Submitted:
29 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials
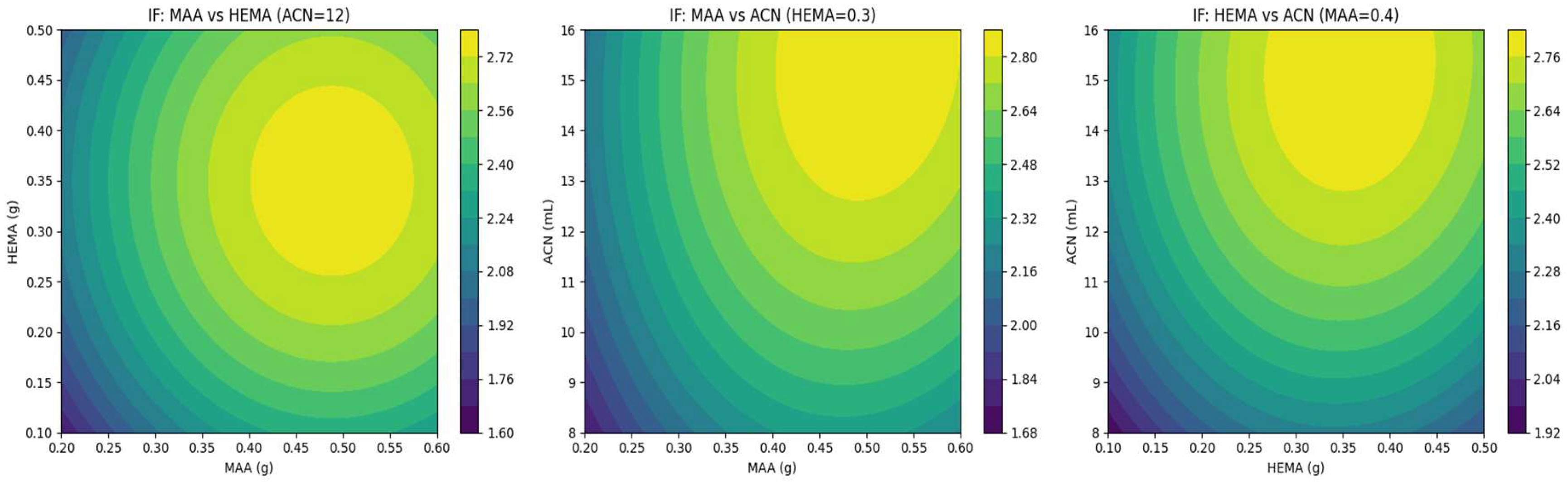
2.2. Experimental Design (Doe)
2.3. Synthesis of Molecularly Imprinted Polymers
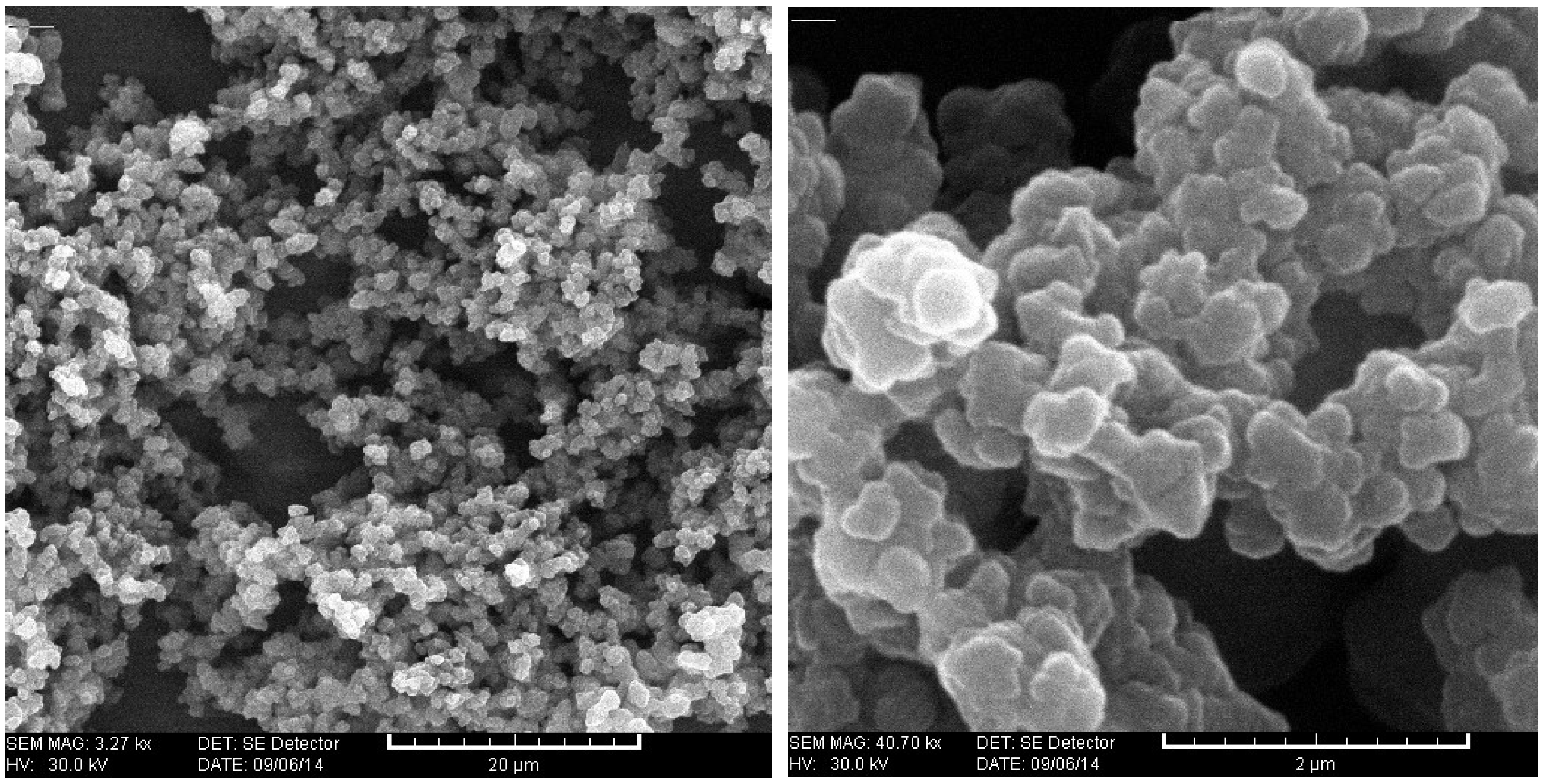
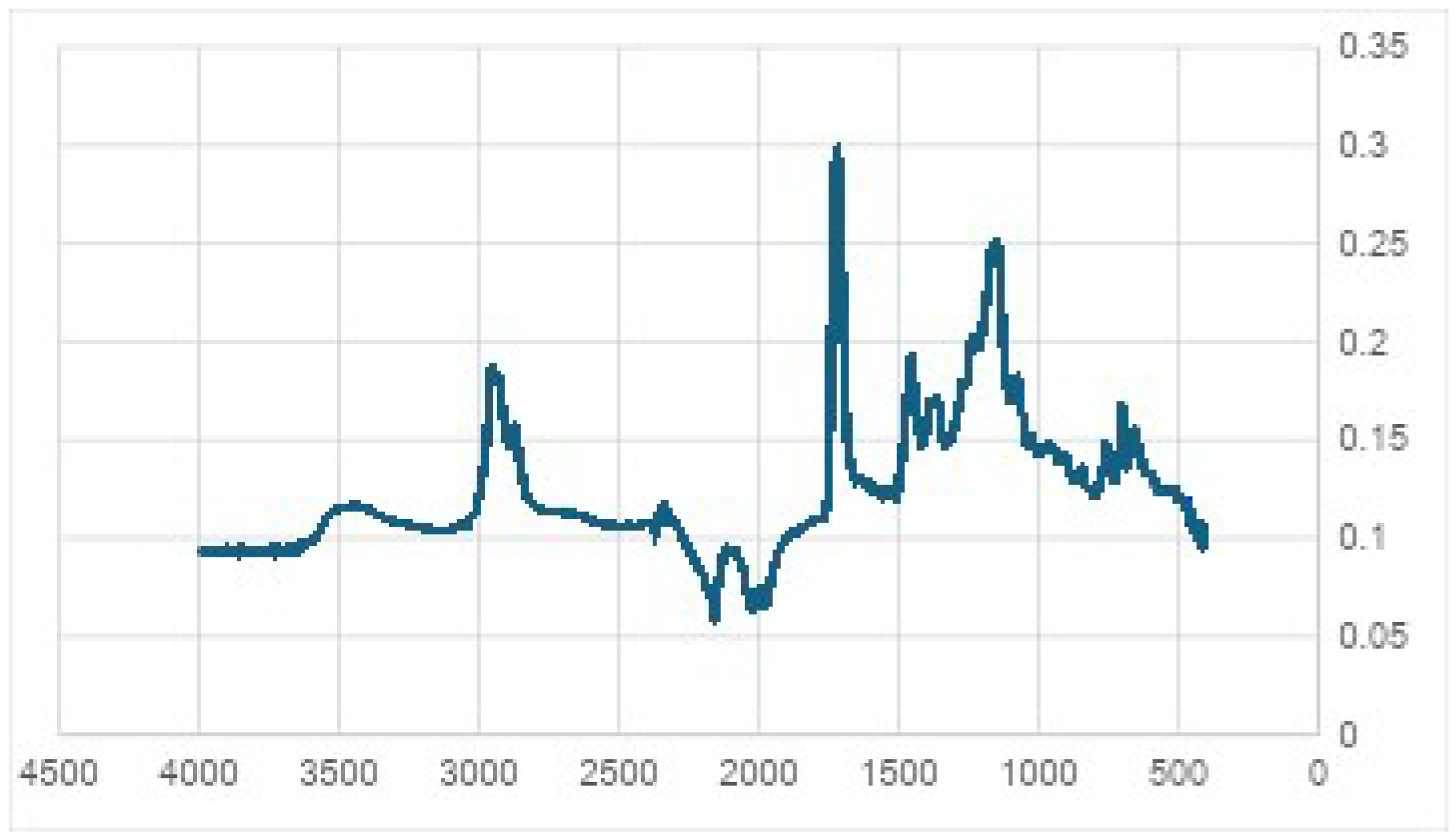
2.4. Characterization
2.5. Adsorption and Release Experiments
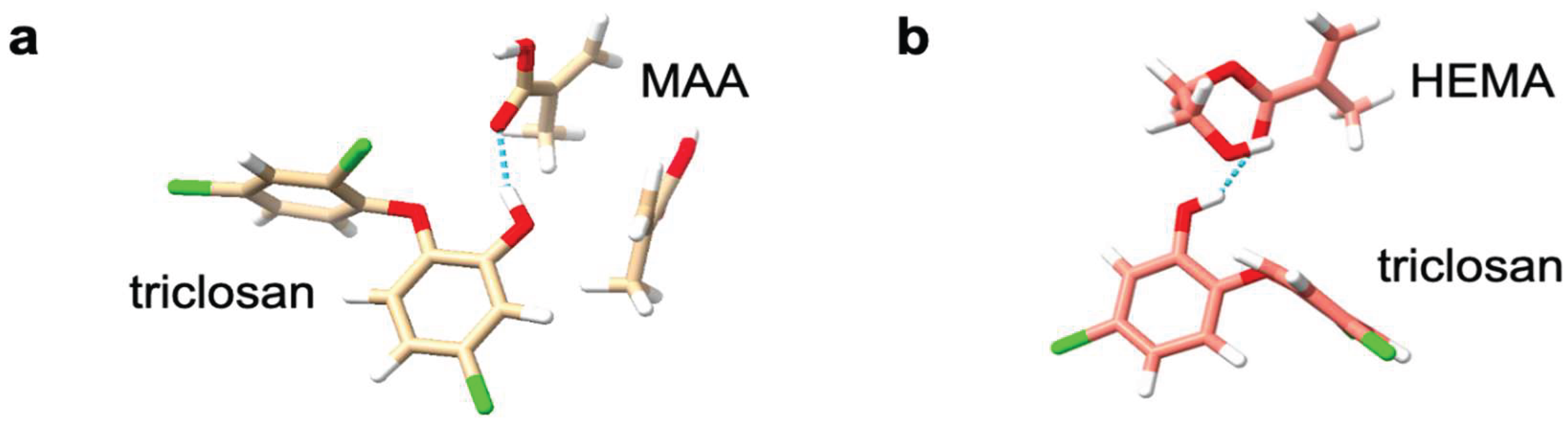
2.6. Molecular Dynamics Simulations
3. Results and Discussion
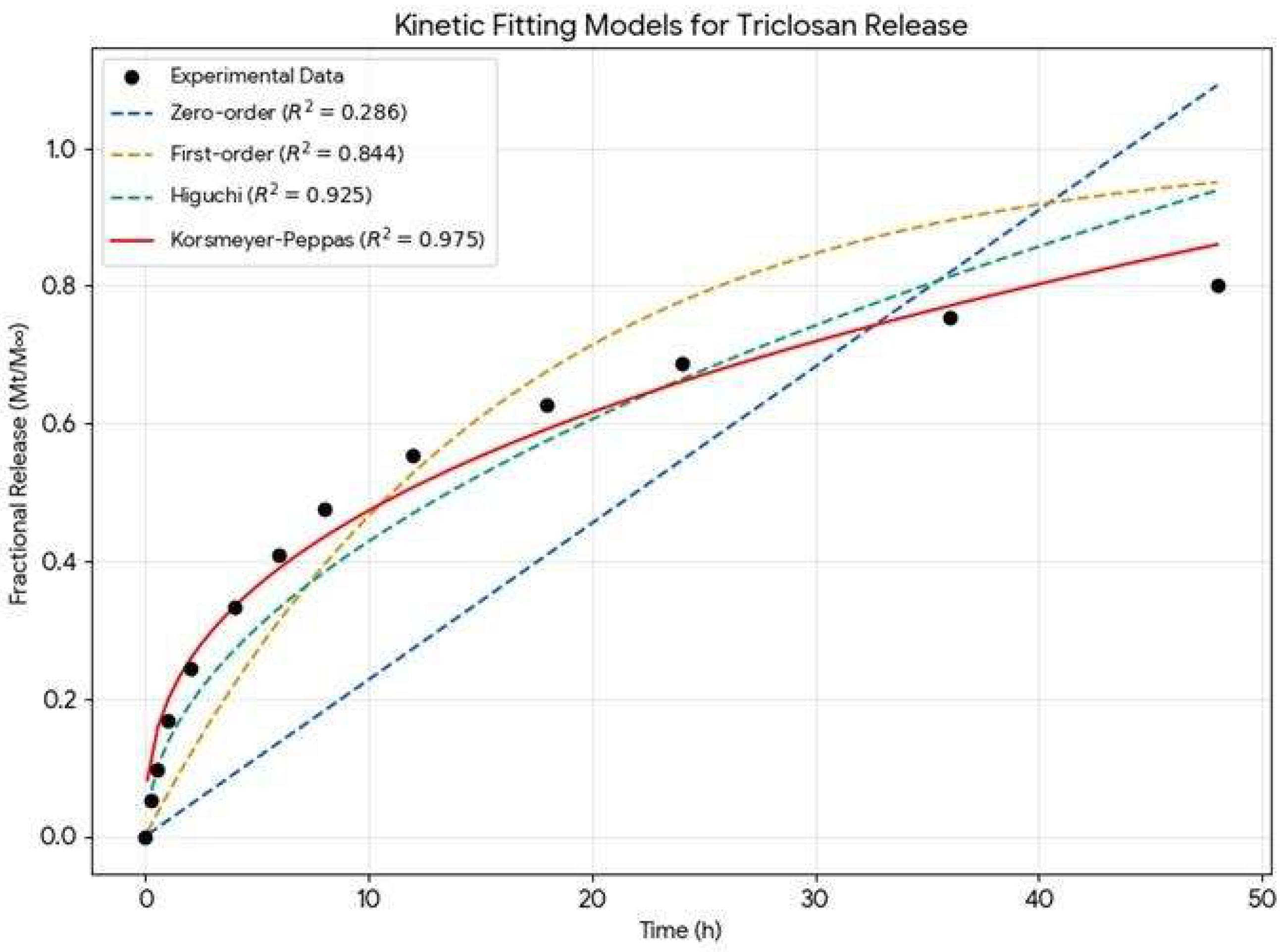
| Model | Equation | R2 | Interpretation |
|---|---|---|---|
| Korsmeyer-Peppas | Q= K.tn | 0.9754 | Best fit; describes complex diffusion. |
| Higuchi | Q= K.t0.5 | 0.925 | High fit; indicates matrix-based diffusion. |
| First-order | Ln(1-Q) = -K.t | 0.8441 | Moderate fit; release depends on concentration. |
| Zero-order | Q=K.t | 0.2864 | Poor fit; release is not at a constant rate. |
4. Conclusion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Davachi, Seyed Mohammad; Kaffashi, Babak. Preparation and characterization of poly L-lactide/triclosan nanoparticles for specific antibacterial and medical applications. Int. J. Polym. Mater. Polym. Biomater. 2015, 64(no. 10), 497–508. [Google Scholar] [CrossRef]
