Submitted:
24 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Experimental
2.1. Tools and Software
2.2. Reagents and Chemicals
2.4. Analysis of Pharmaceutical Preparations
2.5. Methods
2.5.1. Principal Component Regression and Partial Least Squares
2.5.2. Multivariate Curve Resolution-Alternating Least Squares
2.5.3. Artificial Neural Networks
3. Result and Discussion
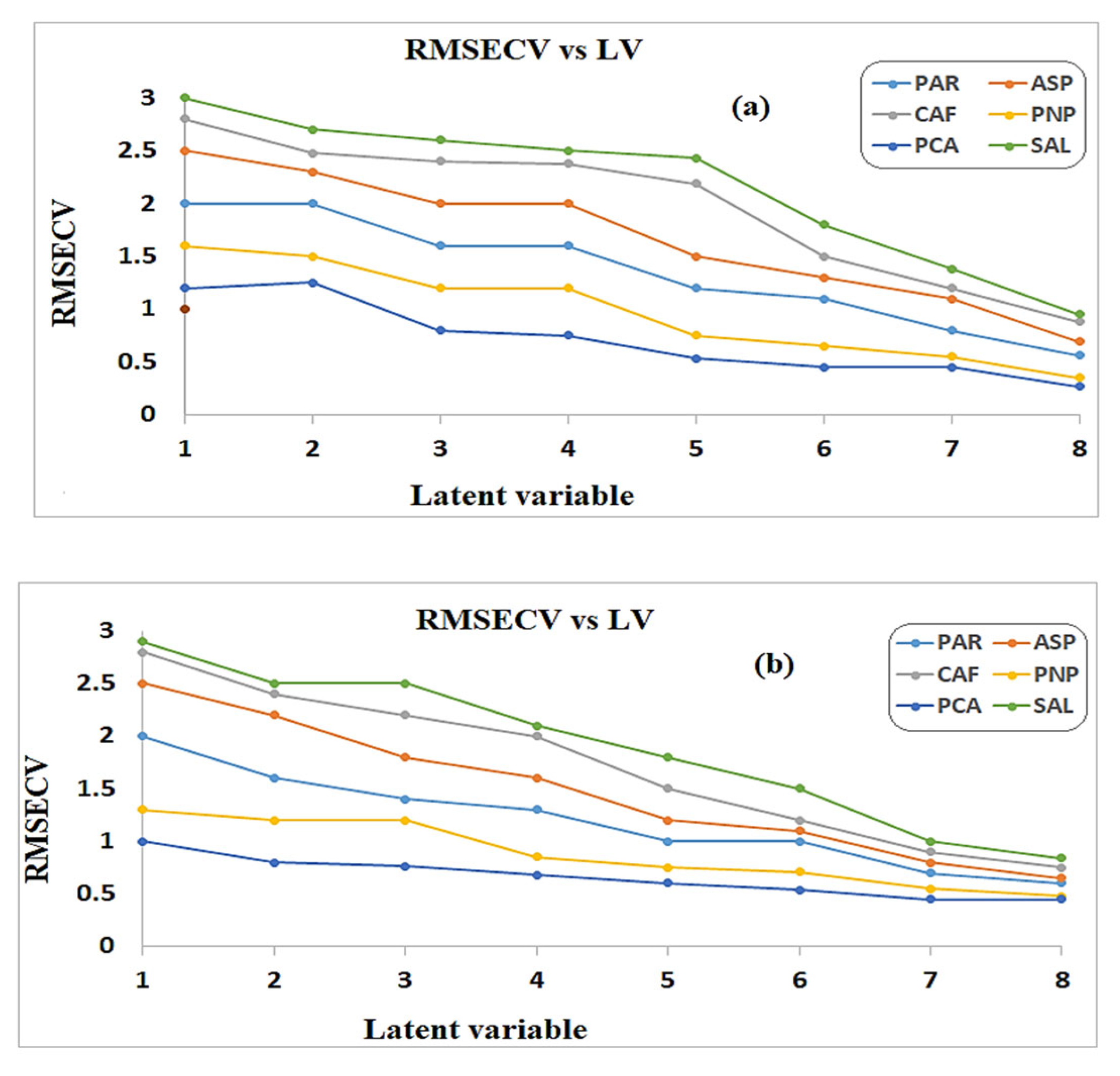
3.1. PCR and PLS Models
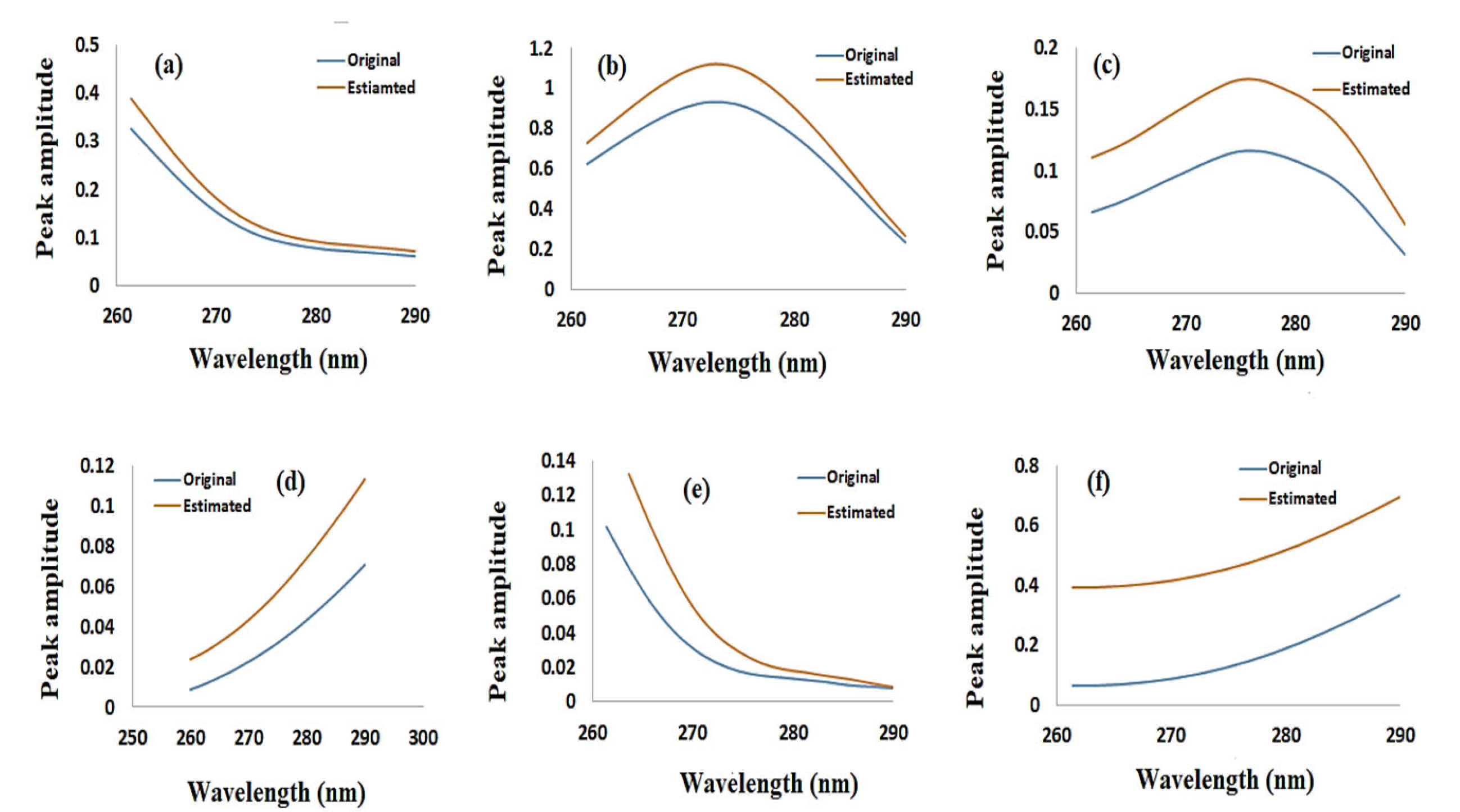
3.2. MCR-ALS Model
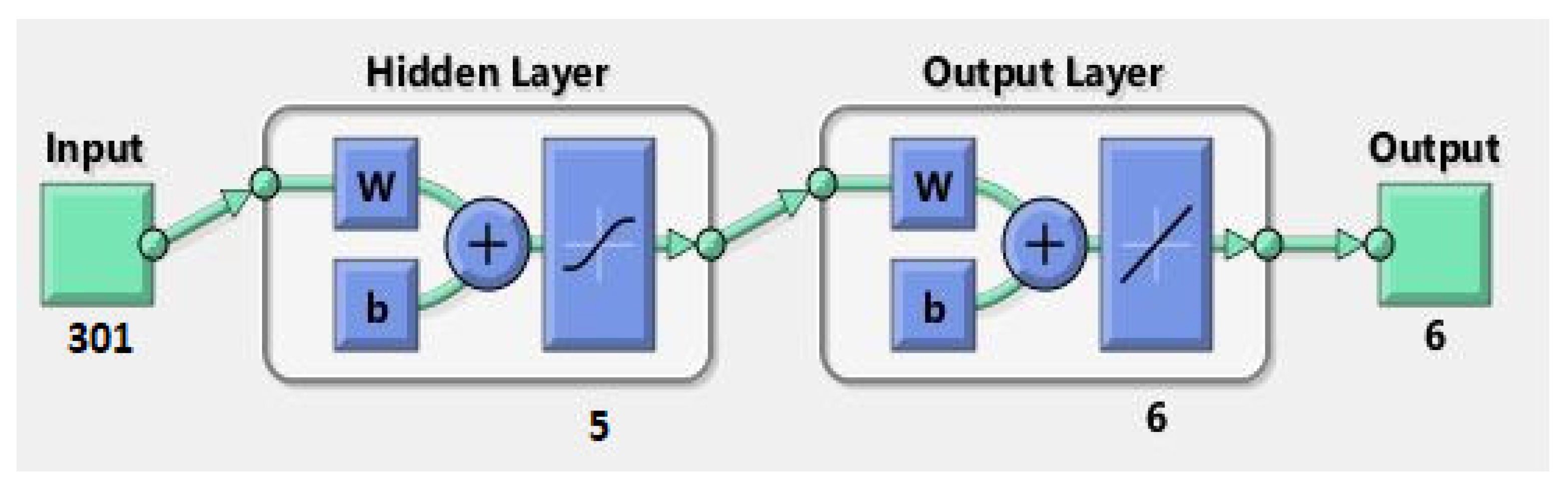
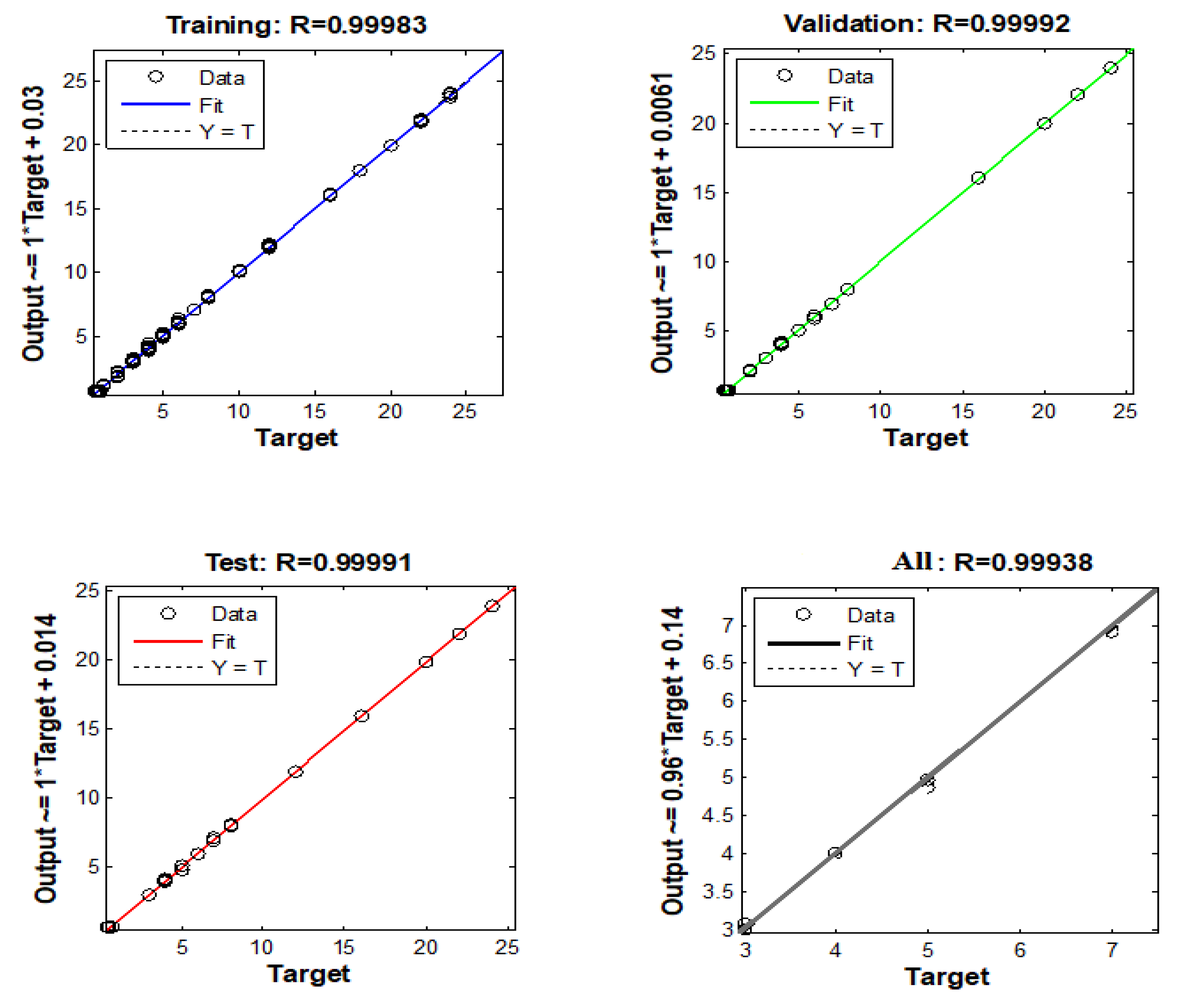
3.3. ANN Model
3.4. Comparative Statistical Study
3.5. Molecular Docking Study
3.6. Evaluation of the Greenness
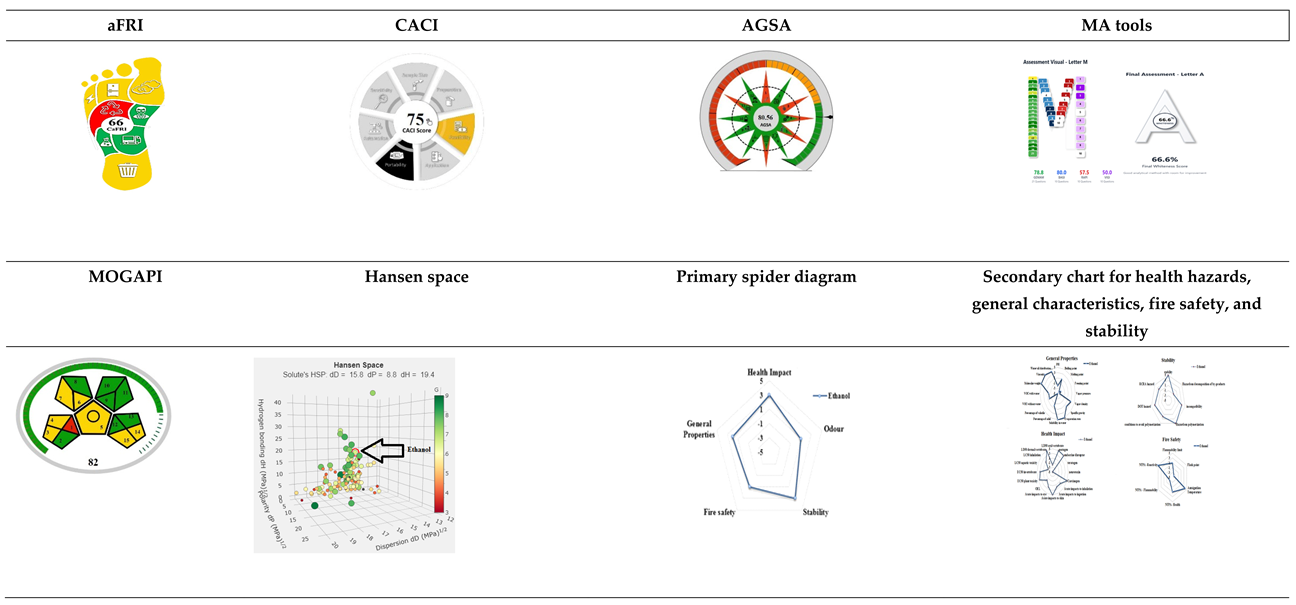
3.6.1. Carbon Footprint Reduction Index (CaFRI)
3.6.2. The Click Analytical Chemistry Index (CACI)
3.6.3. Application of AGSA
3.6.4. Multi-Color Assessment Tool (MA Tool)
3.6.5. Greenness Evaluation with the Modified Green Analytical Procedure Index
3.6.6. Green Solvent Selection Tool
3.6.7. Greenness Index Based on Spider Chart Measurement
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CAF | Caffeine |
| ASP | Directory of open access journals |
| PAR | Three letter acronym |
| SAL | Linear dichroism |
| PNP | P-nitrophenol |
| PCA | P-chloroacetanilide |
| PCR | Principal component regression |
| MCR-ALS | Multivariate curve resolution-alternating least squares |
| ANN | Artificial neural networks |
| PLS | Partial least squares |
References
- Yang, J.; Song, X.; Shi, L.; Zhang, J.; Huang, G.; et al. New insights into the increased risk of migraines from COVID-19 infection and vaccination: A Mendelian randomization study. Front. Neurol. 2024, 15, 1445649. [Google Scholar] [CrossRef]
- Ashina, M.; Buse, D.C.; Ashina, H.; Pozo-Rosich, P.; Peres, M.F.P.; Lee, M.J.; et al. Migraine: Integrated approaches to clinical management and emerging treatments. Lancet 2021, 397, 1505–1518. [Google Scholar] [CrossRef]
- Diener, H.C.; Gaul, C.; Lehmacher, W.; Weiser, T. Aspirin, paracetamol (acetaminophen) and caffeine for the treatment of acute migraine attacks: A systematic review and meta-analysis of randomized placebo-controlled trials. Eur. J. Neurol. 2022, 29, 350–357. [Google Scholar] [CrossRef]
- Magdy, R.; Elmazny, A.; Soliman, S.H.; Elsebaie, E.H.; Ali, S.H.; Abdel Fattah, A.M.; et al. Post-COVID-19 neuropsychiatric manifestations among COVID-19 survivors suffering from migraine: A case–control study. J. Headache Pain. 2022, 23, 101. [Google Scholar] [CrossRef] [PubMed]
- Marzouk, H.M.; Ibrahim, E.A.; Hegazy, M.A.; Saad, S.S. Sustainable liquid chromatographic determination and purity assessment of a possible add-on triple-action over-the-counter pharmaceutical combination in COVID-19. Microchem. J. 2022, 178, 107400. [Google Scholar] [CrossRef] [PubMed]
- El-Adl, S.M.; Mattar, A.A.; El-Abassy, O.M.; Sebaiy, M.M. Development of UV-chemometric techniques for resolving the overlapped spectra of aspirin, caffeine and orphenadrine citrate in their combined pharmaceutical dosage form. BMC Chem. 2025, 19, 75. [Google Scholar] [CrossRef]
