Submitted:
29 April 2026
Posted:
02 May 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
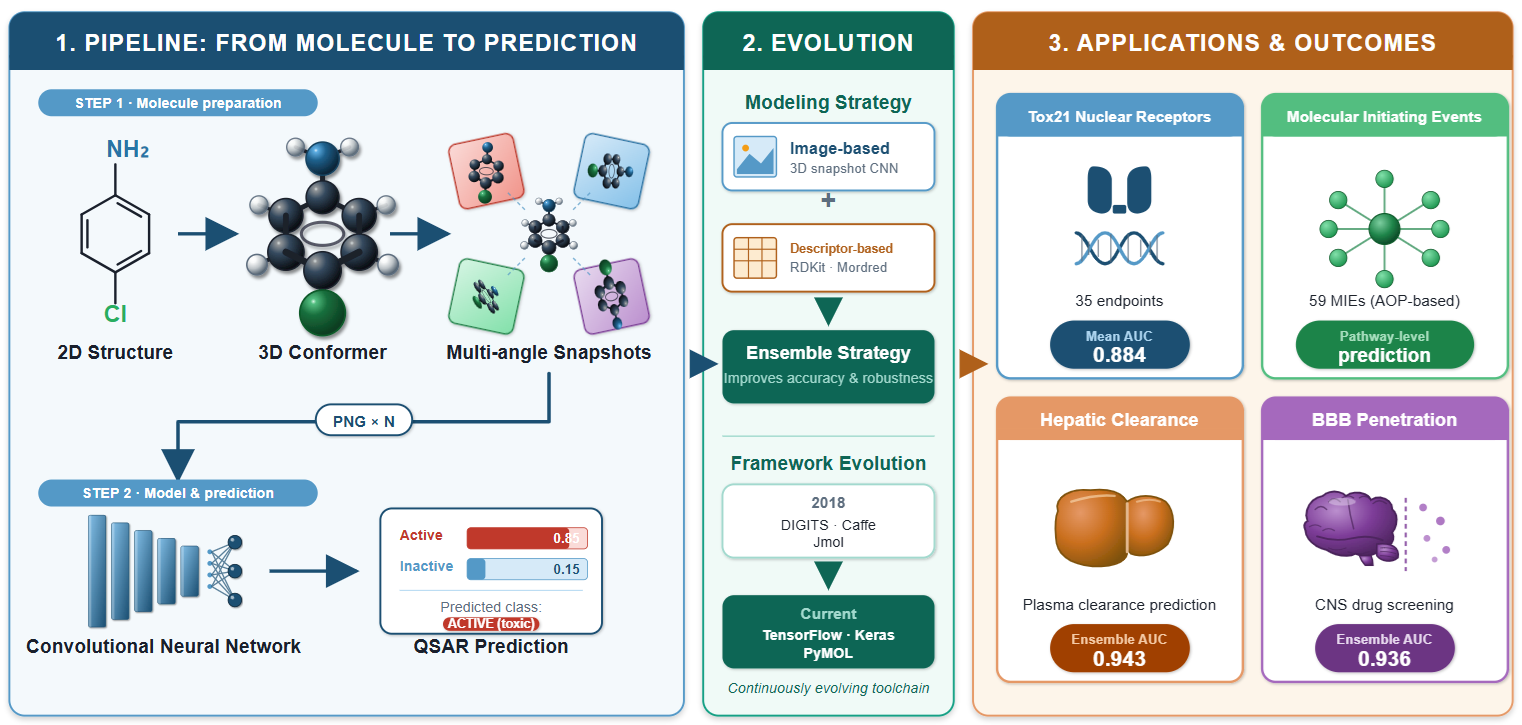
2. Origin and Rationale of DeepSnap
The Inaugural Study
Performance and Comparators
Conceptual Rationale
Early Recognition of Limitations
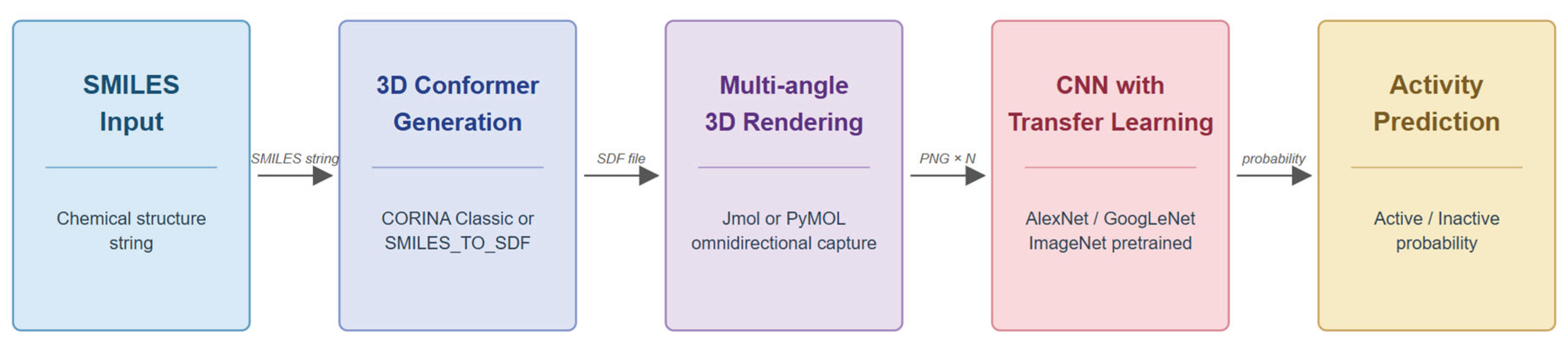
3. Core Rendering and Learning Pipeline
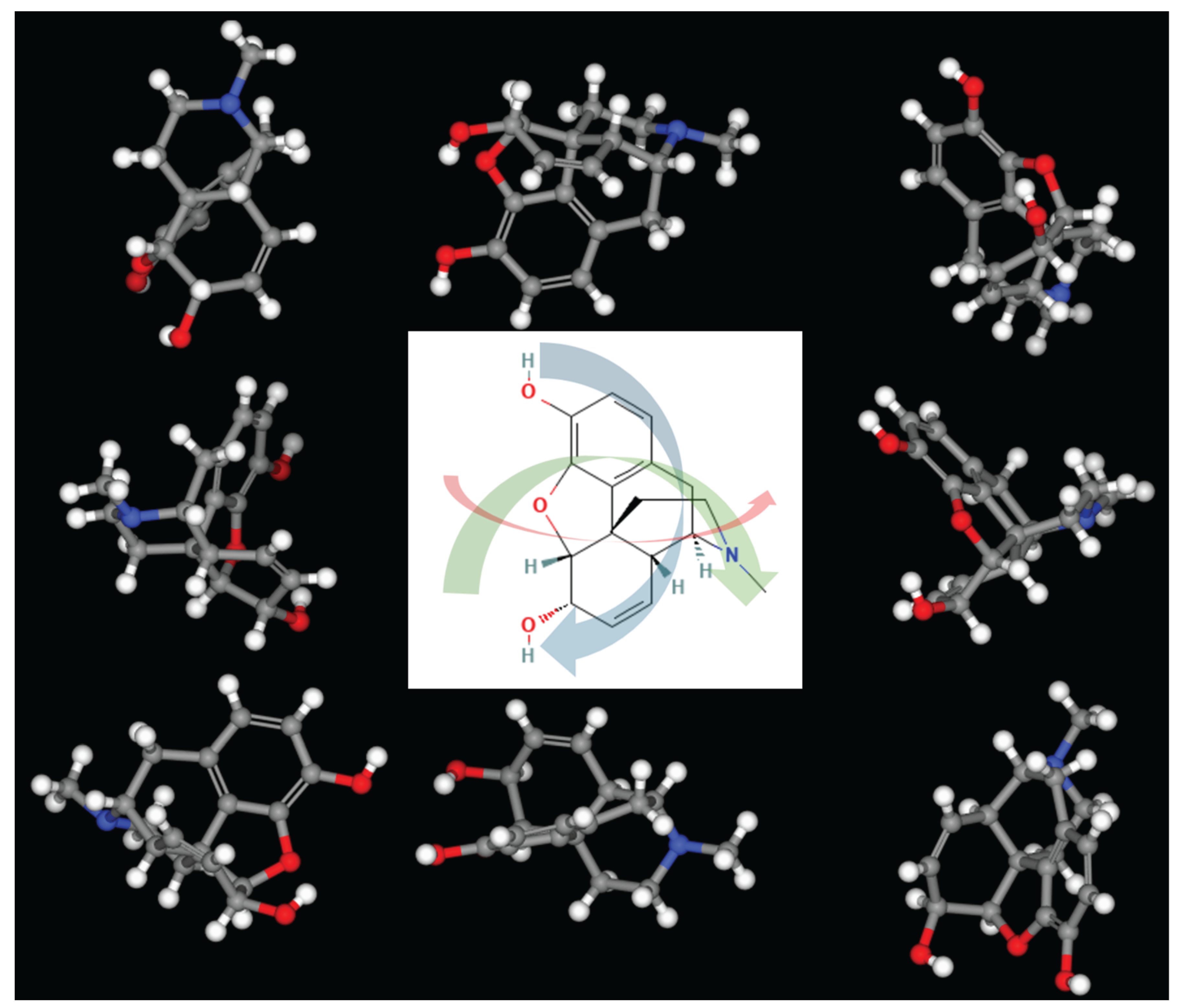
Rendering Parameter Optimization
First-Generation Pipeline
Transition from AlexNet to GoogLeNet
CORINA Wash Conditions
Angle Increment as Key Hyperparameter
Second-Generation Pipeline
Background Color Optimization
Learning Rate, Batch Size, and Epoch Selection
Aggregation, Cut-Off Determination, and Model Validation
4. Applications to Toxicological Targets
| Year | Ref. | Endpoint | Dataset (N) | Pipeline | Best Result | Comparator | Key Limitation |
|---|---|---|---|---|---|---|---|
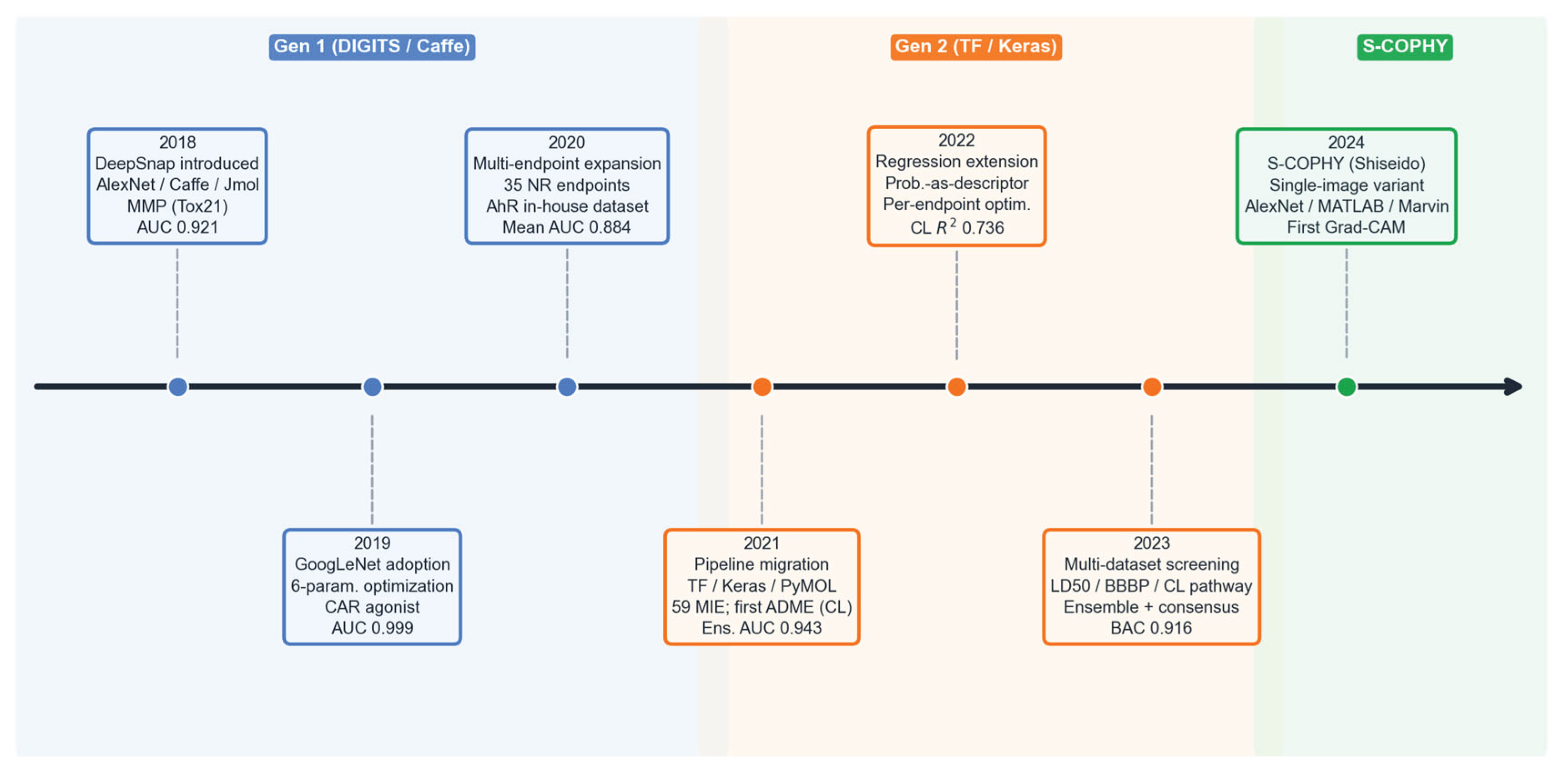
| 2018 | [13] | MMP disruption | Tox21 (7967) | AlexNet / DIGITS | AUC 0.921 | RF+3D desc 0.907 | Single endpoint; no HP tuning |
| 2019 | [21] | CAR param. optim. | Tox21 (9523) | AlexNet / DIGITS | AUC 0.791 | RF+MOE 0.749 | Non-standard threshold; single split |
| 2019 | [23] | CAR agonist | Tox21 (7141) | GoogLeNet / DIGITS | AUC 0.999 | XGB 0.889; RF 0.884 | Near-ceiling AUC; class imbalance |
