Submitted:
01 May 2026
Posted:
04 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
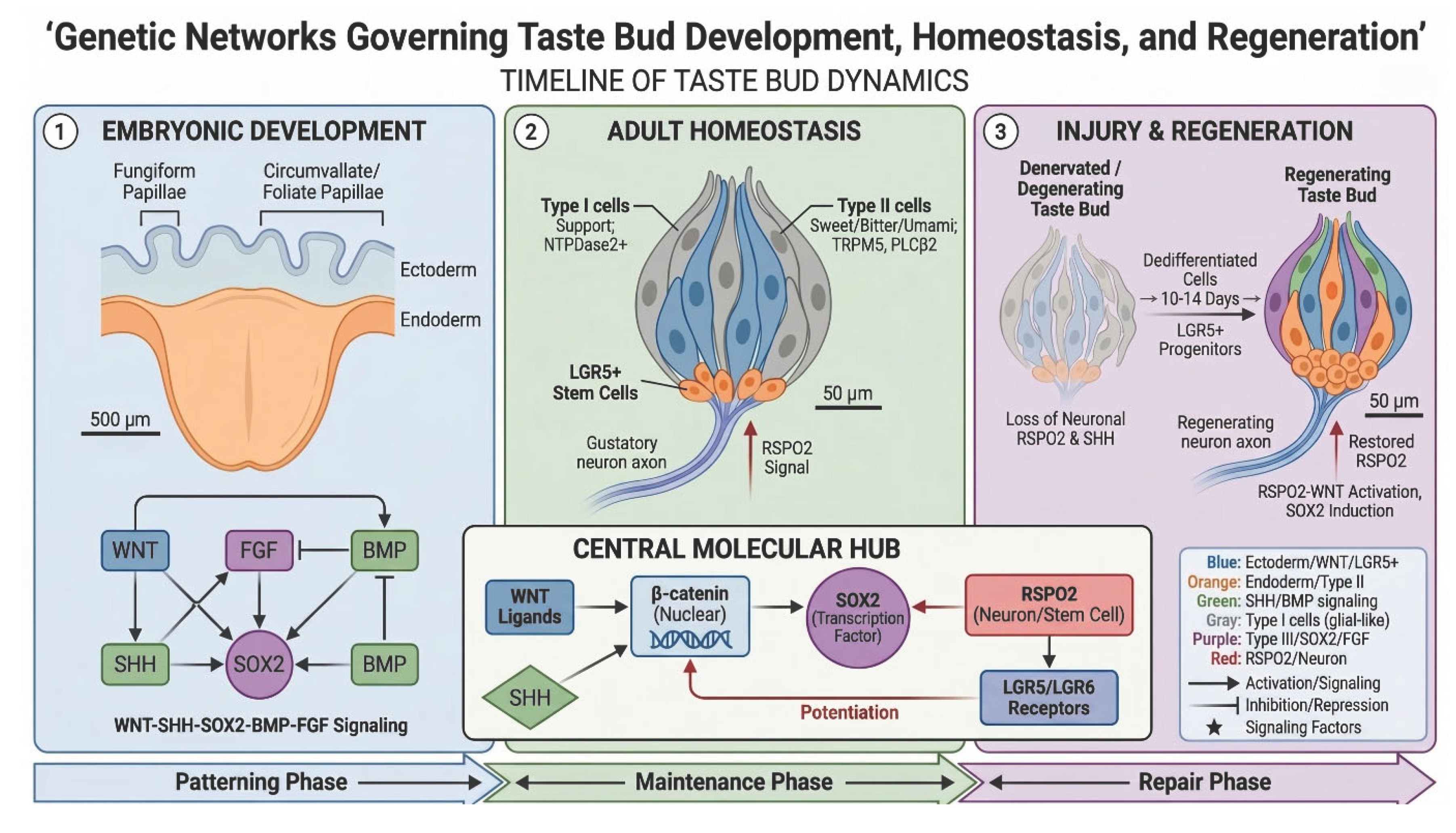
2. Anatomical and Cellular Framework
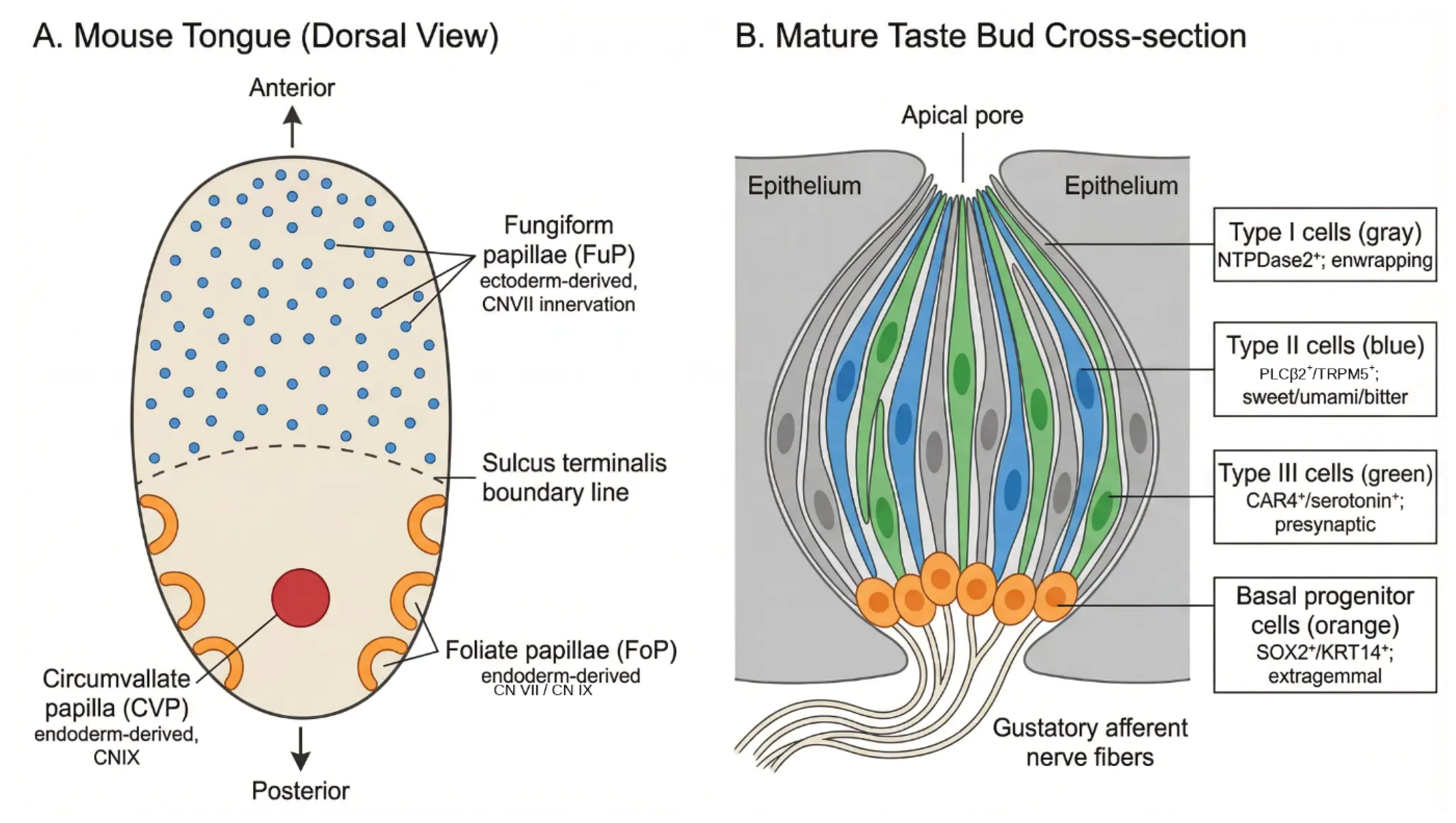
2.1. Taste Fields and Their Embryonic Origins
2.2. Cellular Organization of the Mature Taste Bud
2.3. Animal Models and Experimental Systems
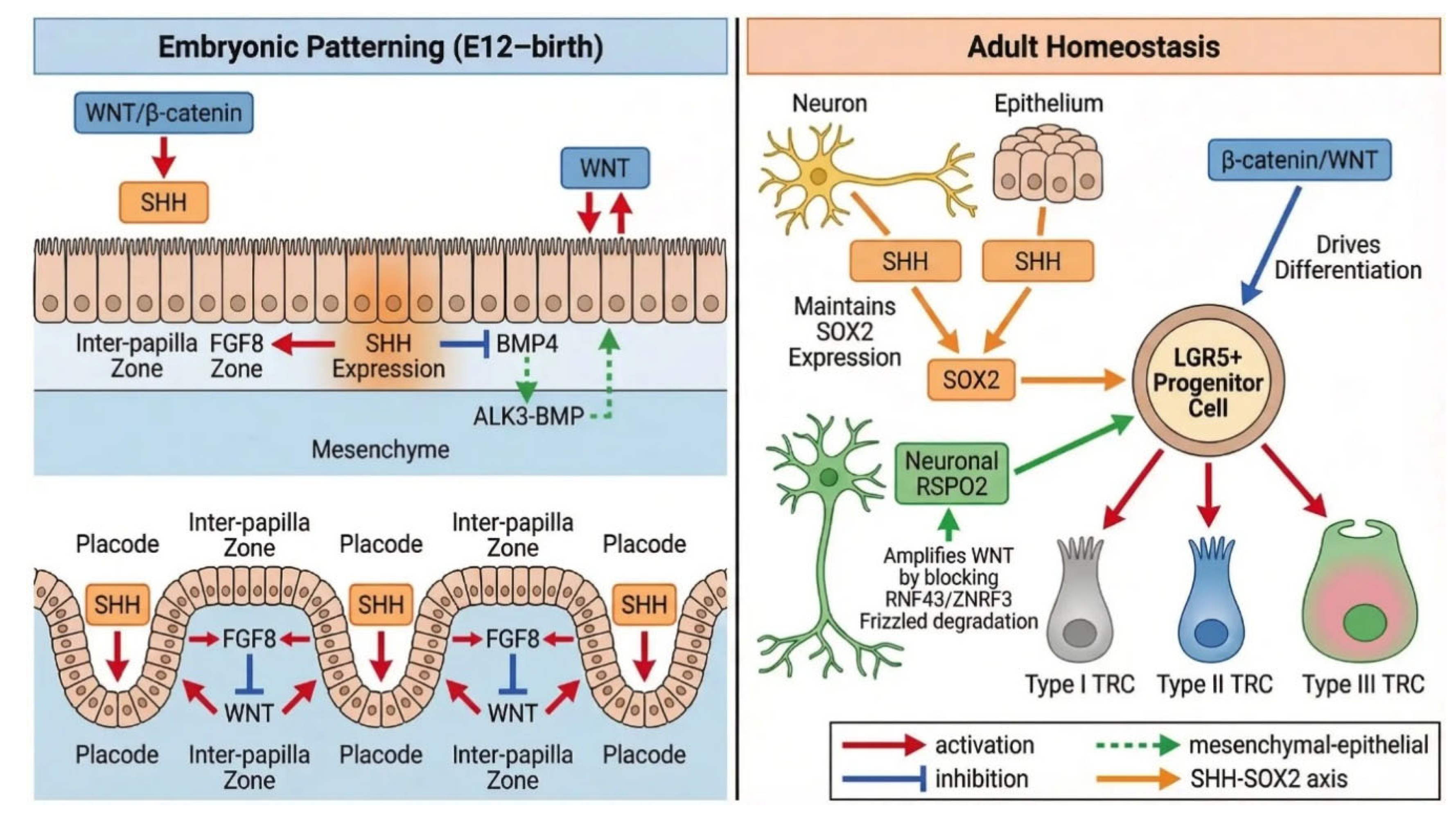
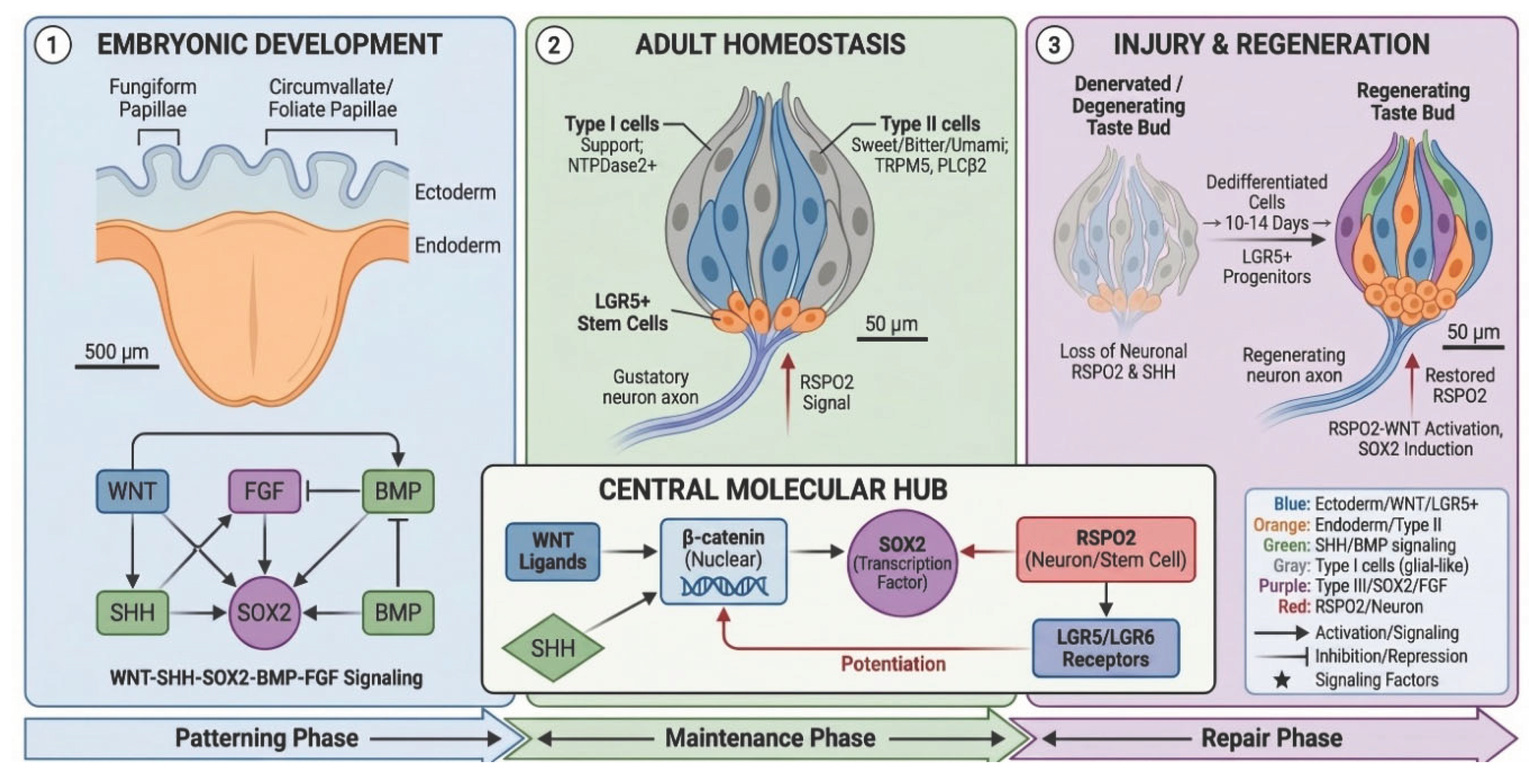
3. Embryonic Development of Taste Papillae
3.1. Early Placode Formation and Papilla Morphogenesis
3.2. WNT–SHH Crosstalk as the Primary Developmental Axis
3.3. SOX2 as a Competence Factor for Taste Fate
3.4. Mesenchymal Control of Epithelial Taste Patterning
3.5. Birth as the Developmental Switch Point
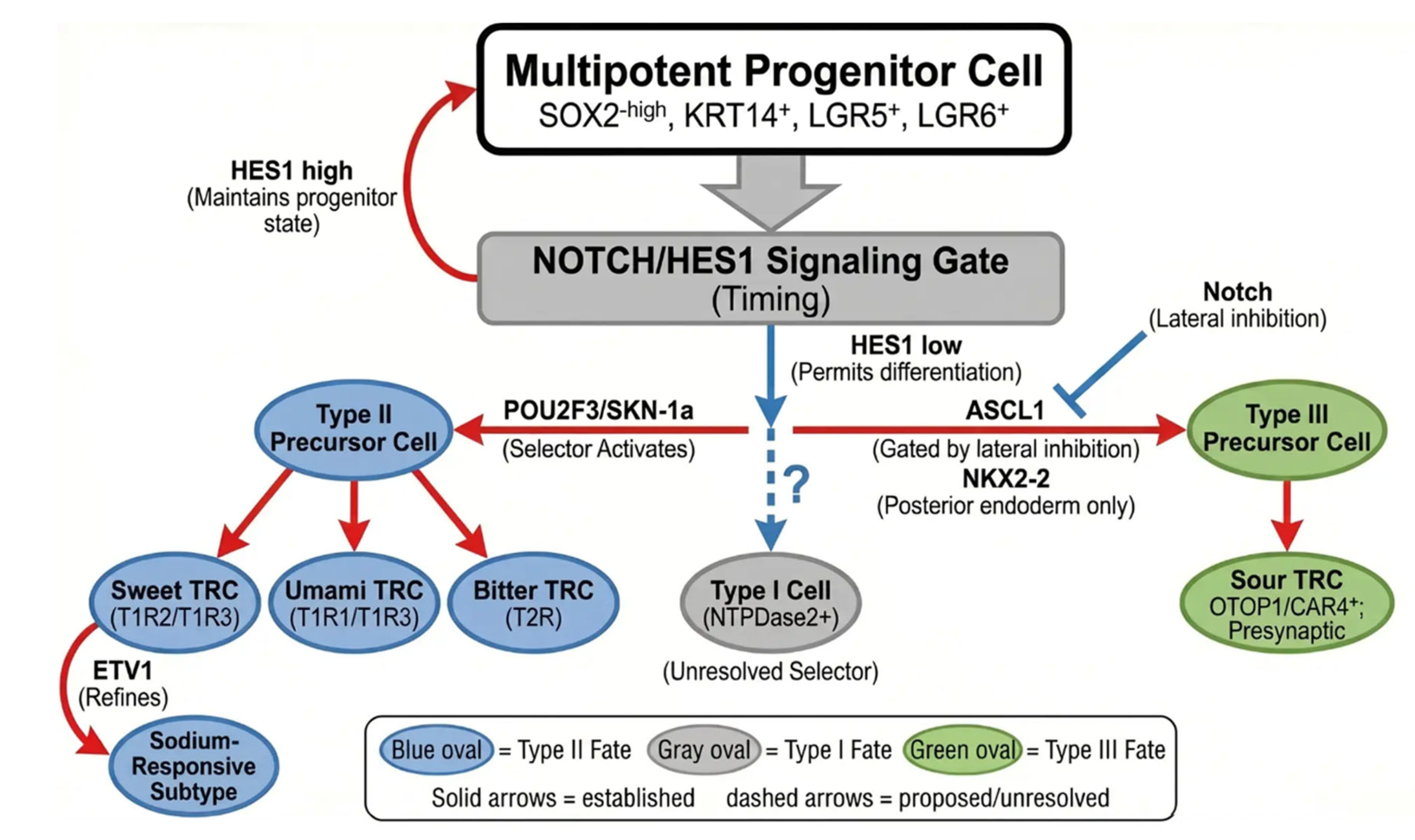
4. Progenitor Cells and Lineage Specification
4.1. KRT14/KRT5 Progenitors and the Extragemmal Origin of New Cells
4.2. LGR5 and LGR6 Stem/Progenitor Compartments
4.3. SOX2-High Competence and Progenitor Heterogeneity
4.4. Type II Lineage Specification: POU2F3 as the Primary Selector
4.5. Type III Lineage Specification: NKX2-2, ASCL1, and Regional Identity.
4.6. Notch/HES Signaling as a Timing and Gating Mechanism
4.7. Emerging Regulators and Unresolved Lineages
5. Adult Homeostasis and Signaling Maintenance
5.1. HH/GLI Signaling as a Homeostatic Requirement
5.2. Downstream of HH in Adult Tissue Maintenance
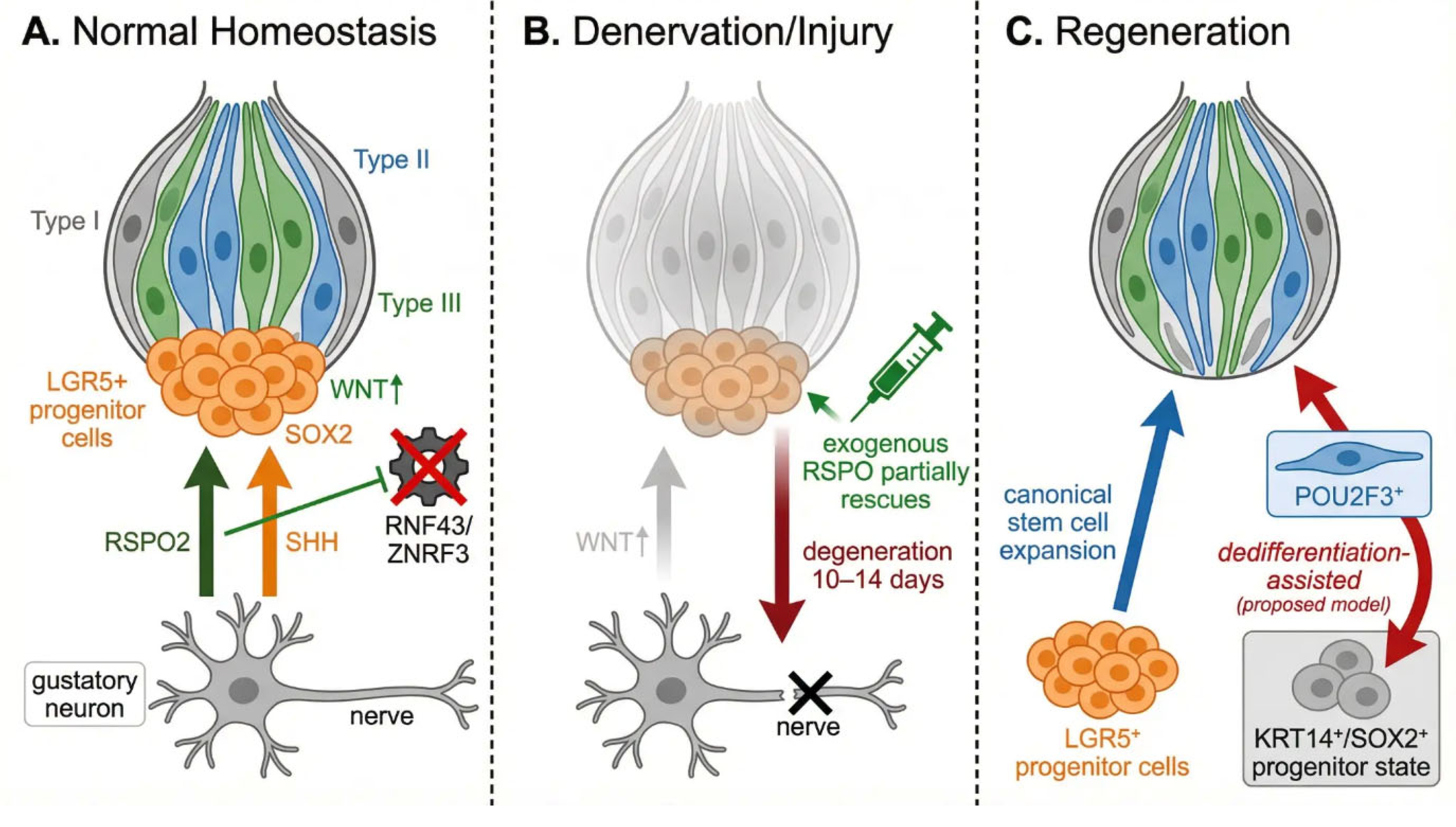
5.3. LGR5/RSPO/WNT Axis and the Neural Niche
6. Injury and Regeneration
6.1. Denervation as the Canonical Regeneration Model
6.2. Adult Epithelial Competence Revealed by Shh Misexpression
6.3. Radiation-Induced Injury and WNT-Mediated Rescue
6.4. Injury-Induced Epithelial Plasticity and Dedifferentiation
7. Comparative and Emerging Perspectives
7.1. Zebrafish: Mechanistic Genetics of Taste Bud Ontogeny
7.2. Cichlid Fish: Coevolution of Taste and Dental Structures
7.3. Sharks: Evolutionary Origin of the SOX2+ Oral Progenitor
7.4. Organoid Platforms: Bridging Genetics and Physiology
7.5. Single-Cell Genomics and Atlas Approaches
7.6. Metabolic Disease, Viral Infection, and Emerging Modulators of Taste Homeostasis
8. Translational Implications
8.1. Taste Dysfunction: Clinical Context and Mechanistic Framework
8.2. Pharmacological Targets for Taste Restoration
8.3. Outstanding Questions
9. Conclusions
Funding
References
- Roper, S.D.; Chaudhari, N. Taste buds: cells, signals and synapses. Nat. Rev. Neurosci. 2017, 18, 485–497. [Google Scholar] [CrossRef]
- Finger, T.E.; Kinnamon, S.C. Taste isn’t just for taste buds anymore. F1000 Biol. Rep. 2011, 3, 20. [Google Scholar] [CrossRef] [PubMed]