- El Sherbiny, D.; Wahba, M.E.K. Analysis of some pharmaceuticals in the presence of their synthetic impurities by applying hybrid micelle liquid chromatography. Open. Chem. 2020, 18, 377–390. [Google Scholar] [CrossRef]
- Farid, J.F.; Mostafa, N.M.; Fayez, Y.M.; Essam, H.M.; ElTanany, B.M. Chemometric quality assessment of paracetamol and phenylephrine hydrochloride with paracetamol impurities; comparative UV-spectrophotometric implementation of four predictive models. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2022, 265, 120308. [Google Scholar] [CrossRef]
- Yiğit, A.; Yardım, Y.; Çelebi, M.; Levent, A.; Şentürk, Z. Graphene/Nafion composite film modified glassy carbon electrode for simultaneous determination of paracetamol, aspirin and caffeine in pharmaceutical formulations. Talanta 2016, 158, 21–29. [Google Scholar] [CrossRef]
- Yiğit, A.; Yardım, Y.; Şentürk, Z. Voltammetric sensor based on boron-doped diamond electrode for simultaneous determination of paracetamol, caffeine, and aspirin in pharmaceutical formulations. IEEE Sens. J. 2015, 16, 1674–1680. [Google Scholar] [CrossRef]
- Sanghavi, B.J.; Srivastava, A.K. Simultaneous voltammetric determination of acetaminophen, aspirin and caffeine using an in situ surfactant-modified multiwalled carbon nanotube paste electrode. Electrochim. Acta 2010, 55, 8638–8648. [Google Scholar] [CrossRef]
- Hajian, R.; Soltaninezhad, A. The spectrophotometric multicomponent analysis of a ternary mixture of paracetamol, aspirin, and caffeine by the double divisor-ratio spectra derivative method. J. Spectrosc. 2013, 2013, 405210. [Google Scholar] [CrossRef]
- Moţ, A.C.; Soponar, F.; Medvedovici, A.; Sârbu, C. Simultaneous spectrophotometric determination of aspirin, paracetamol, caffeine, and chlorphenamine from pharmaceutical formulations using multivariate regression methods. Anal. Lett. 2010, 43, 804–813. [Google Scholar] [CrossRef]
- Özdemir, A.; Dinç, E.; Onur, F. Utilization of multivariate calibration techniques for the spectrophotometric simultaneous determination of paracetamol, aspirin and caffeine in a pharmaceutical formulation. Turk. J. Pharm. Sci. 2004, 1, 139–151. [Google Scholar]
- Bouhsain, Z.; Garrigues, S.; de la Guardia, M. PLS-UV spectrophotometric method for the simultaneous determination of paracetamol, acetylsalicylic acid and caffeine in pharmaceutical formulations. Fresenius J. Anal. Chem. 1997, 357, 973–976. [Google Scholar] [CrossRef]
- Yenda, P.; Katari, N.K.; Ettaboina, S.K.; Satheesh, B.; Muchakayala, S.K.; Gundla, R. An effective and stability-indicating method development and optimization utilizing the Box–Behnken design for the simultaneous determination of acetaminophen, caffeine, and aspirin in tablet formulation. Biomed. Chromatogr. 2023, 37, 55–85. [Google Scholar] [CrossRef]
- Saeed, A.M.; Mohammed, O.J.; Hussein, N.G. Validation of liquid chromatographic analytical method for determination of aspirin, caffeine and paracetamol in some pharmaceutical tablet forms in Iraqi market. Res. J. Pharm. Technol. 2023, 16, 215–220. [Google Scholar] [CrossRef]