| 2020 | [25] | PR antagonist | Tox21 (7582) | GoogLeNet / DIGITS | AUC 0.999 | CB 0.894; LGBM 0.893 | Wash varies by target; imbalance |
| 2020 | [26] | AhR (in-house) | In-house (201) | GoogLeNet / DIGITS | AUC 0.959 | XGB 0.724; RF 0.716 | Small N; no wash optimization |
| 2020 | [14] | 35 NR models | Tox21 (mean 7262) | GoogLeNet / DIGITS | Mean AUC 0.884 | Tox21 Challenge (3/4 exceeded) | n = 2 replicates; class imbalance |
| 2021 | [15] | 59 MIE models | Tox21 (mean 9699) | GoogLeNet / TF-Keras | Mean AUC 0.818 | Prior DIGITS system | Underperforms DIGITS; NFkB failed |
| 2021 | [16] | Rat CL classif. | In-house (1545) | GoogLeNet / DIGITS + ens. | Ens. AUC 0.943 | RF+MD 0.883 | Consensus coverage 69% |
| 2022 | [27] | GR/TRHR/TGFb | Tox21 (7537–7662) | GoogLeNet / TF-Keras | GR AUC 0.983 | DIGITS GR 0.910 | TRHR MCC 0.200; 3 endpoints only |
| 2022 | [30] | Rat CL regression | In-house (1545) | DL prob. + AutoML | R² 0.736 | MD-only R² 0.649 | DL alone R² 0.359; private data |
| 2023 | [17] | LD50/BBBP/CL path. | CATMoS (11886) etc. | GoogLeNet / DIGITS + ens. | Cons. BAC 0.916 | CATMoS 32 orgs 0.87 | Coverage 77–86%; DL < CATMoS |
| 2024 | [18] | Cosmetics vs. pharma | PubChem etc. (2754) | AlexNet / MATLAB | AUC 0.935 | None | Ext. pred. rate 46%; regulatory |
