Submitted:
28 April 2026
Posted:
30 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
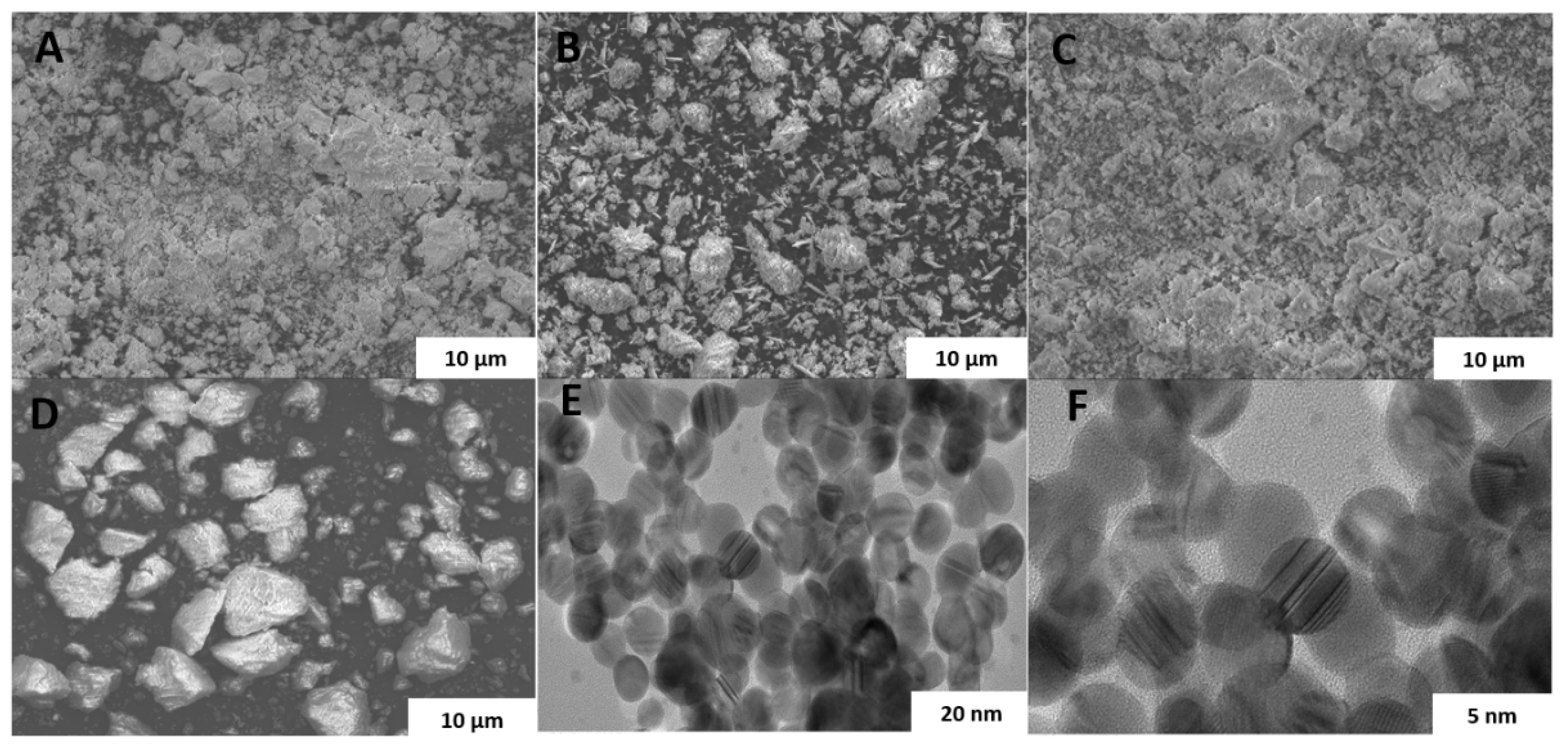
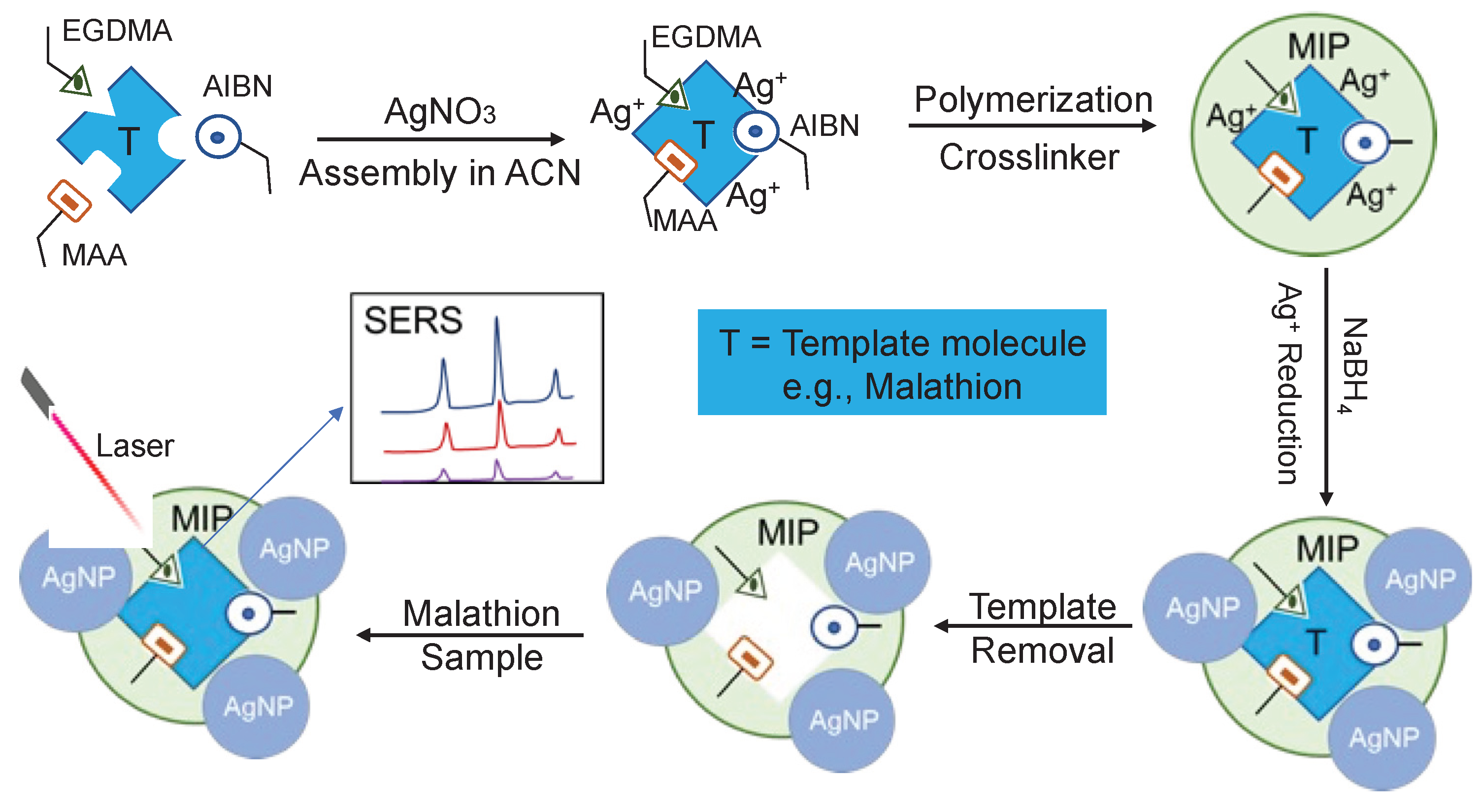
2.1. Synthesis and Characterization of AgNPs@MIPs
2.2. UV-Vis Analysis and AgNPs@MIPs Template Removal Process
2.3. UV-Vis Analysis and AgNPs@MIPs Template Removal Process
2.4. X-Ray Diffraction (XRD) Analysis of AgNPs@MIPs
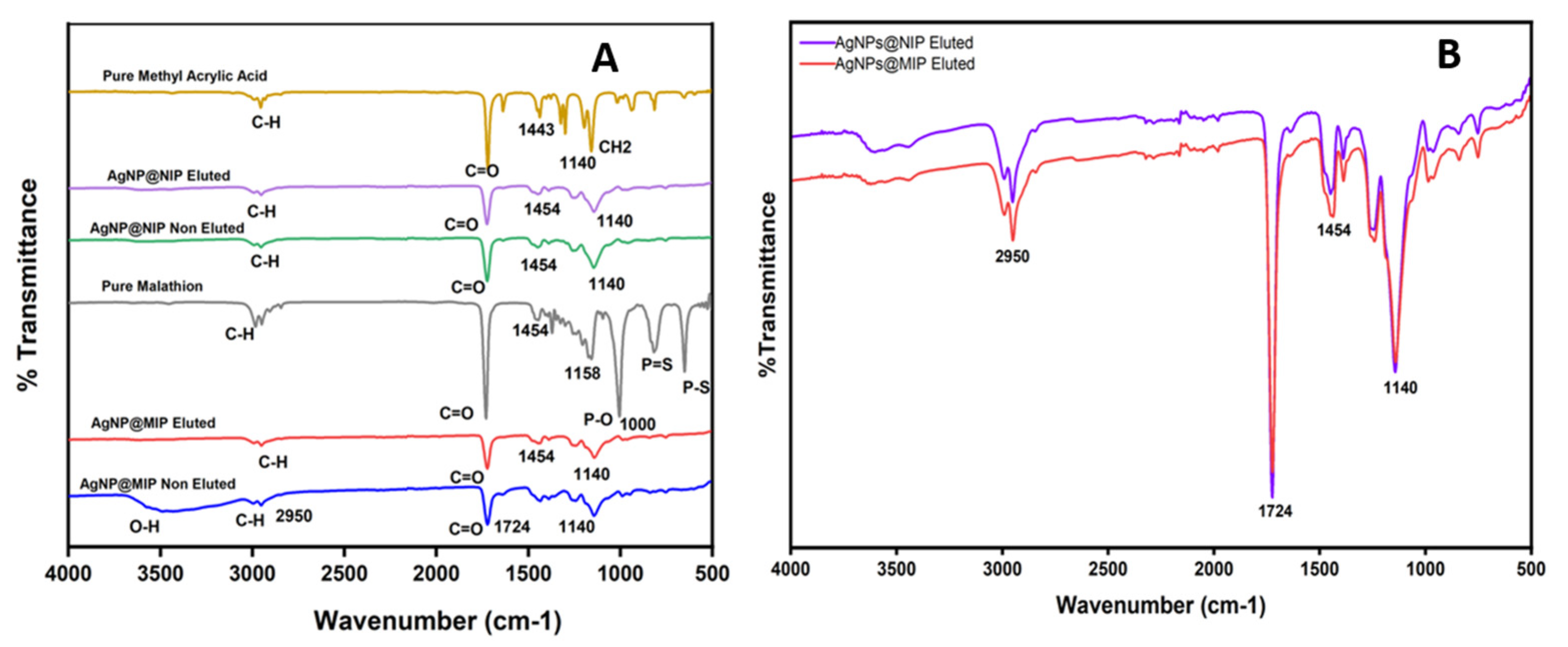
2.5. Fourier Transformed Infrared (FTIR) Analysis of the AgNPs@MIPs
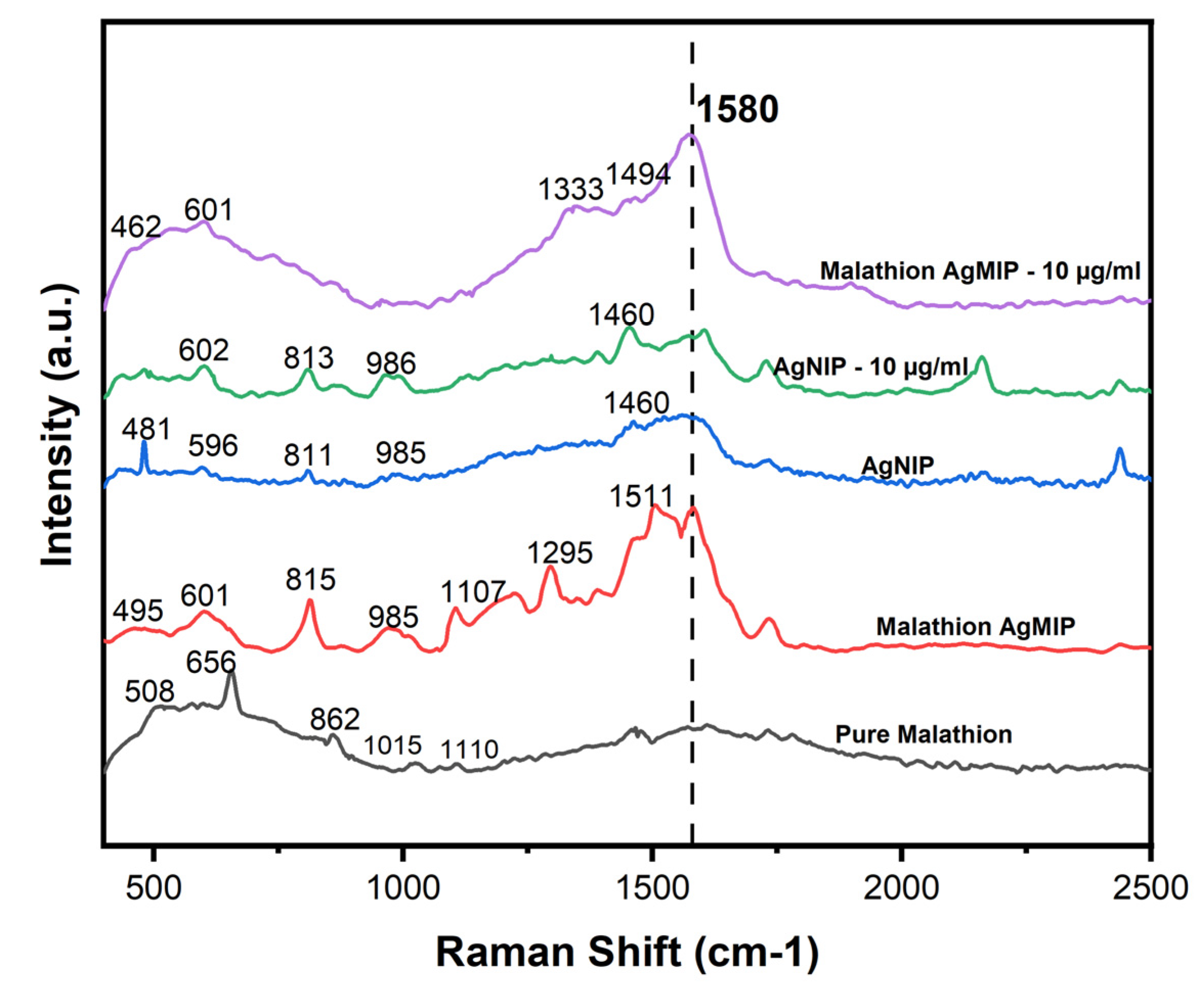
2.6. SERS Analysis of AgNPs@MIPs
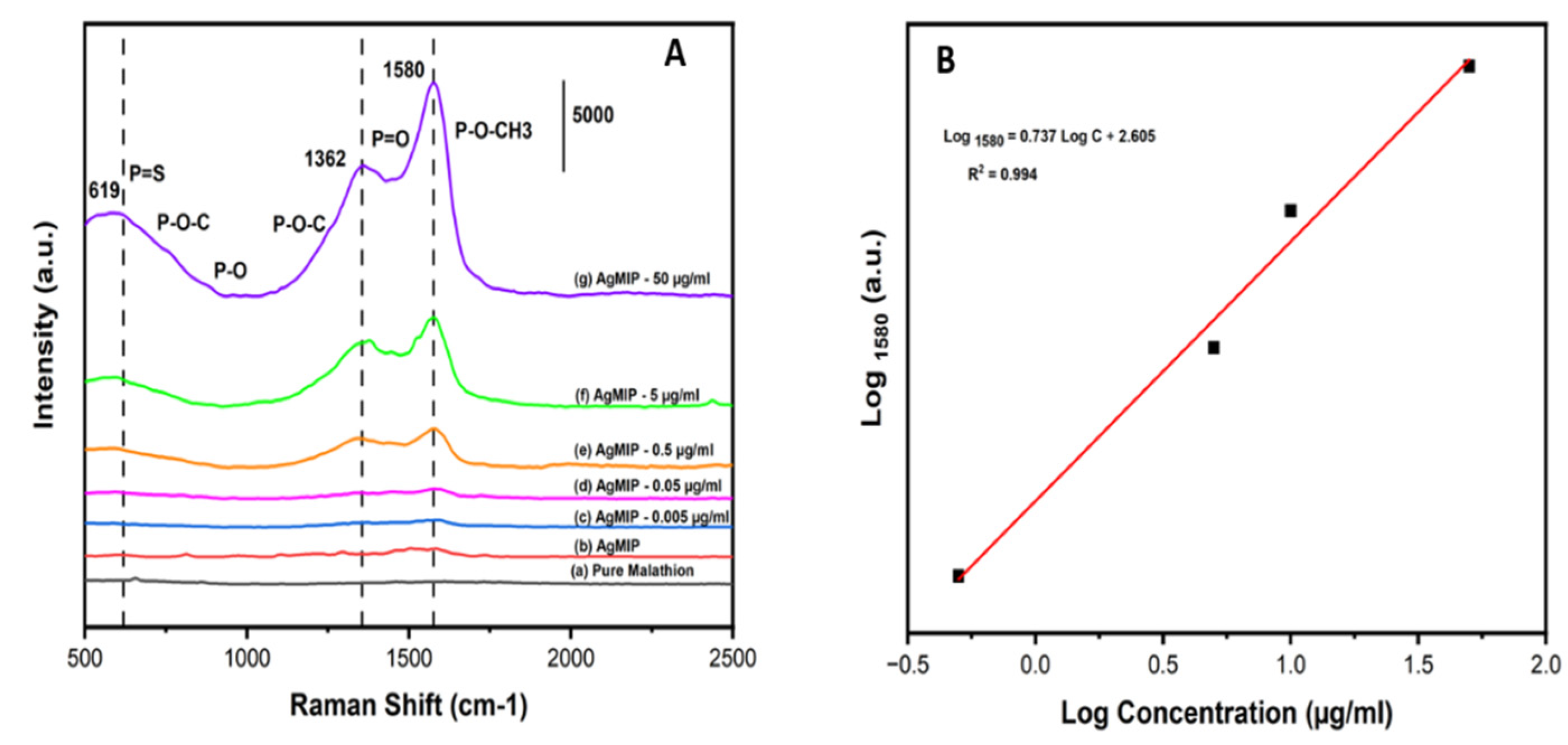
2.7. SERS Assay and Calibration of AgNPs@MIP with Malathion
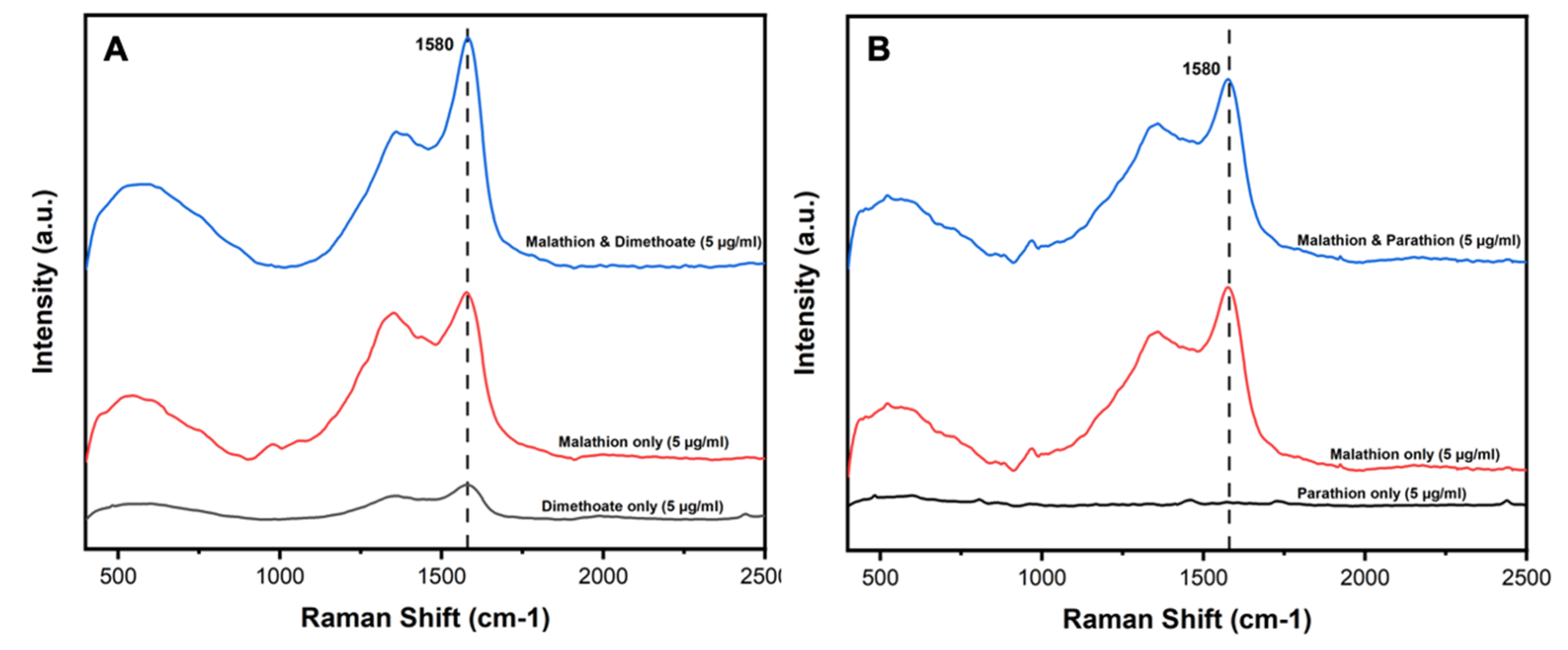
2.8. Selectivity of AgNPs@MIPs
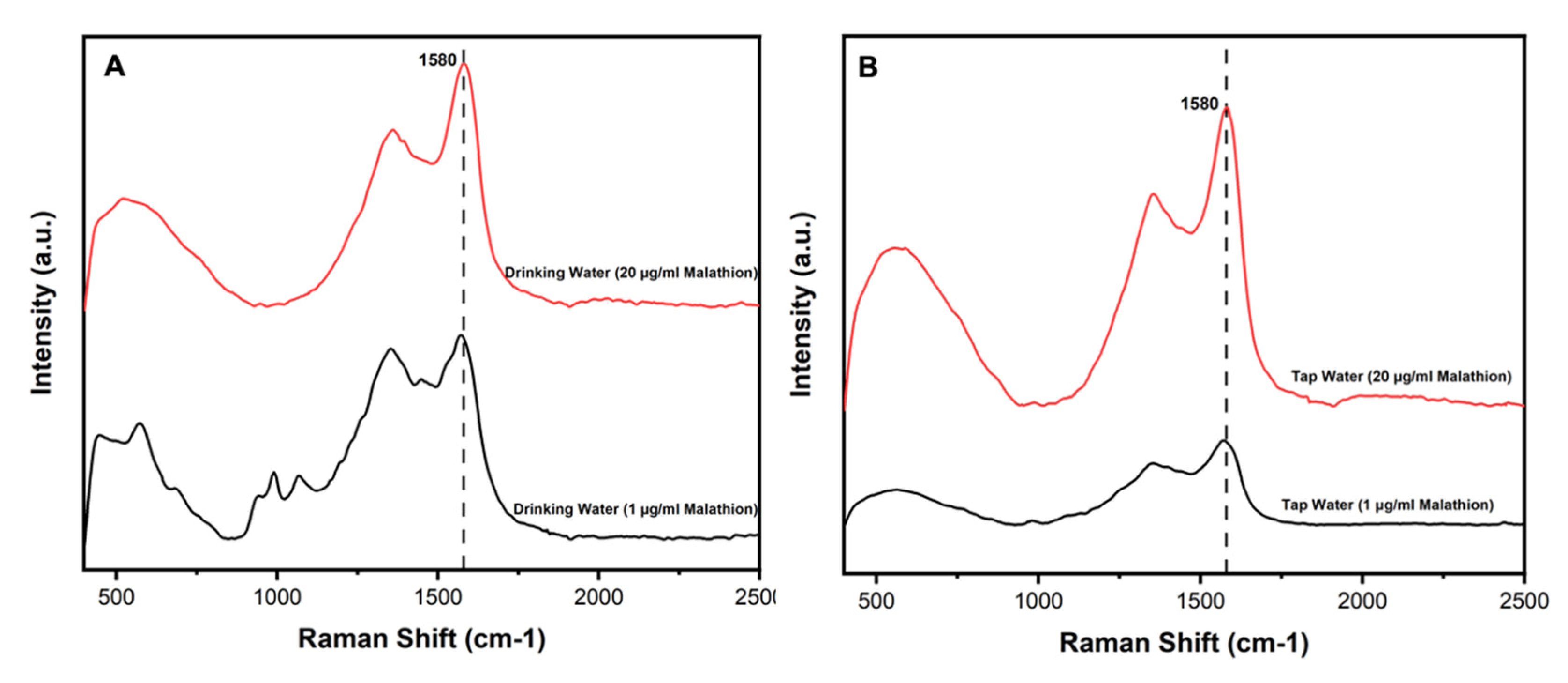
2.9. Detection of Malathion Spiked in Water Samples
3. Materials and Methods
4. Conclusions
5. Patents
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fuller, R.; Landrigan, P. J.; Balakrishnan, K.; Bathan, G.; Bose-O’Reilly, S.; Brauer, M.; Caravanos, J.; Chiles, T.; Cohen, A.; Corra, L.; Cropper, M.; Ferraro, G.; Hanna, J.; Hanrahan, D.; Hu, H.; Hunter, D.; Janata, G.; Kupka, R.; Lanphear, B.; Lichtveld, M.; Martin, K.; Mustapha, A.; Sanchez-Triana, E.; Sandilya, K.; Schaefli, L.; Shaw, J.; Seddon, J.; Suk, W.; Téllez-Rojo, M. M.; Yan, C. Pollution and health: a progress update. Lancet Planet. Health 2022, 6, e535–e547. [Google Scholar] [CrossRef]
- Cao, Z.; Lai, Z.; Bilawal Khaskheli, M.; Wang, L. The Risk of Global Environmental Change to Economic Sustainability and Law: Help from Digital Technology and Governance Regulation. Sustainability 2025, 17, 7094. [Google Scholar] [CrossRef]
- Shetty, S. S.; D, D.; S, H.; Sonkusare, S.; Naik, P. B.; Kumari N, S.; Madhyastha, H. Environmental pollutants and their effects on human health. Heliyon 2023, 9, e19496. [Google Scholar] [CrossRef]
- Anjitha, R.; Antony, A.; Shilpa, O.; Anupama, K. P.; Mallikarjunaiah, S.; Gurushankara, H. P. Malathion induced cancer-linked gene expression in human lymphocytes. Environ. Res. 2020, 182, 109131. [Google Scholar] [CrossRef]
