Submitted:
01 May 2026
Posted:
04 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
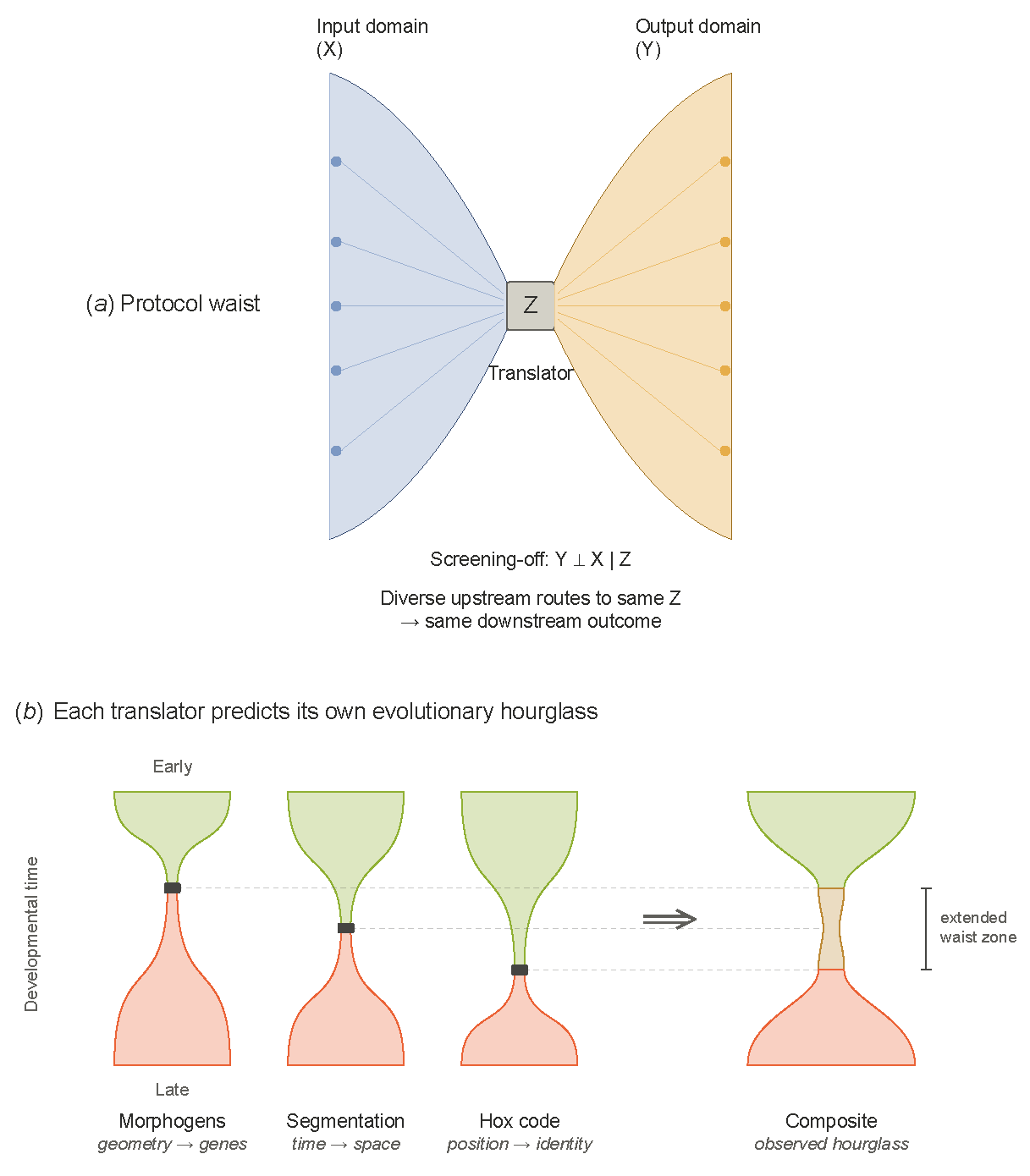
2. Protocol Waists
3. Consequences of Protocol Waists
4. Protocol Waists vs Regulatory Hubs
5. Relation to Prior Theories
6. Four Examples
6.1. Morphogen Gradients: Geometry to Genes
6.1.1. Protocol Waist Properties
6.1.2. Protocol Waist Consequences
6.2. Notch/Delta Lateral Inhibition: Topology to Fate
6.2.1. Protocol Waist Properties
6.2.2. Protocol Waist Consequences
6.3. Vertebrate Segmentation Clock: Time to Space
6.3.1. Protocol Waist Properties
6.3.2. Protocol Waist Consequences
6.4. Hox Genes: Positional History to Identity
6.4.1. Protocol Waist Properties
6.4.2. Protocol Waist Consequences
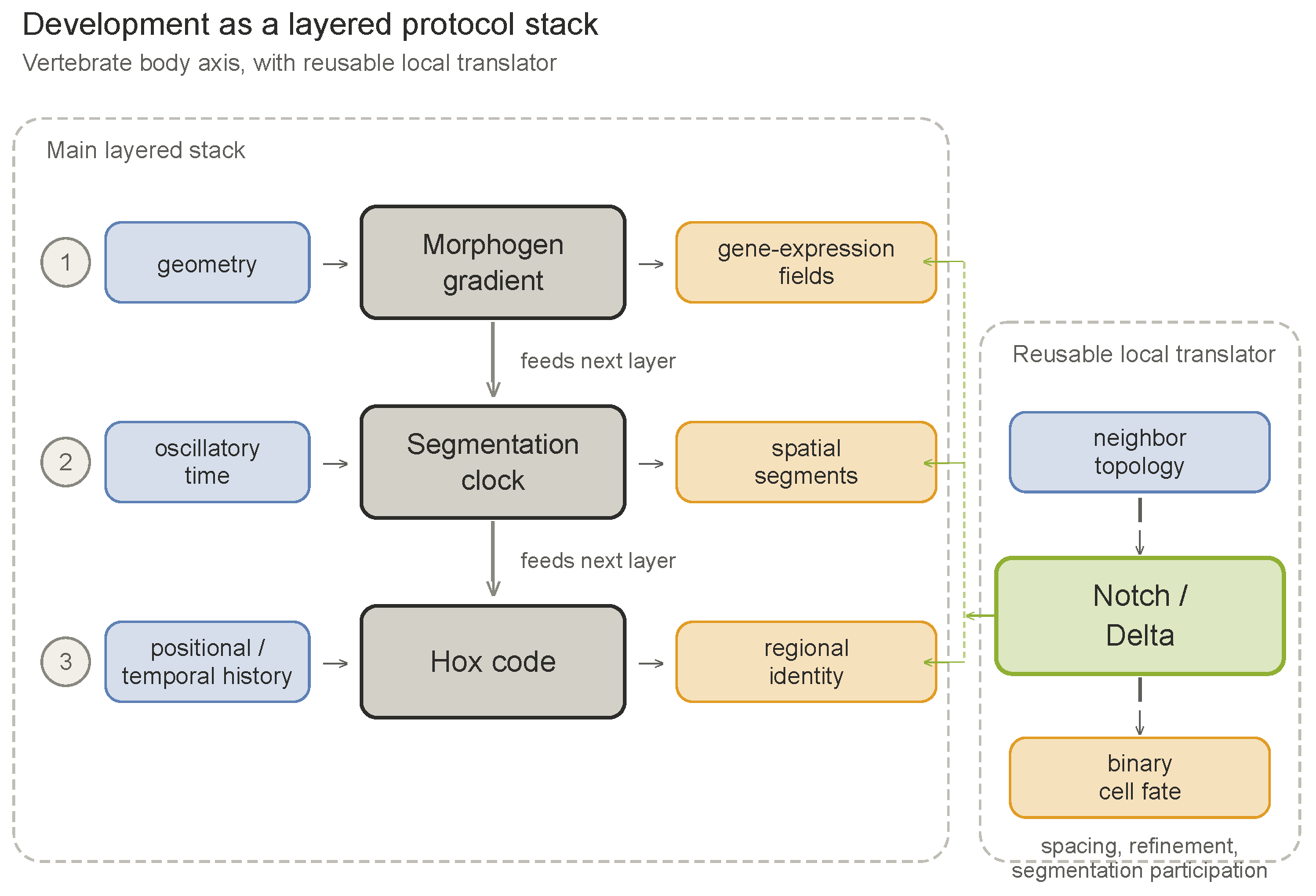
7. The Developmental Protocol Stack
8. Robust Complex Systems
9. Paradox of Robustness
10. Conclusions
Acknowledgments
References
- von Baer, K.E. Über Entwickelungsgeschichte der Thiere: Beobachtung und Reflexion; Gebrüdern Bornträger: Königsberg, 1828. [Google Scholar]
- Sander, K. The evolution of patterning mechanisms: gleanings from insect embryogenesis. In Development and Evolution; Goodwin, B.C., Holder, N., Wylie, C.C., Eds.; The Sixth Symposium of the British Society for Developmental Biology, Cambridge University Press: Cambridge, 1983; pp. 137–159. [Google Scholar]
- Duboule, D. Temporal colinearity and the phylotypic progression: a basis for the stability of a vertebrate Bauplan and the evolution of morphologies through heterochrony. Development 1994, 1994, 135–142. [Google Scholar] [CrossRef]
- Raff, R.A. The Shape of Life: Genes, Development, and the Evolution of Animal Form; University of Chicago Press: Chicago, 1996. [Google Scholar]
- Slack, J.M.W.; Holland, P.W.H.; Graham, C.F. The zootype and the phylotypic stage. Nature 1993, 361, 490–492. [Google Scholar] [CrossRef]
- Hu, H.; Uesaka, M.; Guo, S.; Shimai, K.; Lu, T.M.; Li, F.; Fujimoto, S.; Ishikawa, M.; Liu, S.; Sasagawa, Y.; et al. Constrained vertebrate evolution by pleiotropic genes. Nat. Ecol. Evol. 2017, 1, 1722–1730. [Google Scholar] [CrossRef]
- Irie, N.; Kuratani, S. Comparative transcriptome analysis reveals vertebrate phylotypic period during organogenesis. Nat. Commun. 2011, 2, 248. [Google Scholar] [CrossRef]
- Kalinka, A.T.; Varga, K.M.; Gerrard, D.T.; Preibisch, S.; Corcoran, D.L.; Jarrells, J.; Ohler, U.; Bergman, C.M.; Tomančák, P. Gene expression divergence recapitulates the developmental hourglass model. Nature 2010, 468, 811–814. [Google Scholar] [CrossRef]
- Domazet-Lošo, T.; Tautz, D. A phylogenetically based transcriptome age index mirrors ontogenetic divergence patterns. Nature 2010, 468, 815–818. [Google Scholar] [CrossRef]