- Abdel-Halim, L.M.; Ramadan, N.K.; Rahman, M.K.; Galal, M.M. GC and HPTLC-densitometric methods for simultaneous determination of aspirin, paracetamol, caffeine anhydrous, dextromethorphan hydrobromide and chlorpheniramine maleate: Method validation and application to over-the-counter tablets. Bull. Fac. Pharm. Cairo Univ. 2023, 61, 7. [Google Scholar] [CrossRef]
- Sayed, R.A.; Ibrahim, A.E.; Sharaf, Y.A. Chemometry-assisted UV-spectrophotometric methods for the simultaneous determination of paritaprevir, ritonavir, and ombitasvir in their combined tablet dosage forms: A comparative study. J. Chemom. 2021, 35, 3321. [Google Scholar] [CrossRef]
- Kelani, K.M.; Hegazy, M.A.; Hassan, A.M.; Tantawy, M.A. Univariate versus multivariate spectrophotometric methods for the simultaneous determination of omarigliptin and two of its degradation products. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2022, 271, 120880. [Google Scholar] [CrossRef]
- Yehia, A.M.; Elbalkiny, H.T.; Riad, S.M.; Elsaharty, Y.S. Chemometrics for resolving spectral data of cephalosporins and tracing their residue in wastewater samples. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2019, 219, 436–443. [Google Scholar] [CrossRef]
- Abd El-Hadi, H.R.; Eissa, M.S.; Zaazaa, H.E.; Eltanany, B.M. Development and validation of chemometric-assisted spectrophotometric models for efficient quantitation of a binary mixture of supportive treatments in COVID-19 in the presence of its toxic impurities: A comparative study for eco-friendly assessment. BMC Chem. 2023, 17, 1–16. [Google Scholar] [CrossRef]
- Dong, J.D.; Zhang, Y.Y.; Wang, Y.S.; Wu, M.L.; Zhang, S.; Cai, C.H. Chemometry use in the evaluation of the Sanya Bay water quality. Braz. J. Oceanogr. 2010, 58, 339–352. [Google Scholar] [CrossRef]
- Mouhamed, A.A.; Nadim, A.H.; Mostafa, N.M.; Eltanany, B.M. Application of smart chemometric models for spectra resolution and determination of challenging multi-action quaternary mixture: Statistical comparison with greenness assessment. BMC Chem. 2024, 18, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Wagieh, N.E.; Hegazy, M.A.; Abdelkawy, M.; Abdelaleem, E.A. Quantitative determination of oxybutynin hydrochloride by spectrophotometry, chemometry and HPTLC in presence of its degradation product and additives in different pharmaceutical dosage forms. Talanta 2010, 80, 2007–2015. [Google Scholar] [CrossRef] [PubMed]
- Shaaban, H.; Mostafa, A.; Al-Zahrani, B.; Al-Jasser, B.; Al-Ghamdi, R. Simultaneous determination of drugs affecting central nervous system (CNS) in bulk and pharmaceutical formulations using multivariate curve resolution-alternating least squares (MCR-ALS). J. Anal. Methods Chem. 2020, 2020, 1–8. [Google Scholar] [CrossRef]
- Antunes, M.C.; Simão, J.E.J.; ACD, R.T. Multivariate curve resolution of overlapping voltammetric peaks: Quantitative analysis of binary and quaternary metal mixtures. Analyst 2002, 127, 809–817. [Google Scholar] [CrossRef]
