Submitted:
28 April 2026
Posted:
29 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Strains, Plasmids and Chemicals
2.2. Screening of 4-EG-Producing Strains
2.3. Identification and Physiological Characterization of the 4-EG-Producing Strain
2.4. Cloning and Sequence Analysis of the BcPAD Gene
2.5. Heterologous Expression and Purification of Recombinant BcPAD
2.6. Determination of Recombinant BcPAD Enzyme Activity
2.7. Effects of pH, Temperature, Metal Ions, and Chemical Reagents on the Activity of BcPAD
2.8. Substrate Specificity and Kinetic Parameters of BcPAD
2.9. Structural Analysis of BcPAD
2.10. Statistical Analysis
3. Results and Discussion
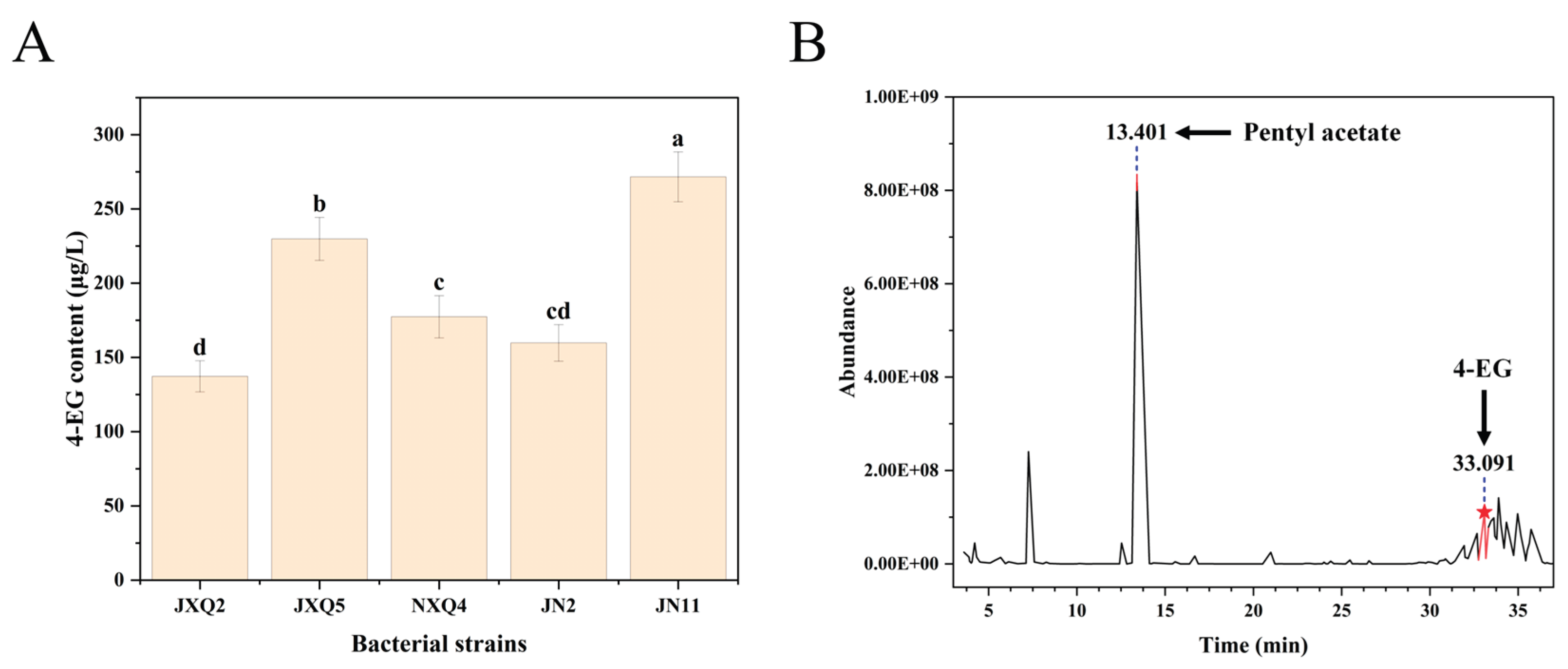
3.1. Screening and Analysis of 4-EG-Producing Strains
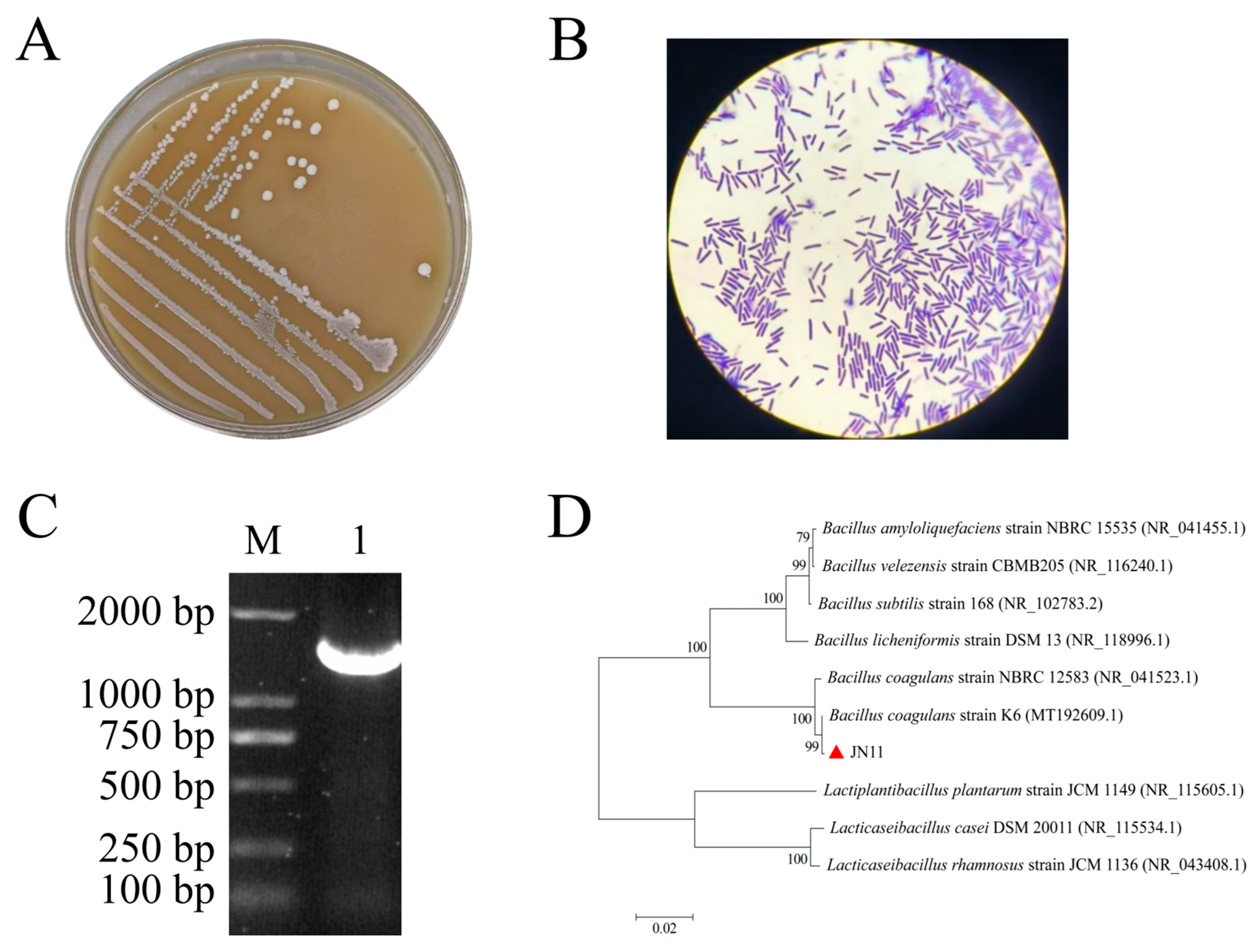
3.2. Identification of JN11 Strain
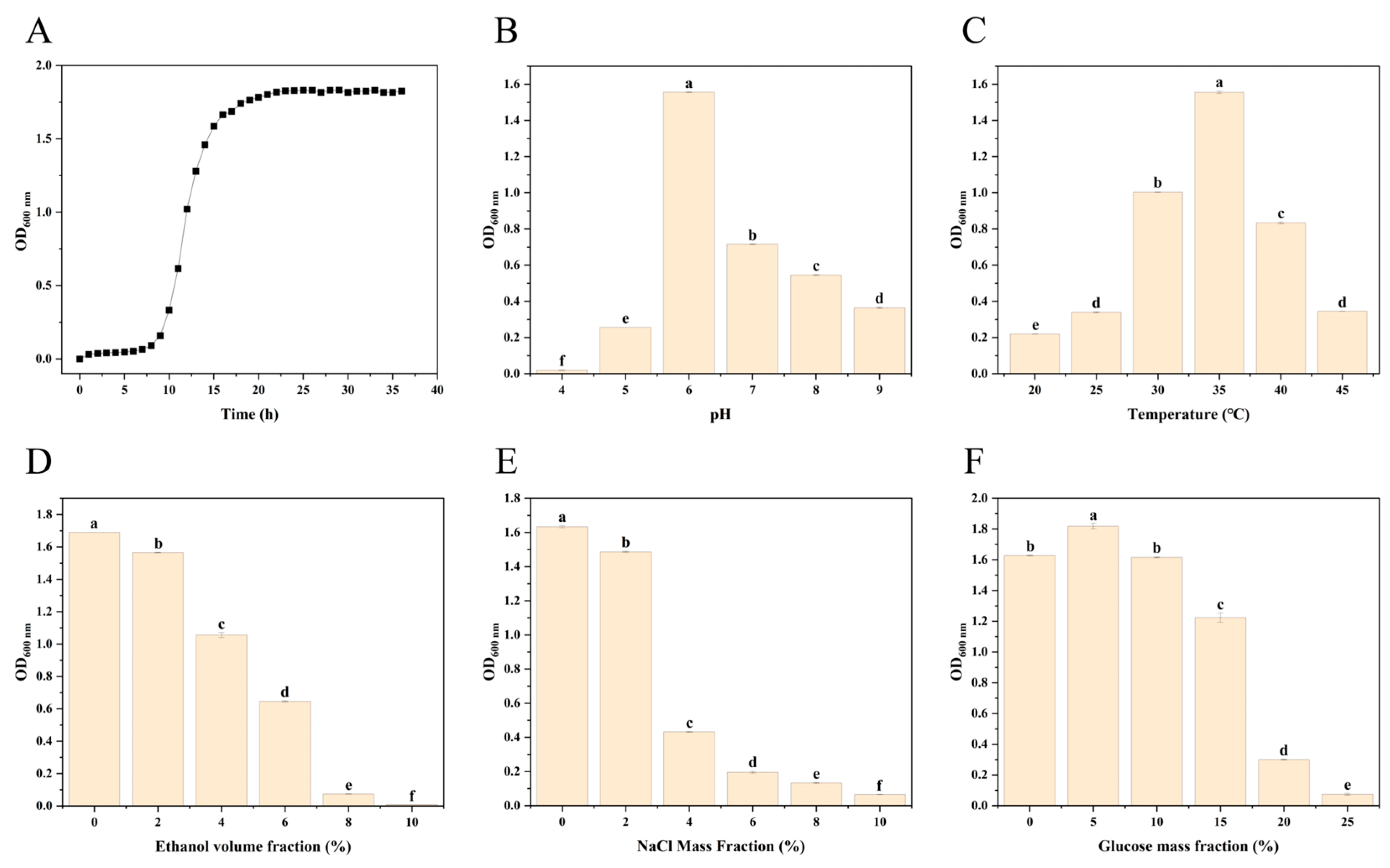
3.3. Physiological Characteristics of Bacillus Coagulans JN11
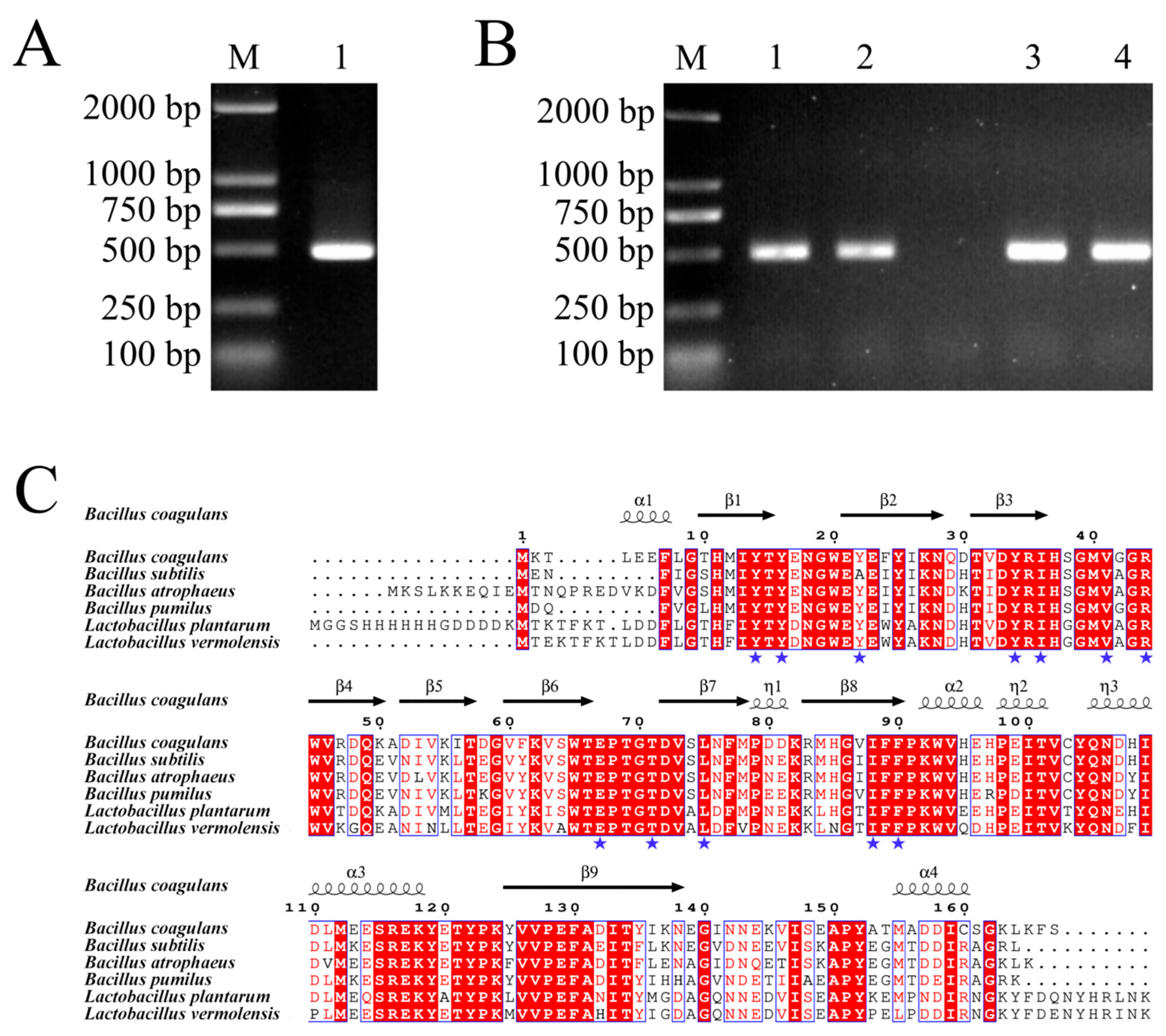
3.4. Gene Cloning and Bioinformatics Analysis of BcPAD
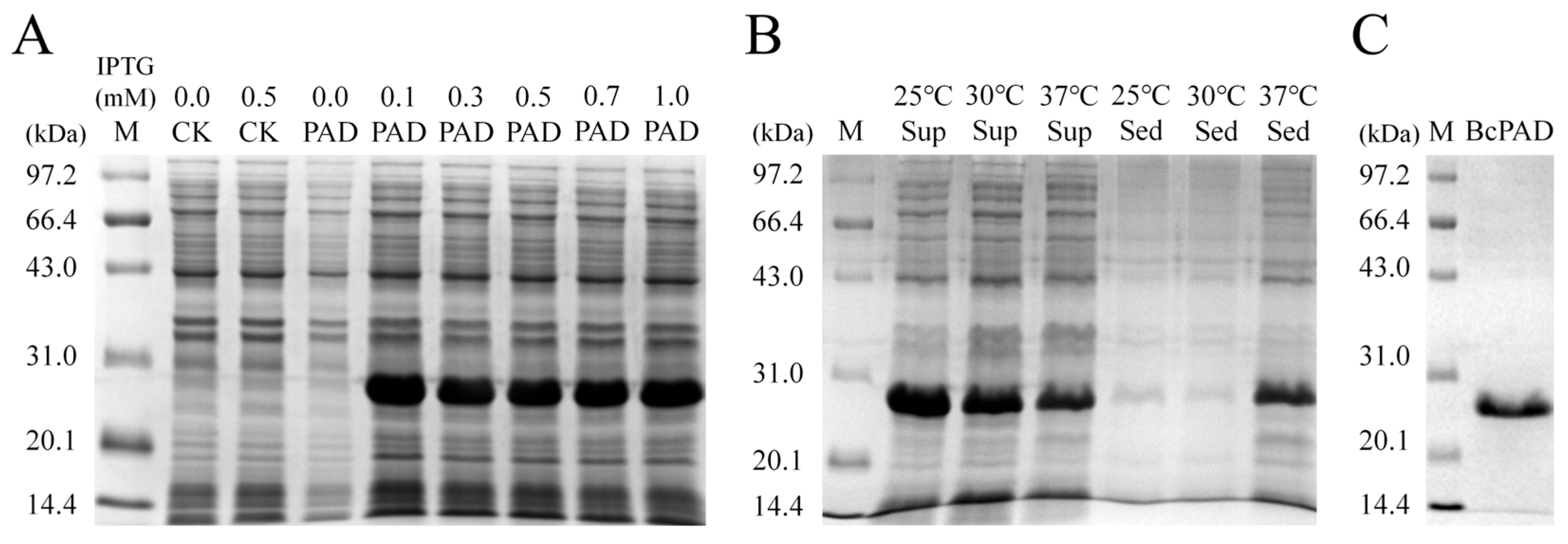
3.5. Expression and Purification of Recombinant BcPAD
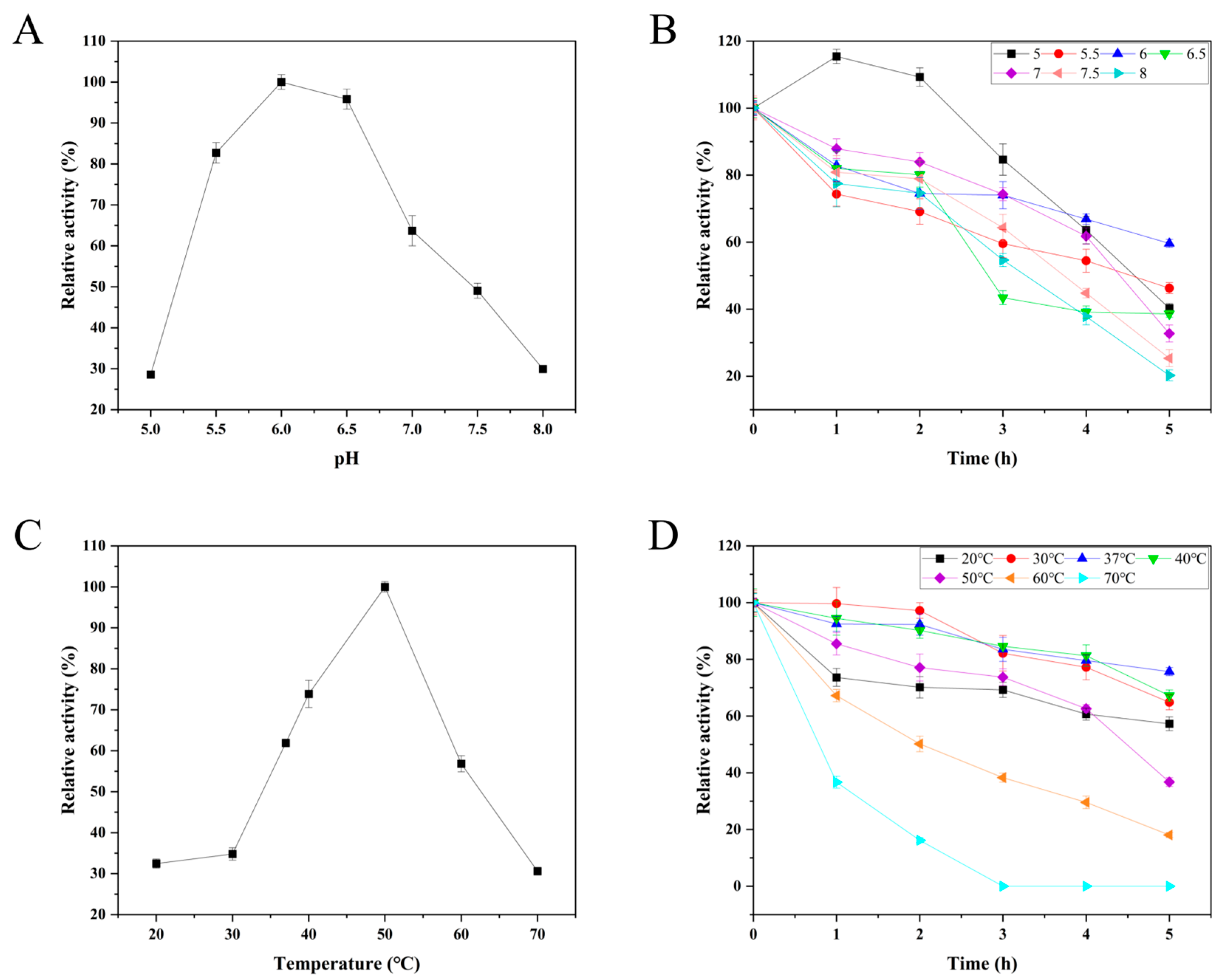
3.6. Effects of pH and Temperature on the Activity and Stability of BcPAD
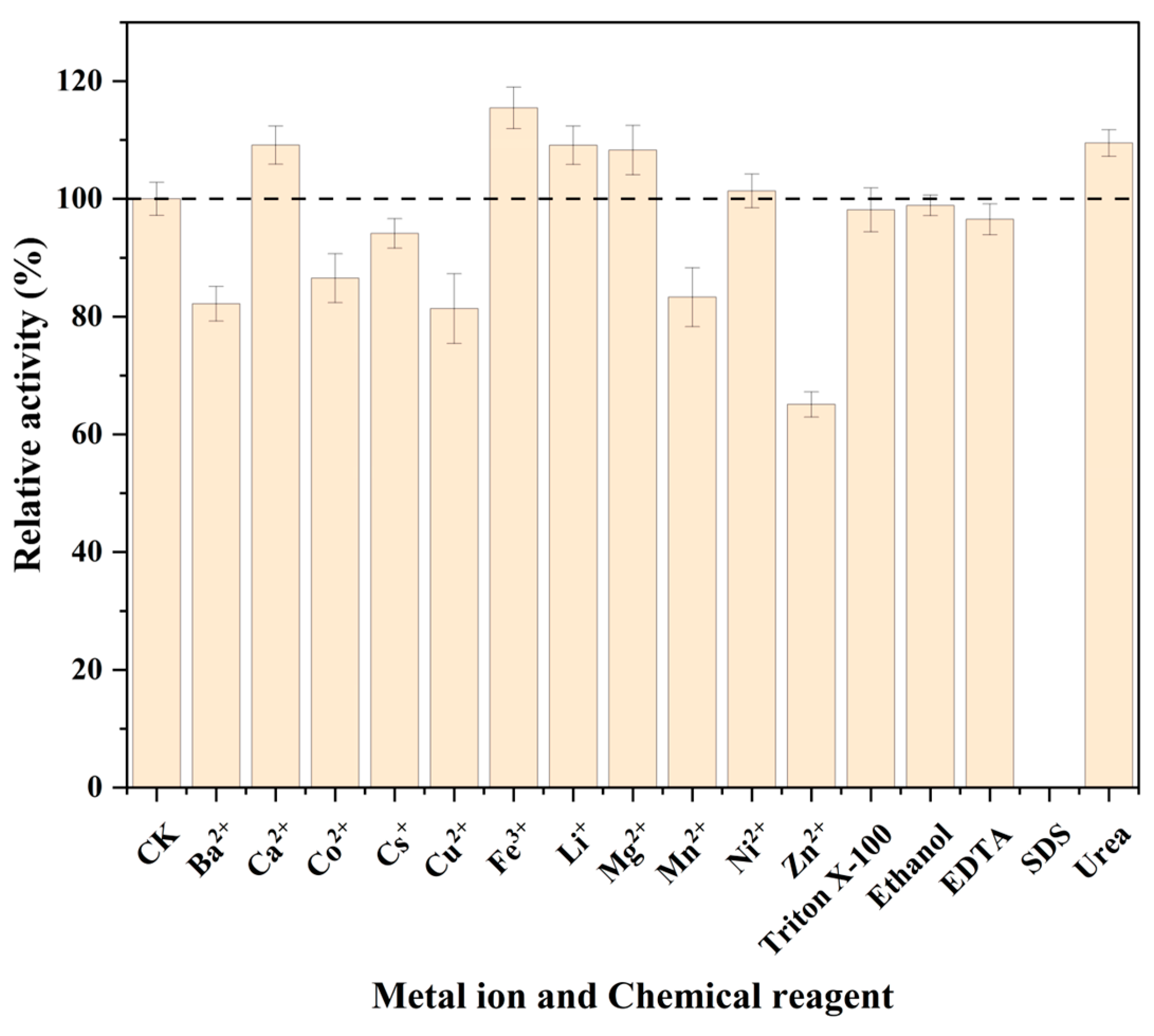
3.7. Effects of Metal Ions and Chemical Reagents on the Activity of BcPAD
3.8. Substrate Specificity and Kinetic Parameters of BcPAD
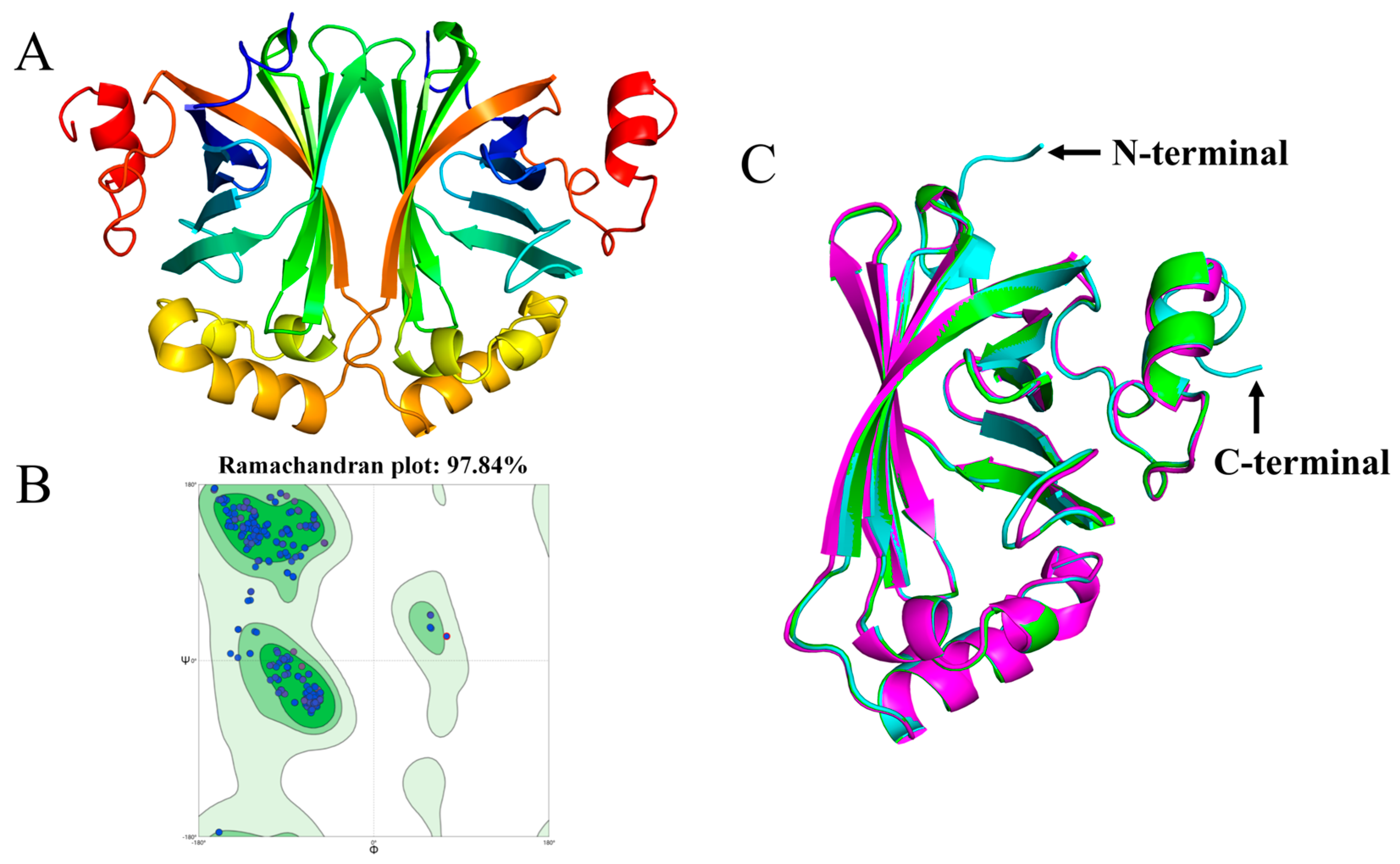
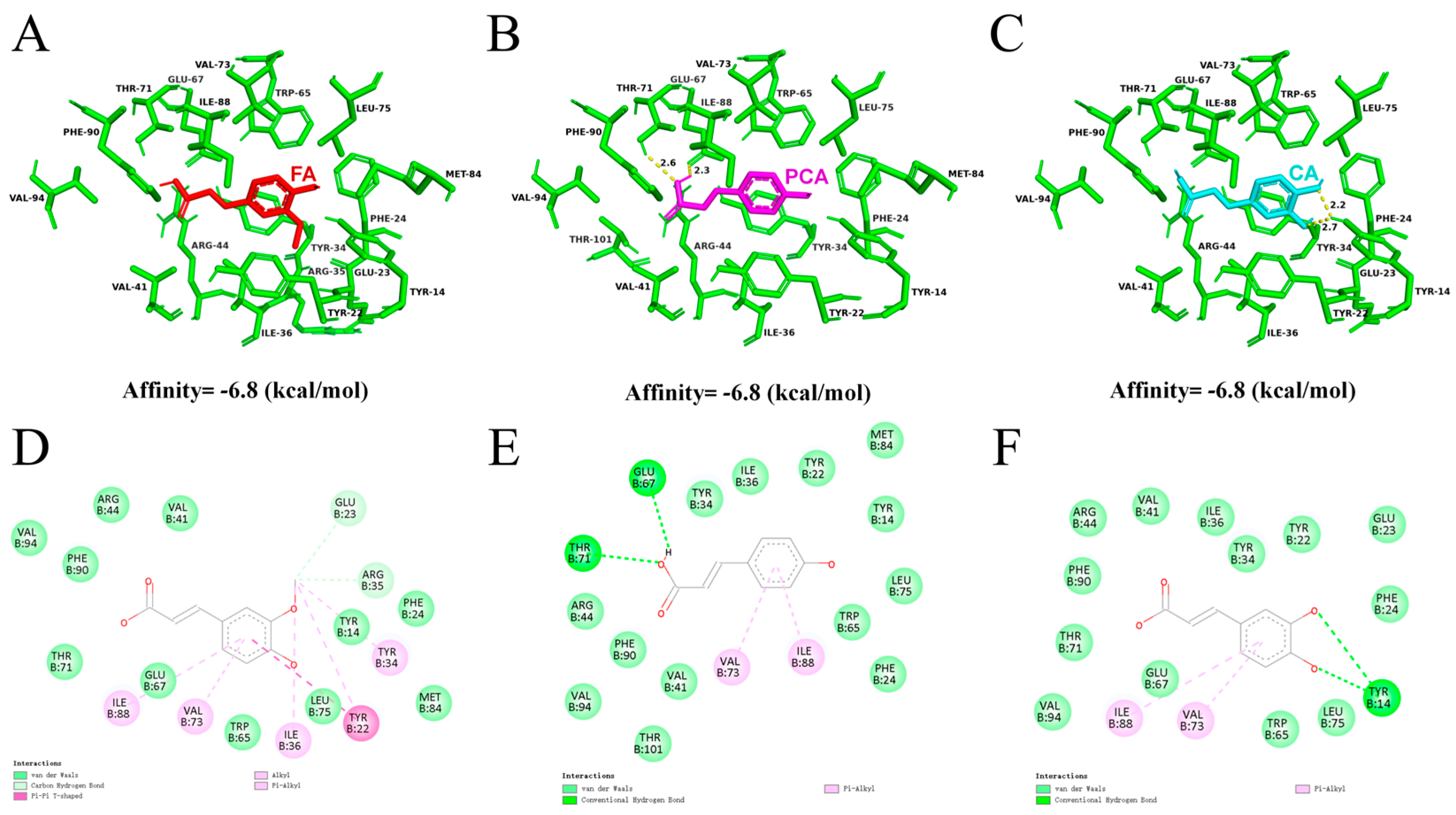
3.9. Structural Analysis of BcPAD
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, Y.; Yu, X.; Zhao, X.; Liu, L.; Huang, Y.; Zhang, J.; Liu, J. Screening and identification of 4-ethylguaiacol-producing strain and its application in soy sauce brewing. China Brew. 2023, 42, 115–121. [Google Scholar]
- Zhao, Y.; Zhu, B.; Zhou, Z.; Wu, Z.; Zhang, W. Cloning and characterization of phenolic acid decarboxylase responsible for aromatic volatile phenols production in Paocai based on metatranscriptomics. Food Biosci. 2023, 55, 102953. [Google Scholar] [CrossRef]
