Submitted:
25 April 2026
Posted:
28 April 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Preliminaries
2.1. Background
2.2. Newton Raphson Based Optimizer
2.2.1. Initialization and Fitness Evaluation
2.2.2. Newton-Raphson Search Rule (NRSR)
2.2.3. Trap Avoidance Strategy (TAS)
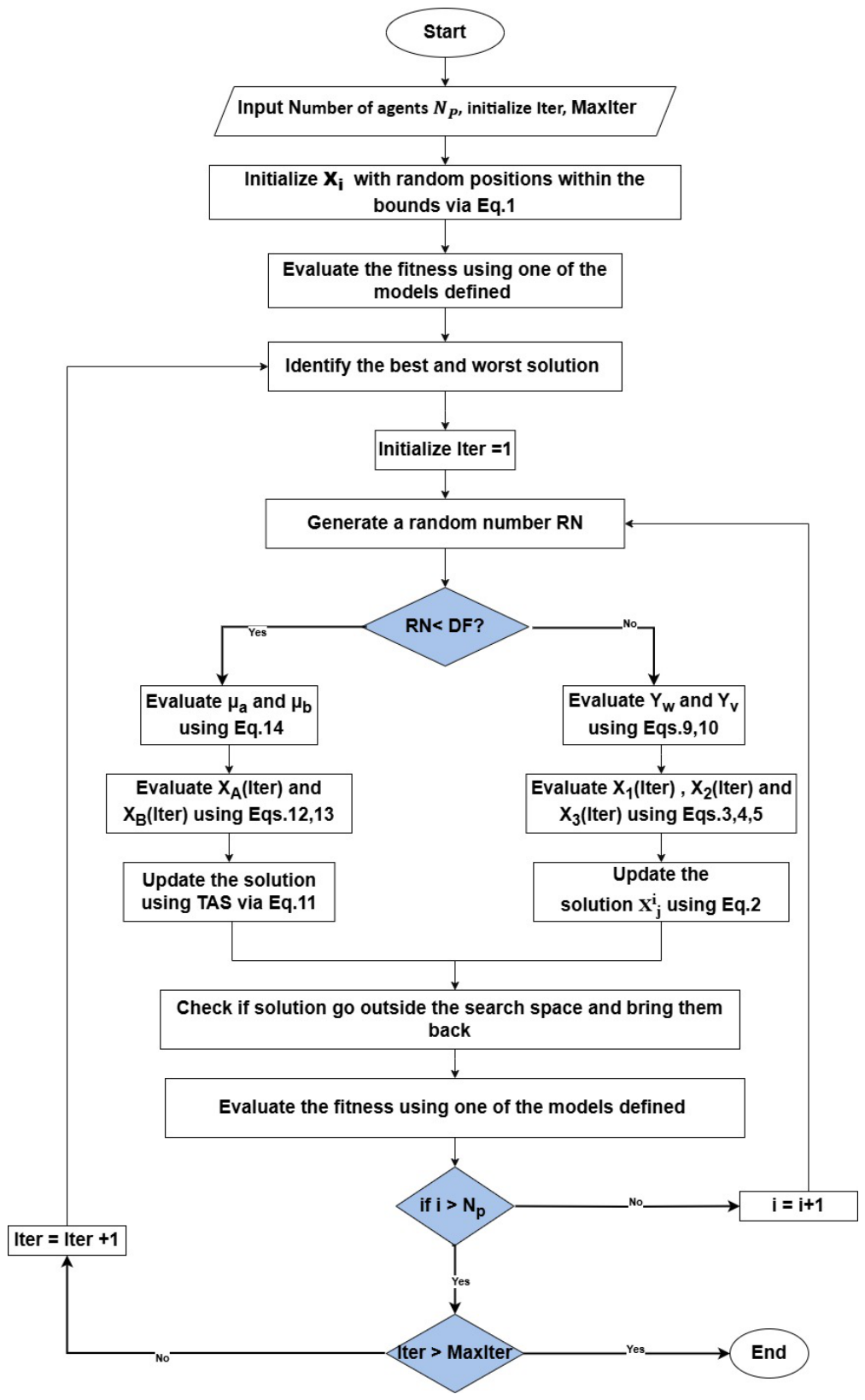
- Step 1.
- Specify the NRBO parameters: .
- Step 2.
- Initialize the solution vectors using Equation (1).
- Step 3.
- For each solution vector, evaluate the fitness function to assess the performance of each solution.
- Step 4.
- Rank the solutions based on their fitness values. Identify the agent with the lowest fitness as the best, and the one with the highest fitness as the worst.
- Step 5.
-
Generate a random number to determine the strategy compaired with Draft factor (DF=0.6) to apply:
- Step 6.
- Re-evaluate the fitness function for the updated design variables.
- Step 7.
- Repeat steps 3–6 until the maximum number of iterations is reached. Once complete, the final position of the best agent is considered the optimal solution.
2.3. Chaotic Maps
2.4. Lennard Potential
3. BCNRBO: Binary Chaotic Newton Raphson Based Optimizer
3.1. Dynamic Potential (DP)
| Algorithm 1 Pseudocode for Dynamic Potential. |
|
3.2. Chaotic Enforcement (CE)
3.3. Chaotic Newton-Raphson Search Rule (CNRSR)
| Algorithm 2 Pseudocode of Binary Chaotic Newton Raphson Based Optimizer (BCNRBO) |
|
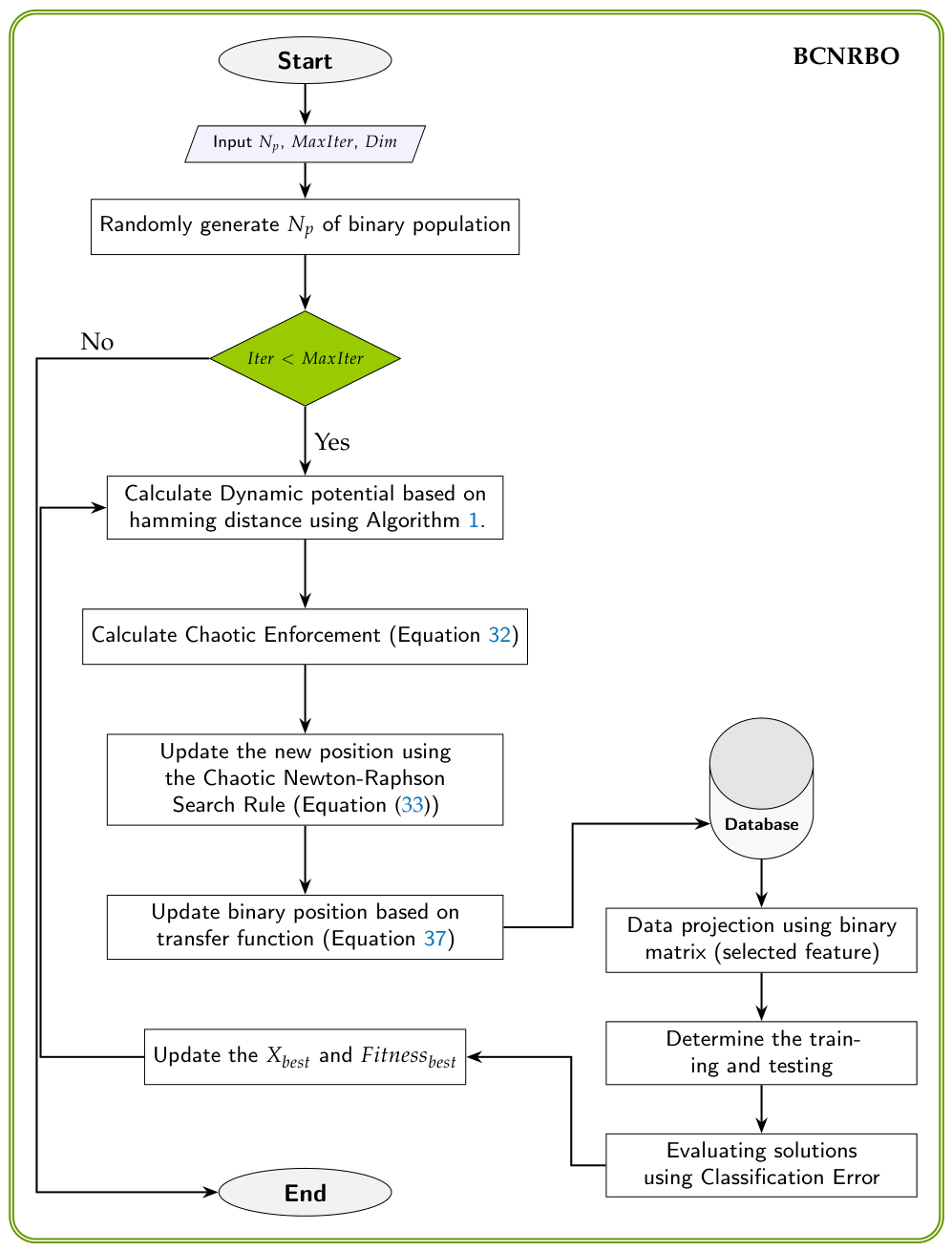
3.4. The Proposed Transfer Function
3.5. Computational Complexity
3.6. Code Availability
4. Computational Experiments
- (a)
- This study focuses on low to medium dimensional large scale datasets, including the Pd Speech dataset with 753 features, to demonstrate the scalability of the proposed algorithm.