- X, A.; Drug, A. Innovative earth-friendly multivariate techniques for quantitative and qualitative analysis of triamterene and xipamide as antihypertensive drug combination. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2024, 285, 121234. [Google Scholar]
- El-Ragehy, N.A.; Yehia, A.M.; Hassan, N.Y.; Tantawy, M.A.; Abdelkawy, M. Chemometrics tools in detection and quantitation of the main impurities present in aspirin/dipyridamole extended-release capsules. J. AOAC Int. 2016, 99, 948–956. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Hadi, H.R.; Eissa, M.S.; Zaazaa, H.E.; Eltanany, B.M. Univariate versus multivariate spectrophotometric data analysis of triamterene and xipamide: A quantitative and qualitative greenly profiled comparative study. BMC Chem. 2023, 17, 1–14. [Google Scholar] [CrossRef]
- Tantawy, M.A.; Michael, A.M. Artificial neural networks versus partial least squares and multivariate resolution-alternating least squares approaches for the assay of ascorbic acid, rutin, and hesperidin in an antioxidant formulation. Spectrosc. Lett. 2019, 52, 1–7. [Google Scholar] [CrossRef]
- Zeid, A.M.; Abdelazim, A.H.; Shahin, M. Simultaneous spectrophotometric quantitative analysis of elbasvir and grazoprevir using assisted chemometric models. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2021, 252, 119505. [Google Scholar] [CrossRef]
- ICH, Q.1A. (R.2). Stability testing of new drug substances and products. International Conference on Harmonization, IFPMA, Geneva, Switzerland, 2003. [Google Scholar]
- Abd El-Hadi, H.R.; Eissa, M.S.; Zaazaa, H.E.; Eltanany, B.M. Chemometric quality assessment of doxylamine succinate with its degradation product; implementation of two predictive models on UV-spectrophotometric data of anti-emetic binary mixture. J. AOAC Int. 2022, 105, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Hadi, H.R.; Eltanany, B.M.; Zaazaa, H.E. Smart spectrophotometric approaches for estimation of difluprednate in existence of its synthetic precursor and alkaline degradation product; comparative statistical studies and analytical ecological appraisal. Bull. Fac. Pharm. Cairo Univ. 2024, 62, 32–43. [Google Scholar] [CrossRef]
- Abd El-Hadi, H.R. Comparative statistical evaluation of greenness, blueness, and whiteness spectrophotometric methods for dexamethasone and chloramphenicol estimation. Sci. Rep. 2025, 15, 1–12. [Google Scholar] [CrossRef]
- Yadav, T.C.; Kumar, N.; Raj, U.; Goel, N.; Vardawaj, P.K.; Prasad, R.; et al. Exploration of interaction mechanism of tyrosol as a potent anti-inflammatory agent. J. Biomol. Struct. Dyn. 2020, 38, 382–397. [Google Scholar] [CrossRef]
- Jahan, I.; Sakib, S.A.; Alam, N.; Majumder, M.; Sharmin, S.; Reza. Pharmacological insights into Chukrasia velutina bark: Experimental and computer-aided approaches. Anim. Model Exp. Med. 2022, 5, 377–388. [Google Scholar] [CrossRef]
- Mansour, F.R.; Nowak, P.M. Introducing the carbon footprint reduction index (CaFRI) as a software-supported tool for greener laboratories in chemical analysis. BMC Chem. 2025, 19, 1–12. [Google Scholar] [CrossRef]