- Wang, C.; Li, L.; Fan, M.; Liu, J.; Yuan, S. Screening, identification and characterization of 4-ethylguaiacol producing Bacillus strains from jiangxiang Daqu. Liquor-Mak. Sci. Technol. 2023, 346, 45–52. [Google Scholar]
- Zhao, D.; Shi, D.; Sun, J.; Li, H.; Zhao, M.; Sun, B. Quantification and cytoprotection by vanillin, 4-methylguaiacol and 4-ethylguaiacol against AAPH-induced abnormal oxidative stress in HepG2 cells. Rsc Adv. 2018, 8, 35474–35484. [Google Scholar] [CrossRef]
- Zhao, D.; Jiang, Y.; Sun, J.; Li, H.; Luo, X.; Zhao, M. Anti-inflammatory mechanism involved in 4-ethylguaiacol-mediated inhibition of LPS-induced inflammation in THP-1 cells. J. Agric. Food. Chem. 2019, 67, 1230–1243. [Google Scholar] [CrossRef]
- Weng, W.; Kuo, P.; Scofield, B.A.; Paraiso, H.C.; Brown, D.A.; Yu, I.; Yen, J. 4-Ethylguaiacol modulates neuroinflammation and promotes heme oxygenase-1 expression to ameliorate brain injury in ischemic stroke. Front. Immunol. 2022, 13, 887000. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, S.; Liao, P.; Chen, L.; Sun, J.; Sun, B.; Zhao, D.; Wang, B.; Li, H. HS-SPME combined with GC-MS/O to analyze the flavor of strong aroma Baijiu Daqu. Foods 2022, 11, 116. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Cao, R.; Shen, F.; Xiang, X.; Yan, S. Screening and identification of a 4-ethyl guaiacol-producing strain from nongxiang Daqu. Liquor-Mak. Sci. Technol. 2018, 287, 48–52. [Google Scholar]
- Hou, R.; Wang, Y.; Yang, J.; Wei, H.; Yang, F.; Jin, Z.; Hu, J.; Gao, M. Bioconversion of waste generated during ethanol production into value-added products for sustainable utilization of rice straw. Biochem. Eng. J. 2020, 164, 107761. [Google Scholar] [CrossRef]
- Landete, J.M.; Rodríguez, H.; Curiel, J.A.; de Las Rivas, B.; Mancheño, J.M.; Muñoz, R. Gene cloning, expression, and characterization of phenolic acid decarboxylase from Lactobacillus brevis RM84. J. Ind. Microbiol. Biotechnol. 2010, 37, 617–624. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Wu, S.; Hua, X.; Fan, Y.; Li, X. Ferulic acid triggering a co-production of 4-vinyl guaiacol and fumaric acid from lignocellulose-based carbon source by Rhizopus oryzae. Food Chem. 2024, 461, 140799. [Google Scholar] [CrossRef]
