Submitted:
01 May 2026
Posted:
05 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Conceptual Framework and Review Approach
2.1. Article Type and Methodological Note
2.2. Evidence Synthesis Workflow and Literature Scope
3. Microbial Communication as a Regulatory Layer in Cocoa Pulp Juice Fermentation
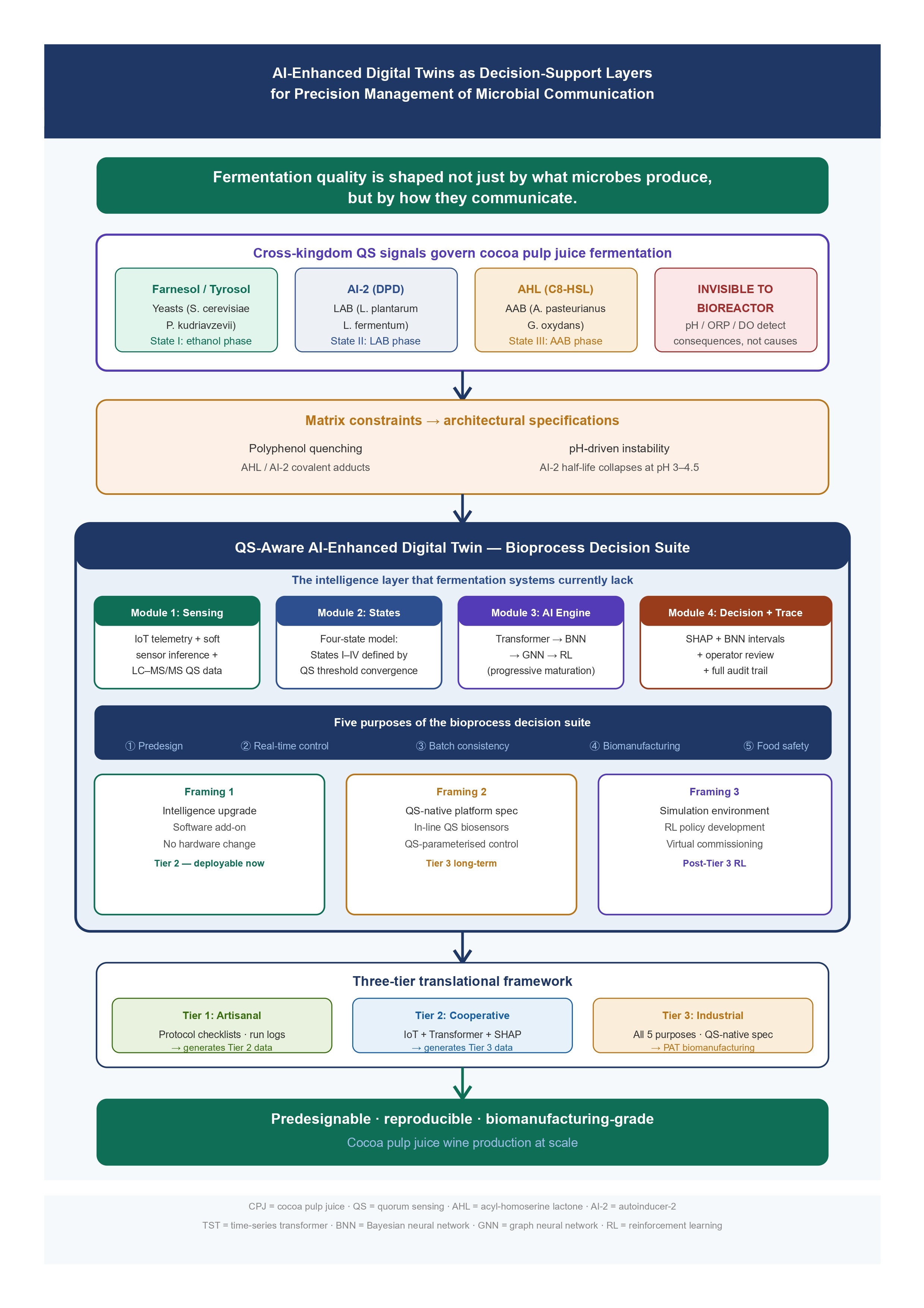
3.1. Quorum Sensing Architectures Relevant to Cocoa Ecosystems
3.2. Cross-Kingdom Signalling, Emergent Community Behaviour, and Quality Governance
3.3. The Informational Lead-Time Advantage over Bioreactor Monitoring
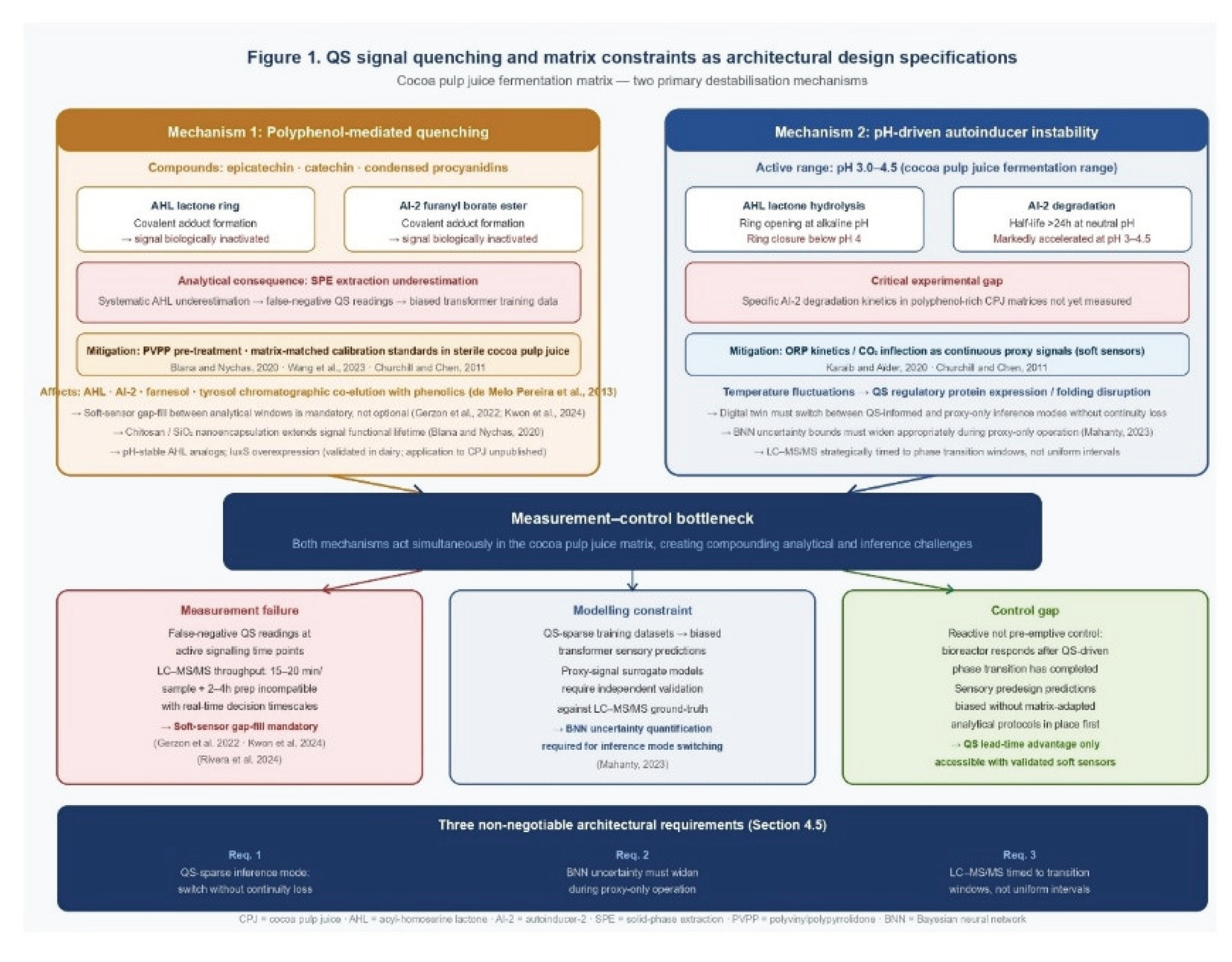
4. Signal Quenching, Matrix Constraints, and the Measurement Gap
4.1. Polyphenol-Mediated Quenching
4.2. pH- and Temperature-Driven Instability
4.3. Throughput Constraints and the Proxy-Signal Imperative
4.4. Mitigation Strategies and their Roles in the Architecture
4.5. Architectural Specifications Derived from Matrix Constraints
5. Molecular and Biosensing Toolkits for QS-Guided Fermentation Control
5.1. The Measurement–Control Interface
5.2. Analytical Platforms
5.2.1. LC–MS/MS as the Reference Standard and Transformer Training Label Source
5.2.2. Whole-Cell Biosensors and In-Line Sensing Prospects
5.2.3. Multi-Omics Integration for Sensory–QS Model Linkage
5.3. Signal Engineering and its Role in the Bioreactor–Digital Twin System
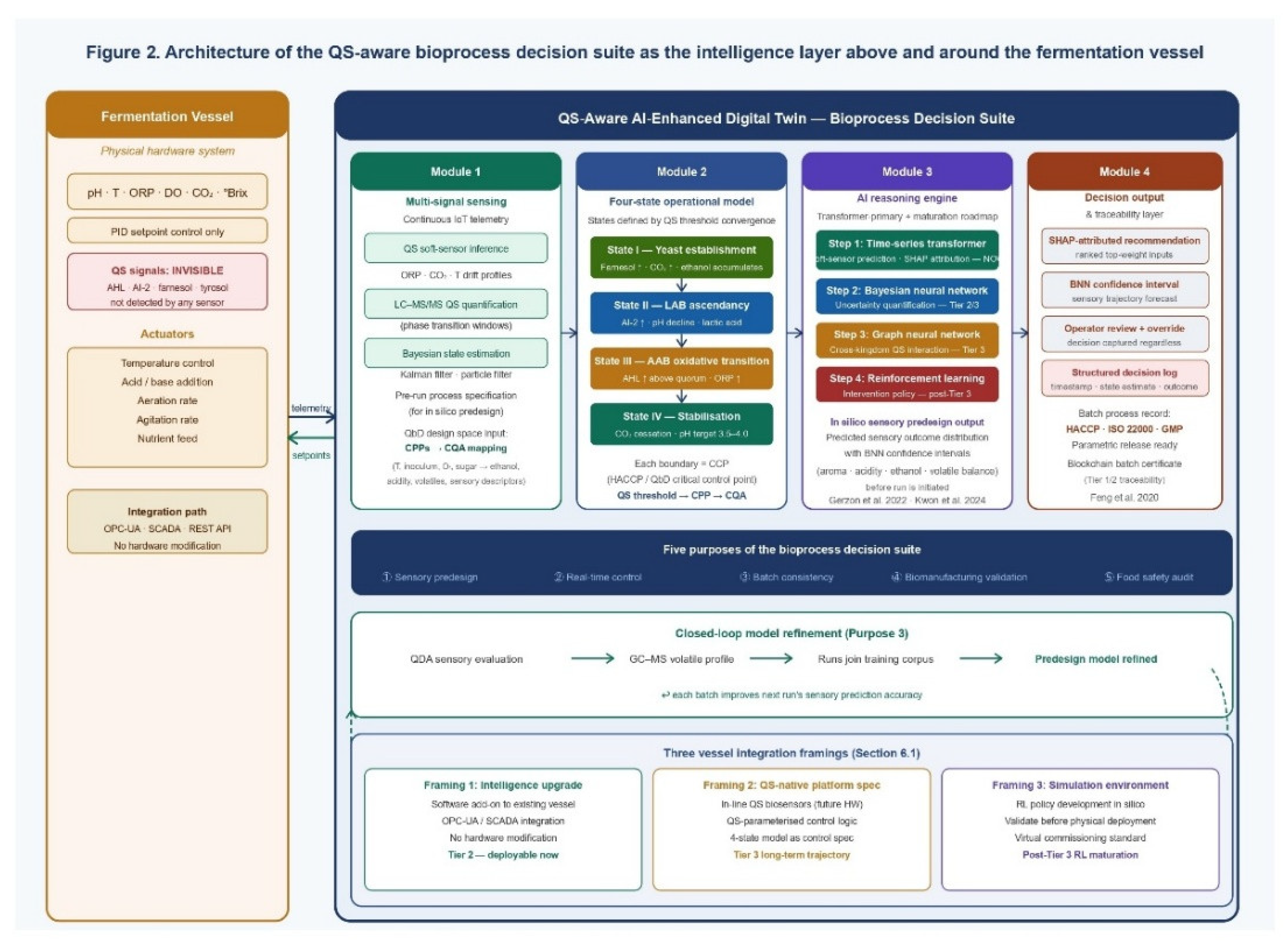
6. The Bioprocess Decision Suite: Architecture, Relationship to Fermentation Vessels, and Operational Purposes
6.1. The Digital Twin and the Bioreactor: A Precise Juxtaposition
6.2. Purpose Architecture: Five Operational Functions of the Bioprocess Decision Suite
6.3. Module Architecture: Four Interdependent Components
6.3.1. Module 1 — Multi-Signal Sensing and Data Ingestion
6.3.2. Module 2 — Four-State Operational Model
- State I — Yeast establishment: ethanol accumulation, CO₂ evolution, rising farnesol. Predesign role: ethanol concentration and ester-alcohol foundation of aroma profile established. Quality risk: insufficient ethanol; early contamination.
- State II — LAB ascendancy: AI-2 accumulation, progressive acidification, lactic acid production. Predesign role: organic acid balance and pH trajectory shaped. Quality risk: under- or over-acidification; insufficient spoilage exclusion.