- Duke, S. O. Pesticide Dose – A Parameter with Many Implications. In Pesticide Dose: Effects on the Environment and Target and Non-Target Organisms; American Chemical Society, 2017; Volume 1249, pp. 1–13. [Google Scholar]
- Gurusamy, L.; Cheng, R.-W.; Anandan, S.; Liu, C.-H.; Wu, J. J. Detection of Environmentally Harmful Malathion Pesticides Using a Bimetallic Oxide of CuO Nanoparticles Dispersed over a 3D ZnO Nanoflower. Materials 2023, 16, 7065. [Google Scholar] [CrossRef] [PubMed]
- Ayivi, R. D.; Obare, S. O.; Wei, J. Molecularly imprinted polymers as chemosensors for organophosphate pesticide detection and environmental applications. TrAC Trends Anal. Chem. 2023, 167, 117231. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhu, Y.; Zeng, Z.; Zeng, G.; Xiao, R.; Wang, Y.; Hu, Y.; Tang, L.; Feng, C. Sensors for the environmental pollutant detection: Are we already there? Coord. Chem. Rev. 2021, 431, 213681. [Google Scholar] [CrossRef]
- Li, H.; Haruna, S. A.; Sheng, W.; Bei, Q.; Ahmad, W.; Zareef, M.; Chen, Q.; Ding, Z. SERS-activated platforms for chemical contaminants in food: Probes, encoding methods, and detection. TrAC Trends Anal. Chem. 2023, 169, 117365. [Google Scholar] [CrossRef]
- Krämer, J.; Kang, R.; Grimm, L. M.; De Cola, L.; Picchetti, P.; Biedermann, F. Molecular Probes, Chemosensors, and Nanosensors for Optical Detection of Biorelevant Molecules and Ions in Aqueous Media and Biofluids. Chem. Rev. 2022, 122, 3459–3636. [Google Scholar] [CrossRef] [PubMed]
- Ayivi, R. D.; Adesanmi, B. O.; McLamore, E. S.; Wei, J.; Obare, S. O. Molecularly Imprinted Plasmonic Sensors as Nano-Transducers: An Effective Approach for Environmental Monitoring Applications. Chemosensors 2023, 11, 203. [Google Scholar] [CrossRef]
- Fu, Q.-B.; Sun, X.; Liu, L.; Zhao, R.-S.; Wang, X. Current Progress of Molecularly Imprinted Polymers for Emerging Pollutants Analysis in Food. J. Agric. Food. Chem. 2025, 73, 25103–25127. [Google Scholar] [CrossRef]