- (b)
- This study simultaneously optimizes classification accuracy (minimum misclassifications error) while minimizing selected features.
- ■
- KNN classifier [78]: The k-nearest neighbor method is a popular classification method in data mining and statistics due to its simplicity and significant classification performance. It uses k nearest neighbors to determine the class of examples, making it a memory-based classification, lazy learning technique. The data were classified with k = 5 for KNN classification, and the training set results were obtained by divide data into training and testing. KNN is still utilized today, proving its accuracy and professionalism [79,80,81].
- ■
- DT classifier [82]: Decision trees is a tree-based technique used in data mining, where a path starts at the root and ends at the leaf node [82]. They are hierarchical exemplifications of knowledge relationships, with nodes representing purposes. Decision trees are widely used in fields like machine learning, image processing, and pattern identification. They unify basic tests efficiently and cohesively, comparing numeric features to threshold values [83]. They are commonly used for grouping purposes and classification models in data mining. Decision trees have found many implementation fields due to their simple analysis and precision on multiple data forms.
- ■
- NB classifier [84]: The Naive Bayes Classifier is a simple yet effective approach based on the Bayesian theorem that is especially well-suited for high-dimensional inputs [84]. It is capable of outperforming more advanced classification algorithms. The prior probability, based on the percentage of Green and Red objects, is used to predict outcomes before they occur, ensuring that new cases are classified accordingly. It was derived from Bayesian Classification is a supervised learning and statistical method that uses probabilistic models to capture uncertainty and solve diagnostic and predictive problems.
4.1. Algorithms, Parameters and Experimental Setup
- Fitness values (using wrapper approach): They are obtained from each approach as reported (classification error: it is obtained by using the selected features on the test dataset). The mean, min, and max fitness values are compared [85]. The average is calculated over 20 independent runs.
- Average selection features: It is the other comparison that has been presented in here.
- value from Wilcoxon’s rank sum test and the mean rank value from Friedman test (Wilcoxon’s rank sum test and Friedman test are non-parametric statistical tests with 5% significance level [86]).
4.2. Datasets Description
4.3. Evaluation and Analysis of the Proposed Algorithm Using Chaotic Maps
5. Experimental Results and Discussion
5.1. Evaluation of BCNRBO with KNN Classifier