- Barlow, L.A. Progress and renewal in gustation: new insights into taste bud development. Development 2015, 142, 3620–3629. [Google Scholar] [CrossRef] [PubMed]
- Witt, M. Anatomy and development of the human taste system. Handb. Clin. Neurol. 2019, 164, 147–171. [Google Scholar] [CrossRef]
- Mistretta, C.M.; Kumari, A. Hedgehog Signaling Regulates Taste Organs and Oral Sensation: Distinctive Roles in the Epithelium, Stroma, and Innervation. Int. J. Mol. Sci. 2019, 20, 1341. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; de Araujo, A.M.; Xi, R.; Lin, X.; Lu, C.; Zhou, M.; Hankenson, K.; Margolskee, R.F.; Matsumoto, I.; de Lartigue, G.; et al. Gustatory-neuron-supplied R-spondin-2 is required for taste bud replenishment. Stem Cell Rep. 2025, 20, 102542. [Google Scholar] [CrossRef]
- Rothova, M.; Thompson, H.; Lickert, H.; Tucker, A.S. Lineage tracing of the endoderm during oral development. Dev. Dyn. 2012, 241, 1183–1191. [Google Scholar] [CrossRef]
- Qin, Y.; Sukumaran, S.K.; Margolskee, R.F. Nkx2-2 expressing taste cells in endoderm-derived taste papillae are committed to the type III lineage. Dev. Biol. 2021, 477, 232–240. [Google Scholar] [CrossRef]
- Chaudhari, N.; Roper, S.D. The cell biology of taste. J. Cell Biol. 2010, 190, 285–296. [Google Scholar] [CrossRef]
- Huang, A.L.; Chen, X.; Hoon, M.A.; Chandrashekar, J.; Guo, W.; Tränkner, D.; Ryba, N.J.P.; Zuker, C.S. The cells and logic for mammalian sour taste detection. Nature 2006, 442, 934–938. [Google Scholar] [CrossRef]
- Okubo, T.; Clark, C.; Hogan, B.L.M. Cell lineage mapping of taste bud cells and keratinocytes in the mouse tongue and soft palate. Stem Cells 2009, 27, 442–450. [Google Scholar] [CrossRef] [PubMed]
- Thirumangalathu, S.; Harlow, D.E.; Driskell, A.L.; Krimm, R.F.; Barlow, L.A. Fate mapping of mammalian embryonic taste bud progenitors. Development 2009, 136, 1519–1528. [Google Scholar] [CrossRef]
- Hansen, A.; Reutter, K.; Zeiske, E. Taste bud development in the zebrafish, Danio rerio. Dev. Dyn. 2002, 223, 483–496. [Google Scholar] [CrossRef]
- Bloomquist, R.F.; Parnell, N.F.; Phillips, K.A.; Fowler, T.E.; Yu, T.Y.; Sharpe, P.T.; Streelman, J.T. Coevolutionary patterning of teeth and taste buds. Proc. Natl. Acad. Sci. USA 2015, 112, E5954–E5962. [Google Scholar] [CrossRef]
- Martin, K.J.; Rasch, L.J.; Cooper, R.L.; Metscher, B.D.; Johanson, Z.; Fraser, G.J. Sox2+ progenitors in sharks link taste development with the evolution of regenerative teeth from denticles. Proc. Natl. Acad. Sci. USA 2016, 113, 14769–14774. [Google Scholar] [CrossRef]