- Bahgat, E.A.; Saleh, H.; Darwish, I.M.; El-Abassy, O.M. Green HPLC technique development for the simultaneous determination of the potential combination of mirabegron and tamsulosin. Sci. Rep. 2025, 15, 1–10. [Google Scholar] [CrossRef]
- Mansour, F.R.; Bedair, A.; Locatelli, M. Click Analytical Chemistry Index as a novel concept and framework, supported with open source software to assess analytical methods. Adv. Sample Prep. 2025, 14, 100164. [Google Scholar] [CrossRef]
- Nabil, M.; Ahmed, D.A.; Abbas, S.S.; Lotfy, H.M.; Marzouk, H.M. Green HPLC strategy for quantification of carvedilol and hydrochlorothiazide in cardiac medications with in-vitro dissolution kinetics and impurity profiling. BMC Chem. 2025, 19, 1–10. [Google Scholar] [CrossRef]
- Mansour, F.R.; Bedair, A.; Belal, F.; Magdy, G.; Locatelli, M. Analytical Green Star Area (AGSA) as a new tool to assess greenness of analytical methods. Sustain. Chem. Pharm. 2025, 46, 102051. [Google Scholar] [CrossRef]
- Abbas, A.E.F.; Al Kamaly, O.; Magdy, G.; Halim, M.K. MA tool—Multi-color assessment platform for white analytical chemistry: Unified evaluation of method greenness, practicality, performance, and innovation. Microchem. J. 2025, 216, 108512. [Google Scholar] [CrossRef]
- El-Kafrawy, D.S.; Issa, A.E.; Beltagy, Y.A.; El-Shoubashy, O.H. A challenging multi-purpose HPLC-DAD concurrent determination of sulfacetamide with two pharmacopeial impurities, three co-formulated drugs and interfering preservative in ophthalmic drops: Triad color appraisal of sustainability. Microchem. J. 2025, 114342. [Google Scholar] [CrossRef]
- Mansour, F.R.; Płotka-Wasylka, J.; Locatelli, M. Modified GAPI (MoGAPI) tool and software for the assessment of method greenness: Case studies and applications. Analytica 2024, 5, 451–457. [Google Scholar] [CrossRef]
- Hafez, H.M.; Mostafa, A.A.; El-Abassy, O.M. Spider diagram with greenness assessment criteria for mathematical UV spectrophotometric methodologies for separating overlaid spectrum signals of diclofenac sodium and lidocaine hydrochloride. Green. Anal. Chem. 2025, 13, 100265. [Google Scholar] [CrossRef]
- Kayali, Z.; Obaydo, R.H.; Sakur, A.A. Spider diagram and sustainability evaluation of UV-methods strategy for quantification of aspirin and sildenafil citrate in the presence of salicylic acid in their bulk and formulation. Heliyon 2023, 9, e14612. [Google Scholar] [CrossRef]
- Larsen, C.; Lundberg, P.; Tang, S.; Ràfols-Ribé, J.; Sandström, A.; Lindh, E.M.; et al. A tool for identifying green solvents for printed electronics. Nat. Commun. 2021, 12, 4510. [Google Scholar] [CrossRef]
- Lotfy, H.M.; Obaydo, R.H.; Nessim, C.K. Spider chart and whiteness assessment of synergistic spectrophotometric strategy for quantification of triple combination recommended in seasonal influenza—Detection of spurious drug. Sustain. Chem. Pharm. 2023, 32, 100980. [Google Scholar] [CrossRef]
- Shen, Y.; Lo, C.; Nagaraj, D.R.; Farinato, R.; Essenfeld, A.; Somasundaran, P. Development of Greenness Index as an evaluation tool to assess reagents: Evaluation based on SDS (Safety Data Sheet) information. Miner. Eng. 2016, 94, 1–9. [Google Scholar] [CrossRef]