- Levin, M.; Anavy, L.; Cole, A.G.; Winter, E.; Mostov, N.; Khair, S.; Senderovich, N.; Kovalev, E.; Silver, D.H.; Feder, M.; et al. The mid-developmental transition and the evolution of animal body plans. Nature 2016, 531, 637–641. [Google Scholar] [CrossRef] [PubMed]
- Ma, F.; Zheng, C. Transcriptome age of individual cell types in Caenorhabditis elegans. Proc. Natl. Acad. Sci. 2023, 120, e2216351120. [Google Scholar] [CrossRef] [PubMed]
- Quint, M.; Drost, H.G.; Gabel, A.; Ullrich, K.K.; Bönn, M.; Grosse, I. A transcriptomic hourglass in plant embryogenesis. Nature 2012, 490, 98–101. [Google Scholar] [CrossRef]
- Cheng, X.; Hui, J.H.L.; Lee, Y.Y.; Law, P.T.W.; Kwan, H.S. A “developmental hourglass” in fungi. Mol. Biol. Evol. 2015, 32, 1556–1566. [Google Scholar] [CrossRef]
- Lotharukpong, J.S.; Zheng, M.; Luthringer, R.; Liesner, D.; Drost, H.G.; Coelho, S.M. A transcriptomic hourglass in brown algae. Nature 2024, 635, 129–135. [Google Scholar] [CrossRef]
- Waddington, C.H. The Strategy of the Genes; George Allen & Unwin: London, 1957. [Google Scholar]
- Wagner, G.P.; Altenberg, L. Complex adaptations and the evolution of evolvability. Evolution 1996, 50, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Galis, F.; Metz, J.A.J. Testing the vulnerability of the phylotypic stage: on modularity and evolutionary conservation. J. Exp. Zool. 2001, 291, 195–204. [Google Scholar] [CrossRef]
- Uchida, Y.; Uesaka, M.; Yamamoto, T.; Takeda, H.; Irie, N. Embryonic lethality is not sufficient to explain hourglass-like conservation of vertebrate embryos. Evodevo 2018, 9, 7. [Google Scholar] [CrossRef] [PubMed]
- Uchida, Y.; Takeda, H.; Furusawa, C.; Irie, N. Stability in gene expression and body-plan development leads to evolutionary conservation. EvoDevo 2023, 14, 4. [Google Scholar] [CrossRef]
- Fraser, H.B.; Hirsh, A.E.; Steinmetz, L.M.; Scharfe, C.; Feldman, M.W. Evolutionary rate in the protein interaction network. Science 2002, 296, 750–752. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.; Mason, S.P.; Barabási, A.L.; Oltvai, Z.N. Lethality and centrality in protein networks. Nature 2001, 411, 41–42. [Google Scholar] [CrossRef]
- Doyle, J.C.; Alderson, D.L.; Li, L.; Low, S.; Roughan, M.; Shalunov, S.; Tanaka, R.; Willinger, W. The “robust yet fragile” nature of the Internet. Proc. Natl. Acad. Sci. 2005, 102, 14497–14502. [Google Scholar] [CrossRef]
- Doyle, J.C.; Csete, M. Architecture, constraints, and behavior. Proc. Natl. Acad. Sci. 2011, 108, 15624–15630. [Google Scholar] [CrossRef]
- Akhshabi, S.; Dovrolis, C. The evolution of layered protocol stacks leads to an hourglass-shaped architecture. In Proceedings of the Proceedings of the ACM SIGCOMM 2011 Conference, New York, NY, 2011; pp. 206–217. [Google Scholar] [CrossRef]
