; Wang, Y.; Huang, Y.; Yu, H.; Huang, Y.; Wu, L.; Huang, L. Up-Regulated miR155 Reverses the Epithelial-Mesenchymal Transition Induced by EGF and Increases Chemo-Sensitivity to Cisplatin in Human Caski Cervical Cancer Cells. PLoS ONE 2012, 7, e52310. [Google Scholar] [CrossRef]
- Abbas, M.; Mehdi, A.; Khan, F.H.; Verma, S.; Ahmad, A.; Khatoon, F.; Raza, S.T.; Afreen, S.; Glynn, S.A.; Mahdi, F. Role of miRNAs in Cervical Cancer: A Comprehensive Novel Approach from Pathogenesis to Therapy. J. Gynecol. Obstet. Hum. Reprod. 2021, 50, 102159. [Google Scholar] [CrossRef]
- Chauhan, P.; Pramodh, S.; Hussain, A.; Elsori, D.; Lakhanpal, S.; Kumar, R.; Alsaweed, M.; Iqbal, D.; Pandey, P.; Al Othaim, A.; et al. Understanding the Role of miRNAs in Cervical Cancer Pathogenesis and Therapeutic Responses. Front Cell Dev. Biol. 2024, 12, 1397945. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.-H.; Qin, Y.-E.; Tang, W.-F.; Tao, J.; Song, H.; Zuo, M. MiR-34a and miR-206 Act as Novel Prognostic and Therapy Biomarkers in Cervical Cancer. Cancer Cell Int. 2017, 17, 63. [Google Scholar] [CrossRef]
- Huang, L.; Lin, J.-X.; Yu, Y.-H.; Zhang, M.-Y.; Wang, H.-Y.; Zheng, M. Downregulation of Six MicroRNAs Is Associated with Advanced Stage, Lymph Node Metastasis and Poor Prognosis in Small Cell Carcinoma of the Cervix. PLoS ONE 2012, 7, e33762. [Google Scholar] [CrossRef]
- Azizmohammadi, S.; Safari, A.; Azizmohammadi, S.; Kaghazian, M.; Sadrkhanlo, M.; Yahaghi, E.; Farshgar, R.; Seifoleslami, M. Molecular Identification of miR-145 and miR-9 Expression Level as Prognostic Biomarkers for Early-Stage Cervical Cancer Detection. QJM An. Int. J. Med. 2017, 110, 11–15. [Google Scholar] [CrossRef] [PubMed]
- Romero-López, M.J.; Jiménez-Wences, H.; Cruz-De la Rosa, M.I.; Román-Fernández, I.V.; Fernández-Tilapa, G. miR-23b-3p, miR-124-3p and miR-218-5p Synergistic or Additive Effects on Cellular Processes That Modulate Cervical Cancer Progression? A Molecular Balance That Needs Attention. Int. J. Mol. Sci. 2022, 23, 13551. [Google Scholar] [CrossRef] [PubMed]
- Dong, C.; Dong, R.; Song, J.; Yu, C.; Zhuang, Y.; Guo, Q. Knockdown of lncRNA EGFR-AS1 Promotes Autophagy-Mediated Ferroptosis in Cervical Cancer via Regulating EGFR Expression through miR-133b. Mol. Cell. Toxicol. 2024, 20, 139–147. [Google Scholar] [CrossRef]