- State III — AAB oxidative transition: AHL accumulation above quorum threshold, ORP rise, acetic acid production, flavour volatile emergence. Predesign role: volatile acidity and aroma complexity determined. Quality risk: over-acidification if QS-mediated gating fails.
- State IV — Stabilisation and endpoint: CO₂ cessation, target pH and volatile profile achieved. Quality decision: accept, continue maturation, or intervene.
6.3.3. Module 3 — Transformer-Primary AI Reasoning Engine with Progressive Maturation Roadmap
6.3.4. Module 4 — Decision Output and Traceability
6.3.5. Worked Example: Predesign-to-Production Cycle with Bioreactor Integration
7. Translational Implementation: From Artisanal Settings to QS-Native Biomanufacturing
7.1. Matching Decision Suite Capability to Production Context and AI Maturation Stage
7.2. Tier 1 — Structured Observation and Data Collection (Artisanal and Smallholder Contexts)
7.3. Tier 2 — Telemetry-Enabled Monitoring with Transformer Intelligence Layer (Cooperative and Semi-industrial Contexts)
7.4. Tier 3 — Full Bioprocess Decision Suite with QS-Informed Biomanufacturing Control and QS-Native Platform Specification
7.5. Producer Data Sovereignty and Microbial Terroir
8. Conclusions
9. Future Perspectives
Author Contributions
Funding
Acknowledgments
Abbreviations
References
- Abdurrahman, O.; Ferrari, R. Digital twin applications in the food industry: a review. Front. Sustain. Food Syst. 2025. [Google Scholar] [CrossRef]
- Albuquerque, P.; Casadevall, A. Quorum sensing in fungi: a review. Med. Mycol. 2012, 50, 337–345. [Google Scholar] [CrossRef]
- Almeida, O.G.G.; Pinto, U.M.; Matos, C.B.; Frazilio, D.A.; Braga, V.F.; von Zeska-Kress, M.R.; De Martinis, E.C.P. Does quorum sensing play a role in microbial shifts along spontaneous fermentation of cocoa beans? An in silico perspective. Food Res. Int. 2020, 131, 109034. [Google Scholar] [CrossRef] [PubMed]
- Almeida, O.G.G.; Vitulo, N.; De Martinis, E.C.P.; Felis, G.E. Pangenome analyses of LuxS-coding genes and enzymatic repertoires in cocoa-related lactic acid bacteria. Genomics 2021, 113, 1659–1670. [Google Scholar] [CrossRef]