- Li, Y.; Xia, T.; Zhang, D.; Zhang, D.; Li, M. Adsorption behavior of triclosan by different microplastics and the impact of water chemistry. J. Contam. Hydrol. 2025, 104815. [Google Scholar] [CrossRef] [PubMed]
- Davachi, S. M.; Kaffashi, B.; Torabinejad, B.; Zamanian, A. In-vitro investigation and hydrolytic degradation of antibacterial nanocomposites based on PLLA/triclosan/nano-hydroxyapatite. Polymer 2016, 83, 101–110. [Google Scholar] [CrossRef]
- Khadem, M.; Faridbod, F.; Norouzi, P.; Rahimi Foroushani, A.; Ganjali, M. R.; Shahtaheri, S. J.; Yarahmadi, R. Modification of carbon paste electrode based on molecularly imprinted polymer for electrochemical determination of diazinon in biological and environmental samples. Electroanalysis 2017, 29(3), 708–715. [Google Scholar] [CrossRef]
- Singhal, A.; Parihar, A.; Xu, C.; Perera, G. S.; Bhaskaran, M.; Sriram, S.; Khan, R. Electropolymerized molecularly imprinted polymer-based simple and stable electrochemical nanosensor for triclosan detection. Biosens. Bioelectron. X 2026, 100782. [Google Scholar] [CrossRef]
- van Wissen, G.; Lowdon, J. W.; Cleij, T. J.; Eersels, K.; van Grinsven, B. Porogenic solvents in molecularly imprinted polymer synthesis: A comprehensive review of current practices and emerging trends. Polymers 2025, 17(8), 1057. [Google Scholar] [CrossRef]