- Kogo, R.; How, C.; Chaudary, N.; Bruce, J.; Shi, W.; Hill, R.P.; Zahedi, P.; Yip, K.W.; Liu, F.-F. The microRNA-218~Survivin Axis Regulates Migration, Invasion, and Lymph Node Metastasis in Cervical Cancer. Oncotarget 2014, 6, 1090–1100. [Google Scholar] [CrossRef]
- Karunakara, S.H.; Eswaran, S.; Mallya, S.; Suresh, P.S.; Chakrabarty, S.; Kabekkodu, S.P. Analysis of miR-497/195 Cluster Identifies New Therapeutic Targets in Cervical Cancer. BMC Res. Notes 2024, 17, 217. [Google Scholar] [CrossRef]
- Santos, J.M.O.; Tavares, V.; Gil da Costa, R.M.; Medeiros, R. MiR-150 and miR-155 Expression Predicts Survival of Cervical Cancer Patients: A Translational Approach to Novel Prognostic Biomarkers. Biomarkers 2023, 28, 617–627. [Google Scholar] [CrossRef]
- Xu, J.; Li, Y.; Wang, F.; Wang, X.; Cheng, B.; Ye, F.; Xie, X.; Zhou, C.; Lu, W. Suppressed miR-424 Expression via Upregulation of Target Gene Chk1 Contributes to the Progression of Cervical Cancer. Oncogene 2013, 32, 976–987. [Google Scholar] [CrossRef]
- Yang, Y.; Song, K.; Chang, H.; Chen, L. Decreased Expression of microRNA-126 Is Associated with Poor Prognosis in Patients with Cervical Cancer. Diagn. Pathol. 2014, 9, 220. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA Signatures in Human Cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef]
- Chakrabortty, A.; Patton, D.J.; Smith, B.F.; Agarwal, P. miRNAs: Potential as Biomarkers and Therapeutic Targets for Cancer. Genes 2023, 14, 1375. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Zhou, M.; Luan, Y.; Li, K.; Wang, Y.; Yang, X. Identification of Key Genes Associated with Cervical Cancer Based on Bioinformatics Analysis. BMC Cancer 2024, 24, 897. [Google Scholar] [CrossRef] [PubMed]
- Rosati, D.; Palmieri, M.; Brunelli, G.; Morrione, A.; Iannelli, F.; Frullanti, E.; Giordano, A. Differential Gene Expression Analysis Pipelines and Bioinformatic Tools for the Identification of Specific Biomarkers: A Review. Comput Struct. Biotechnol. J. 2024, 23, 1154–1168. [Google Scholar] [CrossRef] [PubMed]