- Almeida, O.G.G.; Pereira, M.G.; Bighetti-Trevisan, R.L.; Santos, E.S.; De Campos, E.G.; Felis, G.E.; Guimarães, L.H.S.; Polizeli, M.L.T.M.; De Martinis, B.S.; De Martinis, E.C.P. Investigating luxS gene expression in lactobacilli along lab-scale cocoa fermentations. Food Microbiol. 2024, 119, 104429. [Google Scholar] [CrossRef]
- Blana, V.A.; Lianou, A.; Nychas, G.-J.E. Quorum sensing and microbial ecology of foods. In Quantitative Microbiology in Food Processing: Modeling the Microbial Ecology; Sant’Ana, A.S., Ed.; Wiley: Chichester, 2017; pp. 600–613. [Google Scholar] [CrossRef]
- Campos, S.D.M.; Martínez-Burgos, W.J.; dos Reis, G.A.; Ocán-Torres, D.Y.; dos Santos Costa, G.; Rosas Vega, F.; Lima Serra, J.; Soccol, C.R. The role of microbial dynamics, sensorial compounds, and producing regions in cocoa fermentation. Microbiol. Res. 2025, 16, 75. [Google Scholar] [CrossRef]
- Chen, H.; Fink, G.R. Feedback control of morphogenesis in fungi by aromatic alcohols. Genes Dev. 2006, 20, 1150–1161. [Google Scholar] [CrossRef]
- Churchill, M.E.A.; Chen, L. Structural basis of acyl-homoserine lactone-dependent signaling. Chem. Rev. 2011, 111, 68–85. [Google Scholar] [CrossRef] [PubMed]
- De Vuyst, L.; Weckx, S. The cocoa bean fermentation process: from ecosystem analysis to starter culture development. J. Appl. Microbiol. 2016, 121, 5–17. [Google Scholar] [CrossRef]
- de Melo Pereira, G.V.; Miguel, M.G.C.P.; Ramos, C.L.; Schwan, R.F. Microbiological and physicochemical characterization of small-scale cocoa fermentations and screening of yeast and bacterial strains to develop a defined starter culture. Appl. Environ. Microbiol. 2012, 78, 5395–5405. [Google Scholar] [CrossRef]
- Feng, H.; Wang, X.; Duan, Y.; Zhang, J.; Zhang, X. Applying blockchain technology to improve agri-food traceability: a review of development methods, benefits and challenges. J. Clean. Prod. 2020, 260, 121031. [Google Scholar] [CrossRef]
- Ferrocino, I.; Rantsiou, K.; Cocolin, L. Microbiome and -omics application in food industry. Int. J. Food Microbiol. 2022, 377, 109781. [Google Scholar] [CrossRef]
- Gerzon, G.; Sheng, Y.; Kirkitadze, M. Process analytical technologies – advances in bioprocess integration and future perspectives. J. Pharm. Biomed. Anal. 2022, 207, 114379. [Google Scholar] [CrossRef]