5.1.1. The Statistical Test for KNN Classifier
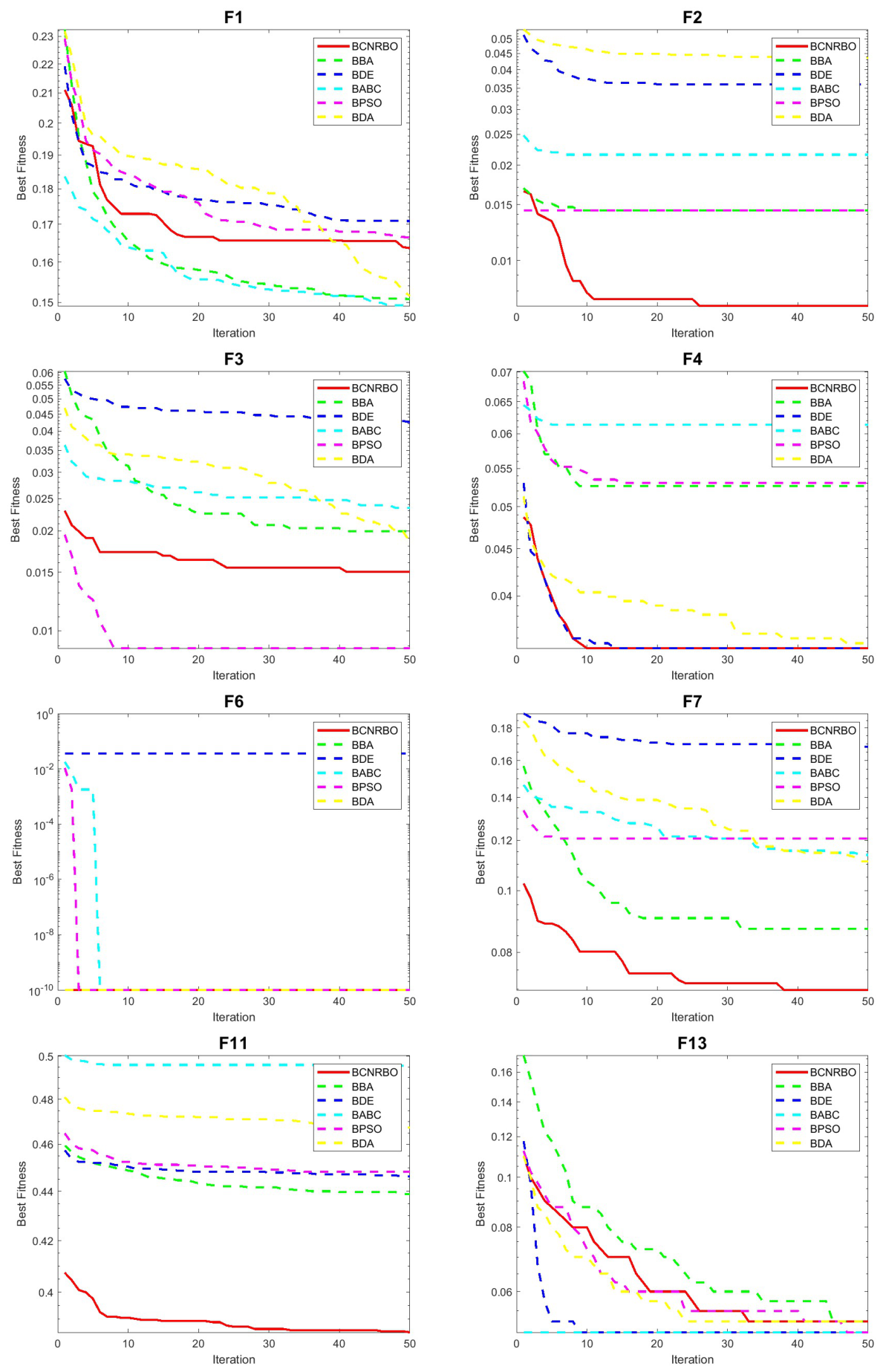
5.1.2. Convergence Graphic for Different Dataset Using KNN Classifier
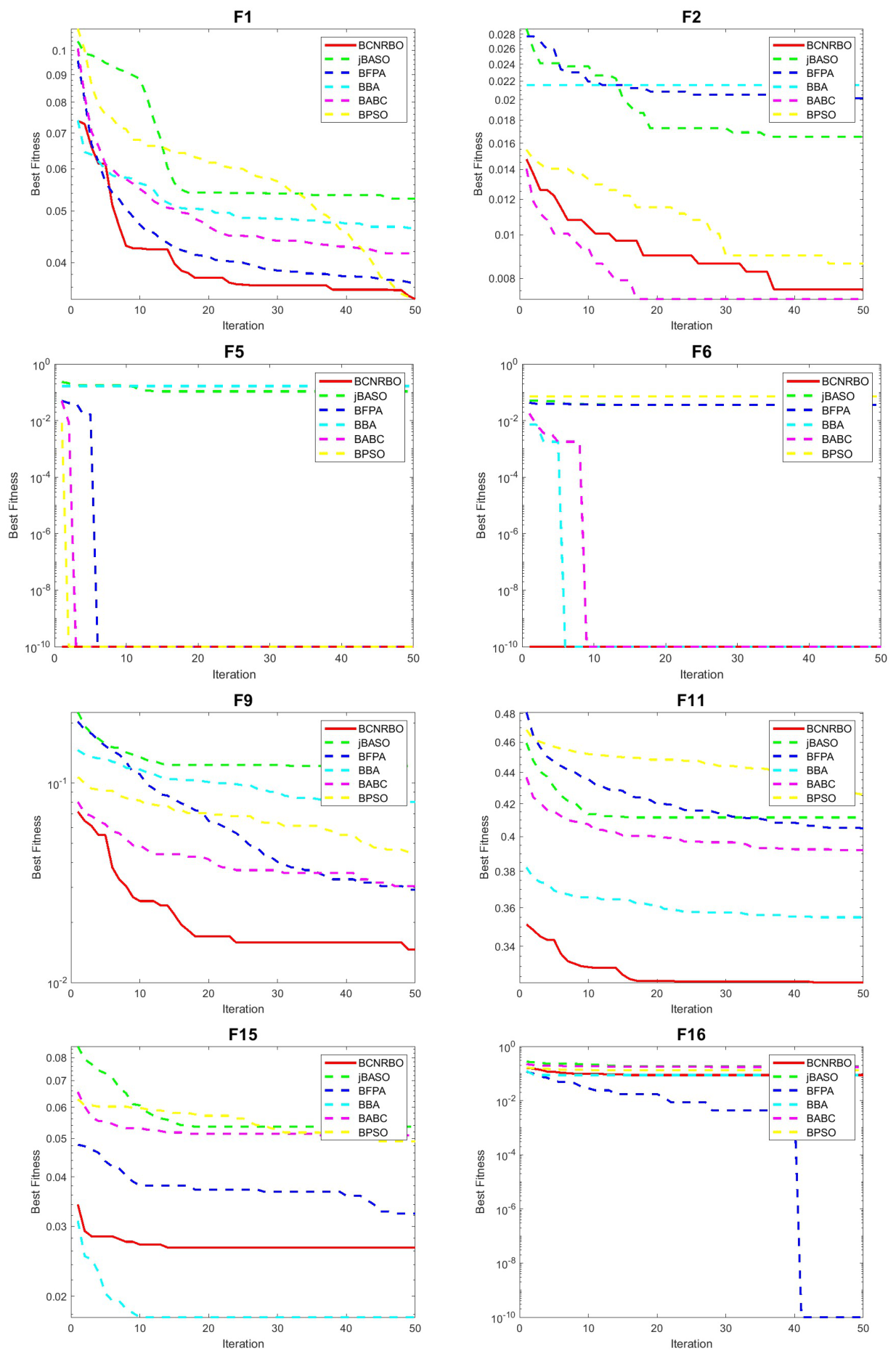
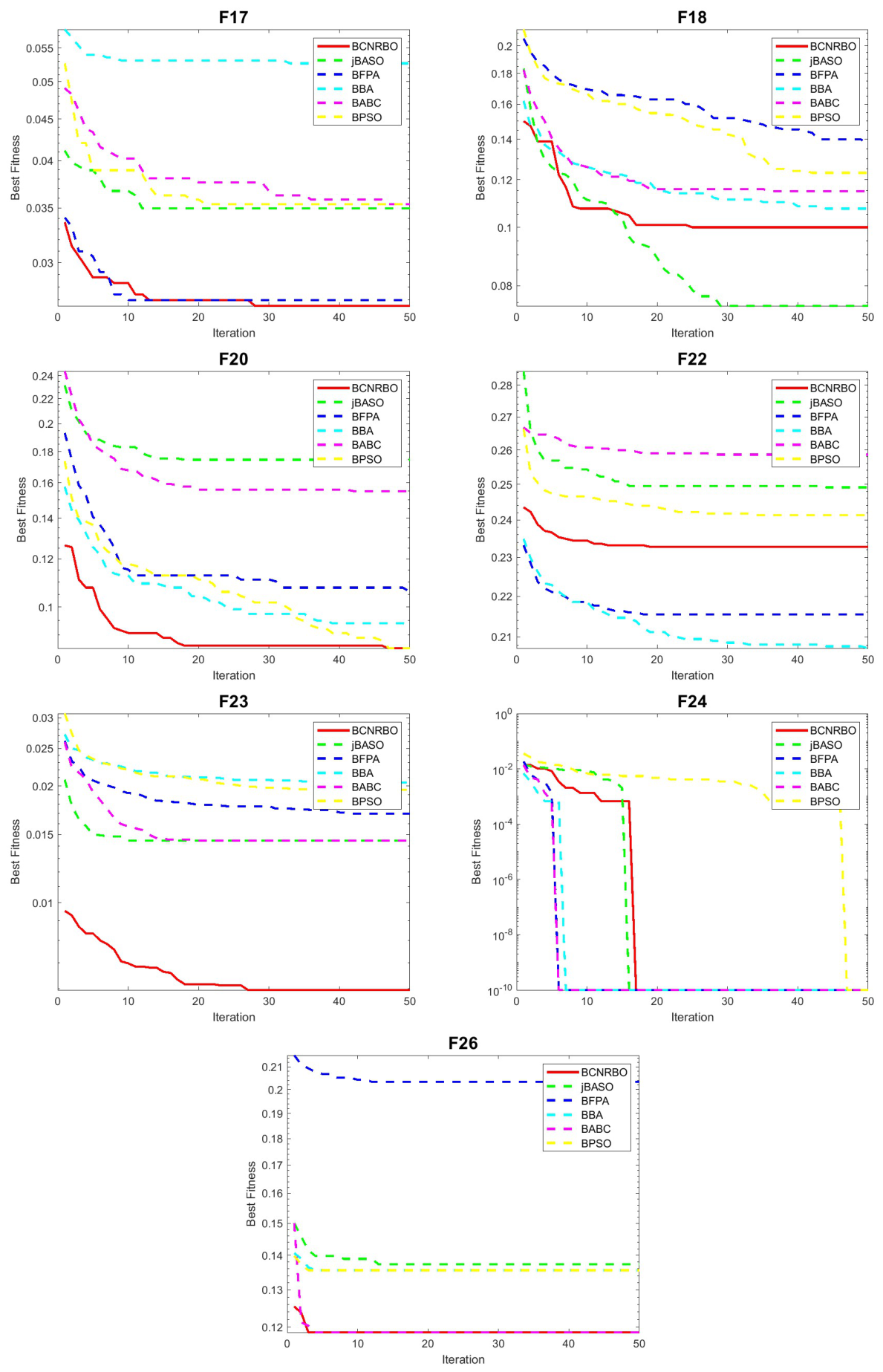
5.2. Evaluation of BCNRBO with DT Classifier
5.2.1. The Statistical Test for DT Classifier
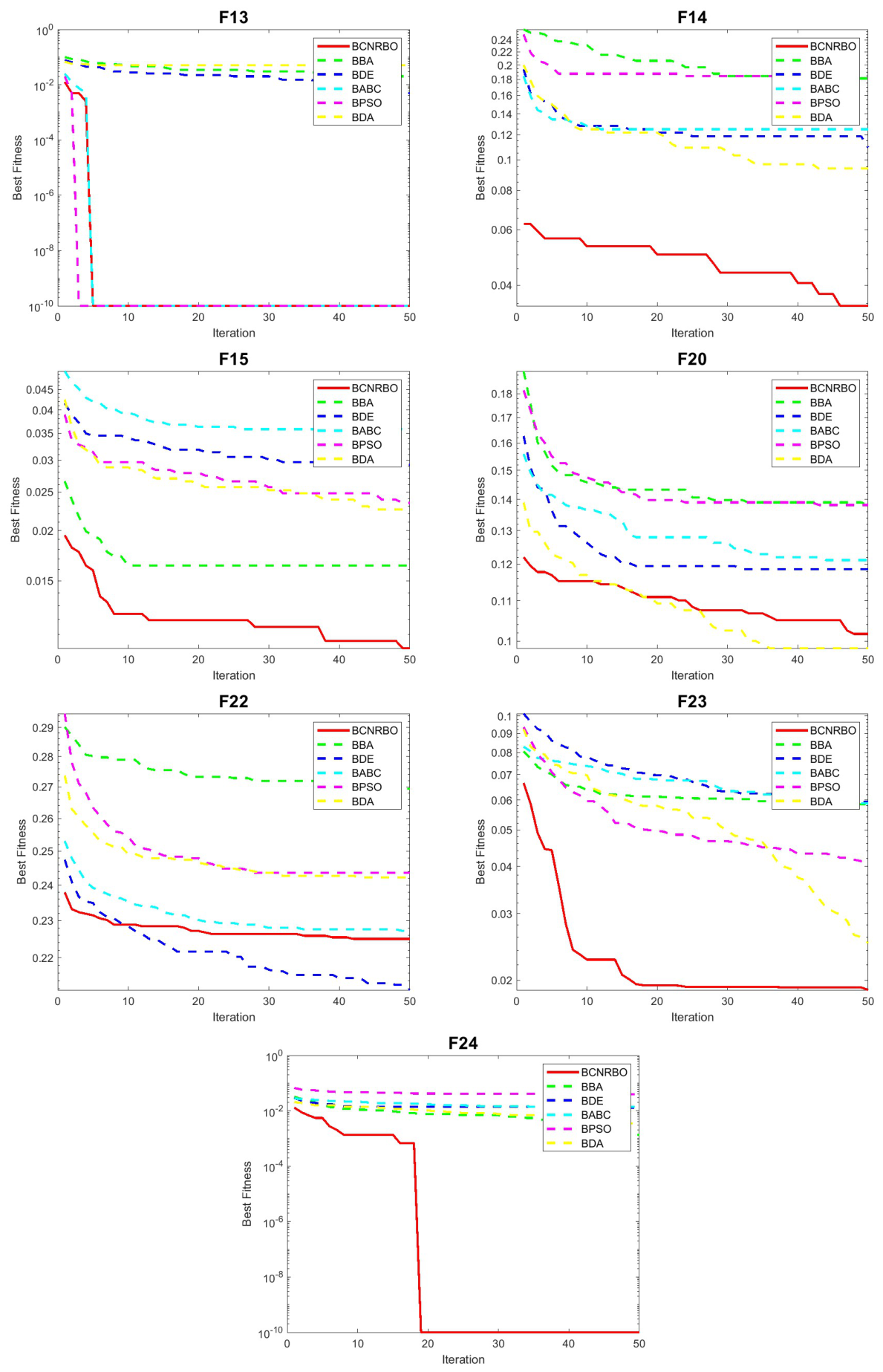
5.2.2. Convergence Graphic for Different Dataset Using DT Classifier
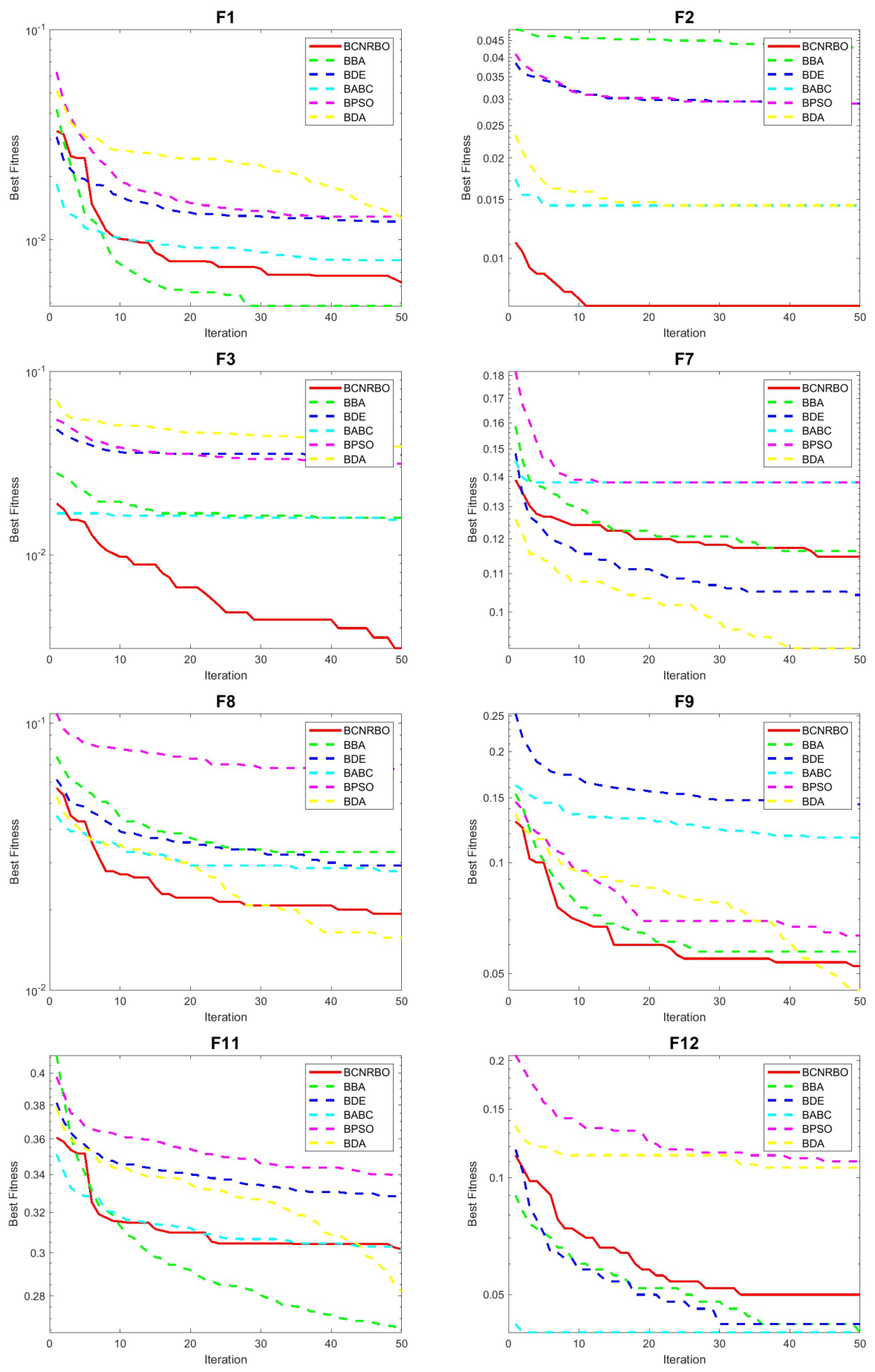
5.3. Evaluation of BCNRBO with NB Classifier
5.3.1. The Statistical Test for NB Classifier
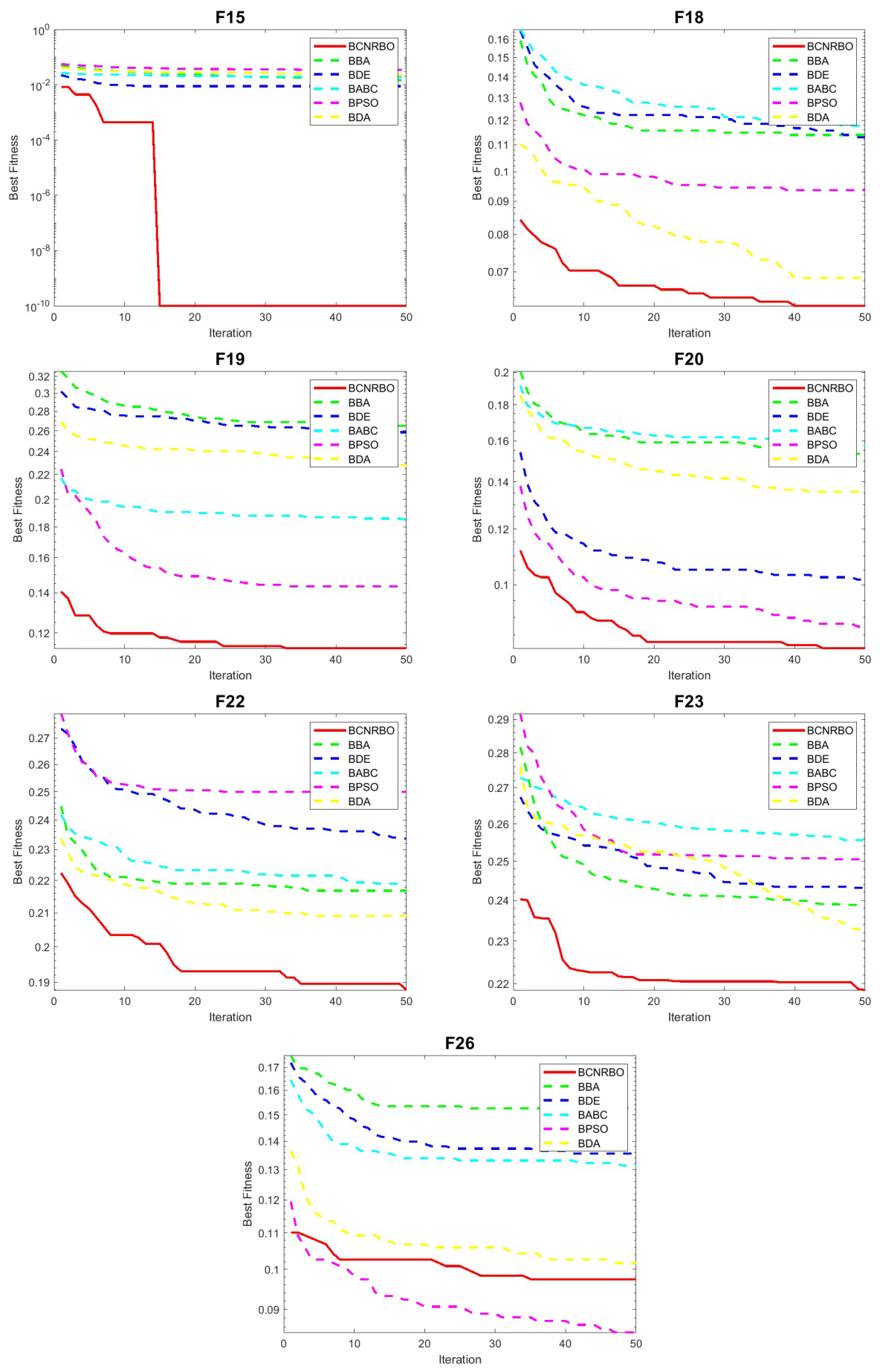
5.3.2. Convergence Graphic for Different Dataset Using NB Classifier
6. Conclusions and Future Work
- The introduction of a binary chaotic NRBO variant capable of effectively handling discrete feature selection problems.
- The incorporation of a dynamic potential mechanism to enhance population diversity and improve exploration–exploitation balance.
- The development of a transfer function mechanism to improve the conversion of continuous solutions into binary space.
- Demonstration of the superior performance of BCNRBO across multiple datasets, classifiers, and benchmark algorithms.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BCNRBO | Binary Chaos-Enhanced Newton-Raphson-Based Optimizer |
| KNN | K-nearest neighbor classifiers |
| DT | decision tree classifiers |
| NB | Naive Bayes classifiers |
| FS | feature selection |
| FSAs | feature selection algorithms |
| WOA | Whale Optimization Algorithm |
| ACO | Ant Colony Optimization |
| BA | Bat Algorithm |
| ABC | Artificial Bee Colony |
| PSO | Particle Swarm Optimization |
| BBO | Biogeography Based Optimization |
| GA | Genetic Algorithm |
| HSA | Harmony Search Algorithm |
| FP | Flower Pollination algorithm |
| GOA | Grasshopper Optimization Algorithm |
| BDF | Binary Dragonfly algorithm |
| BCCSA | Binary Chaotic Crow Search algorithm |
| NRBO | Newton Raphson Based optimizer |
| BAOA | Binary Arithmetic Optimization Algorithm |
| BBA | Binary Bat Algorithm |
| BFPA | Binary Flower Pollination Algorithm |
| BPSO | Binary Particle Swarm Optimization |
| jBASO | Binary Atom Search Optimization |
| BDA | Binary Dwarf Mongoose |
Appendix A. Supplementary Results
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.050078 | 0.076682 | 0.065728 | 0.068858 | 0.050078 | 0.064163 | 0.062598 | 0.051643 | 0.050078 | 0.048513 |
| D 2 | 0.014388 | 0.035971 | 0.021583 | 0.007194 | 0.021583 | 0.021583 | 0.035971 | 0.007194 | 0.007194 | 0.014388 |
| D 3 | 0.053097 | 0.035398 | 0.061947 | 0.061947 | 0.070796 | 0.053097 | 0.061947 | 0.044248 | 0.044248 | 0.017699 |
| D 4 | 0.087719 | 0.096491 | 0.087719 | 0.12281 | 0.096491 | 0.070175 | 0.10526 | 0.087719 | 0.087719 | 0.070175 |
