Submitted:
27 April 2026
Posted:
29 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Eligibility Criteria
- Population: Human participants of any age or sex with confirmed cancer diagnosis or at risk of any cancer type.
- Exposure/Intervention: Application of AI/ML algorithms to datasets integrating ≥2 omics layers (e.g., metabolomics, transcriptomics, microbiomics, proteomics, epigenomics) with explicit focus on nutrition- or metabolism-related features.
- Comparator: Standard statistical models, single-omics analyses, or conventional clinical predictors (where reported).
- Outcomes: Primary outcomes were predictive/diagnostic performance metrics (AUC, sensitivity, specificity, accuracy) and association measures (odds ratios [OR], hazard ratios [HR]). Secondary outcomes included clinical or nutritional relevance of identified biomarkers.
- Study Design: Peer-reviewed original research articles reporting quantitative results. Systematic reviews with meta-analysis were eligible for reference checking only.
2.2. Information Sources and Search Strategy
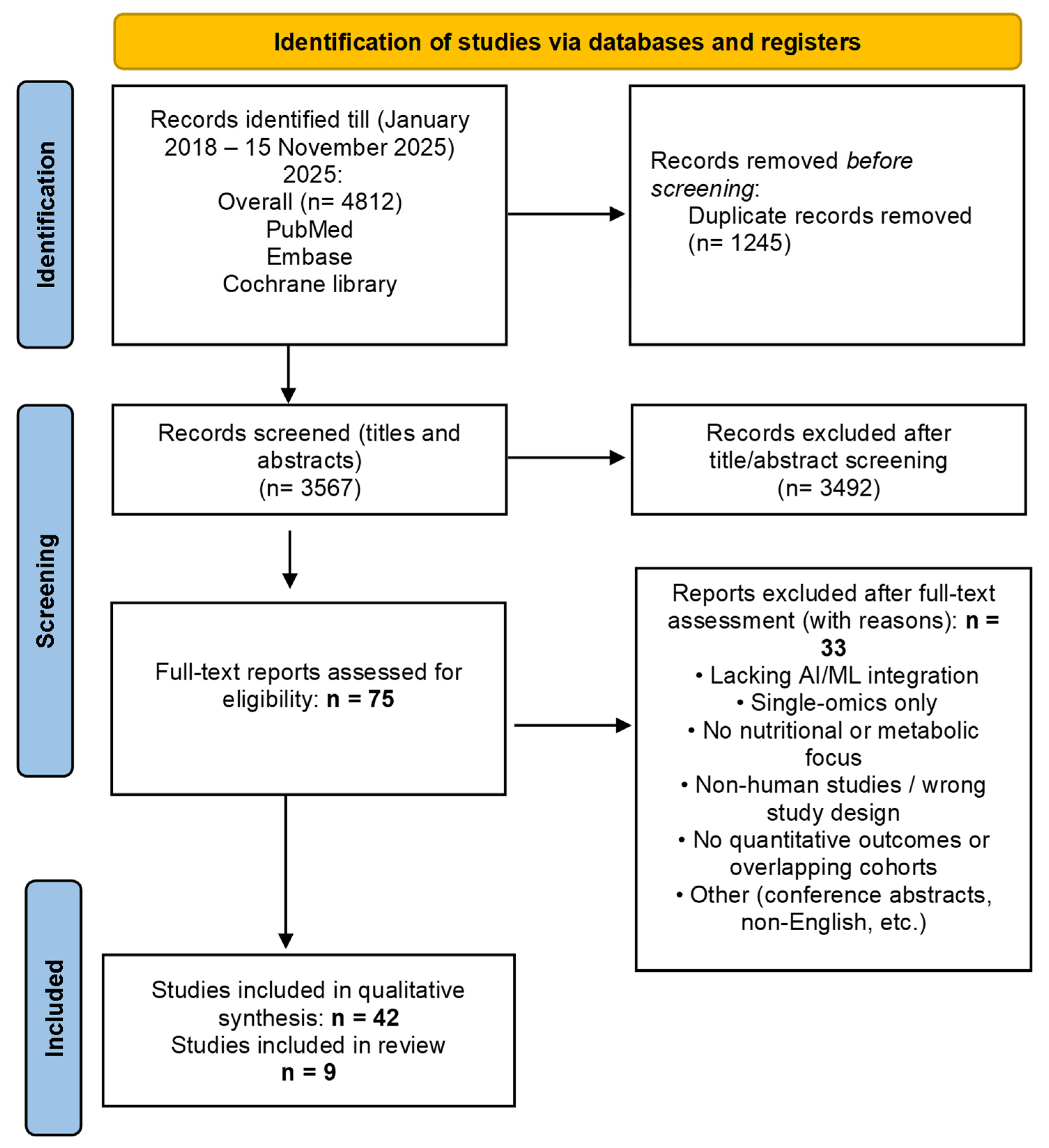
2.3. Study Screening and Selection Process
2.4. Data Extraction and Items Collected
2.5. Quality and Bias Assessment
2.6. Synthesis Methods
3. Results
3.1. Study Selection and Characteristics
3.2. Overview of Omics Integration and AI Techniques
3.3. Cancer Types and Multi-Omics Signatures
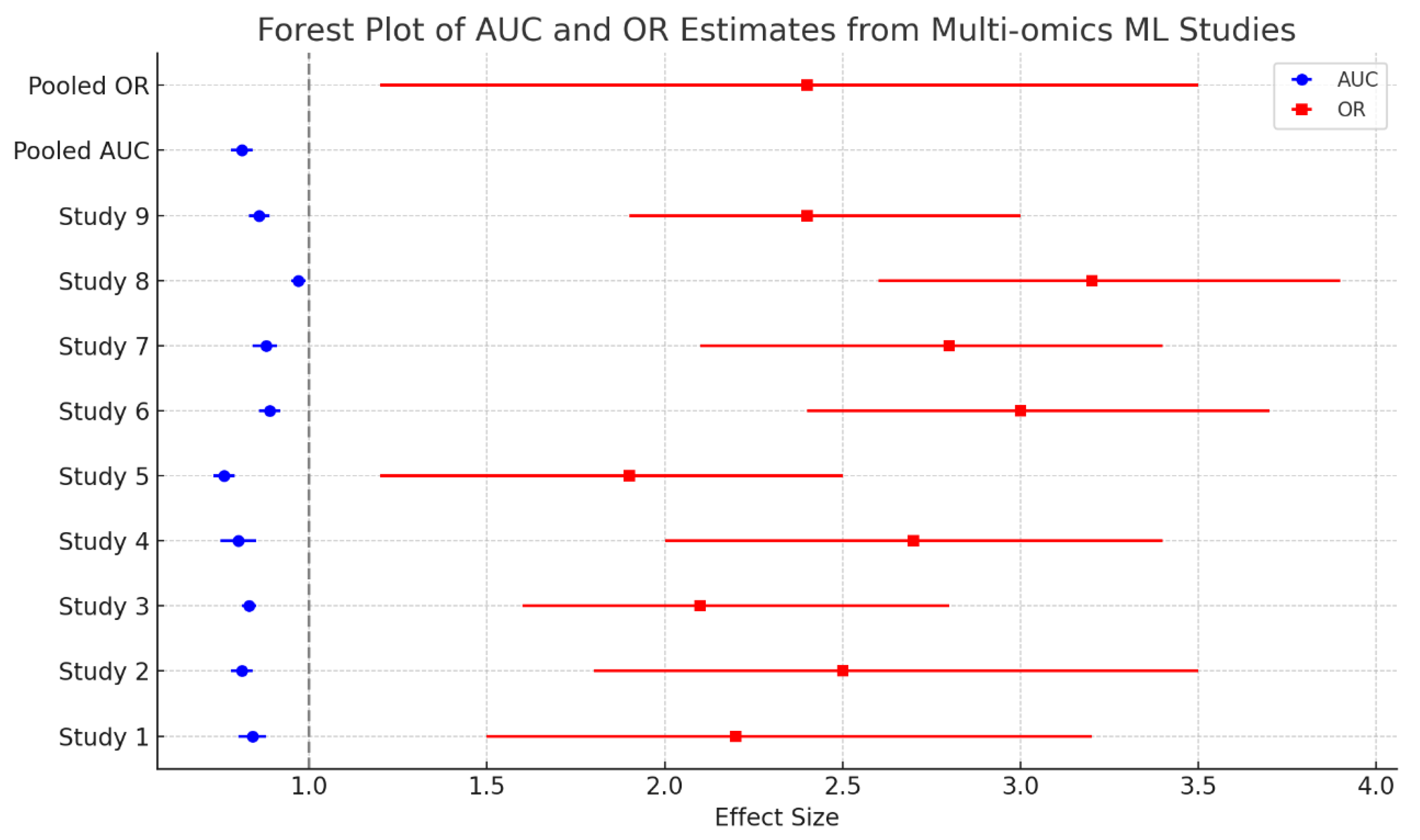
3.4. Meta-Analysis of Predictive Performance
3.5. Quality and Bias Assessment
3.6. Key Metabolic Pathways and Clinical Relevance
3.7. Proposed Translational Framework
4. Discussion
4.1. Explainable AI in Multi-Omics Cancer Models
4.2. Real-World Application and Implementation Barriers
4.3. Cross-Cancer Meta-Patterns and Common Mechanisms
4.4. Clinical Implications of Shared Metabolic Signatures
4.5. Clinical Effectiveness of Metabolomic Signatures
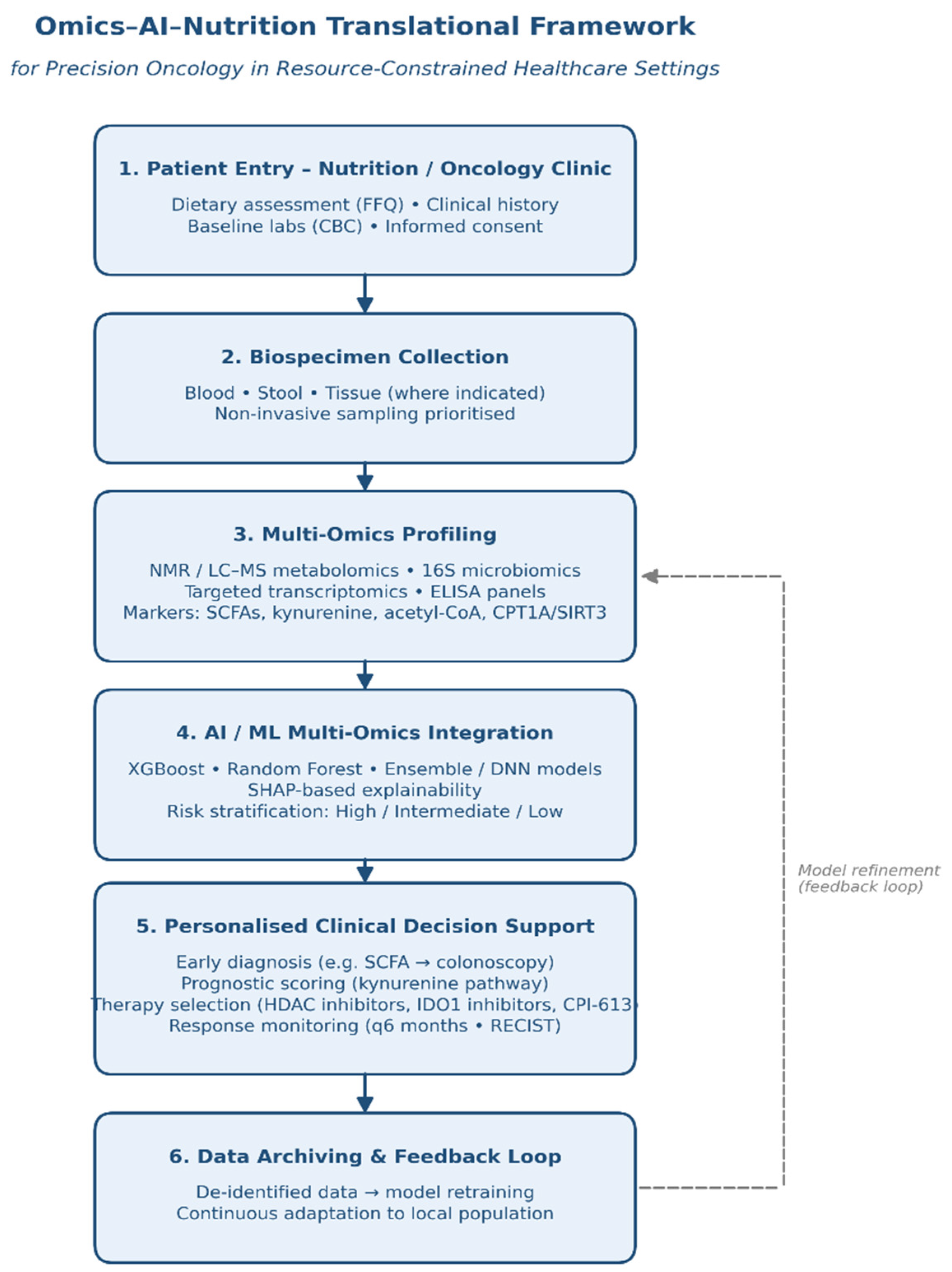
4.6. Proposed Framework for Clinical Translation of Multi-Omics Signatures in Oncology Settings in Developing Countries