| Mix No. | PAR | ASP | CAF | PNP | PCA | SAL |
|---|---|---|---|---|---|---|
| 1 | 6.00 | 10.00 | 11.00 | 4.00 | 0.70 | 10.00 |
| 2 | 6.00 | 6.00 | 3.00 | 6.00 | 0.60 | 14.00 |
| 3 | 2.00 | 6.00 | 19.00 | 3.00 | 0.90 | 10.00 |
| 4 | 2.00 | 14.00 | 7.00 | 6.00 | 0.70 | 8.00 |
| 5 | 10.00 | 8.00 | 19.00 | 4.00 | 0.60 | 8.00 |
| 6 | 4.00 | 14.00 | 11.00 | 3.00 | 0.60 | 12.00 |
| 7 | 10.00 | 10.00 | 7.00 | 3.00 | 0.80 | 14.00 |
| 8 | 6.00 | 8.00 | 7.00 | 5.00 | 0.90 | 12.00 |
| 9 | 4.00 | 8.00 | 15.00 | 6.00 | 0.80 | 10.00 |
| 10 | 4.00 | 12.00 | 19.00 | 5.00 | 0.70 | 14.00 |
| 11 | 8.00 | 14.00 | 15.00 | 4.00 | 0.90 | 14.00 |
| 12 | 10.00 | 12.00 | 11.00 | 6.00 | 0.90 | 6.00 |
| 13 | 8.00 | 10.00 | 19.00 | 6.00 | 0.50 | 12.00 |
| 14 | 6.00 | 14.00 | 19.00 | 2.00 | 0.80 | 6.00 |
| 15 | 10.00 | 14.00 | 3.00 | 5.00 | 0.50 | 10.00 |
| 16 | 10.00 | 6.00 | 15.00 | 2.00 | 0.70 | 12.00 |
| 17 | 2.00 | 12.00 | 3.00 | 4.00 | 0.80 | 12.00 |
| 18 | 8.00 | 6.00 | 11.00 | 5.00 | 0.80 | 8.00 |
| 19 | 2.00 | 10.00 | 15.00 | 5.00 | 0.60 | 6.00 |
| 20 | 6.00 | 12.00 | 15.00 | 3.00 | 0.50 | 8.00 |
| 21 | 8.00 | 12.00 | 7.00 | 2.00 | 0.60 | 10.00 |
| 22 | 8.00 | 8.00 | 3.00 | 3.00 | 0.70 | 6.00 |
| 23 | 4.00 | 6.00 | 7.00 | 4.00 | 0.50 | 6.00 |
| 24 | 2.00 | 8.00 | 11.00 | 2.00 | 0.50 | 14.00 |
| 25 | 4.00 | 10.00 | 3.00 | 2.00 | 0.90 | 8.00 |
| a) Prediction of validation set samples using PCR and PLS models | ||||||||||||||||||||||||||||||||||
| Concentration (µg/mL) | PCR | PLS | ||||||||||||||||||||||||||||||||
| PAR | ASP | CAF | PNP | PCA | SAL | PAR | ASP | CAF | PNP | PCA | SAL | PAR | ASP | CAF | PNP | PCA | SAL | |||||||||||||||||
| 6.00 | 10.00 | 11.00 | 4.00 | 0.70 | 10.00 | 100.58 | 99.05 | 98.79 | 97.45 | 99.54 | 102.10 | 101.03 | 97.01 | 98.40 | 100.7 | 98.11 | 98.42 | |||||||||||||||||
| 2.00 | 6.00 | 19.00 | 3.00 | 0.90 | 10.00 | 99.01 | 98.37 | 99.90 | 100.76 | 100.59 | 98.89 | 99.11 | 98.30 | 98.27 | 99.54 | 101.70 | 99.04 | |||||||||||||||||
| 6.00 | 8.00 | 7.00 | 5.00 | 0.90 | 12.00 | 99.12 | 101.83 | 98.32 | 98.88 | 101.37 | 101.59 | 100.53 | 97.59 | 100.60 | 100.41 | 98.03 | 97.45 | |||||||||||||||||
| 4.00 | 12.00 | 19.00 | 5.00 | 0.70 | 14.00 | 99.88 | 99.01 | 98.21 | 100.24 | 100.06 | 101.70 | 101.08 | 100.40 | 97.32 | 98.81 | 101.49 | 98.73 | |||||||||||||||||
| 2.00 | 12.00 | 3.00 | 4.00 | 0.80 | 12.00 | 99.30 | 97.73 | 99.33 | 98.64 | 98.08 | 101.43 | 101.39 | 97.75 | 98.54 | 97.74 | 99.33 | 101.9 | |||||||||||||||||
| 6.00 | 12.00 | 15.00 | 3.00 | 0.50 | 8.00 | 100.24 | 101.51 | 98.44 | 97.30 | 101.64 | 101.90 | 100.43 | 101.75 | 100.49 | 98.89 | 99.64 | 101.38 | |||||||||||||||||
| 8.00 | 12.00 | 7.00 | 2.00 | 0.60 | 10.00 | 101.25 | 99.791 | 99.21 | 99.93 | 97.35 | 101.37 | 101.25 | 101.20 | 98.18 | 98.80 | 99.02 | 99.67 | |||||||||||||||||
| 8.00 | 8.00 | 3.00 | 3.00 | 0.70 | 6.00 | 101.92 | 100.34 | 98.64 | 99.34 | 99.84 | 102.62 | 102.24 | 97.72 | 99.76 | 100.78 | 98.70 | 99.05 | |||||||||||||||||
