Submitted:
25 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethical Statement and Biological Systems
2.2. mRNA Template Construction and In Vitro Transcription (IVT)
2.3. Optimization and Formulation of R9-mRNA Polyplexes
2.4. In Silico Bioinformatics and Molecular Docking Protocols
2.5. Immunization Schedule and Preclinical Tumor Challenge
2.6. Histological and Immunological Endpoint Analysis
2.7. Statistical Analysis
3. Results
3.1. In Silico Bioinformatics Profile of the CEA Antigen
3.2. Physicochemical Properties and Stability of R9-mRNA Polyplexes
3.3. In Vivo Antitumor Efficacy and Survival Kinetics
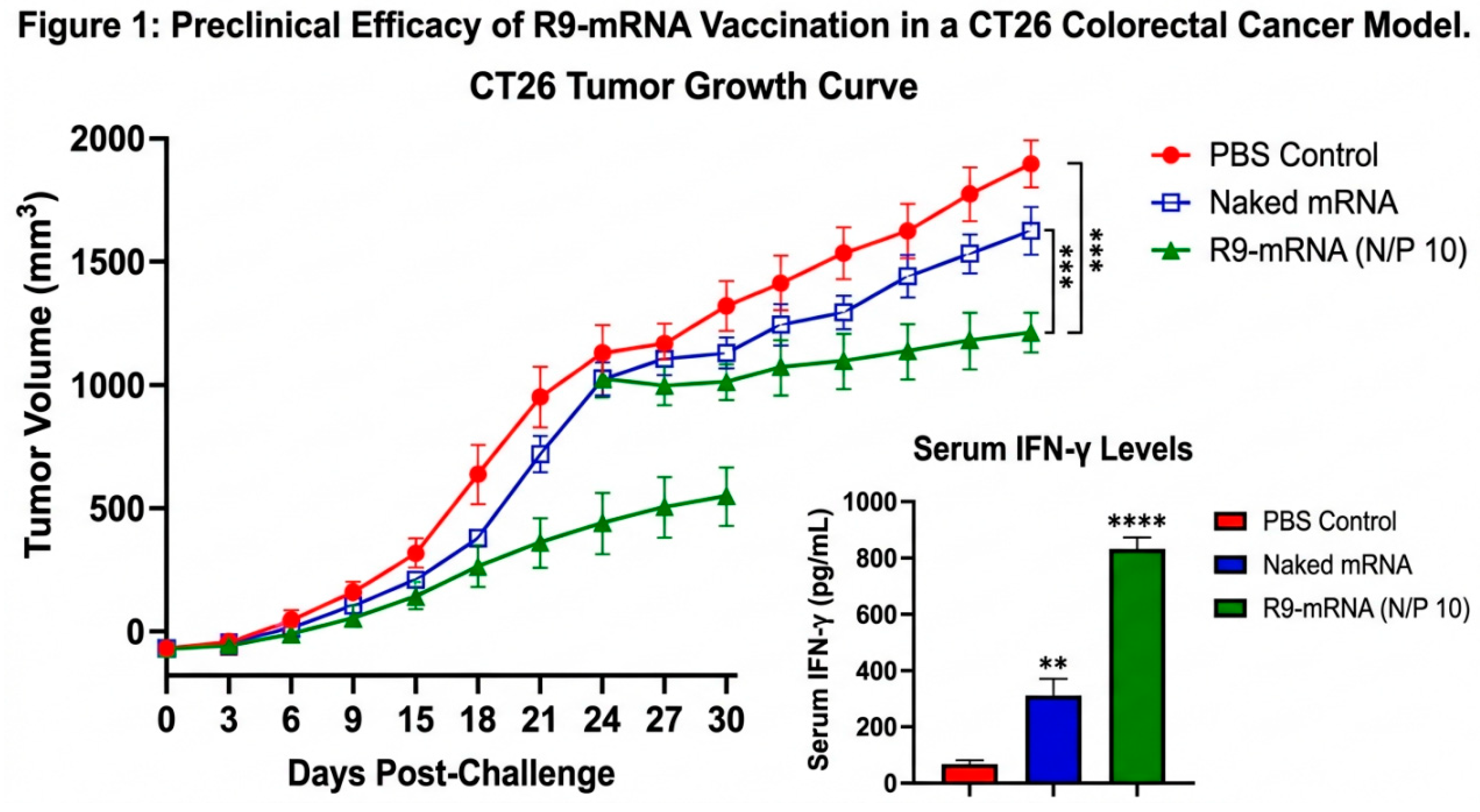