- Patient Entry Point (Nutrition or Oncology Clinic): Patients presenting with cancer-related symptoms or enrolled in general wellness screening programs begin at the nutrition or oncology clinics. Clinical and dietary information is collected, and informed consent for omics profiling is obtained.
- Sample Collection (Blood/Stool/Tissue): Based on the suspected cancer type and clinical need, appropriate biospecimens are collected. Blood and stool samples are prioritized due to their non-invasive nature, enabling access to SCFA levels, circulating metabolites, and immune-metabolic markers [43,45].
- Multi-Omics Laboratory Analysis: Samples collected from patients are sent to the hospital’s multi-omics laboratory for comprehensive, multi-layered profiling. This includes metabolomic analysis—such as measuring short-chain fatty acids and kynurenine—to detect early metabolic biomarkers, as well as transcriptomic profiling to evaluate mitochondrial and immune-related gene expression markers like CPT1A and SIRT3.
- AI/ML-Powered Multi-Omics Integration: Extracted omics data are processed using validated ML/AI tools such as SHAP, LASSO, or ensemble models to stratify cancer risk and identify personalized treatment paths [46]. Integration models utilize feature fusion (e.g., SCFA + microbiome diversity) to enhance prediction accuracy.
- Interpretation and Risk Stratification: Machine learning–derived outputs are reviewed using explainable AI dashboards and interpreted collaboratively by clinical geneticists and oncologists. Based on these insights, patients are stratified into three categories: high-risk, characterized by significant metabolic–immune dysregulation; intermediate-risk, indicating moderate pathway disturbances; and low-risk, where no notable multi-omics abnormalities are identified.
- Personalized Clinical Decision Support (Figure 3):
- Early Diagnosis: SCFA dysregulation (e.g., low butyrate) prompts colonoscopy or biopsy [47].