| Mean (%) | 100.16 | 99.70 | 98.85 | 99.06 | 99.80 | 101.45 | 100.88 | 98.96 | 98.94 | 99.45 | 99.50 | 99.45 | ||||||||||||||||||||||
| RSD (%) | 1.04 | 1.45 | 0.58 | 1.25 | 1.49 | 1.11 | 0.90 | 1.85 | 1.19 | 1.09 | 1.40 | 1.49 | ||||||||||||||||||||||
| b) Prediction of validation set samples using MCR-ALS and ANN models | ||||||||||||||||||||||||||||||||||
| Concentration (µg/mL) | MCR-ALS | ANN | ||||||||||||||||||||||||||||||||
| PAR | ASP | CAF | PNP | PCA | SAL | PAR | ASP | CAF | PNP | PCA | SAL | PAR | ASP | CAF | PNP | PCA | SAL | |||||||||||||||||
| 6.00 | 10.00 | 11.00 | 4.00 | 0.70 | 10.00 | 98.05 | 100.95 | 98.31 | 97.50 | 98.57 | 99.76 | 101.12 | 101.63 | 101.46 | 97.90 | 97.03 | 99.89 | |||||||||||||||||
| 2.00 | 6.00 | 19.00 | 3.00 | 0.90 | 10.00 | 97.53 | 100.33 | 100.91 | 99.46 | 97.77 | 98.48 | 102.69 | 101.93 | 99.68 | 100.53 | 100.01 | 98.93 | |||||||||||||||||
| 6.00 | 8.00 | 7.00 | 5.00 | 0.90 | 12.00 | 98.47 | 99.77 | 98.97 | 97.62 | 98.88 | 98.85 | 99.45 | 99.74 | 98.10 | 98.88 | 97.66 | 98.91 | |||||||||||||||||
| 4.00 | 12.00 | 19.00 | 5.00 | 0.70 | 14.00 | 98.13 | 98.86 | 100.07 | 100.30 | 102.85 | 98.32 | 101.53 | 100.00 | 99.31 | 99.44 | 98.57 | 98.40 | |||||||||||||||||
| 2.00 | 12.00 | 3.00 | 4.00 | 0.80 | 12.00 | 99.55 | 97.06 | 101.36 | 100.11 | 100.50 | 99.80 | 101.48 | 101.92 | 98.93 | 101.10 | 99.42 | 97.73 | |||||||||||||||||
| 6.00 | 12.00 | 15.00 | 3.00 | 0.50 | 8.00 | 101.88 | 98.49 | 98.18 | 100.47 | 98.05 | 98.28 | 97.79 | 97.20 | 101.68 | 98.90 | 100.35 | 98.15 | |||||||||||||||||
| 8.00 | 12.00 | 7.00 | 2.00 | 0.60 | 10.00 | 101.71 | 100.53 | 97.47 | 99.18 | 97.50 | 101.80 | 99.02 | 98.34 | 98.74 | 100.52 | 102.05 | 97.04 | |||||||||||||||||
| 8.00 | 8.00 | 3.00 | 3.00 | 0.70 | 6.00 | 100.25 | 98.26 | 102.81 | 97.53 | 98.57 | 98.70 | 99.21 | 102.00 | 100.31 | 98.54 | 101.60 | 100.71 | |||||||||||||||||
| Mean (%) | 99.44 | 99.28 | 99.76 | 99.02 | 99.08 | 99.24 | 100.28 | 100.34 | 99.77 | 99.47 | 99.58 | 98.72 | ||||||||||||||||||||||
| RSD (%) | 1.69 | 1.33 | 1.84 | 1.28 | 1.77 | 1.19 | 1.65 | 1.84 | 1.28 | 1.12 | 1.78 | 1.17 | ||||||||||||||||||||||
| a) Performance parameters of the calibration and validation sets using PCR and PLS models | |||||||||||||
| Validation parameters | PCR | PLS | |||||||||||
| PAR | ASP | CAF | PNP | PCA | SAL | PAR | ASP | CAF | PNP | PCA | SAL | ||
| Linearity (µg/mL) a | 2.00-10.00 | 6.00-14.00 | 3.00-19.00 | 2.00-6.00 | 0.50-0.90 | 6.00-14.00 | 2.00-10.00 | 6.00-14.00 | 3.00-19.00 | 2.00-6.00 | 0.50-0.90 | 6.00-14.00 | |
| Correlation coefficient (r)a | 0.9996 | 0.9998 | 0.9998 | 0.9998 | 0.9997 | 0.9998 | 0.9997 | 0.9998 | 0.9999 | 0.9990 | 0.9993 | 0.9994 | |
| Slope a | 0.99985 | 0.98775 | 1.003383 | 1.0692 | 0.9955 | 1.0066 | 0.99985 | 0.98275 | 0.971667 | 1.0492 | 0.9755 | 0.9966 | |
| Intercept a | -0.2169 | -0.1463 | -0.30954 | -0.5384 | -0.01005 | -0.395 | -0.1369 | -0.0163 | -0.067 | -0.3584 | -0.00205 | -0.155 | |
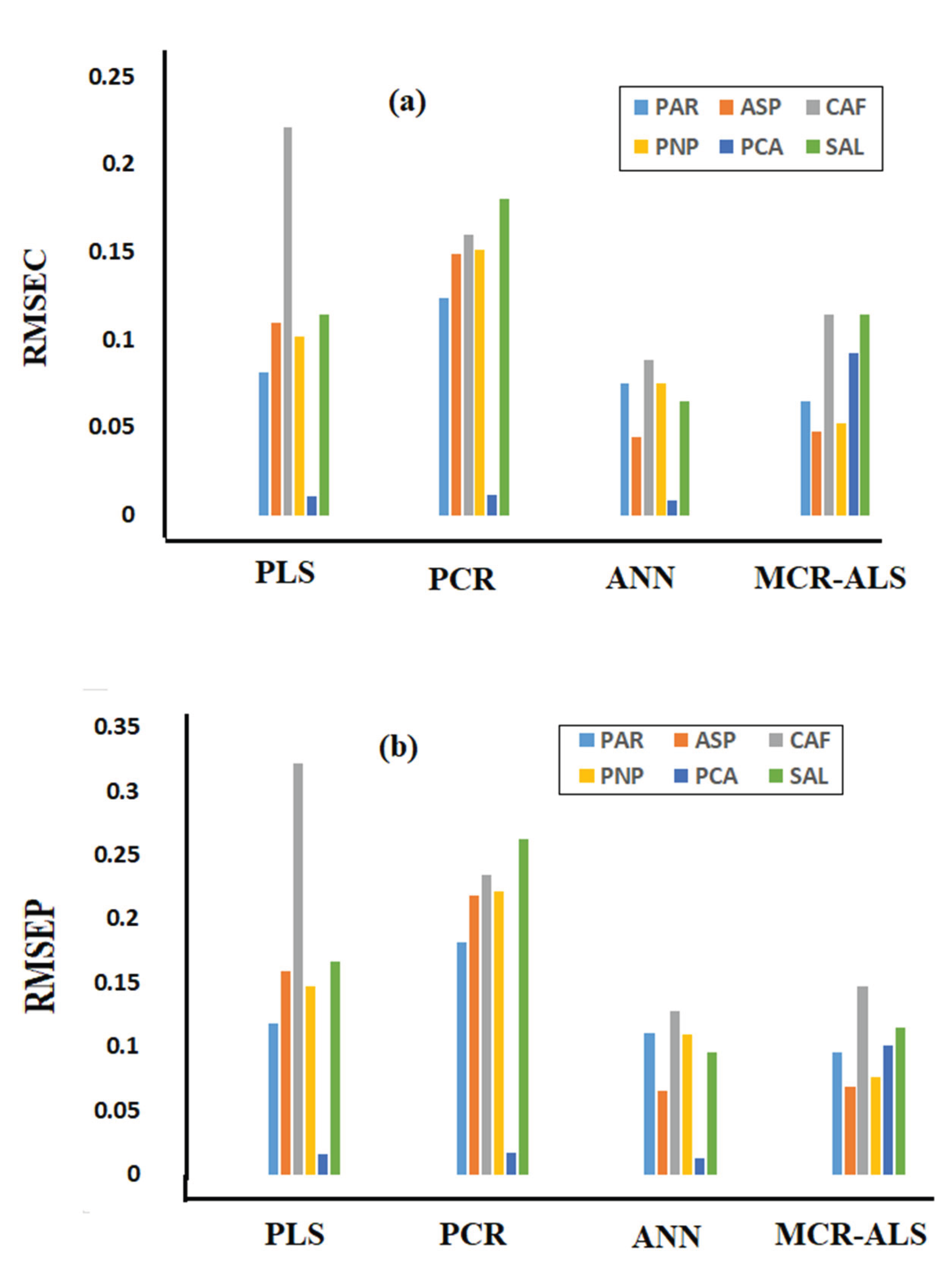
| RMSEC b | 0.124 | 0.149 | 0.16 | 0.151 | 0.0113 | 0.18 | 0.081 | 0.1095 | 0.2206 | 0.1014 | 0.0109 | 0.114 | |