- Ramajayam, K.; Ganesan, S.; Ramesh, P.; Beena, M.; Kokulnathan, T.; Palaniappan, A. Molecularly Imprinted Polymer-Based Biomimetic Systems for Sensing Environmental Contaminants, Biomarkers, and Bioimaging Applications. Biomimetics 2023, 8, 245. [Google Scholar] [CrossRef]
- Wang, Z.; Yan, R.; Liao, S.; Miao, Y.; Zhang, B.; Wang, F.; Yang, H. In situ reduced silver nanoparticles embedded molecularly imprinted reusable sensor for selective and sensitive SERS detection of Bisphenol A. Appl. Surf. Sci. 2018, 457, 323–331. [Google Scholar] [CrossRef]
- Szaniawska, A.; Kudelski, A. Applications of Surface-Enhanced Raman Scattering in Biochemical and Medical Analysis. Front. Chem. 2021, 9, 664134. [Google Scholar] [CrossRef] [PubMed]
- Sultangaziyev, A.; Ilyas, A.; Dyussupova, A.; Bukasov, R. Trends in Application of SERS Substrates beyond Ag and Au, and Their Role in Bioanalysis. Biosensors 2022, 12, 967. [Google Scholar] [CrossRef]
- Feng, S.; Hu, Y.; Ma, L.; Lu, X. Development of molecularly imprinted polymers-surface-enhanced Raman spectroscopy/colorimetric dual sensor for determination of chlorpyrifos in apple juice. Sens. Actuator B-Chem. 2017, 241, 750–757. [Google Scholar] [CrossRef]
- Zhang, X.-F.; Liu, Z.-G.; Shen, W.; Gurunathan, S. Silver Nanoparticles: Synthesis, Characterization, Properties, Applications, and Therapeutic Approaches. Int. J. Mol. Sci. 2016, 17, 1534. [Google Scholar] [CrossRef] [PubMed]
- Sultan, A.; Mohammad, F. Chemical sensing, thermal stability, electrochemistry and electrical conductivity of silver nanoparticles decorated and polypyrrole enwrapped boron nitride nanocomposite. Polymer 2017, 113, 221–232. [Google Scholar] [CrossRef]
- Willner, M. R.; Vikesland, P. J. Nanomaterial enabled sensors for environmental contaminants. J. Nanobiotechnol. 2018, 16, 95. [Google Scholar] [CrossRef]
- Çakır, O.; Bakhshpour, M.; Göktürk, I.; Yılmaz, F.; Baysal, Z. Sensitive and selective detection of amitrole based on molecularly imprinted nanosensor. J. Mol. Recognit. 2021, 34, e2929. [Google Scholar] [CrossRef]