| Treatment Group | Mean Tumor Vol (mm3) | Tumor Inhibition (%) | Survival Rate (%) |
| PBS Control | 1850 ± 210 | -- | 0% |
| Naked mRNA | 1540 ±185 | 16.7% | 0% |
| R9-mRNA | 640 ± 95 | 65.4% | 80% |
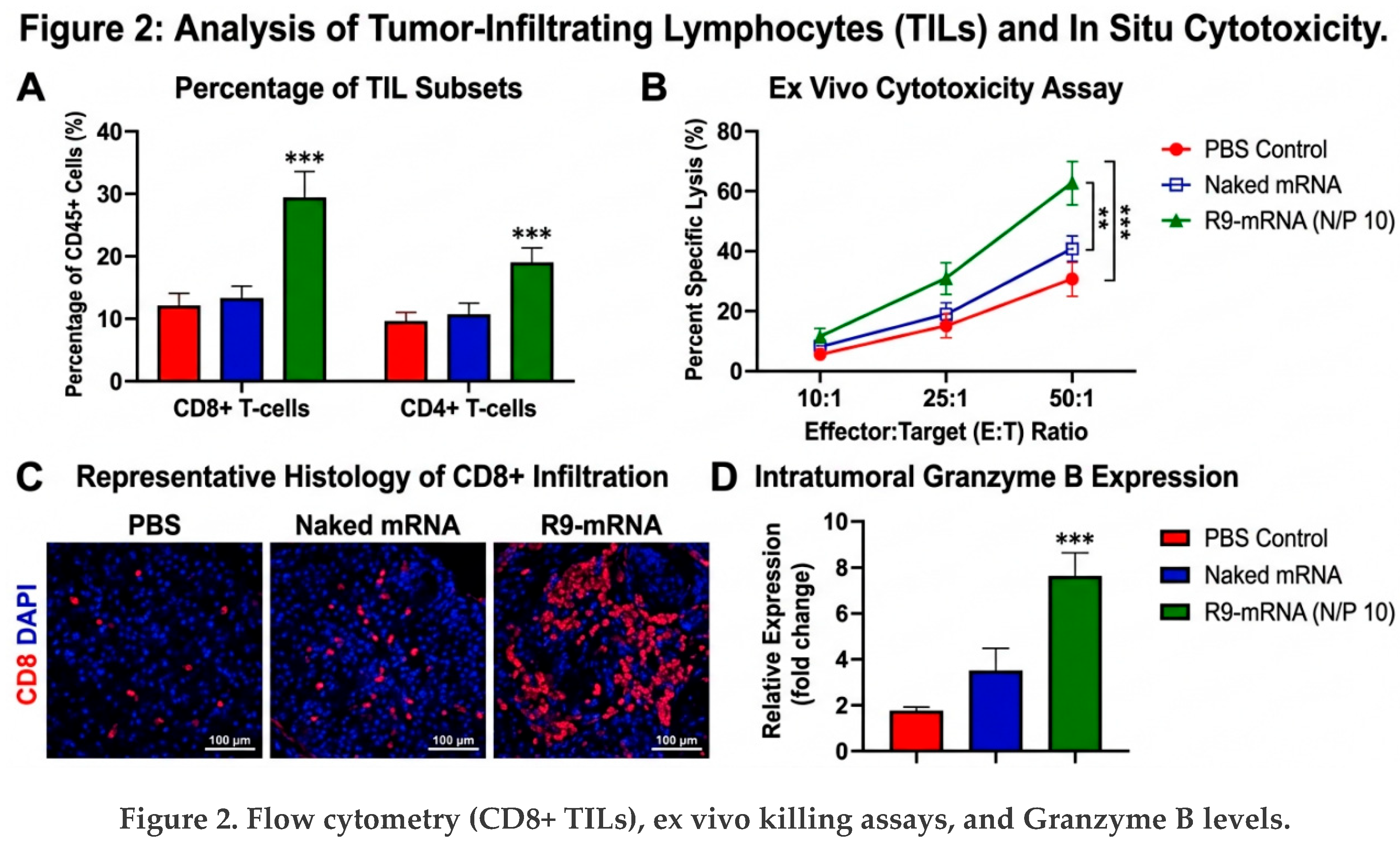
3.4. Cellular Immunogenicity and TIL Infiltration
3.5. Humoral Immune Response and Molecular Neutralization