| D 5 | 0 | 0 | 0.33333 | 0.16667 | 0 | 0 | 0 | 0.16667 | 0 | 0 |
| D 6 | 0 | 0.035714 | 0.035714 | 0.035714 | 0.035714 | 0 | 0 | 0 | 0 | 0.071429 |
| D 7 | 0.15517 | 0.068966 | 0.15517 | 0.086207 | 0.12069 | 0.22414 | 0.068966 | 0.17241 | 0.15517 | 0.13793 |
| D 8 | 0.085714 | 0.042857 | 0.11429 | 0.11429 | 0.14286 | 0.17143 | 0.085714 | 0.1 | 0.071429 | 0.057143 |
| D 9 | 0.02439 | 0.097561 | 0.14634 | 0.12195 | 0.04878 | 0.17073 | 0.19512 | 0.12195 | 0.04878 | 0.073171 |
| D 10 | 0.13793 | 0.13793 | 0.13793 | 0.17241 | 0.13793 | 0.2069 | 0.068966 | 0.10345 | 0.034483 | 0.13793 |
| D 11 | 0.33884 | 0.39669 | 0.43802 | 0.3719 | 0.42975 | 0.46281 | 0.39669 | 0.36364 | 0.40496 | 0.44628 |
| D 12 | 0.32 | 0.28 | 0.28 | 0.32 | 0.44 | 0.32 | 0.48 | 0.2 | 0.48 | 0.24 |
| D 13 | 0.05 | 0.05 | 0.05 | 0.05 | 0 | 0.05 | 0 | 0 | 0 | 0.1 |
| D 14 | 0.25 | 0.125 | 0.125 | 0.1875 | 0.1875 | 0.125 | 0.125 | 0.25 | 0.1875 | 0.125 |
| D 15 | 0.026549 | 0.053097 | 0.088496 | 0.044248 | 0.053097 | 0.088496 | 0.053097 | 0.017699 | 0.053097 | 0.061947 |
| D 16 | 0.086957 | 0.17391 | 0.26087 | 0.13043 | 0 | 0.17391 | 0.086957 | 0.086957 | 0.17391 | 0.13043 |
| D 17 | 0.026549 | 0.053097 | 0.044248 | 0.044248 | 0.035398 | 0.053097 | 0.035398 | 0.053097 | 0.035398 | 0.035398 |
| D 18 | 0.11111 | 0.12963 | 0.074074 | 0.12963 | 0.2037 | 0.33333 | 0.12963 | 0.12963 | 0.12963 | 0.14815 |
| D 19 | 0.22642 | 0.18868 | 0.22642 | 0.24528 | 0.22642 | 0.32075 | 0.22642 | 0.18868 | 0.22642 | 0.15094 |
| D 20 | 0.10169 | 0.15254 | 0.20339 | 0.16949 | 0.15254 | 0.25424 | 0.13559 | 0.10169 | 0.18644 | 0.11864 |
| D 21 | 0.22222 | 0.23529 | 0.20915 | 0.23529 | 0.24837 | 0.26144 | 0.22222 | 0.24183 | 0.20915 | 0.20915 |
| D 22 | 0.23276 | 0.22414 | 0.27586 | 0.26724 | 0.25 | 0.26724 | 0.26724 | 0.21552 | 0.25862 | 0.24138 |
| D 23 | 0.005957 | 0.026383 | 0.014468 | 0.025532 | 0.022128 | 0.02383 | 0.019574 | 0.020426 | 0.014468 | 0.019574 |
| D 24 | 0 | 0 | 0 | 0 | 0 | 0.041096 | 0 | 0 | 0 | 0 |
| D 25 | 0.22517 | 0.21854 | 0.25166 | 0.25166 | 0.2649 | 0.23179 | 0.25166 | 0.25166 | 0.23841 | 0.23179 |
| D 26 | 0.11864 | 0.13559 | 0.15254 | 0.18644 | 0.20339 | 0.15254 | 0.15254 | 0.13559 | 0.11864 | 0.13559 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.006985 | 0.007478 | 0.008375 | 0.005409 | 0.007698 | 0.00105 | 0.005767 | 0.003255 | 0.003605 | 0.005619 |
| D 2 | 0.001609 | 0 | 0.003383 | 0 | 0.002953 | 0.003672 | 0 | 0 | 0 | 0.002953 |
| D 3 | 0.008614 | 0.001979 | 0.007651 | 0.003932 | 0.003958 | 0.005294 | 0.008322 | 0.002724 | 0.004331 | 0 |
| D 4 | 0 | 0.001962 | 0.010052 | 7.12E-17 | 0.003214 | 0.0027 | 4.27E-17 | 0 | 0 | 0 |
| D 5 | 0 | 0 | 0.097857 | 0.051299 | 0 | 0 | 0 | 5.7E-17 | 0 | 0 |
| D 6 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| D 7 | 0.006316 | 0.008106 | 0.012999 | 0.00766 | 7.12E-17 | 0.020534 | 0.0088 | 0.005307 | 0.008666 | 0.0088 |