- Matni, N.; Ames, A.D.; Doyle, J.C. A quantitative framework for layered multirate control: Toward a theory of control architecture. IEEE Control Syst. 2024, 44, 52–94. [Google Scholar] [CrossRef]
- Csete, M.E.; Doyle, J.C. Reverse engineering of biological complexity. Science 2002, 295, 1664–1669. [Google Scholar] [CrossRef]
- Csete, M.; Doyle, J. Bow ties, metabolism and disease. Trends Biotechnol. 2004, 22, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Oda, T.; Elkahloun, A.G.; Pike, B.L.; Okajima, K.; Krantz, I.D.; Genin, A.; Piccoli, D.A.; Meltzer, P.S.; Spinner, N.B.; Collins, F.S.; et al. Mutations in the human Jagged1 gene are responsible for Alagille syndrome. Nat. Genet. 1997, 16, 235–242. [Google Scholar] [CrossRef]
- Li, L.; Krantz, I.D.; Deng, Y.; Genin, A.; Banta, A.B.; Collins, C.C.; Qi, M.; Trask, B.J.; Kuo, W.L.; Cochran, J.; et al. Alagille syndrome is caused by mutations in human Jagged1, which encodes a ligand for Notch1. Nat. Genet. 1997, 16, 243–251. [Google Scholar] [CrossRef]
- Nanni, L.; Ming, J.E.; Bocian, M.; Steinhaus, K.; Bianchi, D.W.; de Die-Smulders, C.; Giannotti, A.; Imaizumi, K.; Jones, K.L.; Del Campo, M.; et al. The mutational spectrum of the Sonic Hedgehog gene in holoprosencephaly: SHH mutations cause a significant proportion of autosomal dominant holoprosencephaly. Hum. Mol. Genet. 1999, 8, 2479–2488. [Google Scholar] [CrossRef] [PubMed]
- Reis, L.M.; Tyler, R.C.; Schilter, K.F.; Abdul-Rahman, O.; Innis, J.W.; Kozel, B.A.; Schneider, A.S.; Bardakjian, T.M.; Lose, E.J.; Martin, D.M.; et al. BMP4 loss-of-function mutations in developmental eye disorders including SHORT syndrome. Hum. Genet. 2011, 130, 495–504. [Google Scholar] [CrossRef]
- Kurban, M.; Wajid, M.; Petukhova, L.; Shimomura, Y.; Christiano, A.M. A nonsense mutation in the HOXD13 gene underlies synpolydactyly with incomplete penetrance. J. Hum. Genet. 2011, 56, 701–706. [Google Scholar] [CrossRef]
- Koenig, S.N.; LaHaye, S.; Feller, J.D.; Rowland, P.; Hor, K.N.; Trask, A.J.; Janssen, P.M.; Radtke, F.; Lilly, B.; Garg, V. Notch1 haploinsufficiency causes ascending aortic aneurysms in mice. JCI Insight 2017, 2, e91353. [Google Scholar] [CrossRef]
- Liu, J.; Viales, R.R.; Khoueiry, P.; Reddington, J.P.; Girardot, C.; Furlong, E.E.M.; Robinson-Rechavi, M. The hourglass model of evolutionary conservation during embryogenesis extends to developmental enhancers with signatures of positive selection. Genome Res. 2021, 31, 1573–1581. [Google Scholar] [CrossRef]
- Zalts, H.; Yanai, I. Developmental constraints shape the evolution of the nematode mid-developmental transition. Nat. Ecol. Evol. 2017, 1, 0113. [Google Scholar] [CrossRef] [PubMed]