- Hoang, V.-T.; Dinh, N. X.; Nhat Trang, N. L.; Khi, N. T.; Quy, N. V.; Tuan, P. A.; Tri, D. Q.; Thang, L. H.; Huy, T. Q.; Le, A.-T. Functionalized silver nanoparticles-based efficient colorimetric platform: Effects of surface capping agents on the sensing response of thiram pesticide in environmental water samples. Mater. Res. Bull. 2021, 139, 111278. [Google Scholar] [CrossRef]
- Jiang, G.-y.; Liu, L.; Wan, Y.-q.; Li, J.-k.; Pi, F.-w. Surface-enhanced Raman scattering based determination on sulfamethazine using molecularly imprinted polymers decorated with silver nanoparticles. Microchim. Acta 2023, 190, 169. [Google Scholar] [CrossRef]
- Hu, R.; Tang, R.; Xu, J.; Lu, F. Chemical nanosensors based on molecularly-imprinted polymers doped with silver nanoparticles for the rapid detection of caffeine in wastewater. Anal. Chim. Acta 2018, 1034, 176–183. [Google Scholar] [CrossRef]
- Neng, J.; Liao, C.; Wang, Y.; Wang, Y.; Yang, K. Rapid and Sensitive Detection of Pentachloronitrobenzene by Surface-Enhanced Raman Spectroscopy Combined with Molecularly Imprinted Polymers. Biosensors 2022, 12, 52. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.; Liu, Y.; Wei, J. Recent advances in surface-enhanced raman spectroscopy (SERS): Finite-difference time-domain (FDTD) method for SERS and sensing applications. TrAC Trends Anal. Chem. 2016, 75, 162–173. [Google Scholar] [CrossRef]
- Reguera, J.; Langer, J.; Jiménez de Aberasturi, D.; Liz-Marzán, L. M. Anisotropic metal nanoparticles for surface enhanced Raman scattering. Chem. Soc. Rev. 2017, 46, 3866–3885. [Google Scholar] [CrossRef] [PubMed]
- Krystosiak, P.; Tomaszewski, W.; Megiel, E. High-density polystyrene-grafted silver nanoparticles and their use in the preparation of nanocomposites with antibacterial properties. J. Colloid Interface Sci. 2017, 498, 9–21. [Google Scholar] [CrossRef]
- Montes-Hernandez, G.; Di Girolamo, M.; Sarret, G.; Bureau, S.; Fernandez-Martinez, A.; Lelong, C.; Eymard Vernain, E. In Situ Formation of Silver Nanoparticles (Ag-NPs) onto Textile Fibers. ACS Omega 2021, 6, 1316–1327. [Google Scholar] [CrossRef]
- Zhang, C.; Chen, S.; Jiang, Z.; Shi, Z.; Wang, J.; Du, L. Highly Sensitive and Reproducible SERS Substrates Based on Ordered Micropyramid Array and Silver Nanoparticles. ACS Appl. Mater. Interfaces 2021, 13, 29222–29229. [Google Scholar] [CrossRef]