5. Applications to ADME Parameters
6. Technical Evolution and Variants
7. Limitations and Unresolved Issues
Parameter Sensitivity and the Absence of Universal Settings
Statistical Evaluation Practices and Class Imbalance
Reproducibility and Data Availability
System Migration and Framework Confounding
Interpretability and Comparison with Alternative Architectures
8. Position Relative to External Methods
| Method Family | Example | Representation | 3D Info | Interpret. | Cost | Benchmark Relation |
|---|---|---|---|---|---|---|
| Descriptor ML | RF/DNN + ECFP [35] | Descriptors, fingerprints | Partial | Medium | Low | Indirect: Tox21 (BA 0.58–0.82) |
| Descriptor DL | DeepTox [7] | ECFP + toxicophores | No | Low | Medium | Indirect: Tox21 Challenge winner |
| GNN | MPNN [8] | Molecular graph | No | Low | Medium | None: QM9 only |
| GNN | D-MPNN [9] | Directed graph + RDKit | No | Low | Medium | Indirect: MoleculeNet scaffold split |
| GNN | AttentiveFP [10] | Graph + attention | No | High | Medium | None: not on DeepSnap endpoints |
| 2D Image | Chemception [11] | 2D structure drawings | No | Low | Low | Indirect: Tox21 different splits |
| 2D Image | ImageMol [12] | 2D images (ResNet-18) | No | Medium | High | Indirect: BBBP overlap |
| 3D Voxel | AtomNet [41] | Voxelized complex | Yes | Low | High | None: structure-based task |