- Piasecka, B.; Lichocki, P.; Moretti, S.; Bergmann, S.; Robinson-Rechavi, M. The hourglass and the early conservation models—co-existing patterns of developmental constraints in vertebrates. PLoS Genet. 2013, 9, e1003476. [Google Scholar] [CrossRef] [PubMed]
- Shubin, N.; Tabin, C.; Carroll, S. Fossils, genes and the evolution of animal limbs. Nature 1997, 388, 639–648. [Google Scholar] [CrossRef]
- De Robertis, E.M. Spemann’s organizer and the self-regulation of embryonic fields. Mech. Dev. 2009, 126, 925–941. [Google Scholar] [CrossRef]
- Driever, W.; Nüsslein-Volhard, C. A gradient of bicoid protein in Drosophila embryos. Cell 1988, 54, 83–93. [Google Scholar] [CrossRef]
- Driever, W.; Nüsslein-Volhard, C. The bicoid protein determines position in the Drosophila embryo in a concentration-dependent manner. Cell 1988, 54, 95–104. [Google Scholar] [CrossRef]
- Reversade, B.; De Robertis, E.M. Regulation of ADMP and BMP2/4/7 at opposite embryonic poles generates a self-regulating morphogenetic field. Cell 2005, 123, 1147–1160. [Google Scholar] [CrossRef] [PubMed]
- Ben-Zvi, D.; Shilo, B.Z.; Fainsod, A.; Barkai, N. Scaling of the BMP activation gradient in Xenopus embryos. Nature 2008, 453, 1205–1211. [Google Scholar] [CrossRef]
- Gregor, T.; Bialek, W.; de Ruyter van Steveninck, R.R.; Tank, D.W.; Wieschaus, E.F. Diffusion and scaling during early embryonic pattern formation. Proc. Natl. Acad. Sci. USA 2005, 102, 18403–18407. [Google Scholar] [CrossRef]
- Ben-Zvi, D.; Barkai, N. Scaling of morphogen gradients by an expansion-repression integral feedback control. Proc. Natl. Acad. Sci. USA 2010, 107, 6924–6929. [Google Scholar] [CrossRef]
- Riddle, R.D.; Johnson, R.L.; Laufer, E.; Tabin, C. Sonic hedgehog mediates the polarizing activity of the ZPA. Cell 1993, 75, 1401–1416. [Google Scholar] [CrossRef] [PubMed]
- Dale, L.; Howes, G.; Price, B.M.J.; Smith, J.C. Bone morphogenetic protein 4: a ventralizing factor in early Xenopus development. Development 1992, 115, 573–585. [Google Scholar] [CrossRef]
- Dosch, R.; Gawantka, V.; Delius, H.; Blumenstock, C.; Niehrs, C. Bmp-4 acts as a morphogen in dorsoventral mesoderm patterning in Xenopus. Development 1997, 124, 2325–2334. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.M.; Dale, L.; Hogan, B.L.M.; Wright, C.V.E.; Smith, J.C. Bone morphogenetic protein-4 (BMP-4) acts during gastrula stages to cause ventralization of Xenopus embryos. Development 1996, 122, 1545–1554. [Google Scholar] [CrossRef] [PubMed]
- Arendt, D.; Nübler-Jung, K. Inversion of dorsoventral axis? Nature 1994, 371, 26. [Google Scholar] [CrossRef] [PubMed]