- Zhou, W.; Zhang, Q.; Huang, K.; Huang, Z.; Ding, W.; Sun, J.; Bai, W. Accelerating the conversion of black chokeberry anthocyanins toward vinylphenolic pyranoanthocyanins by displaying phenolic acid decarboxylase from Lactiplantibacillus plantarum on the surface of Pichia pastoris. Food Chem. 2025, 476, 143408. [Google Scholar] [CrossRef] [PubMed]
- Mathew, S.; Abraham, T.E. Bioconversions of ferulic acid, an hydroxycinnamic acid. Crit. Rev. Microbiol. 2006, 32, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Qin, C.; Li, Q.; Hu, K.; Li, J.; Liu, S. Research progress in phenolic acid decarboxylase derived from microorganisms. Food Sci. 2024, 45, 323–332. [Google Scholar]
- Gu, W.; Li, X.; Huang, J.; Duan, Y.; Meng, Z.; Zhang, K.; Yang, J. Cloning, sequencing, and overexpression in Escherichia coli of the Enterobacter sp. Px6-4 gene for ferulic acid decarboxylase. Appl. Microbiol. Biotechnol. 2011, 89, 1797–1805. [Google Scholar] [CrossRef]
- Cavin, J.F.; Dartois, V.; Diviès, C. Gene cloning, transcriptional analysis, purification, and characterization of phenolic acid decarboxylase from Bacillus subtilis. Appl. Environ. Microbiol. 1998, 64, 1466–1471. [Google Scholar] [CrossRef]
- Huang, H.; Tokashiki, M.; Maeno, S.; Onaga, S.; Taira, T.; Ito, S. Purification and properties of phenolic acid decarboxylase from Candida guilliermondii. J. Ind. Microbiol. Biotechnol. 2012, 39, 55–62. [Google Scholar] [CrossRef]
- Sheng, X.; Lind, M.E.S.; Himo, F. Theoretical study of the reaction mechanism of phenolic acid decarboxylase. Febs J. 2015, 282, 4703–4713. [Google Scholar] [CrossRef]
- Lu, C.; Zhang, J.; Zhao, X.; Zi, Y.; Xiao, X. Biotransformation of phenolic acids in foods: pathways, key enzymes, and technological applications. Foods 2025, 14, 2187. [Google Scholar] [CrossRef] [PubMed]