| 3D Voxel | Ragoza et al. [42] | Voxelized densities | Yes | Low | High | None: structure-based task |
| DeepSnap | DeepSnap-DL [13,14] | 3D molecular images | Yes | Low | Medium | Reference method |
| DeepSnap Ens. | DeepSnap + ML [17] | Hybrid: images + descriptors | Yes | Medium | Medium | Reference method |
9. Future Directions
Expanding the Endpoint Repertoire
Hybrid Image-Graph Models
Systematic Interpretability Analysis
Reducing Hyperparameter Sensitivity
Regression Without Classification Intermediary
Multi-Task Learning
10. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Generative AI Disclosure
References
- Uesawa, Y. AI-based QSAR Modeling for Prediction of Active Compounds in MIE/AOP. Yakugaku Zasshi 2020, 140, 499–505. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. Deep learning using molecular image of chemical structure. In Cheminformatics, QSAR and Machine Learning Applications for Novel Drug Development; Roy, K., Ed.; Elsevier: Amsterdam, The Netherlands, 2023; Chapter 18, pp. 473–501. [CrossRef]
- Huang, R.; Xia, M.; Nguyen, D.-T.; Zhao, T.; Sakamuru, S.; Zhao, J.; Shahane, S.A.; Rossoshek, A.; Simeonov, A. Tox21 Challenge to Build Predictive Models of Nuclear Receptor and Stress Response Pathways as Mediated by Exposure to Environmental Chemicals and Drugs. Front. Environ. Sci. 2016, 3, 85. [CrossRef]