- De Robertis, E.M.; Sasai, Y. A common plan for dorsoventral patterning in Bilateria. Nature 1996, 380, 37–40. [Google Scholar] [CrossRef]
- Holley, S.A.; Jackson, P.D.; Sasai, Y.; Lu, B.; De Robertis, E.M.; Hoffmann, F.M.; Ferguson, E.L. A conserved system for dorsal-ventral patterning in insects and vertebrates involving sog and chordin. Nature 1995, 376, 249–253. [Google Scholar] [CrossRef]
- Briscoe, J.; Small, S. Morphogen rules: design principles of gradient-mediated embryo patterning. Development 2015, 142, 3996–4009. [Google Scholar] [CrossRef]
- Beddington, R.S.P. Induction of a second neural axis by the mouse node. Development 1994, 120, 613–620. [Google Scholar] [CrossRef]
- Bachiller, D.; Klingensmith, J.; Kemp, C.; Belo, J.A.; Anderson, R.M.; May, S.R.; McMahon, J.A.; McMahon, A.P.; Harland, R.M.; Rossant, J.; et al. The organizer factors Chordin and Noggin are required for mouse forebrain development. Nature 2000, 403, 658–661. [Google Scholar] [CrossRef]
- De Robertis, E.M. Spemann’s organizer and self-regulation in amphibian embryos. Nat. Rev. Mol. Cell Biol. 2006, 7, 296–302. [Google Scholar] [CrossRef]
- Struhl, G.; Johnston, P.; Lawrence, P.A. Control of Drosophila body pattern by the hunchback morphogen gradient. Cell 1992, 69, 237–249. [Google Scholar] [CrossRef]
- Dessaud, E.; McMahon, A.P.; Briscoe, J. Pattern formation in the vertebrate neural tube: a sonic hedgehog morphogen-regulated transcriptional network. Development 2008, 135, 2489–2503. [Google Scholar] [CrossRef]
- Frohnhöfer, H.G.; Nüsslein-Volhard, C. Organization of anterior pattern in the Drosophila embryo by the maternal gene bicoid. Nature 1986, 324, 120–125. [Google Scholar] [CrossRef]
- Roessler, E.; Belloni, E.; Gaudenz, K.; Jay, P.; Berta, P.; Scherer, S.W.; Tsui, L.C.; Muenke, M. Mutations in the human Sonic Hedgehog gene cause holoprosencephaly. Nat. Genet. 1996, 14, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Artavanis-Tsakonas, S.; Rand, M.D.; Lake, R.J. Notch signaling: cell fate control and signal integration in development. Science 1999, 284, 770–776. [Google Scholar] [CrossRef] [PubMed]
- Simpson, P. Lateral inhibition and the development of the sensory bristles of the adult peripheral nervous system of Drosophila. Development 1990, 109, 509–519. [Google Scholar] [CrossRef]
- Heitzler, P.; Simpson, P. The choice of cell fate in the epidermis of Drosophila. Cell 1991, 64, 1083–1092. [Google Scholar] [CrossRef]
- Collier, J.R.; Monk, N.A.M.; Maini, P.K.; Lewis, J.H. Pattern formation by lateral inhibition with feedback: a mathematical model of Delta-Notch intercellular signalling. J. Theor. Biol. 1996, 183, 429–446. [Google Scholar] [CrossRef]