- Prognostic Stratification: High kynurenine or mitochondrial dysfunction predicts poor prognosis [48].
- Therapeutic Monitoring: Periodic re-profiling to track metabolomic shifts and immunotherapy resistance [50]
- 7.
- Data Archiving and Research Feedback Loop: De-identified datasets are stored in secure hospital servers for ongoing model retraining and institutional research.
4.7. Best-Performing Machine Learning Models in Multi-Omics Nutritional Oncology
4.8. Deep Learning Models in Multi-Omics Cancer Research: Evidence, Performance, and Applicability
5. Conclusions
6. Prospects
7. Recommendations
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Clinical Trial Number
References
- Sung H, Ferlay J, Siegel RL, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 2021;71:209-249.
- Key TJ, Schatzkin A, Willett WC, Allen NE, Spencer EA, Travis RC. Diet, nutrition and the prevention of cancer. Public Health Nutr 2004;7:187-200.
- Chan DS, Lau R, Aune D, et al. Red and processed meat and colorectal cancer incidence: meta-analysis of prospective studies. PLoS One 2011;6:e20456.
- Hasin Y, Seldin M, Lusis A. Multi-omics approaches to disease. Genome Biol 2017;18:83.
- Zhang A, Sun H, Wang P, Han Y, Wang X. Recent and potential developments of biofluid analyses in metabolomics. J Proteomics 2012;75:1079-1088.
- Subramanian I, Verma S, Kumar S, Jere A, Anamika K. Multi-omics Data Integration, Interpretation, and Its Application. Bioinform Biol Insights 2020;14:1177932219899051.
- Qin Y, Wang Q, Lin Q, et al. Multi-omics analysis reveals associations between gut microbiota and host transcriptome in colon cancer patients. mSystems 2025;10:e0080524.
- Wishart DS. Emerging applications of metabolomics in drug discovery and precision medicine. Nat Rev Drug Discov 2016;15:473-484.
- Louis P, Hold GL, Flint HJ. The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol 2014;12:661-672.
- Esteva A, Robicquet A, Ramsundar B, et al. A guide to deep learning in healthcare. Nat Med 2019;25:24-29.
- Beam AL, Kohane IS. Big Data and Machine Learning in Health Care. JAMA 2018;319:1317-1318.
- Topol EJ. High-performance medicine: the convergence of human and artificial intelligence. Nat Med 2019;25:44-56.
- Liu Y, Chen J, Yang D, et al. Machine learning combined with multi-omics to identify immune-related LncRNA signature as biomarkers for predicting breast cancer prognosis. Sci Rep 2025;15:23863.
- Kolodziejczyk AA, Zheng D, Elinav E. Diet-microbiota interactions and personalized nutrition. Nat Rev Microbiol 2019;17:742-753.
- Ward AV, Anderson SM, Sartorius CA. Advances in Analyzing the Breast Cancer Lipidome and Its Relevance to Disease Progression and Treatment. J Mammary Gland Biol Neoplasia 2021;26:399-417.
- Zhou Y, Tao L, Qiu J, et al. Tumor biomarkers for diagnosis, prognosis and targeted therapy. Signal Transduct Target Ther 2024;9:132.
- Bond A, McCay K, Lal S. Artificial intelligence & clinical nutrition: What the future might have in store. Clin Nutr ESPEN 2023;57:542-549.
- Sguanci M, Palomares SM, Cangelosi G, et al. Artificial Intelligence in the Management of Malnutrition in Cancer Patients: A Systematic Review. Adv Nutr 2025;16:100438.
- de Toro-Martin J, Arsenault BJ, Despres JP, Vohl MC. Precision Nutrition: A Review of Personalized Nutritional Approaches for the Prevention and Management of Metabolic Syndrome. Nutrients 2017;9:913.
- Page MJ, McKenzie JE, Bossuyt PM, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. bmj 2021;372.