- Elaine, A. A.; Krisyanto, S. I.; Hasanah, A. N. Dual-functional monomer MIPs and their comparison to mono-functional monomer MIPs for SPE and as sensors. Polymers 2022, 14(17), 3498. [Google Scholar] [CrossRef]
- Madineh, H.; Ebrahimi, N. G.; Zarrintaj, P. Engineering PEG-alginate microcarriers for precision drug Delivery: Optimized design and controlled release. Carbohydr. Res. 2025, 554, 109546. [Google Scholar] [CrossRef]
- Huo, Y.; Ristic, R.; Savoie, M.; Muhlack, R.; Herderich, M.; Wilkinson, K. Adsorption properties of molecularly imprinted polymers designed for removal of smoke taint compounds from wine. Food Res. Int. 2025, 206, 116048. [Google Scholar] [CrossRef]
- Song, X.; Wang, J.; Zhu, J. Effect of porogenic solvent on selective performance of molecularly imprinted polymer for quercetin. Mater. Res. 2009, 12(3), 299–304. [Google Scholar] [CrossRef]
- Landrum, G.; Tosco, P.; Kelly, B.; Rodriguez, R.; Cosgrove, D.; Vianello, R.; Gedeck, P.; Jones, G.; Kawashima, E.; Schneider, N.; Probst, D. (2015). RDKit [Software].
- Halgren, T. A. Merck molecular force field. I. Basis, form, scope, parameterization, and performance of MMFF94. J. Comput. Chem. 1996, 17(5–6), 490–519. [Google Scholar] [CrossRef]