- Mu, Y.; Huang, J.; Zhou, R.; Mao, F.; Pan, Q.; Chen, S.; Lu, Z.; Du, L.; Xie, F. Exploring the response patterns of strong-flavor baijiu brewing microecosystem to fortified Daqu under different pit ages. Food Res. Int. 2022, 155, 111062. [Google Scholar] [CrossRef]
- Li, H.; Liu, S.; Liu, Y.; Hui, M.; Pan, C. Functional microorganisms in baijiu Daqu: research progress and fortification strategy for application. Front. Microbiol. 2023, 14, 1119675. [Google Scholar] [CrossRef]
- Szambelan, K.; Jeleń, H.; Nowak, J.; Frankowski, J.; Szwengiel, A. The promising prospective technology for biofuel production with mixed cultures using Sorghum grain (Sorghum bicolor (L.) Moench) grown in a temperate climate: characteristics of fermentation factors and volatile compounds analysis. J. Agric. Food. Chem. 2025, 73, 14408–14419. [Google Scholar] [CrossRef]
- Zhang, Y.; Sun, Q.; Liu, X.; Basit, R.A.; Ma, J.; Fu, Z.; Cheng, L.; Fan, G.; Teng, C. Screening, identification, and fermentation condition optimization of a high-yield 3-methylthiopropanol yeast and its aroma-producing characteristics. Foods 2024, 13, 418. [Google Scholar] [CrossRef]
- Kumar, A.; Singh, R.; Yadav, A.; Giri, D.D.; Singh, P.K.; Pandey, K.D. Isolation and characterization of bacterial endophytes of Curcuma longa L. 3 Biotech. 2016, 6, 60. [Google Scholar] [CrossRef]
- Zhang, Y.; Feng, Z.; Xiang, H.; Zhang, X.; Yang, L. Characterization of feruloyl esterase from Klebsiella oxytoca Z28 and its application in the release of ferulic acid from de-starching wheat bran. Microorganisms 2023, 11, 989. [Google Scholar] [CrossRef]
- Noskova, Y.; Pentekhina, I.; Seitkalieva, A.; Nedashkovskaya, O.; Goroshkova, Y.; Balabanova, L. Characterization of an atypical GH19 family chitinase from Vibrio jasicida KMM 6832. Microorganisms 2026, 14, 443. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Liu, L.; Hu, G.; Wang, S.; Shan, L.; Chen, J. Effect of non-Saccharomyces yeasts derived from traditional fermented foods on beer fermentation characteristics and flavor profiles. Foods 2025, 14, 1395. [Google Scholar] [CrossRef]