- Giaouris, E.; Heir, E.; Desvaux, M.; Hébraud, M.; Møretrø, T.; Langsrud, S.; et al. Intra- and inter-species interactions within biofilms of important foodborne bacterial pathogens. Front. Microbiol. 2015, 6, 841. [Google Scholar] [CrossRef]
- Jin, X.; Liu, C.; Xu, T.; Su, L.; Zhang, X. Artificial intelligence biosensors: challenges and prospects. Biosens. Bioelectron. 2020, 165, 112412. [Google Scholar] [CrossRef] [PubMed]
- Kareb, O.; Aïder, M. Quorum sensing circuits in the communicating mechanisms of bacteria and its implication in the biosynthesis of bacteriocins by lactic acid bacteria: a review. Probiotics Antimicrob. Proteins 2020, 12, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.J.; Shiu, J.H.; Yamakawa, C.K.; Rivera, E.C. Enhancing fermentation process monitoring through data-driven modeling and synthetic time series generation. Bioengineering 2024, 11, 803. [Google Scholar] [CrossRef] [PubMed]
- Mahanty, B. Hybrid modeling in bioprocess dynamics: structural variabilities, implementation strategies, and practical challenges. Biotechnol. Bioeng. 2023, 120, 2072–2091. [Google Scholar] [CrossRef]
- Niyigaba, T.; Küçüköz, K.; Kołożyn-Krajewska, D.; Królikowski, T.; Trząskowska, M. Advances in fermentation technology: a focus on health and safety. Appl. Sci. 2025, 15, 3001. [Google Scholar] [CrossRef]
- Rivera, E.C.; Yamakawa, C.K.; Rossell, C.E.V.; Nolasco, J.; Kwon, H.J. Prediction of intensified ethanol fermentation of sugarcane using a deep learning soft sensor and process analytical technology. J. Chem. Technol. Biotechnol. 2024, 99, 207–216. [Google Scholar] [CrossRef]
- Rutherford, S.T.; Bassler, B.L. Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harb. Perspect. Med. 2012, 2, a012427. [Google Scholar] [CrossRef] [PubMed]
- Schwan, R.F.; Wheals, A.E. The microbiology of cocoa fermentation and its role in chocolate quality. Crit. Rev. Food Sci. Nutr. 2004, 44, 205–221. [Google Scholar] [CrossRef] [PubMed]
- Treloar, N.J.; Fedorec, A.J.H.; Ingalls, B.; Barnes, C.P. Deep reinforcement learning for the control of microbial co-cultures in bioreactors. PLoS Comput. Biol. 2020, 16, e1007783. [Google Scholar] [CrossRef]
- Valastyan, J.S.; Kraml, C.M.; Pelczer, I.; Ferrante, T.; Bassler, B.L. Saccharomyces cerevisiae requires CFF1 to produce 4-hydroxy-5-methylfuran-3(2H)-one, a mimic of the bacterial quorum-sensing autoinducer AI-2. mBio 2021, 12, e03303-20. [Google Scholar] [CrossRef]