- Ankley, G.T.; Bennett, R.S.; Erickson, R.J.; Hoff, D.J.; Hornung, M.W.; Johnson, R.D.; Mount, D.R.; Nichols, J.W.; Russom, C.L.; Schmieder, P.K.; Serrano, J.A.; Tietge, J.E.; Villeneuve, D.L. Adverse Outcome Pathways: A Conceptual Framework to Support Ecotoxicology Research and Risk Assessment. Environ. Toxicol. Chem. 2010, 29, 730–741. [CrossRef]
- Cherkasov, A.; Muratov, E.N.; Fourches, D.; Varnek, A.; Baskin, I.I.; Cronin, M.; Dearden, J.; Gramatica, P.; Martin, Y.C.; Todeschini, R.; Consonni, V.; Kuz’min, V.E.; Cramer, R.; Benigni, R.; Yang, C.; Rathman, J.; Terfloth, L.; Gasteiger, J.; Richard, A.; Tropsha, A. QSAR Modeling: Where Have You Been? Where Are You Going To? J. Med. Chem. 2014, 57, 4977–5010. [CrossRef]
- Rogers, D.; Hahn, M. Extended-Connectivity Fingerprints. J. Chem. Inf. Model. 2010, 50, 742–754. [CrossRef]
- Mayr, A.; Klambauer, G.; Unterthiner, T.; Hochreiter, S. DeepTox: Toxicity Prediction using Deep Learning. Front. Environ. Sci. 2016, 3, 80. [CrossRef]
- Gilmer, J.; Schoenholz, S.S.; Riley, P.F.; Vinyals, O.; Dahl, G.E. Neural Message Passing for Quantum Chemistry. In Proceedings of the 34th International Conference on Machine Learning (ICML 2017), PMLR 70:1263–1272, 2017. arXiv:1704.01212.