| D 8 | 0.015001 | 0.00864 | 0.015701 | 0.006991 | 0.020454 | 0.010364 | 0.006717 | 0.011794 | 0.008794 | 0.010467 |
| D 9 | 0.012259 | 0.021344 | 0.020936 | 0.017472 | 0.01276 | 0.019823 | 0.021085 | 0.021085 | 0.010836 | 0.018175 |
| D 10 | 0.014151 | 0.016874 | 0.023995 | 0.017601 | 0.011188 | 0.035555 | 0.016212 | 0 | 0 | 0.01804 |
| D 11 | 0.008893 | 0.009843 | 0.018693 | 0.00592 | 0.014439 | 0.014003 | 0.007988 | 0.007806 | 0.009843 | 0.014314 |
| D 12 | 0.014654 | 0.02393 | 0.045837 | 0.018806 | 0.1126 | 0.016416 | 0.035303 | 8.54E-17 | 0.026833 | 0.014654 |
| D 13 | 0 | 0.025521 | 0 | 0.018317 | 0 | 0.025521 | 0 | 0 | 0 | 0.01539 |
| D 14 | 0.019237 | 0 | 0 | 0.031414 | 0.057711 | 0.025649 | 0 | 0.036696 | 0 | 0 |
| D 15 | 0 | 0.003932 | 0.018931 | 0 | 0.016562 | 0.012671 | 0.005196 | 0 | 0.003932 | 0.008359 |
| D 16 | 4.27E-17 | 0.022304 | 0.027768 | 5.7E-17 | 0 | 0.051396 | 4.27E-17 | 4.27E-17 | 5.7E-17 | 5.7E-17 |
| D 17 | 0 | 0.005196 | 0.012992 | 0.002724 | 0.001979 | 0.004161 | 0.003932 | 0.001979 | 0 | 0 |
| D 18 | 0.009308 | 0.014839 | 2.85E-17 | 0.011827 | 0.032019 | 0.045944 | 5.7E-17 | 0.014218 | 0.0076 | 0.017283 |
| D 19 | 0.017206 | 0.021117 | 0.023129 | 0.019622 | 0.025681 | 0.031154 | 0.017821 | 0.01608 | 0.011288 | 0.018237 |
| D 20 | 0.00379 | 0.016126 | 0.014653 | 0.005217 | 0.020502 | 0.027219 | 0.015035 | 0.010288 | 0.008294 | 0.018628 |
| D 21 | 5.7E-17 | 0 | 0.005807 | 0.001462 | 0.011473 | 0.019258 | 5.7E-17 | 0 | 5.7E-17 | 5.7E-17 |
| D 22 | 2.85E-17 | 0.001928 | 0.010046 | 0.00383 | 0.017013 | 0.005867 | 0.009475 | 0.001928 | 1.14E-16 | 1.42E-16 |
| D 23 | 0 | 0.001598 | 0 | 0.000312 | 0.004778 | 0.001802 | 0.000478 | 0 | 0 | 0 |
| D 24 | 0 | 0 | 0 | 0 | 0 | 0.011675 | 0 | 0 | 0 | 0 |
| D 25 | 1.42E-16 | 0.014509 | 0.021322 | 0 | 0.027775 | 0.003114 | 0.005382 | 0.002426 | 0.015683 | 0.027756 |
| D 26 | 7.12E-17 | 5.7E-17 | 0.005217 | 0 | 5.7E-17 | 0.021227 | 0.007969 | 5.7E-17 | 7.12E-17 | 5.7E-17 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 18.35 | 16.55 | 18.2 | 23.45 | 19.8 | 35.35 | 22.05 | 23.7 | 20.45 | 21.95 |
| D 2 | 4.95 | 5.9 | 5.85 | 6.6 | 6 | 5.4 | 6.2 | 5.45 | 3.65 | 5.05 |
| D 3 | 6.3 | 12.9 | 10.8 | 18.9 | 14.1 | 20.9 | 16.45 | 17.9 | 13 | 13 |
| D 4 | 4.05 | 5.6 | 5.4 | 5.65 | 4.75 | 7.55 | 6 | 5.6 | 5.75 | 6 |
| D 5 | 8.35 | 27.65 | 20.8 | 34.8 | 26.85 | 27.85 | 35 | 28.4 | 27.25 | 27.95 |
| D 6 | 2 | 2.75 | 5.3 | 3.35 | 2.75 | 4.85 | 3.95 | 2.45 | 3 | 3.35 |
| D 7 | 4.9 | 5.05 | 5.7 | 6.2 | 6.45 | 5.8 | 6.45 | 6.25 | 5.45 | 5.05 |
| D 8 | 9.5 | 13.85 | 4.35 | 20.6 | 13.05 | 14.8 | 19.65 | 15.8 | 13.7 | 13.2 |
| D 9 | 23.2 | 24.75 | 12.95 | 37 | 25.65 | 29.35 | 37.45 | 34.75 | 29.55 | 27.6 |
| D 10 | 5 | 8.15 | 6.55 | 12.35 | 8.8 | 9.5 | 10.3 | 11.9 | 9.8 | 9.25 |