- Terra Machado, D.; Bernardes Brustolini, O.J.; Côrtes Martins, Y.; Grivet Mattoso Maia, M.A.; Ribeiro de Vasconcelos, A.T. Inference of Differentially Expressed Genes Using Generalized Linear Mixed Models in a Pairwise Fashion. PeerJ 2023, 11, e15145. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma Powers Differential Expression Analyses for RNA-Sequencing and Microarray Studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Hsu, S.-D.; Lin, F.-M.; Wu, W.-Y.; Liang, C.; Huang, W.-C.; Chan, W.-L.; Tsai, W.-T.; Chen, G.-Z.; Lee, C.-J.; Chiu, C.-M.; et al. miRTarBase: A Database Curates Experimentally Validated microRNA-Target Interactions. Nucleic Acids Res. 2011, 39, D163–169. [Google Scholar] [CrossRef]
- Yi, Y.; Liu, Y.; Wu, W.; Wu, K.; Zhang, W. The Role of miR-106p-5p in Cervical Cancer: From Expression to Molecular Mechanism. Cell Death Discov. 2018, 4, 1–15. [Google Scholar] [CrossRef]
- Xu, M.; Huseinovic, A.; Jaspers, A.; Yuan, L.; Steenbergen, R.D.M. Downregulation of miR-193a/b-3p during HPV-Induced Cervical Carcinogenesis Contributes to Anchorage-Independent Growth through PI3K-AKT Pathway Regulators. J. Med. Virol. 2023, 95, e28589. [Google Scholar] [CrossRef] [PubMed]
- Burk, R.D.; Chen, Z.; Saller, C.; Tarvin, K.; Carvalho, A.L.; Scapulatempo-Neto, C.; Silveira, H.C.; Fregnani, J.H.; Creighton, C.J.; Anderson, M.L.; et al. Integrated Genomic and Molecular Characterization of Cervical Cancer. Nature 2017, 543, 378–384. [Google Scholar] [CrossRef] [PubMed]
- Biswas, N.; Chakrabarti, S. Artificial Intelligence (AI)-Based Systems Biology Approaches in Multi-Omics Data Analysis of Cancer. Front Oncol. 2020, 10, 588221. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Ding, J.; Luo, J.; Lu, L.; Yang, F.; Tan, X. Seven Protective miRNA Signatures for Prognosis of Cervical Cancer. Oncotarget 2016, 7, 56690–56698. [Google Scholar] [CrossRef]
- Afshar, S.; Afshar, S.; Warden, E.; Manochehri, H.; Saidijam, M. Application of Artificial Neural Network in miRNA Biomarker Selection and Precise Diagnosis of Colorectal Cancer. Iran. BioMed J. 2019, 23, 175–183. [Google Scholar] [CrossRef]
- Ozaki, Y.; Broughton, P.; Abdollahi, H.; Valafar, H.; Blenda, A.V. Integrating Omics Data and AI for Cancer Diagnosis and Prognosis. Cancers 2024, 16, 2448. [Google Scholar] [CrossRef]