- Yang, K.; Swanson, K.; Jin, W.; Coley, C.; Eiden, P.; Gao, H.; Guzman-Perez, A.; Hopper, T.; Kelley, B.; Mathea, M.; Palmer, A.; Settels, V.; Jaakkola, T.; Jensen, K.; Barzilay, R. Analyzing Learned Molecular Representations for Property Prediction. J. Chem. Inf. Model. 2019, 59, 3370–3388. [CrossRef]
- Xiong, Z.; Wang, D.; Liu, X.; Zhong, F.; Wan, X.; Li, X.; Li, Z.; Luo, X.; Chen, K.; Jiang, H.; Zheng, M. Pushing the Boundaries of Molecular Representation for Drug Discovery with the Graph Attention Mechanism. J. Med. Chem. 2020, 63, 8749–8760. [CrossRef]
- Goh, G.B.; Hodas, N.O.; Siegel, C.; Vishnu, A. Chemception: A Deep Neural Network with Minimal Chemistry Knowledge Matches the Performance of Expert-developed QSAR/QSPR Models. arXiv preprint 2017, arXiv:1706.06689.
- Zeng, X.; Xiang, H.; Yu, L.; Wang, J.; Li, K.; Nussinov, R.; Cheng, F. Accurate prediction of molecular properties and drug targets using a self-supervised image representation learning framework. Nat. Mach. Intell. 2022, 4, 1004–1016. [CrossRef]
- Uesawa, Y. Quantitative structure-activity relationship analysis using deep learning based on a novel molecular image input technique. Bioorg. Med. Chem. Lett. 2018, 28, 3400–3403. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. Molecular Image-Based Prediction Models of Nuclear Receptor Agonists and Antagonists Using the DeepSnap-Deep Learning Approach with the Tox21 10K Library. Molecules 2020, 25, 2764. [CrossRef]
- Matsuzaka, Y.; Totoki, S.; Handa, K.; Shiota, T.; Kurosaki, K.; Uesawa, Y. Prediction Models for Agonists and Antagonists of Molecular Initiation Events for Toxicity Pathways Using an Improved Deep-Learning-Based Quantitative Structure-Activity Relationship System. Int. J. Mol. Sci. 2021, 22, 10821. [CrossRef]
- Mamada, H.; Nomura, Y.; Uesawa, Y. Prediction Model of Clearance by a Novel Quantitative Structure-Activity Relationship Approach, Combination DeepSnap-Deep Learning and Conventional Machine Learning. ACS Omega 2021, 6, 23570–23577. [CrossRef]
- Mamada, H.; Takahashi, M.; Ogino, M.; Nomura, Y.; Uesawa, Y. Predictive Models Based on Molecular Images and Molecular Descriptors for Drug Screening. ACS Omega 2023, 8, 37186–37195. [CrossRef]
- Hisaki, T.; Yoshida, K.; Nukaga, T.; Iwanaga, S.; Mori, M.; Uesawa, Y.; Sekine, S.; Tamura, A. S-COPHY: A deep learning model for predicting the chemical class of compounds as cosmetics or pharmaceuticals based on single 3D molecular images. Comput. Toxicol. 2024, 30, 100311. [CrossRef]
- Uesawa, Y. A Novel Molecular Information Input Method for Deep Learning: Deep Snap. CICSJ Bull. 2018, 36, 51. [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems 25 (NeurIPS 2012); Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q., Eds.; Curran Associates: Red Hook, NY, 2012; pp. 1097–1105.
- Matsuzaka, Y.; Uesawa, Y. Optimization of a Deep-Learning Method Based on the Classification of Images Generated by Parameterized Deep Snap a Novel Molecular-Image-Input Technique for Quantitative Structure-Activity Relationship (QSAR) Analysis. Front. Bioeng. Biotechnol. 2019, 7, 65. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. Computational Models That Use a Quantitative Structure-Activity Relationship Approach Based on Deep Learning. Processes 2023, 11, 1296. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. Prediction Model with High-Performance Constitutive Androstane Receptor (CAR) Using DeepSnap-Deep Learning Approach from the Tox21 10K Compound Library. Int. J. Mol. Sci. 2019, 20, 4855. [CrossRef]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going Deeper with Convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 1–9. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. DeepSnap-Deep Learning Approach Predicts Progesterone Receptor Antagonist Activity With High Performance. Front. Bioeng. Biotechnol. 2020, 7, 485. [CrossRef]