- Ren, W.; Lewandowski, B.C.; Watson, J.; Aihara, E.; Iwatsuki, K.; Bachmanov, A.A.; Margolskee, R.F.; Jiang, P. Single Lgr5- or Lgr6-expressing taste stem/progenitor cells generate taste bud cells ex vivo. Proc. Natl. Acad. Sci. USA 2014, 111, 16401–16406. [Google Scholar] [CrossRef]
- Adpaikar, A.A.; Zhang, S.; Kim, H.Y.; Kim, K.W.; Moon, S.J.; Lee, J.M.; Jung, H.S. Fine-tuning of epithelial taste bud organoid to promote functional recapitulation of taste reactivity. Cell. Mol. Life Sci. 2022, 79, 211. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Verweij, L.H.G.; Lin, L.; van Es, J.H.; Slack, J.; Winkel, C.; et al. Organoid Modeling of Mouse Anterior Tongue Epithelium Reveals Regional and Cellular Identities. Adv. Sci. 2025, 12, e06738. [Google Scholar] [CrossRef] [PubMed]
- Mistretta, C.M.; Liu, H.X. Development of fungiform papillae: patterned lingual gustatory organs. Arch. Histol. Cytol. 2006, 69, 199–208. [Google Scholar] [CrossRef]
- Jung, H.S.; Oropeza, V.; Thesleff, I. Shh, Bmp-2, Bmp-4 and Fgf-8 are associated with initiation and patterning of mouse tongue papillae. Mech. Dev. 1999, 81, 179–182. [Google Scholar] [CrossRef]
- Hall, J.M.H.; Bell, M.L.; Finger, T.E. Disruption of sonic hedgehog signaling alters growth and patterning of lingual taste papillae. Dev. Biol. 2003, 255, 263–277. [Google Scholar] [CrossRef]
- Liu, H.X.; Ermilov, A.; Grachtchouk, M.; Li, L.; Gumucio, D.L.; Dlugosz, A.A.; Mistretta, C.M. Multiple Shh signaling centers participate in fungiform papilla and taste bud formation and maintenance. Dev. Biol. 2013, 382, 82–97. [Google Scholar] [CrossRef] [PubMed]
- Iwatsuki, K.; Liu, H.X.; Gröner, A.; Singer, M.A.; Lane, T.F.; Grosschedl, R.; Mistretta, C.M.; Margolskee, R.F. Wnt signaling interacts with Shh to regulate taste papilla development. Proc. Natl. Acad. Sci. USA 2007, 104, 2253–2258. [Google Scholar] [CrossRef]
- Okubo, T.; Pevny, L.H.; Hogan, B.L.M. Sox2 is required for development of taste bud sensory cells. Genes Dev. 2006, 20, 2654–2659. [Google Scholar] [CrossRef] [PubMed]
- Castillo-Azofeifa, D.; Seidel, K.; Gross, L.; Golden, E.J.; Jacquez, B.; Klein, O.D.; Barlow, L.A. SOX2 regulation by hedgehog signaling controls adult lingual epithelium homeostasis. Development 2018, 145, dev164889. [Google Scholar] [CrossRef]
- Shechtman, L.A.; Scott, J.K.; Larson, E.D.; Isner, T.J.; Johnson, B.J.; Gaillard, D.; Dempsey, P.J.; Barlow, L.A. High Sox2 expression predicts taste lineage competency of lingual progenitors in vitro. Development 2023, 150, dev201375. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.X.; Komatsu, Y.; Mishina, Y.; Mistretta, C.M. Neural crest contribution to lingual mesenchyme, epithelium and developing taste papillae and taste buds. Dev. Biol. 2012, 368, 294–303. [Google Scholar] [CrossRef]