- Wang, D.; Fan, F.; Qin, Y.; et al. Quorum-quenching enzymes: promising bioresources and their opportunities and challenges as alternative bacteriostatic agents in food industry. Compr. Rev. Food Sci. Food Saf. 2023, 22, 1104–1127. [Google Scholar] [CrossRef]
- Yee, C.S.; Zahia-Azizan, N.A.; Abd Rahim, M.H.; Mohd Zaini, N.A.; Raja-Razali, R.B.; Ushidee-Radzi, M.A.; Ilham, Z.; Wan-Mohtar, W.A.A.Q.I. Smart fermentation technologies: microbial process control in traditional fermented foods. Fermentation 2025, 11, 323. [Google Scholar] [CrossRef]
- Zhao, S.G.; Jiao, T.; Adade, S.Y.S.S.; Wang, Z.; Ouyang, Q.; Chen, Q. Digital twin for predicting and controlling food fermentation: a case study of kombucha fermentation. J. Food Eng. 2025, 393, 112467. [Google Scholar] [CrossRef]
| Signal Class | Primary Producer(s) | Key Regulated Behaviours | Quality Consequence in Cocoa Wine | Invisible to Bioreactor’s Standard Sensor Suite? | Fermentation State |
| AHLs (C8-HSL, C10-HSL) | A. pasteurianus, G. oxydans | Oxidative metabolism onset, biofilm at O₂ interface, alcohol dehydrogenase upregulation | Acetic acid concentration; volatile acidity profile; over-acidification risk | Yes — ORP registers consequence hours later | III–IV |
| AI-2 (DPD) | L. plantarum, L. fermentum | Acid stress response, biofilm, substrate switching, bacteriocin induction | Lactic acid profile; pH trajectory; spoilage suppression | Yes — pH registers consequence after LAB expansion | II–III |
| Peptide autoinducers | L. plantarum, Leuconostoc spp. | Bacteriocin production, competitive exclusion of spoilage taxa | Microbiological safety; spoilage-free shelf life | Yes — no physicochemical correlate exists | II |
| Farnesol | S. cerevisiae, P. kudriavzevii | Morphogenesis suppression, cross-kingdom competitive signalling | Ethanol yield; ester-alcohol balance; yeast-to-AAB transition timing | Yes — ethanol accumulation is the delayed consequence | I |
| Tyrosol | S. cerevisiae, P. kudriavzevii | Oxidative stress tolerance, metabolic rate modulation | LAB readiness; flavour precursor accumulation rate | Yes — no physicochemical correlate exists | I–II |