- Gazave, E.; Lapébie, P.; Richards, G.S.; Brunet, F.; Ereskovsky, A.V.; Degnan, B.M.; Borchiellini, C.; Vervoort, M.; Renard, E. Origin and evolution of the Notch signalling pathway: an overview from eukaryotic genomes. BMC Evol. Biol. 2009, 9, 249. [Google Scholar] [CrossRef] [PubMed]
- Hara, K.; Kajita, R.; Torii, K.U.; Bergmann, D.C.; Kakimoto, T. The secretory peptide gene EPF1 enforces the stomatal one-cell-spacing rule. Genes Dev. 2007, 21, 1720–1725. [Google Scholar] [CrossRef]
- Weng, A.P.; Ferrando, A.A.; Lee, W.; Morris, IV, J.P.; Silverman, L.B.; Sanchez-Irizarry, C.; Blacklow, S.C.; Look, A.T.; Aster, J.C. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 2004, 306, 269–271. [Google Scholar] [CrossRef]
- De Joussineau, C.; Soulé, J.; Martin, M.; Anguille, C.; Montcourrier, P.; Alexandre, D. Delta-promoted filopodia mediate long-range lateral inhibition in Drosophila. Nature 2003, 426, 555–559. [Google Scholar] [CrossRef]
- Cooke, J.; Zeeman, E.C. A clock and wavefront model for control of the number of repeated structures during animal morphogenesis. J. Theor. Biol. 1976, 58, 455–476. [Google Scholar] [CrossRef]
- Palmeirim, I.; Henrique, D.; Ish-Horowicz, D.; Pourquié, O. Avian hairy gene expression identifies a molecular clock linked to vertebrate segmentation and somitogenesis. Cell 1997, 91, 639–648. [Google Scholar] [CrossRef] [PubMed]
- Oates, A.C.; Morelli, L.G.; Ares, S. Patterning embryos with oscillations: structure, function and dynamics of the vertebrate segmentation clock. Development 2012, 139, 625–639. [Google Scholar] [CrossRef]
- Riedel-Kruse, I.H.; Müller, C.; Oates, A.C. Synchrony dynamics during initiation, failure, and rescue of the segmentation clock. Science 2007, 317, 1911–1915. [Google Scholar] [CrossRef]
- Uriu, K.; Liao, B.K.; Oates, A.C.; Morelli, L.G. From local resynchronization to global pattern recovery in the zebrafish segmentation clock. eLife 2021, 10, e61358. [Google Scholar] [CrossRef] [PubMed]
- Sonnen, K.F.; Lauschke, V.M.; Uraji, J.; Falk, H.J.; Petersen, Y.; Funk, M.C.; Beaupeux, M.; François, P.; Merten, C.A.; Aulehla, A. Modulation of phase shift between Wnt and Notch signaling oscillations controls mesoderm segmentation. Cell 2018, 172, 1079–1090.e12. [Google Scholar] [CrossRef]
- Sanchez, P.G.L.; Mochulska, V.; Mauffette Denis, C.; Mönke, G.; Tomita, T.; Tsuchida-Straeten, N.; Petersen, Y.; Sonnen, K.; François, P.; Aulehla, A. Arnold tongue entrainment reveals dynamical principles of the embryonic segmentation clock. eLife 2022, 11, e79575. [Google Scholar] [CrossRef] [PubMed]
- Bessho, Y.; Sakata, R.; Komatsu, S.; Shiota, K.; Yamada, S.; Kageyama, R. Dynamic expression and essential functions of Hes7 in somite segmentation. Genes Dev. 2001, 15, 2642–2647. [Google Scholar] [CrossRef]