- Ishan, M.; Wang, Z.; Zhao, P.; Yao, Y.; Stice, S.L.; Wells, L.; Mishina, Y.; Liu, H.X. Taste papilla cell differentiation requires the regulation of secretory protein production by ALK3-BMP signaling in the tongue mesenchyme. Development 2023, 150, dev201838. [Google Scholar] [CrossRef]
- Golden, E.J.; Larson, E.D.; Shechtman, L.A.; Trahan, G.D.; Gaillard, D.; et al. Onset of taste bud cell renewal starts at birth and coincides with a shift in SHH function. eLife 2021, 10, e64013. [Google Scholar] [CrossRef]
- Lin, X.; Lu, C.; Ohmoto, M.; Choma, K.; Margolskee, R.F.; Matsumoto, I.; Jiang, P. R-spondin substitutes for neuronal input for taste cell regeneration in adult mice. Proc. Natl. Acad. Sci. USA 2021, 118, e2001833118. [Google Scholar] [CrossRef]
- Yee, K.K.; Li, Y.; Redding, K.M.; Iwatsuki, K.; Margolskee, R.F.; Jiang, P. Lgr5-EGFP marks taste bud stem/progenitor cells in posterior tongue. Stem Cells 2013, 31, 992–1000. [Google Scholar] [CrossRef] [PubMed]
- Takeda, N.; Jain, R.; Li, D.; Li, L.; Lu, M.M.; Epstein, J.A. Lgr5 identifies progenitor cells capable of taste bud regeneration after injury. PLoS ONE 2013, 8, e66314. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Seo, D.W.; Shim, J.; Lee, J.S.; Choi, S.H.; Kim, D.H.; Moon, S.J.; Jung, H.S.; Jeong, Y.T. Reassessing the genetic lineage tracing of lingual Lgr5+ and Lgr6+ cells in vivo. Anim. Cells Syst. 2024, 28, 353–366. [Google Scholar] [CrossRef]
- Verweij, L.H.G.; Kim, S.Y.; Laskaris, D.; Lin, L.; van Son, G.J.F.; et al. Tripotent Lgr5 stem cells in the posterior tongue generate lingual, taste, and salivary gland lineages. Nat. Commun. 2025, 16, 10266. [Google Scholar] [CrossRef]
- Matsumoto, I.; Ohmoto, M.; Narukawa, M.; Yoshihara, Y.; Abe, K. Skn-1a (Pou2f3) specifies taste receptor cell lineage. Nat. Neurosci. 2011, 14, 685–687. [Google Scholar] [CrossRef] [PubMed]
- Ohmoto, M.; Jyotaki, M.; Yee, K.K.; Matsumoto, I. A Transcription Factor Etv1/Er81 Is Involved in the Differentiation of Sweet, Umami, and Sodium Taste Cells. eNeuro 2023 2023, 10, ENEURO.0236–22. [Google Scholar] [CrossRef]
- Seta, Y.; Oda, M.; Kataoka, S.; Toyono, T.; Toyoshima, K. Mash1 is required for the differentiation of AADC-positive type III cells in mouse taste buds. Dev. Dyn. 2011, 240, 775–784. [Google Scholar] [CrossRef]
- Seta, Y.; Seta, C.; Barlow, L.A. Notch-associated gene expression in embryonic and adult taste papillae and taste buds suggests a role in taste cell lineage decisions. J. Comp. Neurol. 2003, 464, 49–61. [Google Scholar] [CrossRef]