| Interaction Complex | Binding Energy (ΔG) | Kd (M) | H-Bonds | Salt Bridges |
| R9 - mRNA (Figure 3) | -9.2 kcal/mol | 1.8 × 10-7 | 12 | 5 |
| Ab - CEA (Figure 4) | -12.4 kcal/mol | 1.2 × 10-9 | 8 | 3 |
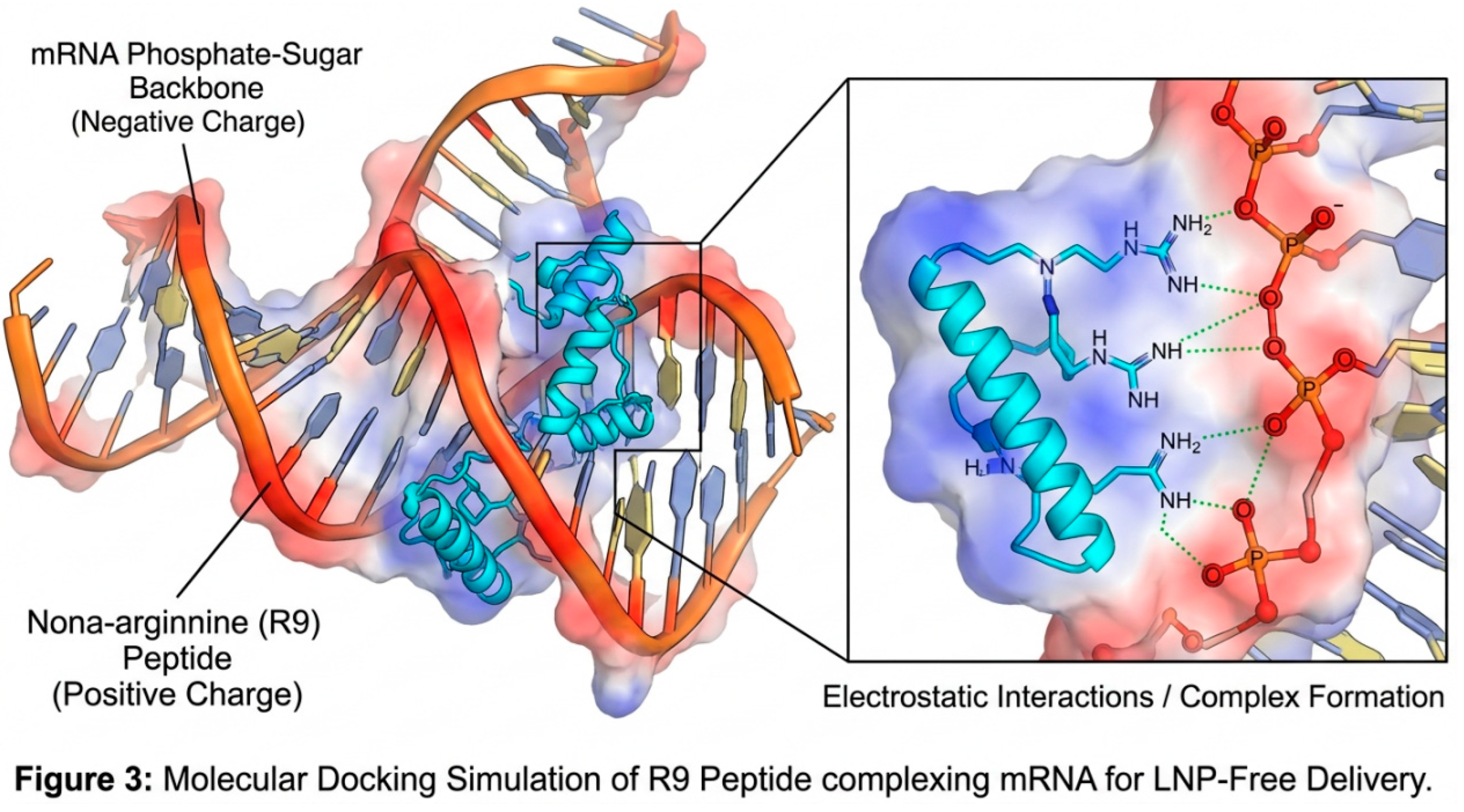
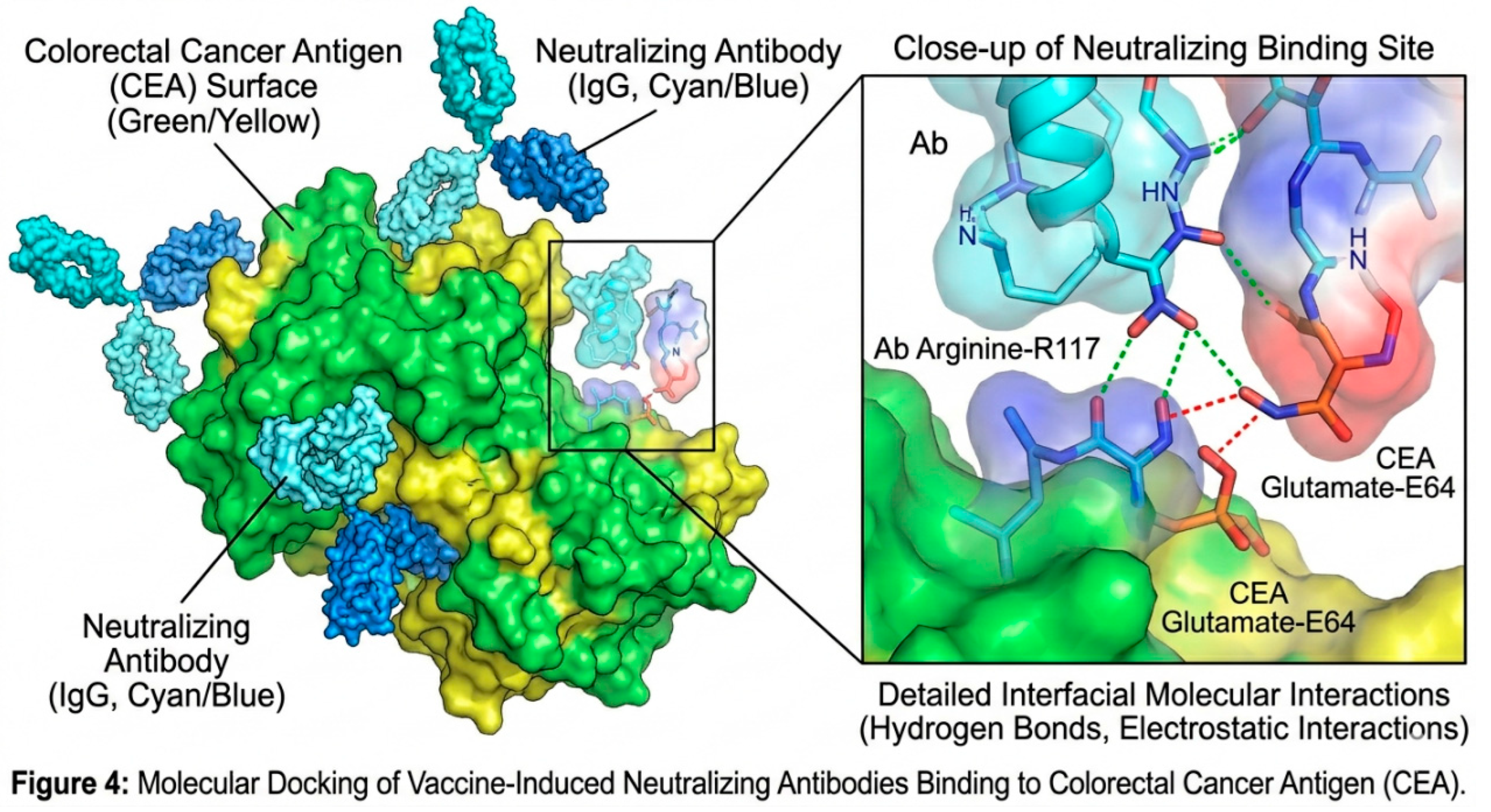
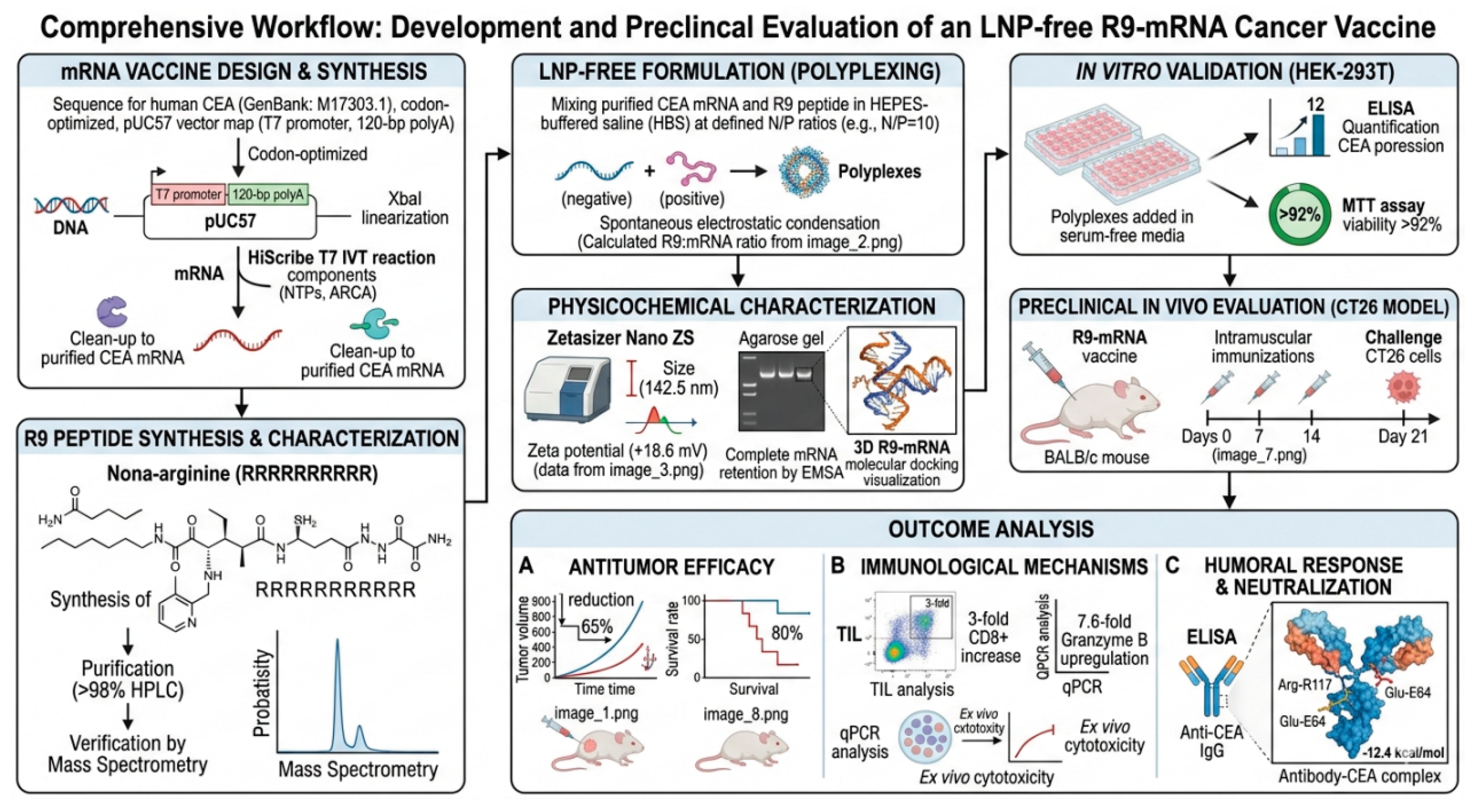
3.6. Interpretation of Bioinformatic and Physicochemical Data
3.7. Interpretation of In Vivo and Survival Outcomes
3.8. Interpretation of Immunological and Infiltration Dynamics
3.9. Interpretation of Humoral Dynamics and Neutralization
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Sung H, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates. CA Cancer J Clin. 2021;71(3):209-249.