- Hirata, H.; Bessho, Y.; Kokubu, H.; Masamizu, Y.; Yamada, S.; Lewis, J.; Kageyama, R. Instability of Hes7 protein is crucial for the somite segmentation clock. Nat. Genet. 2004, 36, 750–754. [Google Scholar] [CrossRef] [PubMed]
- Dubrulle, J.; McGrew, M.J.; Pourquié, O. FGF signaling controls somite boundary position and regulates segmentation clock control of spatiotemporal Hox gene activation. Cell 2001, 106, 219–232. [Google Scholar] [CrossRef]
- Krol, A.J.; Roellig, D.; Dequéant, M.L.; Tassy, O.; Glynn, E.; Hattem, G.; Mushegian, A.; Oates, A.C.; Pourquié, O. Evolutionary plasticity of segmentation clock networks. Development 2011, 138, 2783–2792. [Google Scholar] [CrossRef]
- Gomez, C.; Özbudak, E.M.; Wunderlich, J.; Baumann, D.; Lewis, J.; Pourquié, O. Control of segment number in vertebrate embryos. Nature 2008, 454, 335–339. [Google Scholar] [CrossRef]
- Turnpenny, P.D.; Alman, B.; Cornier, A.S.; Giampietro, P.F.; Offiah, A.; Tassy, O.; Pourquié, O.; Kusumi, K.; Dunwoodie, S. Abnormal vertebral segmentation and the notch signaling pathway in man. Dev. Dyn. 2007, 236, 1456–1474. [Google Scholar] [CrossRef]
- McGinnis, W.; Krumlauf, R. Homeobox genes and axial patterning. Cell 1992, 68, 283–302. [Google Scholar] [CrossRef]
- de Rosa, R.; Grenier, J.K.; Andreeva, T.; Cook, C.E.; Adoutte, A.; Akam, M.; Carroll, S.B.; Balavoine, G. Hox genes in brachiopods and priapulids and protostome evolution. Nature 1999, 399, 772–776. [Google Scholar] [CrossRef]
- Deschamps, J.; Duboule, D. Embryonic timing, axial stem cells, chromatin dynamics, and the Hox clock. Genes Dev. 2017, 31, 1406–1416. [Google Scholar] [CrossRef] [PubMed]
- Noordermeer, D.; Leleu, M.; Schorderet, P.; Joye, E.; Chabaud, F.; Duboule, D. Temporal dynamics and developmental memory of 3D chromatin architecture at Hox gene loci. eLife 2014, 3, e02557. [Google Scholar] [CrossRef]
- Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 276, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Kessel, M.; Gruss, P. Homeotic transformations of murine vertebrae and concomitant alteration of Hox codes induced by retinoic acid. Cell 1991, 67, 89–104. [Google Scholar] [CrossRef]
- Malicki, J.; Schughart, K.; McGinnis, W. Mouse Hox-2.2 specifies thoracic segmental identity in Drosophila embryos and larvae. Cell 1990, 63, 961–967. [Google Scholar] [CrossRef]
- Lutz, B.; Lu, H.C.; Eichele, G.; Miller, D.; Kaufman, T.C. Rescue of Drosophila labial null mutant by the chicken ortholog Hoxb-1 demonstrates that the function of Hox genes is phylogenetically conserved. Genes Dev. 1996, 10, 176–184. [Google Scholar] [CrossRef]