- Jakalian, A.; Bush, B. L.; Jack, D. B.; Bayly, C. I. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: I. Method. J. Comput. Chem. 2000, 21(2), 132–146. [Google Scholar] [CrossRef]
- Jakalian, A.; Jack, D. B.; Bayly, C. I. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J. Comput. Chem. 2002, 23(16), 1623–1641. [Google Scholar] [CrossRef]
- Wang, L.; Behara, P. K.; Thompson, M. W.; Gokey, T.; Wang, Y.; Wagner, J. R.; Cole, D.; Gilson, M. K.; Shirts, M. R.; Mobley, D. L. The Open Force Field Initiative: Open software and open science for molecular modeling. J. Phys. Chem. B 2024, 128(29), 7043–7067. [Google Scholar] [CrossRef]
- Mobley, D. L.; Bannan, C. C.; Rizzi, A.; Bayly, C. I.; Chodera, J. D.; Lim, V. T.; Lim, N. M.; Beauchamp, K. A.; Slochower, D. R.; Shirts, M. R.; Gilson, M. K.; Eastman, P. K.; Wang, L.-P. Escaping atom types in force fields using direct chemical perception. J. Chem. Theory Comput. 2018, 14(11), 6076–6092. [Google Scholar] [CrossRef]
- Boothroyd, S.; Behara, P. K.; Madin, O.; Hahn, D.; Jang, H.; Gapsys, V.; Wagner, J.; Horton, J.; Dotson, D.; Thompson, M.; Maat, J.; Gokey, T.; Wang, L.-P.; Cole, D.; Gilson, M. K.; Chodera, J. D.; Bayly, C. I.; Shirts, M. R.; Mobley, D. L. Development and benchmarking of Open Force Field 2.0.0: The Sage small molecule force field. J. Chem. Theory Comput. 2023, 19(11), 3251–3275. [Google Scholar] [CrossRef]
- Eastman, P.; Friedrichs, M. S.; Chodera, J. D.; Radmer, R. J.; Bruns, C. M.; Ku, J. P.; Beauchamp, K. A.; Lane, T. J.; Wang, L.-P.; Shukla, D.; Tye, T.; Houston, M.; Stich, T.; Klein, C.; Shirts, M. R.; Pande, V. S. OpenMM 4: A reusable, extensible, hardware independent library for high performance molecular simulation. J. Chem. Theory Comput. 2013, 9(1), 461–469. [Google Scholar] [CrossRef]
- Eastman, P.; Swails, J.; Chodera, J. D.; McGibbon, R. T.; Zhao, Y.; Beauchamp, K. A.; Wang, L.-P.; Simmonett, A. C.; Harrigan, M. P.; Stern, C. D.; Wiewiora, R. P.; Brooks, B. R.; Pande, V. S. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLoS Comput. Biol. 2017, 13(7), e1005659. [Google Scholar] [CrossRef]
- McGibbon, R. T.; Beauchamp, K. A.; Harrigan, M. P.; Klein, C.; Swails, J. M.; Hernández, C. X.; Schwantes, C. R.; Wang, L.-P.; Lane, T. J.; Pande, V. S. MDTraj: A modern open library for the analysis of molecular dynamics trajectories. Biophys. J. 2015, 109(8), 1528–1532. [Google Scholar] [CrossRef]