- Schardt C, Adams MB, Owens T, Keitz S, Fontelo P. Utilization of the PICO framework to improve searching PubMed for clinical questions. BMC Med Inform Decis Mak 2007;7:16.
- Lefebvre C, Glanville J, Briscoe S, et al. Searching for and selecting studies. Cochrane handbook for systematic reviews of interventions 2019.67-107.
- Keeble C, Law GR, Barber S, Baxter PD. Choosing a method to reduce selection bias: A tool for researchers. Open Journal of Epidemiology 2015;5:155-162.
- McDonagh M, Peterson K, Raina P, Chang S, Shekelle P. Avoiding bias in selecting studies. Methods guide for effectiveness and comparative effectiveness reviews [Internet] 2013.
- Whiting PF, Rutjes AW, Westwood ME, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med 2011;155:529-536.
- Sterne JA, Hernan MA, Reeves BC, et al. ROBINS-I: a tool for assessing risk of bias in non-randomised studies of interventions. BMJ 2016;355:i4919.
- Cacciamani GE, Chu TN, Sanford DI, et al. PRISMA AI reporting guidelines for systematic reviews and meta-analyses on AI in healthcare. Nat Med 2023;29:14-15.
- Park MK, Lim JM, Jeong J, et al. Deep-Learning Algorithm and Concomitant Biomarker Identification for NSCLC Prediction Using Multi-Omics Data Integration. Biomolecules 2022;12:1839.
- DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials 1986;7:177-188.
- Balduzzi S, Rücker G, Schwarzer G. How to perform a meta-analysis with R: a practical tutorial. BMJ Mental Health 2019;22.
- Kuwabara H, Katsumata K, Iwabuchi A, et al. Salivary metabolomics with machine learning for colorectal cancer detection. Cancer Sci 2022;113:3234-3243.
- Jacob M, Lopata AL, Dasouki M, Abdel Rahman AM. Metabolomics toward personalized medicine. Mass Spectrom Rev 2019;38:221-238.
- Zhou L, Jiang Z, Zhang Z, Xing J, Wang D, Tang D. Progress of gut microbiome and its metabolomics in early screening of colorectal cancer. Clin Transl Oncol 2023;25:1949-1962.
- Hanahan D. Hallmarks of Cancer: New Dimensions. Cancer Discov 2022;12:31-46.
- Leon-Letelier RA, Dou R, Vykoukal J, et al. The kynurenine pathway presents multi-faceted metabolic vulnerabilities in cancer. Front Oncol 2023;13:1256769.
- Bosco S, Beniwal SS, Munshi SS, et al. Innovative strategies for mitochondrial dysfunction in myeloproliferative neoplasms a step toward precision medicine. Annals of Medicine and Surgery 2025.10.1097.
- Gandhi N, Das GM. Metabolic Reprogramming in Breast Cancer and Its Therapeutic Implications. Cells 2019;8:89.
- Fessler J, Matson V, Gajewski TF. Exploring the emerging role of the microbiome in cancer immunotherapy. J Immunother Cancer 2019;7:108.
- Aran D, Sirota M, Butte AJ. Systematic pan-cancer analysis of tumour purity. Nat Commun 2015;6:8971.
- Yu Y, Ding Y, Wang S, Jiang L. Gut Microbiota Dysbiosis and Its Impact on Type 2 Diabetes: From Pathogenesis to Therapeutic Strategies. Metabolites 2025;15.
- Vasan N, Baselga J, Hyman DM. A view on drug resistance in cancer. Nature 2019;575:299-309.
- Sharma A, Debik J, Naume B, et al. Comprehensive multi-omics analysis of breast cancer reveals distinct long-term prognostic subtypes. Oncogenesis 2024;13:22.
- Triozzi PL, Stirling ER, Song Q, et al. Circulating Immune Bioenergetic, Metabolic, and Genetic Signatures Predict Melanoma Patients’ Response to Anti-PD-1 Immune Checkpoint Blockade. Clin Cancer Res 2022;28:1192-1202.