- Xu, K.; Zhou, R.; Takei, K.; Hong, M. Toward Flexible Surface-Enhanced Raman Scattering (SERS) Sensors for Point-of-Care Diagnostics. Adv. Sci. 2019, 6, 1900925. [Google Scholar] [CrossRef]
- Kadhem, A. J.; Gentile, G. J.; Fidalgo de Cortalezzi, M. M. Molecularly Imprinted Polymers (MIPs) in Sensors for Environmental and Biomedical Applications: A Review. Molecules 2021, 26, 6233. [Google Scholar] [CrossRef] [PubMed]
- Susanti, I.; Hasanah, A. N. How to develop molecularly imprinted mesoporous silica for selective recognition of analytes in pharmaceutical, environmental, and food samples. Polym. Adv. Technol. 2021, 32, 1965–1980. [Google Scholar] [CrossRef]
- Kimani, M.; Kislenko, E.; Gawlitza, K.; Rurack, K. Fluorescent molecularly imprinted polymer particles for glyphosate detection using phase transfer agents. Sci. Rep. 2022, 12, 14151. [Google Scholar] [CrossRef]
- Amarh, F. A.; Kangmennaa, A.; Agorku, E. S.; Voegborlo, R. B. A state-of-the-art review of trends in molecularly imprinted polymers in the clean-up of pesticides in environmental samples. Sustain. Environ. 2024, 10, 2298067. [Google Scholar] [CrossRef]
- Ali, M. H.; Azad, M. A. K.; Khan, K. A.; Rahman, M. O.; Chakma, U.; Kumer, A. Analysis of Crystallographic Structures and Properties of Silver Nanoparticles Synthesized Using PKL Extract and Nanoscale Characterization Techniques. ACS Omega 2023, 8, 28133–28142. [Google Scholar] [CrossRef]
- Oliveira, A. E. F.; Pereira, A. C.; de Resende, M. A. C.; Ferreira, L. F. Synthesis of a silver nanoparticle ink for fabrication of reference electrodes. Talanta Open. 2022, 5, 100085. [Google Scholar] [CrossRef]
- Kalogiouri, N. P.; Kabir, A.; Furton, K. G.; Samanidou, V. F. A tutorial on the synthesis and applications of molecularly imprinted polymers in analytical chemistry. J. Chromatogr. Open. 2025, 8, 100244. [Google Scholar] [CrossRef]
- Gong, Y.; Chen, X.; Wu, W. Application of fourier transform infrared (FTIR) spectroscopy in sample preparation: Material characterization and mechanism investigation. Adv. Sample Prep. 2024, 11, 100122. [Google Scholar] [CrossRef]