- Ota, M.S.; Kaneko, Y.; Kondo, K.; Ogishima, S.; Tanaka, H.; Eto, K.; Kondo, T. Combined In Silico and In Vivo Analyses Reveal Role of Hes1 in Taste Cell Differentiation. PLoS Genet. 2009, 5, e1000443. [Google Scholar] [CrossRef]
- Castillo, D.; Seidel, K.; Salcedo, E.; Ahn, C.; de Sauvage, F.J.; Klein, O.D.; Barlow, L.A. Induction of ectopic taste buds by SHH reveals the competency and plasticity of adult lingual epithelium. Development 2014, 141, 2993–3002. [Google Scholar] [CrossRef]
- Castillo-Azofeifa, D.; Losacco, J.T.; Salcedo, E.; Golden, E.J.; Finger, T.E.; Barlow, L.A. Sonic hedgehog from both nerves and epithelium is a key trophic factor for taste bud maintenance. Development 2017, 144, 3054–3065. [Google Scholar] [CrossRef]
- Miura, H.; Scott, J.K.; Harada, S.; Barlow, L.A. Sonic hedgehog-expressing basal cells are general post-mitotic precursors of functional taste receptor cells. Dev. Dyn. 2014, 243, 1286–1297. [Google Scholar] [CrossRef]
- Ohmoto, M.; Lei, W.; Yamashita, J.; Hirota, J.; Jiang, P.; Matsumoto, I. SOX2 regulates homeostasis of taste bud cells and lingual epithelial cells in posterior tongue. PLoS ONE 2020, 15, e0240848. [Google Scholar] [CrossRef]
- Gaillard, D.; Bowles, S.G.; Salcedo, E.; Xu, M.; Millar, S.E.; Barlow, L.A. β-catenin is required for taste bud cell renewal and behavioral taste perception in adult mice. PLoS Genet. 2017, 13, e1006990. [Google Scholar] [CrossRef] [PubMed]
- Piarowski, C.M.; Isner, T.J.; Barlow, L.A. Developing and Regenerating a Sense of Taste. Curr. Top. Dev. Biol. 2025, 165, 353–404. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Lin, X.; Yamashita, J.; Xi, R.; Zhou, M.; Zhang, Y.V.; et al. RNF43/ZNRF3 negatively regulates taste tissue homeostasis. Stem Cell Rep. 2022, 17, 369–383. [Google Scholar] [CrossRef]
- Nguyen, H.M.; Reyland, M.E.; Barlow, L.A. Mechanisms of taste bud cell loss after head and neck irradiation. J. Neurosci. 2012, 32, 3474–3484. [Google Scholar] [CrossRef]
- Gaillard, D.; Shechtman, L.A.; Millar, S.E.; Barlow, L.A. Fractionated head and neck irradiation impacts taste progenitors, differentiated taste cells, and Wnt/β-catenin signaling in adult mice. Sci. Rep. 2019, 9, 17934. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Zhang, H.; Li, J.; Zheng, X.; Jia, X.; Xie, Q.; et al. LiCl Promotes Recovery of Radiation-Induced Oral Mucositis and Dysgeusia. J. Dent. Res. 2021, 100, 754–763. [Google Scholar] [CrossRef]