| Protein | miRNAs | Expression | Target | Cell Process | Ref. |
| E5 | miR-146a | Up | ZNF813 | Cell adhesion and cell cycle | [37] |
| E5 | miR-203 | Down | p63 | Cell junction, cell migration, and cell motility | [37] |
| E5 | miR-324-5p | Down | CDH2, CTNNB1 | Transendothelial migration | [37] |
| E6 | miR-409-3p | Down | PI3K/AKT | Proliferation and migration | [38] |
| E6 | miR-143 | Up | HIF-1a, PD-L1 | Immune system | [39] |
| E7 | miR-27b | Up | PLK2 | Cell proliferation, invasion and apoptosis | [40] |
| E7 | miR-21 | Up | No identified | Cell growth, proliferation and invasion | [41] |
| E6/E7 | miR-17-5p, miR-186-5p, miR-378a-3p, miR-378f, miR-629-5p, and miR-7-5p (intracellular) | Down | No identified | Cell proliferation and apoptosis | [42] |
| E6/E7 | miR-143-3p, miR-23a-3p, miR-23b-3p, and miR-27b-3p (intracellular) | Up | No identified | Cell proliferation and apoptosis | [42] |
| E6/E7 | let-7d-5p, miR-20a-5p, miR-378a-3p, miR-423-3p, miR-7-5p, miR-92a-3p (exosomal) | Down | p21 | Cell proliferation and apoptosis | [42] |
| E6/E7 | miR-21-5p (exosomal) | Up | p21 | Cell proliferation and apoptosis | [42] |
| E6/E7 | miR-142-5p | Down | PD-L1 | Immune system | [43] |
| E6/E7 | miR-331-3p | Up | NRP2 | Cell proliferation, cell cycle, apoptosis | [44] |
| E6/E7 | miR-199b-5p and miR-190a-5p | Down | No identified | Cell proliferation | [45] |
| E5/E6/E7 | miR-148a-3p | Down | No identified | Cell proliferation | [45] |
| This table summarizes the role of HPV oncoproteins (E5, E6, and E7) in regulating microRNAs (miRNAs) involved in cervical cancer progression. The "Protein" column lists the HPV oncoproteins responsible for miRNA regulation. The "miRNAs" column shows the specific miRNAs affected by these oncoproteins. The "Regulation" column indicates whether the miRNA is upregulated (increased) or downregulated (decreased). The "Target" column identifies the key molecular targets of the regulated miRNAs. The "Cell Process" column describes the biological processes influenced, such as cell proliferation, migration, apoptosis, and immune evasion. The "Ref." column references the studies that reported these interactions. | |||||