- Royani, I.; Widayani; Abdullah, M.; Khairurrijal. An Atrazine Molecularly Imprinted Polymer Synthesized Using a Cooling-Heating Method with Repeated Washing: Its Physicochemical Characteristics and Enhanced Cavities. Int. J. Electrochem. Sci. 2014, 9, 5651–5662. [Google Scholar] [CrossRef]
- Askari, F.; Zandi, M.; Shokrolahi, P.; Tabatabaei, M. H.; Hajirasoliha, E. Reduction in protein absorption on ophthalmic lenses by PEGDA bulk modification of silicone acrylate-based formulation. Prog. Biomater. 2019, 8, 169–183. [Google Scholar] [CrossRef]
- Roland, R. M.; Bhawani, S. A.; Ibrahim, M. N. M. Synthesis of molecularly imprinted polymer by precipitation polymerization for the removal of ametryn. BMC Chem. 2023, 17, 165. [Google Scholar] [CrossRef] [PubMed]
- Roland, R. M.; Bhawani, S. A.; Ibrahim, M. N. M. Synthesis of molecularly imprinted polymer for the removal of cyanazine from aqueous samples. Chem. Biol. Technol. Agric. 2023, 10, 92. [Google Scholar] [CrossRef]
- Aheto, J. H.; Huang, X.; Tian, X.; Zhang, X.; Zhang, W.; Yu, S. Activated carbon@silver nanoparticles conjugates as SERS substrate for capturing malathion analyte molecules for SERS detection. J. Food Saf. 2023, 43, e13072. [Google Scholar] [CrossRef]
- Al’Abri, A. M.; Abdul Halim, S. N.; Abu Bakar, N. K.; Saharin, S. M.; Sherino, B.; Rashidi Nodeh, H.; Mohamad, S. Highly sensitive and selective determination of malathion in vegetable extracts by an electrochemical sensor based on Cu-metal organic framework. J. Environ. Sci. Health B 2019, 54, 930–941. [Google Scholar] [CrossRef] [PubMed]
- Roland, R. M.; Bhawani, S. A. Synthesis and Characterization of Molecular Imprinting Polymer Microspheres of Piperine: Extraction of Piperine from Spiked Urine. J. Anal. Methods Chem. 2016, 2016, 5671507. [Google Scholar] [CrossRef]
- Joshi, N.; Jain, N.; Pathak, A.; Singh, J.; Prasad, R.; Upadhyaya, C. P. Biosynthesis of silver nanoparticles using Carissa carandas berries and its potential antibacterial activities. J. Sol.-Gel Sci. Technol. 2018, 86, 682–689. [Google Scholar] [CrossRef]