- Guo, Q.; Chen, S.; Rao, X.; Li, Y.; Pan, M.; Fu, G.; Yao, Y.; Gao, X.; Tang, P.; Zhou, Y.; et al. Inhibition of SIRT1 promotes taste bud stem cell survival and mitigates radiation-induced oral mucositis in mice. Am. J. Transl. Res. 2019, 11, 4789–4799. [Google Scholar] [PubMed]
- Adpaikar, A.A.; Lee, J.M.; Lee, D.J.; Cho, H.Y.; Ohshima, H.; Moon, S.J.; Jung, H.S. Epithelial plasticity enhances regeneration of committed taste receptor cells following nerve injury. Exp. Mol. Med. 2023, 55, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Kapsimali, M.; Kaushik, A.L.; Gibon, G.; Dirian, L.; Ernest, S.; Rosa, F.M. Fgf signaling controls pharyngeal taste bud formation through miR-200 and Delta-Notch activity. Development 2011, 138, 3473–3484. [Google Scholar] [CrossRef] [PubMed]
- Ren, W.; Aihara, E.; Lei, W.; Gheewala, N.; Uchiyama, H.; Margolskee, R.F.; Iwatsuki, K.; Jiang, P. Transcriptome analyses of taste organoids reveal multiple pathways involved in taste cell generation. Sci. Rep. 2017, 7, 4004. [Google Scholar] [CrossRef]
- Ren, W.; Li, W.; Cha, X.; Wang, S.; Cai, B.; Wang, T.; et al. Single-cell transcriptomic atlas of taste papilla aging. Aging Cell 2024, 23, e14308. [Google Scholar] [CrossRef]
- Kaufman, A.; Choo, E.; Koh, A.; Dando, R. Inflammation arising from obesity reduces taste bud abundance and inhibits renewal. PLoS Biol. 2018, 16, e2001959. [Google Scholar] [CrossRef]
- Doval-Caballero, J.L.E.; Ferreira-Hermosillo, A.; Eugenio-Ponce, G.D.; García-Sáenz, M.R.; Ibarra-Salce, R.; Tenorio-Rojo, A.P.; Luna-Avila, E.S.; Gete-Palacios, P.C.; Pérez-Hernández, F.; Rojas-Milán, E.; et al. Potential role of glucagon-like peptide-1 in taste receptors. Front. Endocrinol. 2026, 16, 1683419. [Google Scholar] [CrossRef]
- Yao, Q.; Doyle, M.E.; Liu, Q.R.; Appleton, A.; O'Connell, J.F.; Weng, N.P.; Egan, J.M. Long-Term Dysfunction of Taste Papillae in SARS-CoV-2. NEJM Evid. 2023, 2, EVIDoa2300046. [Google Scholar] [CrossRef]
- Morad, H.; Vanhala, T.; Kisiel, M.A.; Andreason, A.; Li, M.; Andersson, G.; Laurell, G.; Finger, T.E.; Hellekant, G. Taste dysfunction in long COVID. Chem. Senses 2026, 51, bjaf068. [Google Scholar] [CrossRef] [PubMed]
- Seubert, A.C.; Krafft, M.; Bopp, S.; Helal, M.; Bhandare, P.; Wolf, E.; Alemany, A.; Riedel, A.; Kretzschmar, K. Spatial transcriptomics reveals molecular cues underlying the site specificity of the adult mouse oral mucosa and its stem cell niches. Stem Cell Rep. 2024, 19, 1706–1719. [Google Scholar] [CrossRef]
- Matsu-Ura, T.; Nasu, A.; Lee, S.; Yoshida, N. Circadian clock–gated cell renewal controls time-dependent changes in taste sensitivity. Proc. Natl. Acad. Sci. USA 2025, 122, e2421421122. [Google Scholar] [CrossRef]