- Matsuzaka, Y.; Hosaka, T.; Ogaito, A.; Yoshinari, K.; Uesawa, Y. Prediction Model of Aryl Hydrocarbon Receptor Activation by a Novel QSAR Approach, DeepSnap-Deep Learning. Molecules 2020, 25, 1317. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. A Deep Learning-Based Quantitative Structure-Activity Relationship System Construct Prediction Model of Agonist and Antagonist with High Performance. Int. J. Mol. Sci. 2022, 23, 2141. [CrossRef]
- Kurosaki, K.; Wu, R.; Uesawa, Y. A Toxicity Prediction Tool for Potential Agonist/Antagonist Activities in Molecular Initiating Events Based on Chemical Structures. Int. J. Mol. Sci. 2020, 21, 7853. [CrossRef]
- Mansouri, K.; Karmaus, A.; Fitzpatrick, J.; Patlewicz, G.; Pradeep, P.; et al. CATMoS: Collaborative Acute Toxicity Modeling Suite. Environ. Health Perspect. 2021, 129, 47013. [CrossRef]
- Mamada, H.; Nomura, Y.; Uesawa, Y. Novel QSAR Approach for a Regression Model of Clearance That Combines DeepSnap-Deep Learning and Conventional Machine Learning. ACS Omega 2022, 7, 17055–17062. [CrossRef]
- Matsuzaka, Y.; Uesawa, Y. Ensemble Learning, Deep Learning-Based and Molecular Descriptor-Based Quantitative Structure-Activity Relationships. Molecules 2023, 28, 2410. [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [CrossRef]
- Ministry of Economy, Trade and Industry (METI). Technology Evaluation Report (Final Evaluation) for the Project on the Development of Evaluation Technologies for Energy-Saving Electronic Device Materials (Development of High-Speed, High-Efficiency Safety Evaluation Technologies Supporting the Social Implementation of Functional Materials) [in Japanese]; METI: Tokyo, Japan, 2023. Available online: https://www.meti.go.jp/policy/tech_evaluation/e00/03/r04/600.pdf (accessed on 6 April 2026).
- Ministry of Economy, Trade and Industry (METI). Supplementary Explanatory Materials for the Final Evaluation of the Project on the Development of Evaluation Technologies for Energy-Saving Electronic Device Materials (Development of High-Speed, High-Efficiency Safety Evaluation Technologies Supporting the Social Implementation of Functional Materials) [in Japanese]; METI: Tokyo, Japan, 2023. Available online: https://www.meti.go.jp/shingikai/sankoshin/sangyo_gijutsu/kenkyu_innovation/hyoka_wg/pdf/064_h02_00.pdf (accessed on 6 April 2026).
- Capuzzi, S.J.; Politi, R.; Isayev, O.; Farag, S.; Tropsha, A. QSAR Modeling of Tox21 Challenge Stress Response and Nuclear Receptor Signaling Toxicity Assays. Front. Environ. Sci. 2016, 4, 3. [CrossRef]
- Wu, Z.; Ramsundar, B.; Feinberg, E.N.; Gomes, J.; Geniesse, C.; Pappu, A.S.; Leswing, K.; Pande, V. MoleculeNet: a benchmark for molecular machine learning. Chem. Sci. 2018, 9, 513–530. [CrossRef]
- Chen, D.; Gao, K.; Nguyen, D.D.; Chen, X.; Jiang, Y.; Wei, G.-W.; Pan, F. Algebraic Graph-Assisted Bidirectional Transformers for Molecular Property Prediction. Nat. Commun. 2021, 12, 3521. [CrossRef]
- Sheridan, R.P. Time-Split Cross-Validation as a Method for Estimating the Goodness of Prospective Prediction. J. Chem. Inf. Model. 2013, 53, 783–790. [CrossRef]
- Kaboudi, N.; Shayanfar, A. Predicting the Drug Clearance Pathway with Structural Descriptors. Eur. J. Drug Metab. Pharmacokinet. 2022, 47, 363–369. [CrossRef]
- Lombardo, F.; Obach, R.S.; Varma, M.V.; Stringer, R.; Berellini, G. Clearance Mechanism Assignment and Total Clearance Prediction in Human Based upon in Silico Models. J. Med. Chem. 2014, 57, 4397–4405. [CrossRef]
- Wallach, I.; Dzamba, M.; Heifets, A. AtomNet: A Deep Convolutional Neural Network for Bioactivity Prediction in Structure-based Drug Discovery. arXiv preprint 2015, arXiv:1510.02855.
- Ragoza, M.; Hochuli, J.; Idrobo, E.; Sunseri, J.; Koes, D.R. Protein-Ligand Scoring with Convolutional Neural Networks. J. Chem. Inf. Model. 2017, 57, 942–957. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