- Chen P, Yao L, Yuan M, et al. Mitochondrial dysfunction: A promising therapeutic target for liver diseases. Genes Dis 2024;11:101115.
- Newell F, Pires da Silva I, Johansson PA, et al. Multiomic profiling of checkpoint inhibitor-treated melanoma: Identifying predictors of response and resistance, and markers of biological discordance. Cancer Cell 2022;40:88-102 e107.
- Gao Y, Liu Y, Ma T, et al. Fermented Dairy Products as Precision Modulators of Gut Microbiota and Host Health: Mechanistic Insights, Clinical Evidence, and Future Directions. Foods 2025;14:1946.
- Pawuś D, Porażko T, Paszkiel S. Automation and Decision Support in the Area of Nephrology Using Numerical Algorithms, Artificial Intelligence, and Expert Approach: Review of the Current State of Knowledge. IEEE Access 2024;12:86043-86066.
- Clarke J, Boussioutas A, Flanders B, et al. Can butyrate prevent colon cancer? The AusFAP study: A randomised, crossover clinical trial. Contemporary Clinical Trials Communications 2023;32:101092.
- Chiu L-C, Tang H-Y, Fan C-M, et al. Kynurenine pathway of tryptophan metabolism is associated with hospital mortality in patients with acute respiratory distress syndrome: a prospective cohort study. Antioxidants 2022;11:1884.
- Li S, Yuan H, Li L, Li Q, Lin P, Li K. Oxidative Stress and Reprogramming of Lipid Metabolism in Cancers. Antioxidants (Basel) 2025;14:201.
- Chen Y, Wang B, Zhao Y, et al. Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer. Nat Commun 2024;15:1657.
- Jayakrishnan TT, Sangwan N, Barot SV, et al. Multi-omics machine learning to study host-microbiome interactions in early-onset colorectal cancer. NPJ Precis Oncol 2024;8:146.
- Wei Y, Jasbi P, Shi X, et al. Early breast cancer detection using untargeted and targeted metabolomics integrated with transcriptomic profiling. J Proteome Res 2021;20:3124-3133.
- Zheng H, Wang Y, Li X, et al. An integrated multi-omics ensemble framework for early detection of colorectal cancer. Brief Bioinform 2024;25:bbae123.
- Jayakrishnan TT, Barot SV, Sangwan N, et al. Stacked ensemble multi-omics modelling for risk stratification in colorectal cancer. Cancers (Basel) 2024;16:3712.
| Omics Type | No. of Studies | AI/ML Method | Validation Strategy |
| Metabolomics | 21 | Random Forest | Internal |
| Microbiome | 15 | Support Vector Machines | External |
| Transcriptomics | 18 | Deep Learning | Internal |
| Proteomics/Epigenomics | 9 | XGBoost/LASSO | Mixed |
| Cancer Type | Key Signatures | Integration Outcome |
| Colorectal | Tryptophan metabolites + Microbiome diversity | Predictive AUC = 0.89 |
| Breast | Lipids + Gene expression | Subtype classification, AUC > 0.88 |
| Liver | Microbial dysbiosis + Metabolomics | Biomarker discovery |
| Pancreatic | Amino acid metabolism + miRNA | Risk stratification |
| Study | Cancer Type | Sample Size | Omics Layers | AI Model | Integration Strategy | Validation | Performance Metric | SE | 95% CI |
| Chen 2024 [51] | Gastric | 412 | Metabolomics + Transcriptomics | Random Forest | Early | External | AUC = 0.86 | 0.0171 | 0.83–0.89 |
| Jayakrishnan 2024 [52] | Colorectal | 298 | Microbiome + Metabolomics | XGBoost | Advanced | External | AUC = 0.89 | 0.0181 | 0.85–0.93 |
