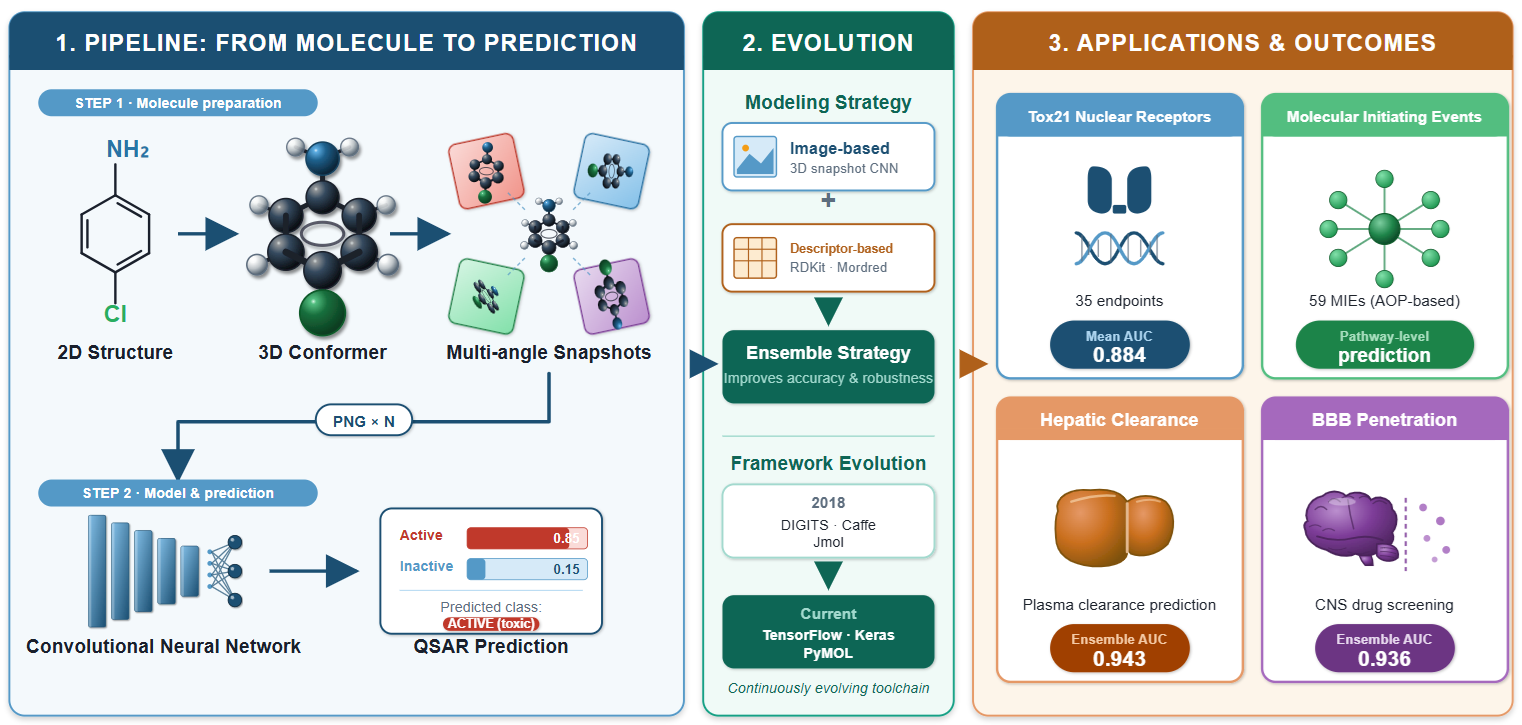
Quantitative structure–activity relationship (QSAR) modeling has traditionally relied on expert-designed molecular descriptors to encode chemical structures. DeepSnap is a descriptor-free QSAR approach that converts three-dimensional molecular structures into image representations and feeds them directly into convolutional neural networks for activity prediction. The method generates a conformer for each molecule, renders it as a color-coded molecular image, and captures omnidirectional snapshots from systematically varied viewing angles. This review traces DeepSnap from its introduction in 2018 to its current state. The method has been applied to 35 nuclear receptor endpoints from the Tox21 10K library (mean AUC 0.884), 59 molecular initiating event models spanning the full Tox21 target panel, rat hepatic clearance (ensemble AUC 0.943), and blood–brain barrier penetration (ensemble AUC 0.936). An ensemble strategy combining image-based and descriptor-based predictions has consistently outperformed either approach alone. The computational pipeline has evolved from a DIGITS/Caffe/Jmol system to a TensorFlow/Keras/PyMOL framework. Limitations include endpoint-dependent parameter sensitivity, class imbalance effects, the absence of direct comparisons with graph neural networks, and an interpretability gap addressed in part by CAM-family visualization in the AI-SHIPS platform and S-COPHY. Future directions include systematic application of explainable AI methods, automated hyperparameter optimization, and integration with graph-based approaches.