- Wang, M.; Zheng, X.; Bian, X.; Ren, J.; Bai, L.; Yao, Y.; Dong, B.; Zhao, G. Bacillus licheniformis and Wickerhamiella versatilis: sources of the pleasant smoky and fruity flavors of soybean paste. Food Chem. 2025, 477, 143218. [Google Scholar] [CrossRef] [PubMed]
- Suezawa, Y.; Suzuki, M. Bioconversion of ferulic acid to 4-vinylguaiacol and 4-ethylguaiacol and of 4-vinylguaiacol to 4-ethylguaiacol by halotolerant yeasts belonging to the genus Candida. Biosci. Biotech. Bioch 2007, 71, 1058–1062. [Google Scholar] [CrossRef]
- Hu, H.; Li, L.; Ding, S. An organic solvent-tolerant phenolic acid decarboxylase from Bacillus licheniformis for the efficient bioconversion of hydroxycinnamic acids to vinyl phenol derivatives. Appl. Microbiol. Biotechnol. 2015, 99, 5071–5081. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.; Eborall, W.; Hyde, R.; Hart, S.; Turkenburg, J.P.; Grogan, G. Mutational analysis of phenolic acid decarboxylase from Bacillus subtilis (BsPAD), which converts bio-derived phenolic acids to styrene derivatives. Catal. Sci. Technol. 2012, 2, 1568–1574. [Google Scholar] [CrossRef]
- Li, L.; Long, L.; Ding, S. Bioproduction of high-concentration 4-vinylguaiacol using whole-cell catalysis harboring an organic solvent-tolerant phenolic acid decarboxylase from Bacillus atrophaeus. Front. Microbiol. 2019, 10, 1798. [Google Scholar] [CrossRef]
- Matte, A.; Grosse, S.; Bergeron, H.; Abokitse, K.; Lau, P.C.K. Structural analysis of Bacillus pumilus phenolic acid decarboxylase, a lipocalin-fold enzyme. Acta Crystallogr. Sect. F. 2010, 66, 1407–1414. [Google Scholar] [CrossRef]
- Rodríguez, H.; Angulo, I.; de Las Rivas, B.; Campillo, N.; Páez, J.A.; Muñoz, R.; Mancheño, J.M. p-Coumaric acid decarboxylase from Lactobacillus plantarum: structural insights into the active site and decarboxylation catalytic mechanism. Proteins Struct. Funct. Bioinform. 2010, 78, 1662–1676. [Google Scholar] [CrossRef]