| Oxylipins | Minor fungal populations | Fungal sporulation; potential bacterial QS interference | Sporadic off-flavour risk; microbial community instability | Yes — not detectable by any standard bioreactor instrument | Sporadic |
| Dimension | Conventional Bioreactor / Fermentation Vessel | Digital Twin (Bioprocess Decision Suite) |
| What it is | Physical vessel: hardware system providing and regulating the fermentation environment | Software intelligence layer: computational reasoning system operating above the vessel |
| What it senses | pH, temperature, DO, ORP, CO₂, agitation — macroscopic physicochemical outcomes | QS signal kinetics (via LC–MS/MS + proxy surrogates), physicochemical telemetry, and historical run patterns |
| What it acts on | Deviations from physicochemical setpoints — reactive feedback control | QS-driven community state transitions — predictive, pre-emptive intervention |
| Intelligence type | Setpoint-based PID control; no awareness of biological communication | Transformer-based temporal reasoning; attention-attributed explainability; BNN uncertainty quantification |
| Relationship to QS | Unaware of QS signals; monitors only their downstream physicochemical consequences | Reads QS proxy signals; interprets community communication states; predicts quality trajectory from QS kinetics |
| Sensory prediction | None — quality is assessed only after the batch is complete | In silico predesign: predicts sensory outcome distribution before the run is initiated |
| Batch consistency | Manual parameter replication; no learning between batches | Closed-loop learning: each run refines the process–QS–sensory coupling model |
| Governance | Process log; setpoint records; deviation reports | Full audit trail: QS state estimates, AI predictions, XAI attribution, operator decisions, observed outcomes |
| Near-term integration | Existing hardware; no modification required | Software add-on: integrates with existing bioreactor sensor and control interfaces |
| Long-term trajectory | Unchanged hardware paradigm | Functional specification for QS-native bioreactor with in-line QS sensing and QS-parameterised control logic |
| Feature | Tier 1: Artisanal / Smallholder | Tier 2: Cooperative / Semi-Industrial | Tier 3: Industrial / Biomanufacturing |