- Guo, X.; Li, J.; Arabi, M.; Wang, X.; Wang, Y.; Chen, L. Molecular-Imprinting-Based Surface-Enhanced Raman Scattering Sensors. ACS Sens. 2020, 5, 601–619. [Google Scholar] [CrossRef]
- Aboualigaledari, N.; Jayapalan, A.; Tukur, P.; Liu, M.; Tukur, F.; Zhang, Y.; Ducatte, G.; Verma, M.; Tarus, J.; Hunyadi Murph, S. E.; Wei, J. Surface-enhanced Raman scattering enhancement using a hybrid gold nanoparticles@carbon nanodot substrate for herbicide detection. Analyst 2024, 149, 5277–5286. [Google Scholar] [CrossRef]
- Tukur, F.; Tukur, P.; Hunyadi Murph, S. E.; Wei, J. Advancements in mercury detection using surface-enhanced Raman spectroscopy (SERS) and ion-imprinted polymers (IIPs): a review. Nanoscale 2024, 16, 11384–11410. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Aljuhani, W.; Zhang, Y.; Ye, Z.; Li, C.; Bell, S. E. J. A practical approach to quantitative analytical surface-enhanced Raman spectroscopy. Chem. Soc. Rev. 2025, 54, 62–84. [Google Scholar] [CrossRef]
- Tarannum, N.; Kumar, D.; Yadav, A.; Yadav, A. K. Raman spectroscopy-based molecularly imprinted polymer sensor for sensitive detection of lysophosphatidic acid in serum. J. Raman Spectrosc. 2024, 55, 809–818. [Google Scholar] [CrossRef]
- Lazarević-Pašti, T.; Jocić, A.; Milanković, V.; Tasić, T.; Batalović, K.; Breitenbach, S.; Unterweger, C.; Fürst, C.; Pašti, I. A. Investigating the Adsorption Kinetics of Dimethoate, Malathion and Chlorpyrifos on Cellulose-Derived Activated Carbons: Understanding the Influence of Physicochemical Properties. C 2023, 9, 103. [Google Scholar] [CrossRef]
- Wu, H.; Luo, Y.; Huang, Y.; Dong, Q.; Hou, C.; Huo, D.; Zhao, J.; Lei, Y. A Simple SERS-Based Trace Sensing Platform Enabled by AuNPs-Analyte/AuNPs Double-Decker Structure on Wax-Coated Hydrophobic Surface. Front. Chem. 2018, 6, 482. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Johnson, E. R.; Giammar, D. E. Lead Phosphate Particles in Tap Water: Challenges for Point-of-Use Filters. Environ. Sci. Technol. Lett. 2021, 8, 244–249. [Google Scholar] [CrossRef]