| D 11 | 43.2 | 38.75 | 16.9 | 60.6 | 47.3 | 49.65 | 63.35 | 64.6 | 46.3 | 45.25 |
| D 12 | 116.2 | 141.1 | 59.75 | 197.25 | 144.6 | 164.6 | 198.25 | 189.25 | 151.05 | 132.65 |
| D 13 | 4.75 | 7.95 | 9.4 | 10.15 | 8.45 | 9.95 | 10.35 | 9.45 | 9.05 | 10.1 |
| D 14 | 4.2 | 9.25 | 8.5 | 9.75 | 7.9 | 8.35 | 11.15 | 9.5 | 9.6 | 10.15 |
| D 15 | 5.85 | 13.15 | 8.85 | 18.85 | 14.05 | 16.85 | 17.6 | 17.5 | 13.2 | 13.65 |
| D 16 | 4 | 5.65 | 3.55 | 4.6 | 5.1 | 4.05 | 5.5 | 7.55 | 5.25 | 5.6 |
| D 17 | 3.95 | 13.9 | 13.55 | 20.5 | 14.25 | 20.85 | 18.5 | 15.55 | 15.15 | 15.25 |
| D 18 | 5.45 | 4.8 | 7.95 | 7.1 | 5.8 | 5.45 | 7.75 | 7.25 | 5 | 4.7 |
| D 19 | 9.5 | 6.5 | 12.05 | 13.75 | 9.1 | 10.6 | 12.2 | 12.3 | 11.55 | 10.65 |
| D 20 | 3.4 | 6.6 | 4.45 | 7.5 | 6.3 | 6.1 | 6.45 | 6.25 | 4.85 | 5.45 |
| D 21 | 4 | 4.9 | 4.4 | 3.05 | 5.35 | 4.75 | 5.1 | 2.55 | 5.15 | 5.45 |
| D 22 | 3 | 4.2 | 2.7 | 4.4 | 5.55 | 5.55 | 3.1 | 4.05 | 5.35 | 3.95 |
| D 23 | 3.05 | 6.3 | 8.8 | 11.15 | 6.5 | 8.95 | 10.2 | 9 | 5.75 | 6.85 |
| D 24 | 11.5 | 17.05 | 16.8 | 22.25 | 19.6 | 18.25 | 21.4 | 21.6 | 18.5 | 19.5 |
| D 25 | 302.15 | 310.1 | 111.9 | 491.35 | 374.55 | 372.45 | 482.1 | 398.55 | 379.7 | 368.35 |
| D 26 | 2 | 5.65 | 4.3 | 5.5 | 6.45 | 5.85 | 6.35 | 8 | 6 | 5.35 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.010955 | 0.050078 | 0.015649 | 0.010955 | 0.007825 | 0.017214 | 0.014085 | 0.00939 | 0.017214 | 0.018779 |
| D 2 | 0.007194 | 0.014388 | 0.043165 | 0.028777 | 0.05036 | 0.043165 | 0.035971 | 0.014388 | 0.035971 | 0.014388 |
| D 3 | 0.00885 | 0.061947 | 0.026549 | 0.026549 | 0.017699 | 0.044248 | 0.035398 | 0.017699 | 0.044248 | 0.044248 |
| D 4 | 0.078947 | 0.078947 | 0.087719 | 0.04386 | 0.087719 | 0.078947 | 0.061404 | 0.026316 | 0.070175 | 0.070175 |
| D 5 | 0.333333 | 0.166667 | 0.166667 | 0.166667 | 0.166667 | 0.166667 | 0 | 0.166667 | 0 | 0.166667 |
| D 6 | 0 | 0.107143 | 0.071429 | 0.035714 | 0.035714 | 0.035714 | 0.035714 | 0.035714 | 0.071429 | 0.035714 |
| D 7 | 0.12069 | 0.172414 | 0.086207 | 0.155172 | 0.137931 | 0.137931 | 0.12069 | 0.137931 | 0.137931 | 0.103448 |
| D 8 | 0.028571 | 0.057143 | 0.057143 | 0.042857 | 0.057143 | 0.071429 | 0.042857 | 0.028571 | 0.085714 | 0.028571 |
| D 9 | 0.073171 | 0.097561 | 0.146341 | 0.146341 | 0.097561 | 0.243902 | 0.170732 | 0.146341 | 0.097561 | 0.097561 |
| D 10 | 0.137931 | 0.137931 | 0.103448 | 0.068966 | 0.068966 | 0.206897 | 0.103448 | 0.034483 | 0.137931 | 0.137931 |
| D 11 | 0.322314 | 0.371901 | 0.355372 | 0.338843 | 0.297521 | 0.396694 | 0.347107 | 0.322314 | 0.355372 | 0.31405 |
| D 12 | 0.12 | 0.08 | 0.12 | 0.12 | 0.04 | 0.24 | 0.08 | 0.04 | 0.16 | 0.12 |
| D 13 | 0 | 0 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0 | 0 | 0.05 |