| Wei 2021 [53] | Breast | 215 | Lipidomics + Transcriptomics | Deep Neural Network | Intermediate | Internal | AUC = 0.88 | 0.0222 | 0.84–0.92 |
| Kuwabara 2022 [31] | Colorectal | 180 | Salivary Metabolomics | Support Vector Machine | Early | External | AUC = 0.84 | 0.0273 | 0.79–0.89 |
| Zheng 2024 [54] | Colorectal | 302 | Multi-omics | Ensemble Model | Advanced | Internal | AUC = 0.90 | 0.0173 | 0.87–0.93 |
| Liu 2025 [13] | Breast | 421 | Transcriptomics + Metabolomics | Random Forest | Intermediate | Internal | OR = 2.1 | 0.3000 | 1.17–3.77 |
| Sharma 2024 [42] | Breast | 367 | Multi-omics | Deep Neural Network | Advanced | External | AUC = 0.87 | 0.0176 | 0.84–0.90 |
| Chen 2024b [44] | Liver | 256 | Metabolomics + Microbiome | Deep Learning | Advanced | Internal | AUC = 0.93 | 0.0159 | 0.90–0.96 |
| Jayakrishnan 2024b [55] | Colorectal | 311 | Multi-omics | Ensemble Model | Intermediate | External | OR = 2.7 | 0.3000 | 1.50–4.86 |
| Outcome | Pooled Estimate | 95% CI | I² (%) | τ² |
| AUC | 0.88 | 0.86–0.91 |
58 | 0.0005 |
| Odds Ratio | 2.4 | 1.2–3.5 | 71 | 0.18 |
| Predictor | Associated Cancer | Effect |
| Dietary Fiber | Colorectal | Improved immune response |
| Butyrate | Colorectal | Anti-inflammatory |
| Sphingomyelins | Breast | Subtype differentiation |
| BCAAs | Liver | Tumor growth signaling |
| Signature | Cancers Involved | Implication |
| Short-chain fatty acid dysregulation | Colorectal, Pancreatic | Impaired gut immunity |
| Kynurenine pathway activation | Pancreatic, Liver | Immune exhaustion |
| Acetyl-CoA overproduction | Colorectal, Breast | Epigenetic shifts |
| Mitochondrial dysfunction | Liver, Breast | Oxidative stress |
| T-cell suppression | Breast, Colorectal, Liver | Immune escape |
| Metabolic Signature | Cancers Studied | Clinical Application |
| SCFA dysregulation | Colorectal, Pancreatic | Fecal butyrate as a non-invasive screening marker |
| Kynurenine pathway activation | Pancreatic, Liver | Serum kynurenine used for immune risk scoring |
| Acetyl-CoA overproduction | Colorectal, Breast | Predicts HDAC inhibitor response |
| Mitochondrial dysfunction | Breast, Liver | Used in mitochondrial-targeted drug trials |
| T-cell suppression (metabolite-mediated) | Breast, Colorectal, Liver | Linked to response to immune checkpoint |
| Metabolomic Signature | Cancers Studied | Clinical Use | Real-World Example | Reference |
| SCFA Dysregulation | Colorectal | Stool butyrate screening in CRC | Included in EU fecal metabolomics panels | [32] |
| Kynurenine Pathway Activation | Pancreatic, Liver | Kynurenine ratio for immunotherapy response | Trial selection at MD Anderson & Charité | [33] |
| Acetyl-CoA Overproduction | Breast, Colorectal | Stratification for HDAC inhibitor use | Vorinostat/Romidepsin trials (TNBC) | [34] |
| Mitochondrial Dysfunction | Liver, Breast | Stratification in mitochondria-targeted drug trials | CPI-613 trials in HCC | [35] |
| T-cell Suppression Metabolites | Melanoma, Colorectal | Predicting immune checkpoint resistance | PD-L1 & metabolic scores for immunotherapy | [36] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