- Myrtollari, K.; Calderini, E.; Kracher, D.; Schöngaßner, T.; Galušić, S.; Slavica, A.; Taden, A.; Mokos, D.; Schrüfer, A.; Wirnsberger, G.; Gruber, K.; Daniel, B.; Kourist, R. Stability increase of phenolic acid decarboxylase by a combination of protein and solvent engineering unlocks applications at elevated temperatures. ACS Sustain. Chem. Eng. 2024, 12, 3575–3584. [Google Scholar] [CrossRef]
- Zhang, L.; Shen, J.; Ren, M.; Xu, Y.; Lin, S.; Bao, S.; Wang, L.; Fan, R.; Jin, L.; Quan, C. Characterization and modification of phenolic acid decarboxylase from marine bacteria for the synthesis of 4-vinyl phenol derivatives. BMC Microbiol. 2025, 26, 31. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, L.; Yang, S.; Liu, G.; Zeng, Q.; Liu, Y. Molecular characterization and functional analysis of a pathogenesis-related β-1,3-glucanase gene in spruce (Picea asperata). Eur. J. Plant Pathol. 2022, 164, 177–192. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, L.; Yang, C.; Han, S.; Yang, S.; Liu, G.; Zeng, Q.; Liu, Y. Molecular identification and antifungal activity of a defensin (PaDef) from spruce. J. Plant Growth Regul. 2022, 41, 494–506. [Google Scholar] [CrossRef]
- Maeda, M.; Tokashiki, M.; Tokashiki, M.; Uechi, K.; Ito, S.; Taira, T. Characterization and induction of phenolic acid decarboxylase from Aspergillus luchuensis. J. Biosci. Bioeng. 2018, 126, 162–168. [Google Scholar] [CrossRef]
- Arcus, V.L.; van der Kamp, M.W.; Pudney, C.R.; Mulholland, A.J. Enzyme evolution and the temperature dependence of enzyme catalysis. Curr. Opin. Struct. Biol. 2020, 65, 96–101. [Google Scholar] [CrossRef]
- Ni, J.; Wu, Y.; Tao, F.; Peng, Y.; Xu, P. A coenzyme-free biocatalyst for the value-added utilization of lignin-derived aromatics. J. Am. Chem. Soc. 2018, 140, 16001–16005. [Google Scholar] [CrossRef]
- Li, Q.; Xia, Y.; Zhao, T.; Gong, Y.; Fang, S.; Chen, M. Improving the catalytic characteristics of phenolic acid decarboxylase from Bacillus amyloliquefaciens by the engineering of N-terminus and C-terminus. BMC Biotechnol. 2021, 21, 44. [Google Scholar] [CrossRef]