- Wang, J.; Liu, R.; Ma, H.; Zhang, W. The Pathogenesis of COVID-19–Related Taste Disorder and Treatments. J. Dent. Res. 2023, 102, 1191–1198. [Google Scholar] [CrossRef]
- Nosrat, C.A.; Blomlöf, J.; ElShamy, W.M.; Ernfors, P.; Olson, L. Lingual deficits in BDNF and NT3 mutant mice leading to gustatory and somatosensory disturbances, respectively. Development 1997, 124, 1333–1342. [Google Scholar] [CrossRef] [PubMed]
- Vercauteren Drubbel, A.; Beck, B. Single-cell transcriptomics uncovers the differentiation of a subset of murine esophageal progenitors into taste buds in vivo. Sci. Adv. 2023, 9, eadd9135. [Google Scholar] [CrossRef] [PubMed]
- Ki, S.Y.; Jang, J.H.; Kim, D.H.; Jeong, Y.T. c-Kit signaling confers damage-resistance to sweet taste cells upon nerve injury. Int. J. Oral. Sci. 2025, 17, 57. [Google Scholar] [CrossRef] [PubMed]
- Piarowski, C.M.; Barlow, L.A. Tyrosine kinase inhibitors affect sweet taste and dysregulate fate selection of specific taste bud cell subtypes via KIT inhibition. Development 2026, 153, dev205259. [Google Scholar] [CrossRef]
| Organoid System | Starting Cell(s) | Species | Culture Conditions | Key Findings | Ref. |
| LGR5/LGR6-derived taste organoids |
Single LGR5⁺ or LGR6⁺ cells from CVP |
Mouse | Matrigel; WNT3a, RSPO1, Noggin, EGF |
First demonstration that single taste stem cells generate all TRC types ex vivo; established clonal organoid culture | [16] |
| Anterior tongue epithelial organoids |
Anterior tongue epithelium (non-taste) |
Mouse | Modified intestinal organoid protocol |
Revealed regional and cellular identities; anterior tongue epithelium can form organoids distinct from CVP-derived ones | [18] |
| Fine-tuned taste bud organoids |
CVP epithelial cells | Mouse | Optimized with BMP inhibition, FGF10 |
Enhanced functional recapitulation of taste reactivity; improved Type II and Type III cell differentiation | [17] |
| Transcriptome-profiled taste organoids |
LGR5⁺/LGR6⁺ cells | Mouse | Standard taste organoid medium |
RNA-seq revealed multiple pathways (WNT, Notch, SHH) involved in taste cell generation within organoids | [53] |
| SOX2-high progenitor organoids |
SOX2-high lingual progenitors |
Mouse | WNT/RSPO-based medium |
High SOX2 expression predicts taste lineage competency in vitro; SOX2-low cells form non-taste epithelium | [26] |
| Tripotent posterior tongue organoids |
Single LGR5⁺ cells from posterior tongue |
Mouse | WNT/RSPO-enriched medium |
Single LGR5⁺ stem cell generates taste bud, salivary gland, and lingual epithelial lineages | [34] |
| Gene/Molecule | Class | Stage | Function in Taste | Key Ref. |
|---|---|---|---|---|