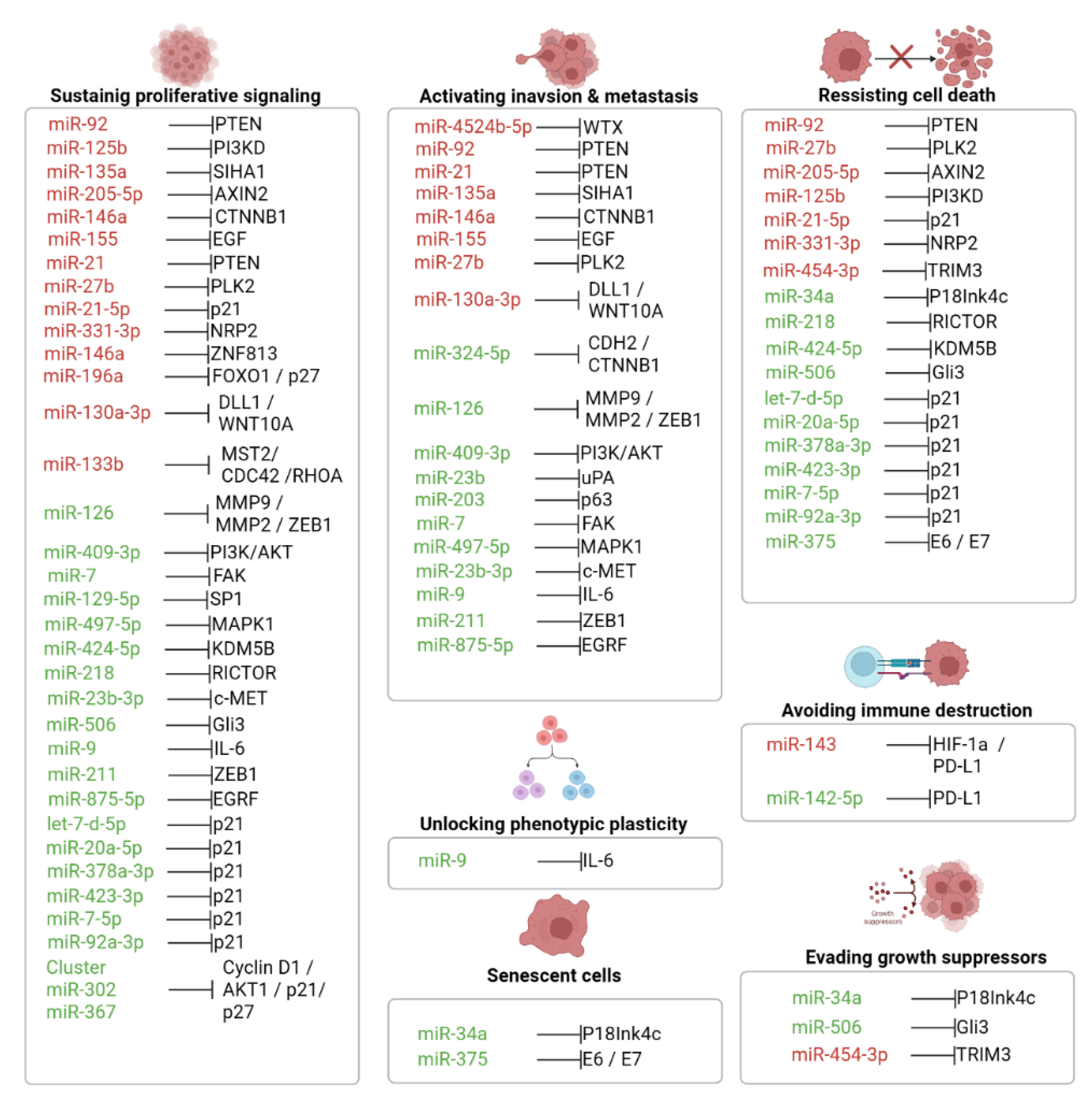
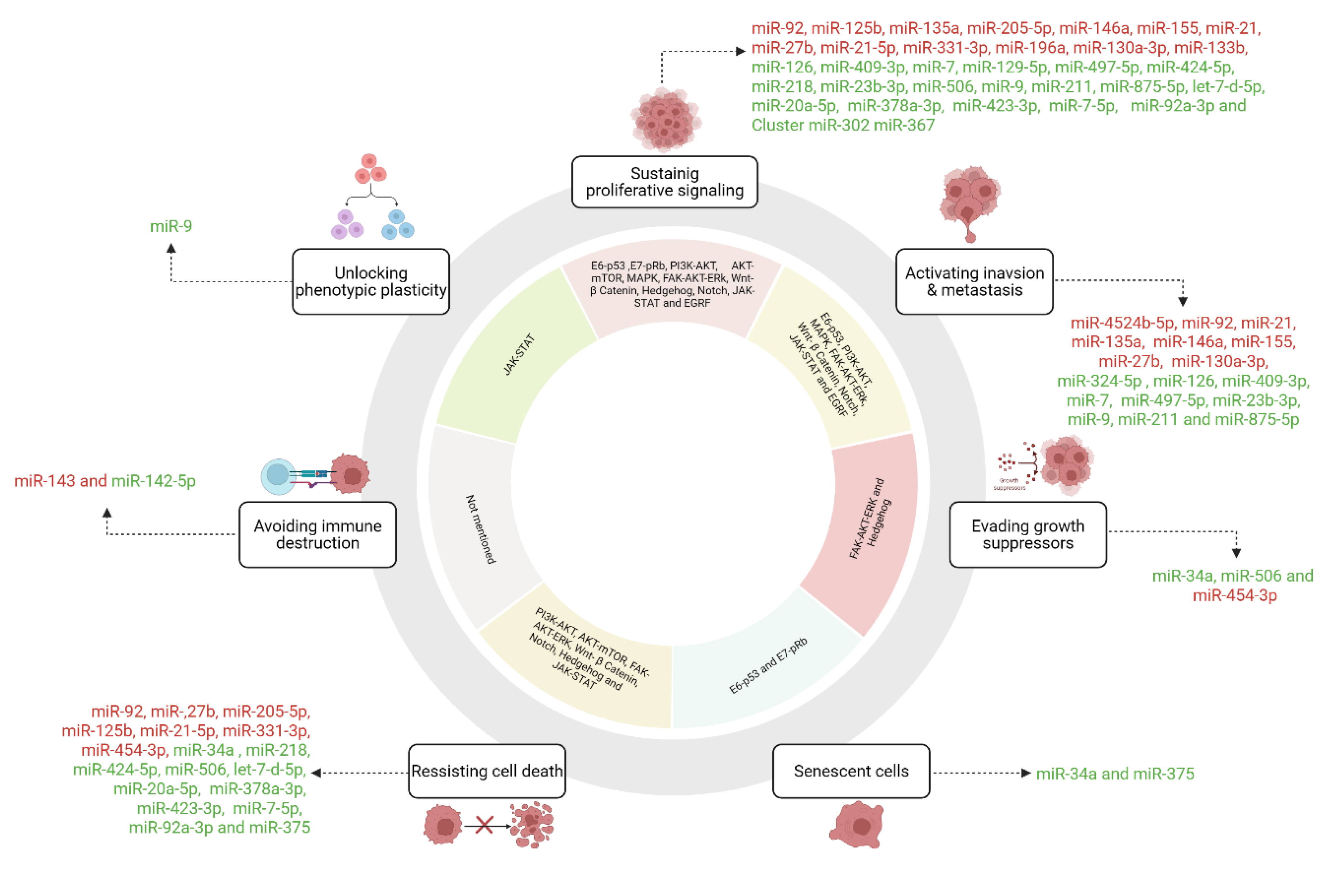
| miRNA | Target | Signaling pathways | Hallmarks of cancer | Ref. | ||||
| miR-34a | p18Ink4c | E6/p53 | Senescent cells, evading growth suppressors and resisting cell death | [51,52] | ||||
| miR-23b | uPA | E6/p53 | Activating invasion & metastasis | [53] | ||||
| miR-375 | E6 and E7 | E6/p53 and E7/pRb | senescent cells and resisting cell death | [54] | ||||
| miR-129-5p | SP1 | E6/p53 and E7/pRb | Sustaining proliferative signaling | [56] | ||||
| miR-92 | PTEN | PI3K/AKT | Sustaining proliferative signaling, activating invasion & metastasis and resisting cell death | [57] | ||||
| Cluster miR-302 and miR-367 | Cyclin D1, AKT1, and p27 | PI3K/AKT | Sustaining proliferative signaling | [58] | ||||
| miR-196a | FOXO1 and p27 | PI3K/AKT | Sustaining proliferative signaling | [59] | ||||
| miR-125b | PIK3CD | PI3K/AKT/mTOR | Sustaining proliferative signaling and resisting cell death | [60] | ||||
| miR-133b | MST2, CDC42 and RHOA | AKT/ERK | Sustaining proliferative signaling | [61] | ||||
| miR-218 | RICTOR | AKT/mTOR | Sustaining proliferative signaling and resisting cell death | [62] | ||||
| miR-23b-3p | c-Met | FAK/AKT/ERK | Sustaining proliferative signaling and activating invasion & metastasis | [63] | ||||
| miR-21 | PTEN | FAK/AKT/ERK | Sustaining proliferative signaling and activating invasion & metastasis | [64] | ||||
| miR-7 | FAK | FAK/AKT/ERK | Sustaining proliferative signaling and activating invasion & metastasis | [65] | ||||
| miR-497-5p | MAPK1 | MAPK | Sustaining proliferative signaling and activating invasion & metastasis | [66] | ||||
| miR-454-3p | TRIM3 | MAPK | Evading growth suppressors and resisting cell death | [67] | ||||
| miR-135a | SIAH1 | Wnt/β-catenin | Activating invasion & metastasis and sustaining proliferative signaling | [68] | ||||
| miR-205-5p | AXIN2 | Wnt/β-catenin | Resisting cell death and sustaining proliferative signaling | [69] | ||||
| miR-146a | CTNNB1 | Wnt | Activating invasion & metastasis and sustaining proliferative signaling | [70] | ||||
| miR-4524b-5p | WTX | Wnt/β-catenin | Activating invasion & metastasis | [71] | ||||