- Goddard, T. D.; Huang, C. C.; Meng, E. C.; Pettersen, E. F.; Couch, G. S.; Morris, J. H.; Ferrin, T. E. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 2018, 27(1), 14–25. [Google Scholar] [CrossRef]
- Pettersen, E. F.; Goddard, T. D.; Huang, C. C.; Meng, E. C.; Couch, G. S.; Croll, T. I.; Morris, J. H.; Ferrin, T. E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30(1), 70–82. [Google Scholar] [CrossRef]
- Li, Z.; Day, M.; Ding, J.; Faid, K. Synthesis and characterization of functional methacrylate copolymers and their application in molecular imprinting. Macromolecules 2005, 38(7), 2620–2625. [Google Scholar] [CrossRef]
- Tunc, Y.; Hasirci, N.; Yesilada, A.; Ulubayram, K. Comonomer effects on binding performances and morphology of acrylate-based imprinted polymers. Polymer 2006, 47(20), 6931–6940. [Google Scholar] [CrossRef]
- Kuru, C. I.; Ulucan, F.; Kuşat, K.; Akgöl, S. A model study by using polymeric molecular imprinting nanomaterials for removal of penicillin G. Environ. Monit. Assess. 2020, 192(6), 367. [Google Scholar] [CrossRef]