| SHH | Morphogen | Embryonic → Adult | Placode marker; papilla patterning; postnatal switch to pro-differentiation; adult homeostatic requirement; maintains SOX2 | [21,22,29,41] |
| WNT/β-catenin | Signaling pathway |
Embryonic → Adult | Upstream of placodal Shh expression; embryonic papilla patterning; adult progenitor renewal; β-catenin required for taste cell turnover | [23,44] |
| SOX2 | HMG-box TF | Embryonic → Adult | Required for taste bud sensory cell development; maintained by SHH in adults; SOX2-high state predicts taste competency; regulates posterior tongue homeostasis | [24,25,26] |
| LGR5 | WNT target GPCR |
Postnatal → Adult | Marks taste bud stem/progenitor cells in posterior tongue; capable of taste bud regeneration after injury; tripotent in posterior tongue | [31,32,34] |
| LGR6 | WNT target GPCR |
Postnatal → Adult | Marks taste stem/progenitor cells; generates taste bud cells ex vivo alongside LGR5; anterior tongue organoid formation | [16,18] |
| RSPO2 | WNT amplifier |
Adult | Gustatory-neuron-supplied niche signal; required for taste bud replenishment; substitutes for neuronal input in regeneration | [6,30] |
| RNF43/ZNRF3 | Ub ligases | Adult | Negative regulators of WNT; loss leads to expanded taste tissue; RSPO blocks their Frizzled-degrading activity to amplify WNT | [46] |
| POU2F3/ SKN-1a |
POU TF | Postnatal → Adult | Master selector for Type II (sweet/umami/bitter) taste receptor cell lineage; absence eliminates all Type II cells | [35] |
| NKX2-2 | HD TF | Postnatal → Adult | Expressed in endoderm-derived (CVP/FoP) taste cells; committed to Type III lineage; posterior endoderm only | [8] |
| ASCL1/MASH1 | bHLH TF | Postnatal → Adult | Type III differentiation; proneural; absence reduces AADC⁺ Type III cells; gated by Notch lateral inhibition | [37] |
| HES1 | Notch target | All stages | Timing gate; maintains progenitor state and constrains premature differentiation; may influence POU2F3/ASCL1 balance | [38,39] |
| BMP4/ALK3 | BMP/receptor | Embryonic | Mesenchymal BMP signaling via ALK3 required for taste papilla cell differentiation; regulates secretory protein production | [28] |
| FGF8 | Growth factor | Embryonic | Expressed in inter-papilla zones; inhibited by SHH; contributes to placode spacing pattern | [20] |
| ETV1 | ETS TF | Postnatal → Adult | Refines Type II subtype identity; involved in differentiation of sweet, umami, and sodium taste cells; downstream of POU2F3 | [36] |
| FOXA1/FOXA2 | Forkhead TF | Birth → Adult | Expressed in postmitotic taste precursors at birth; may contribute to early differentiation programs | [29] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