| miR-424-5p | KDM5B | Notch | Sustaining proliferative signaling and resisting cell death | [72] | ||||
| miR-34a | Notch1 and Jagged1 | Notch | Activating invasion & metastasis | [73] | ||||
| miR-628-5p | Jagged1 | Notch | Sustaining proliferative signaling and resisting cell death | [74] | ||||
| miR-130a-3p | DLL1 e WNT10A | Notch/Wnt | Activating invasion & metastasis and sustaining proliferative signaling | [75] | ||||
| miR-506 | Gli3 | Hedgehog | Sustaining proliferative signaling, evading growth suppressors and resisting cell death | [76] | ||||
| miR-9 | IL-6 | JAK/STAT | Sustaining proliferative signaling, activating invasion & metastasis and unlocking phenotypic plasticity | [77] | ||||
| miR-126 | MMP2, MMP9 and ZEB1 | JAK/STAT | Sustaining proliferative signaling and activating invasion & metastasis | [78] | ||||
| miR-211 | ZEB1 | JAK/STAT | Sustaining proliferative signaling and activating invasion & metastasis | [79] | ||||
| miR-875-5p | EGFR | EGFR | Sustaining proliferative signaling and activating invasion & metastasis | [80] | ||||
| miR-155 | EGF | EGFR | Sustaining proliferative signaling, activating invasion & metastasis | [81] | ||||
| miRNAs | Expression | Outcome | Ref. |
| miR-34a | Downregulated | Shorter overall survival | [84] |
| miR-23b | Downregulated | Shorter overall survival | [87] |
| miR-125b | Downregulated | Shorter overall survival | [85] |
| miR-133b | Upregulated | Shorter overall survival | [88] |
| miR-218 | Downregulated | Shorter overall survival | [89] |
| miR-196a | Upregulated | Shorter overall survival | [59] |
| miR-497-5p | Downregulated | Poor survival | [90] |
| miR-21 | Upregulated | Shorter overall survival | [91] |
| miR-205-5p | Upregulated | Poor survival | [69] |
| miR-146a | Downregulated | Shorter overall survival | [91] |
| miR-424-5p | Downregulated | Poor prognosis | [92] |
| miR-9 | Upregulated | Shorter overall survival | [86] |
| miR-126 | Downregulated | Shorter overall survival | [93] |
| miR-875-5p | Downregulated | Shorter overall survival | [80] |
| miR-155 | Downregulated | Shorter overall survival | [91] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