| Fermentation vessel type | Open or closed artisanal tank; ceramic or food-grade vessel | Closed cooperative tank; small bioreactor; pilot-scale fermenter | Industrial stirred-tank bioreactor; commercial-scale closed fermentation system |
| Vessel integration | No integration — digital twin exists as protocol checklist and run log | Software add-on to existing vessel control interface; no hardware modification required | PAT integration via OPC-UA/SCADA; bioreactor telemetry as primary data feed; setpoint recommendations communicated to control system |
| Primary sensing | Manual pH, T, °Brix; visual/olfactory; standardised run logs | Continuous IoT telemetry: pH, T, ORP, CO₂, °Brix; cloud logging | Full telemetry + periodic LC–MS/MS QS quantification; GC–MS volatile profiling; QDA sensory data at run completion |
| AI maturation stage | None — human protocol intelligence; generates Tier 2 TST training data | Step 1: TST classifier + Kalman uncertainty; Step 2: BNN when multi-season dataset available | Steps 1–2: TST + BNN; Step 3: GNN when multi-omics corpus available; Step 4: RL post-validation → QS-native platform specification |
| Decision suite purposes | Purpose 3 (nascent): manual batch consistency tracking | Purposes 1–3 at proxy-signal resolution; Purpose 5 (partial) | All 5 purposes fully operational; biomanufacturing-grade process validation and food safety audit trail |
| Sensory predesign capability | Empirical: best-batch pattern identification from run log history | Transformer-based: predicted sensory trajectory from process parameters + telemetry | QS-informed: full sensory outcome distribution with BNN confidence intervals from QS-parameterised model |
| Governance / quality alignment | Blockchain batch certificates; cooperative quality protocol | TST alert governance policy; HACCP-compatible CCP monitoring records; cloud audit trail | Full PAT implementation; HACCP/ISO 22000/GMP process validation documentation; parametric release records |
| Long-term vessel trajectory | Unchanged vessel hardware; digital twin remains protocol-level | Vessel gains QS-aware intelligence layer through software integration | Digital twin provides functional specification for QS-native bioreactor with in-line QS sensing and QS-parameterised control logic |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