| RMSEP c | 0.18 | 0.21 | 0.23 | 0.22 | 0.01 | 0.26 | 0.11 | 0.15 | 0.32 | 0.14 | 0.01 | 0.16 | |
| LOD d | 0.32 | 0.38 | 0.46 | 0.12 | 0.02 | 0.29 | 0.26 | 0.37 | 0.11 | 0.34 | 0.04 | 0.66 | |
| LOQ d | 0.97 | 1.15 | 1.40 | 0.36 | 0.08 | 0.89 | 0.81 | 1.14 | 0.36 | 1.04 | 0.14 | 2.00 | |
| b) Performance parameters of the calibration and validation sets using MCR-ALS and ANN models | |||||||||||||
| Validation parameters | MCR-ALS | ANN | |||||||||||
| PAR | ASP | CAF | PNP | PCA | SAL | PAR | ASP | CAF | PNP | PCA | SAL | ||
|
Linearity (µg/mL) a |
2.00-10.00 | 6.00-14.00 | 3.00-19.00 | 2.00-6.00 | 0.50-0.90 | 6.00-14.00 | 2.00-10.00 | 6.00-14.00 | 3.00-19.00 | 2.00-6.00 | 0.50-0.90 | 6.00-14.00 | |
| Correlation coefficient (r)a | 0.999871 | 0.999976 | 0.999952 | 0.999288 | 0.999232 | 0.99944 | 0.999873 | 0.999877 | 0.999969 | 0.999148 | 0.999272 | 0.999961 | |
| Slope a | 1.00525 | 0.98715 | 0.991374 | 1.0054 | 1.02 | 0.9966 | 0.98975 | 0.99715 | 0.997701 | 0.99604 | 1.016 | 1.0193 | |
| Intercept a | -0.1249 | 0.0393 | -0.10984 | -0.101 | -0.024 | -0.155 | -0.0679 | -0.0407 | -0.13178 | -0.11088 | -0.0182 | -0.2992 | |
| RMSEC b | 0.06 | 0.04 | 0.11 | 0.05 | 0.09 | 0.11 | 0.07 | 0.04 | 0.08 | 0.07 | 0.01 | 0.06 | |
| RMSEP c | 0.09 | 0.06 | 0.14 | 0.07 | 0.100 | 0.16 | 0.11 | 0.06 | 0.12 | 0.11 | 0.01 | 0.09 | |
| LOD d | 0.20 | 0.13 | 0.22 | 0.25 | 0.05 | 0.66 | 0.20 | 0.31 | 0.18 | 0.33 | 0.05 | 0.17 | |
| LOQ d | 0.61 | 0.41 | 0.68 | 0.92 | 0.16 | 2.00 | 0.60 | 0.94 | 0.55 | 1.01 | 0.15 | 0.52 | |
| Parameters | PAR | ASP | CAF | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Proposed methods | Reported Method a |
Proposed methods | Reported Method a |
Proposed methods | Reported Method a |
||||||||||
| PCR | PLS | MCR-ALS | ANN | HPLC | PCR | PLS | MCR-ALS | ANN | HPLC | PCR | PLS | MCR-ALS | ANN | HPLC | |
| Mean b (%) | 100.31 | 100.00 | 99.94 | 100.06 | 101.56 | 100.81 | 100.09 | 99.93 | 99.67 | 100.89 | 100.42 | 100.42 | 101.09 | 100.62 | 101.58 |
| SD | 1.260 | 1.529 | 1.603 | 1.639 | 0.906 | 1.011 | 1.047 | 1.121 | 1.065 | 1.378 | 1.289 | 1.041 | 0.757 | 0.871 | 1.648 |
| Variance | 1.58 | 2.33 | 2.56 | 2.68 | 0.82 | 1.02 | 1.09 | 1.25 | 1.13 | 1.89 | 1.66 | 1.08 | 0.57 | 0.75 | 2.71 |
| n | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 6 |
|
Student’s t-test (2.23)c |
1.97 |
2.16 |
2.16 |
1.96 |
- | 0.12 |
1.14 |
1.32 |
1.72 |
- | 1.35 |
1.45 |
0.66 |
1.26 |
- |
|
F-value c (5.05)c |
1.93 |
2.85 |
3.13 |
3.27 |
- | 1.86 |
1.73 |
1.51 |
1.67 |
- | 1.63 |
2.51 |
4.74 |
3.58 |
- |
| Source of variation | SS | df | MS | f | P-value | F-crit | |
|---|---|---|---|---|---|---|---|
| PAR | Between groups | 4.08 | 4 | 1.02 | 0.54 | 0.70 | 2.75 |
| Within groups | 46.88 | 25 | 1.87 | - | - | - | |
| ASP | Between groups | 5.25 | 4 | 1.31 | 1.36 | 0.27 | 2.75 |
| Within groups | 24.11 | 25 | 0.96 | - | - | - | |
| CAF | Between groups | 2.77 | 4 | 0.69 | 0.75 | 0.56 | 2.75 |
| Within groups | 22.86 | 25 | 0.91 | - | - | - |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