| D 14 | 0.0625 | 0.1875 | 0.25 | 0.125 | 0.25 | 0.25 | 0.125 | 0.125 | 0.1875 | 0.125 |
| D 15 | 0.017699 | 0.053097 | 0.035398 | 0.035398 | 0.017699 | 0.035398 | 0.035398 | 0.044248 | 0.035398 | 0.026549 |
| D 16 | 0.173913 | 0.086957 | 0.086957 | 0.086957 | 0.217391 | 0.347826 | 0.130435 | 0.217391 | 0 | 0.173913 |
| D 17 | 0.026549 | 0.026549 | 0.044248 | 0.026549 | 0.035398 | 0.070796 | 0.035398 | 0.017699 | 0.00885 | 0 |
| D 18 | 0.185185 | 0.166667 | 0.12963 | 0.185185 | 0.092593 | 0.185185 | 0.092593 | 0.148148 | 0.055556 | 0.092593 |
| D 19 | 0.245283 | 0.188679 | 0.226415 | 0.226415 | 0.188679 | 0.283019 | 0.188679 | 0.169811 | 0.226415 | 0.207547 |
| D 20 | 0.118644 | 0.135593 | 0.135593 | 0.135593 | 0.152542 | 0.186441 | 0.118644 | 0.135593 | 0.152542 | 0.118644 |
| D 21 | 0.281046 | 0.281046 | 0.235294 | 0.235294 | 0.261438 | 0.261438 | 0.24183 | 0.215686 | 0.248366 | 0.248366 |
| D 22 | 0.232759 | 0.241379 | 0.267241 | 0.25 | 0.301724 | 0.310345 | 0.232759 | 0.232759 | 0.25 | 0.25 |
| D 23 | 0.035745 | 0.084255 | 0.085106 | 0.058723 | 0.075745 | 0.098723 | 0.07234 | 0.065532 | 0.051064 | 0.051064 |
| D 24 | 0 | 0.013699 | 0.027397 | 0.027397 | 0.013699 | 0.041096 | 0.013699 | 0.013699 | 0.041096 | 0.013699 |
| D 25 | 0.086093 | 0.119205 | 0.125828 | 0.10596 | 0.086093 | 0.198675 | 0.13245 | 0.152318 | 0.086093 | 0.099338 |
| D 26 | 0.118644 | 0.152542 | 0.118644 | 0.118644 | 0.135593 | 0.186441 | 0.118644 | 0.067797 | 0.067797 | 0.118644 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.002154 | 0.007421 | 0.001788 | 0.001984 | 0.0007 | 0 | 0.00194 | 0.001124 | 0.00232 | 0.001893 |
| D 2 | 0 | 0.001609 | 0.007041 | 0 | 0.005462 | 0.005006 | 0.001609 | 0 | 0.001609 | 0 |
| D 3 | 0.004331 | 0.006594 | 0.004331 | 0.006074 | 0.003632 | 0.006734 | 0.001979 | 0.003932 | 0.006718 | 0.006023 |
| D 4 | 0 | 0.005324 | 0.007129 | 0 | 0.001961 | 0.013311 | 0 | 0 | 0.003923 | 0 |
| D 5 | 0.068399 | 0.061058 | 0.061058 | 5.7E-17 | 0.051299 | 0.061058 | 0 | 0.061058 | 0 | 5.7E-17 |
| D 6 | 0 | 4.27E-17 | 0.017951 | 0 | 0 | 0.007986 | 0 | 0 | 0 | 0 |
| D 7 | 0.008437 | 0.007911 | 4.27E-17 | 0.008437 | 0.014013 | 0.011734 | 0.003855 | 5.7E-17 | 5.7E-17 | 0.008106 |
| D 8 | 0.009583 | 0.010234 | 0.013027 | 0.00864 | 0.008161 | 0.010968 | 0.003194 | 0.003194 | 0.009385 | 0.004397 |
| D 9 | 0.014321 | 0.018175 | 0.029048 | 0.015014 | 0.018175 | 0.027298 | 0.010908 | 0.018726 | 0.024262 | 0.028832 |
| D 10 | 0.016212 | 0.015421 | 0 | 0.010614 | 0.021227 | 0.026237 | 0 | 0 | 0.025998 | 0.007711 |
| D 11 | 0.013272 | 0.013567 | 0.012655 | 0.009888 | 0.020578 | 0.006823 | 0.009988 | 0.011533 | 0.010548 | 0.01584 |
| D 12 | 0.025547 | 0.01777 | 0.02285 | 0.008944 | 0 | 0.030435 | 0.008944 | 0 | 0.036419 | 0.019574 |
| D 13 | 0 | 0 | 0.02052 | 0.01118 | 0.025131 | 0.022213 | 0.01539 | 0 | 0 | 0 |