- Maeda, M.; Motosoko, M.; Tokashiki, T.; Tokashiki, J.; Mizutani, O.; Uechi, K.; Goto, M.; Taira, T. Phenolic acid decarboxylase of Aspergillus luchuensis plays a crucial role in 4-vinylguaiacol production during awamori brewing. J. Biosci. Bioeng. 2020, 130, 352–359. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, X.; Yuan, S.; Liu, J. Molecular identification and functional characterization of a chitinase gene in Picea asperata. Discov. Plan. 2025, 2, 145. [Google Scholar] [CrossRef]
- Godoy, L.; Martínez, C.; Carrasco, N.; Ganga, M.A. Purification and characterization of a p-coumarate decarboxylase and a vinylphenol reductase from Brettanomyces bruxellensis. Int. J. Food Microbiol. 2008, 127, 6–11. [Google Scholar] [CrossRef]
- Gu, W.; Yang, J.; Lou, Z.; Liang, L.; Sun, Y.; Huang, J.; Li, X.; Cao, Y.; Meng, Z.; Zhang, K. Structural basis of enzymatic activity for the ferulic acid decarboxylase (FADase) from Enterobacter sp. Px6-4. PLoS ONE 2011, 6, e16262. [Google Scholar] [CrossRef]
- Mphahlele, M.J.; Magwaza, N.M.; Gildenhuys, S.; Setshedi, I.B. Synthesis, α-glucosidase inhibition and antioxidant activity of the 7-carbo-substituted 5-bromo-3-methylindazoles. Bioorganic Chem. 2020, 97, 103702. [Google Scholar] [CrossRef]
- Yang, D.; Chen, X.; Liu, X.; Han, N.; Liu, Z.; Li, S.; Zhai, J.; Yin, J. Antioxidant and α-glucosidase inhibitory activities guided isolation and identification of components from mango seed kernel. Oxidative Med. Cell. Longev. 2020, 2020, 8858578. [Google Scholar] [CrossRef]
- Chen, T.; Chen, Z.; Wang, N.; Chu, J.; Fan, B.; Cheng, C.; Qin, S.; He, B. Highly regioselective and efficient biosynthesis of polydatin by an engineered UGTBL1-AtSuSy cascade reaction. J. Agric. Food. Chem. 2021, 69, 8695–8702. [Google Scholar] [CrossRef]