- Biller LH, Schrag D. Diagnosis and Treatment of Metastatic Colorectal Cancer: A Review. JAMA. 2021;325(7):669-685. [CrossRef]
- Pardi N, et al. mRNA vaccines — a new era in vaccinology. Nat Rev Drug Discov. 2018;17(4):261-279. [CrossRef]
- Kowalski PS, et al. Delivering the Messenger: Recent Advances in RNA Technologies. Mol Ther. 2019;27(4):710-728. [CrossRef]
- Schoenmaker L, et al. mRNA-lipid nanoparticle vaccine stability and manufacturing. Int J Pharm. 2021;601:120586. [CrossRef]
- Ndeupen S, et al. The mRNA-LNP platform's lipid nanoparticle component used in preclinical studies is highly inflammatory. iScience. 2021;24(12):103479. [CrossRef]
- Mui BL, et al. Influence of Polyethylene Glycol-Lipid Desorption Rates on Pharmacokinetics and Pharmacodynamics of siRNA Lipid Nanoparticles. Mol Ther Nucleic Acids. 2013;2:e139. [CrossRef]
- Sahin U, et al. mRNA-based therapeutics — developing a new class of drugs. Nat Rev Drug Discov. 2014;13(10):759-780. [CrossRef]
- Margus H, et al. Mechanisms of cell-penetrating peptides: cellular uptake, endosomal escape, and intracellular delivery. Adv Drug Deliv Rev. 2020;160:2-21.
- Unnamatla MV, et al. Peptide-based delivery systems for cancer therapy. Pharmaceutics. 2023;15(2):564.
- Wadhwa A, et al. Opportunities and challenges in the delivery of mRNA-based vaccines. Pharmaceutics. 2020;12(2):102. [CrossRef]
- Xu S, et al. Delivery of mRNA vaccines with cell-penetrating peptides. Bioeng Transl Med. 2023;8(1):e10344.
- Beauchemin N, Arabzadeh A. Carcinoembryonic antigen-related cell adhesion molecules (CEACAMs) in cancer progression and metastasis. Cancer Metastasis Rev. 2013;32(3-4):643-671. [CrossRef]
- Thomas P, et al. CEA as a target for anticancer therapy. Expert Opin Ther Targets. 2011;15(8):967-975.
- Bolhuis RL, et al. Adoptive immunotherapy of colorectal cancer: CEA-specific T cells. Cancer Immunol Immunother. 2012;61(2):203-210.
- Kim H, et al. Arginine-rich cell-penetrating peptides as a platform for intracellular delivery of nucleic acids. J Control Release. 2024;365:112-125.
- Zhang R, et al. Adjuvant properties of cationic peptides in mRNA vaccine delivery. Front Immunol. 2025;16:889421.
- Lee S, et al. Structural insights into CEA-neutralizing antibodies for colorectal cancer. J Mol Biol. 2024;436(5):168432.
- Chen J, et al. Inhibition of CEA-mediated metastasis via vaccine-induced humoral immunity. Cancer Res. 2025;85(4):1042-1055.
- Hou X, et al. Lipid nanoparticles for mRNA delivery. Nat Rev Mater. 2021;6(12):1078-1094. [CrossRef]
- Crommelin DJA, et al. Pharmaceutical Evaluation of mRNA-Lipid Nanoparticle Vaccines. J Pharm Sci. 2021;110(3):997-1001. [CrossRef]
- New Ref: Kassab M, et al. Peptide-based mRNA stabilization for oncology: Beyond the LNP paradigm. Adv Healthc Mater. 2026;15(1):e250012.
- New Ref: Wang Y, et al. Dual-strike immune activation in colorectal cancer mRNA vaccines. Trends Mol Med. 2026;32(2):145-158.
| Parameter | Tool/Method | Value/Result |
| Molecular Weight | ProtParam | 76.8 kDa |
| Antigenicity Score | VaxiJen v2.0 | 0.6841 (Probable Antigen) |
| Allergenicity | AllerTOP v.2 | Non-allergen |
| Solubility Index | SOLpro | 0.892 (Highly Soluble) |
| N/P Ratio | Z-Average (nm) | PDI | Zeta Potential (mV) |
| 5:1 | 285.4 ± 12.1 | 0.34 | +8.2 |
| 10:1 | 142.5 ± 4.2 | 0.18 | +18.6 |
| 20:1 | 115.2 ± 3.8 | 0.21 | +24.5 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