- Galván-Romero, V.; Gonzalez-Salazar, F.; Vargas-Berrones, K.; Flores, F. C.; Mendoza, B. N. Z.; de Loera Carrera, D. A.; Flores-Ramírez, R. Molecularly imprinted polymer systems for the controlled release of naphthoquinone derivatives with antimicrobial activity. React. Funct. Polym. 2025, 215, 106388. [Google Scholar] [CrossRef]
- Liu, Z.; Xu, Z.; Wang, D.; Yang, Y.; Duan, Y.; Ma, L.; Liu, H. A review on molecularly imprinted polymers preparation by computational simulation-aided methods. Polymers 2021, 13(16), 2657. [Google Scholar] [CrossRef]
- Mohsenzadeh, E.; Ratautaite, V.; Brazys, E.; Ramanavicius, S.; Zukauskas, S.; Plausinaitis, D.; Ramanavicius, A. Design of molecularly imprinted polymers (MIP) using computational methods: A review of strategies and approaches. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2024, 14(3), e1713. [Google Scholar] [CrossRef]
- Nicholls, I. A.; Golker, K.; Wiklander, J. G. The evolution of molecular dynamics as a tool for the study and development of molecularly imprinted materials: Status quo, quo vadis? Trends Anal. Chem. 194 2026, 118533. [Google Scholar] [CrossRef]
- Nicholls, I. A.; Golker, K.; Olsson, G. D.; Suriyanarayanan, S.; Wiklander, J. G. The use of computational methods for the development of molecularly imprinted polymers. Polymers 2021, 13(17), 2841. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