- Parada-Fabián, J.C.; Hernández-Sánchez, H.; Méndez-Tenorio, A. Substrate specificity of the phenolic acid decarboxylase from Lactobacillus plantarum and related bacteria analyzed by molecular dynamics and docking. J. Plant Biochem. Biotechnol. 2019, 28, 91–104. [Google Scholar] [CrossRef]
| Strains/plasmids | Description | Source |
| JXQ2 | The strain was isolated from the high-temperature Daqu | Lab stock |
| JXQ5 | The strain was isolated from the high-temperature Daqu | Lab stock |
| NXQ4 | The strain was isolated from the medium-temperature Daqu | Lab stock |
| JN2 | The strain was isolated from the mud of the liquor cellar | Lab stock |
| JN11 | The strain was isolated from the mud of the liquor cellar | Lab stock |
| E. coli BL21(DE3) | Host for gene expression | TransGen |
| E. coli BL21(DE3)/pET-28a | Expression strain carrying plasmid pET-28a | This work |
| E. coli BL21(DE3)/pET-28a-BcPAD | Expression strain carrying plasmid pET-28a-BcPAD | This work |
| pET-28a(+) | E. coli expression vector, Kanr | TaKaRa |
| pET-28a-BcPAD | pET-28a(+) derivative harboring BcPAD gene, Kanr | This work |
| Biochemical Test | Test Result | Biochemical Test | Test Result |
| Glucose fermentation | + | Gelatin hydrolysis | + |
| Sucrose fermentation | + | Starch hydrolysis | + |
| Lactose fermentation | + | Nitrate reduction | + |
| Mannitol fermentation | - | Citrate utilization | - |
| Catalase activity | + | VP test | + |
| Cellulase activity | + | Indole test | - |
| Proteinase activity | + | Anaerobic growth | + |
| Substrates | Specific activity (U/mg) | Relative activity (%) |
| Ferulic acid | 34.46 ± 1.11 | 7.48 |
| p-Coumaric acid | 460.55 ± 18.30 | 100 |
| Caffeic acid | 145.88 ± 2.93 | 31.68 |
| Sinapic acid | ND | ND |
| Kinetic Parameters | Substrates | ||
| Ferulic acid | p-Coumaric acid | Caffeic acid | |
| Km (mM) | 4.72 ± 0.68 | 26.70 ± 3.17 | 10.16 ± 1.36 |
| Vmax (U/mg) | 35.28 ± 0.61 | 837.06 ± 10.70 | 155.99 ± 3.42 |
| Kcat (s−1) | 7.04 ± 0.12 | 167.12 ± 2.14 | 31.14 ± 0.68 |
| Kcat/Km (mM−1 • s−1) | 1.49 ± 0.22 | 6.26 ± 0.75 | 3.06 ± 0.41 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