- Jereb, G.; Poljšak, B.; Eržen, I. Contribution of Drinking Water Softeners to Daily Phosphate Intake in Slovenia. Int. J. Environ. Res. Public Health 2017, 14, 1186. [Google Scholar] [CrossRef]
- Alizadeh, T. High Selective Parathion Voltammetric Sensor Development by Using an Acrylic Based Molecularly Imprinted Polymer-Carbon Paste Electrode. Electroanalysis 2009, 21, 1490–1498. [Google Scholar] [CrossRef]
- Yao, G.-H.; Liang, R.-P.; Huang, C.-F.; Wang, Y.; Qiu, J.-D. Surface Plasmon Resonance Sensor Based on Magnetic Molecularly Imprinted Polymers Amplification for Pesticide Recognition. Anal. Chem. 2013, 85, 11944–11951. [Google Scholar] [CrossRef]
- Wang, Y.-L.; Gao, Y.-L.; Wang, P.-P.; Shang, H.; Pan, S.-Y.; Li, X.-J. Sol–gel molecularly imprinted polymer for selective solid phase microextraction of organophosphorous pesticides. Talanta 2013, 115, 920–927. [Google Scholar] [CrossRef]
- Wang, F.-R.; Lee, G.-J.; Haridharan, N.; Wu, J. J. Electrochemical Sensor Using Molecular Imprinting Polymerization Modified Electrodes to Detect Methyl Parathion in Environmental Media. Electrocatalysis 2018, 9, 1–9. [Google Scholar] [CrossRef]

| Sample | Spiked (µg/mL) | Mesured (µg/mL) | Recovery (± RSD) |
|---|---|---|---|
| Drinking Water | 1 | 0.93 | 93.00% ± 0.05% |
| Drinking Water | 20 | 19.88 | 99.40% ± 0.43% |
| Tap Water | 1 | 0.99 | 98.00 ± 0.71% |
| Tap Water | 20 | 20.10 | 100.50 ± 0.35% |
| Polymerization Technique | MIP Sensing Technique | Template /Analyte | Sample Source | LOD (µg/mL) | Linear range (µg/mL) | Ref. |
|---|---|---|---|---|---|---|
| Precipitation | Electrochemical | Parathion | Cabbage | 1.46 × 10−4 | 4.95 × 10−4 – 0.262 | [57] |
| Magnetic Imprinting | SPR | Chlorpyriphos | Apple Juice | 2.66 × 10−4 | 3.51 × 10−4 – 3.51 | [58] |
| Sol-Gel | SPME-GC | Diazinon | Vegetable | 4.8 × 10−5 | 1.7 × 10−5 – 7.7 × 10−4 | [59] |
| Surface Imprinting | Electrochemical | Methyl Parathion | Tangerine Juice | 2.63 × 10−3 | 0.013 – 3.95 | [60] |
| Precipitation | Surface Enhanced Raman | Malathion | Tap / Drinking Water | 0.005 | 0.005 – 50 | This Work |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