- Goodman, F.R. Limb malformations and the human HOX genes. Am. J. Med. Genet. 2002, 112, 256–265. [Google Scholar] [CrossRef]
- Spitz, F.; Herkenne, C.; Morris, M.A.; Duboule, D. Inversion-induced disruption of the Hoxd cluster leads to the partition of regulatory landscapes. Nat. Genet. 2005, 37, 889–893. [Google Scholar] [CrossRef]
- Wolpert, L. Positional information and the spatial pattern of cellular differentiation. J. Theor. Biol. 1969, 25, 1–47. [Google Scholar] [CrossRef]
- Simsek, M.F.; Özbudak, E.M. Spatial fold change of FGF signaling encodes positional information for segmental determination in zebrafish. Cell Rep. 2018, 24, 66–78. [Google Scholar] [CrossRef] [PubMed]
- Nüsslein-Volhard, C.; Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 1980, 287, 795–801. [Google Scholar] [CrossRef] [PubMed]
- Bray, S.J. Notch signalling: a simple pathway becomes complex. Nat. Rev. Mol. Cell Biol. 2006, 7, 678–689. [Google Scholar] [CrossRef] [PubMed]
- Frank, S.A. Robustness and complexity. Cell Syst. 2023, 14, 1015–1020. [Google Scholar] [CrossRef]
- Friedlander, T.; Mayo, A.E.; Tlusty, T.; Alon, U. Evolution of bow-tie architectures in biology. PLoS Comput. Biol. 2015, 11, e1004055. [Google Scholar] [CrossRef] [PubMed]
- Schröter, C.; Herrgen, L.; Cardona, A.; Brouhard, G.J.; Feldman, B.; Oates, A.C. Dynamics of zebrafish somitogenesis. Dev. Dyn. 2008, 237, 545–553. [Google Scholar] [CrossRef]
- Dequéant, M.L.; Glynn, E.; Gaudenz, K.; Wahl, M.; Chen, J.; Mushegian, A.; Pourquié, O. A complex oscillating network of signaling genes underlies the mouse segmentation clock. Science 2006, 314, 1595–1598. [Google Scholar] [CrossRef]
- Lewis, J. Autoinhibition with transcriptional delay: a simple mechanism for the zebrafish somitogenesis oscillator. Curr. Biol. 2003, 13, 1398–1408. [Google Scholar] [CrossRef] [PubMed]
- Frank, S.A. Maladaptation and the paradox of robustness in evolution. PLoS ONE 2007, 2, e1021. [Google Scholar] [CrossRef] [PubMed]
- Rutherford, S.L.; Lindquist, S. Hsp90 as a capacitor for morphological evolution. Nature 1998, 396, 336–342. [Google Scholar] [CrossRef]
- Stoltzfus, A. On the possibility of constructive neutral evolution. J. Mol. Evol. 1999, 49, 169–181. [Google Scholar] [CrossRef]
- Spencer, H. Essays: Scientific, Political, and Speculative. In D; Appleton: New York, 1907; Vol. 3. [Google Scholar]
- Duboule, D.; Morata, G. Colinearity and functional hierarchy among genes of the homeotic complexes. Trends Genet. 1994, 10, 358–364. [Google Scholar] [CrossRef]
- Yekta, S.; Shih, I.H.; Bartel, D.P. MicroRNA-directed cleavage of HOXB8 mRNA. Science 2004, 304, 594–596. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