| D 14 | 0.031901 | 0.022897 | 0.038474 | 0.030585 | 0.030585 | 0.047986 | 0.034382 | 0 | 0.019237 | 0.032062 |
| D 15 | 0.003242 | 0.005294 | 0.006074 | 0.006974 | 0.004331 | 0.009894 | 0.004161 | 0.001979 | 0.006594 | 0.004517 |
| D 16 | 5.7E-17 | 4.27E-17 | 4.27E-17 | 4.27E-17 | 0.019316 | 0.046518 | 5.7E-17 | 0.015928 | 0 | 5.7E-17 |
| D 17 | 0.005814 | 0.005294 | 0.006959 | 0.003632 | 0.008566 | 0.006718 | 0.004448 | 0.001979 | 0.001979 | 0 |
| D 18 | 0.013799 | 0.018904 | 0.008707 | 0.008282 | 0 | 0.013799 | 0 | 0.0095 | 0 | 0.009452 |
| D 19 | 0.010778 | 0.014838 | 0.018967 | 0.014225 | 0.013516 | 0.012841 | 0.011288 | 0.016196 | 0.017821 | 0.016051 |
| D 20 | 0.007777 | 0.014445 | 0.009525 | 0.012142 | 0.006209 | 0.017798 | 7.12E-17 | 0.006209 | 0.006209 | 0.008867 |
| D 21 | 5.7E-17 | 0.004294 | 0.003336 | 0 | 0.005846 | 0.01112 | 0 | 5.7E-17 | 0.001461 | 0.001461 |
| D 22 | 0.002653 | 0.005917 | 0.008518 | 0 | 0.012151 | 0.018736 | 0.008609 | 0.004219 | 0.00383 | 0.002653 |
| D 23 | 0.004463 | 0.012468 | 0.023872 | 0.004434 | 0.023522 | 0.009328 | 0.005743 | 0.006664 | 0.006399 | 0.010421 |
| D 24 | 0 | 0 | 0.006441 | 0.005018 | 0.003063 | 0.008777 | 0.003063 | 0 | 0.004216 | 0.006086 |
| D 25 | 0.009433 | 0.007474 | 0.010802 | 0.008754 | 0.011625 | 0.01615 | 0.007126 | 0.005028 | 0.00603 | 0.010438 |
| D 26 | 7.12E-17 | 0 | 0.006209 | 0.01263 | 0.005217 | 0.022988 | 7.12E-17 | 0.00753 | 0 | 7.12E-17 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 22.2 | 19.95 | 28.65 | 28.3 | 27.55 | 36 | 27.9 | 28.8 | 25.25 | 26.1 |
| D 2 | 4 | 5.45 | 3.35 | 4.05 | 4.4 | 4.6 | 4.8 | 5.15 | 5.25 | 4.6 |
| D 3 | 7.85 | 11.6 | 10.4 | 16.75 | 14.75 | 14.6 | 18.6 | 17.1 | 13.7 | 11.4 |
| D 4 | 5 | 4.9 | 5.6 | 5.55 | 5 | 5.3 | 6 | 6.65 | 6 | 6.15 |
| D 5 | 12.5 | 27.7 | 26.1 | 37.45 | 28.7 | 27.9 | 34 | 26.75 | 27.15 | 26.45 |
| D 6 | 2 | 3.7 | 3.8 | 4 | 4.25 | 4.05 | 5 | 2.7 | 2.75 | 3.65 |
| D 7 | 6.1 | 5.8 | 6.15 | 8.4 | 5.15 | 5.45 | 7.7 | 9.7 | 5.45 | 7.15 |
| D 8 | 11.25 | 13.9 | 14.95 | 20.85 | 16.4 | 17.85 | 22.2 | 19.7 | 14.4 | 18 |
| D 9 | 20 | 27 | 23.8 | 39.05 | 28.85 | 30.05 | 39.6 | 36.65 | 30.3 | 26.45 |
| D 10 | 6 | 7.45 | 9.5 | 8.4 | 8.65 | 9.15 | 11.65 | 11.95 | 8.15 | 8.45 |
| D 11 | 45.9 | 40.7 | 26.2 | 62.65 | 45.75 | 74.85 | 64.25 | 62.2 | 48.55 | 50.85 |
| D 12 | 124.65 | 139.25 | 105 | 196.7 | 152.25 | 151.95 | 198.8 | 173.95 | 150.6 | 155.05 |
| D 13 | 3.45 | 7.5 | 8.5 | 9.5 | 8.05 | 7.95 | 9.85 | 10.35 | 8.2 | 8.45 |
| D 14 | 4.15 | 7.05 | 5.25 | 9.35 | 6.9 | 9.45 | 11.15 | 10.6 | 8.6 | 7.6 |
| D 15 | 7.9 | 11.6 | 12.25 | 17.1 | 14.5 | 15.15 | 17.7 | 16.9 | 12.55 | 13.45 |
| D 16 | 3 | 6.75 | 4.7 | 5.8 | 4.25 | 5.5 | 4.45 | 3.05 | 5.6 | 4.75 |
| D 17 | 8.15 | 12.9 | 7.75 | 18.05 | 12.4 | 14.8 | 18 | 16.4 | 15.65 | 14.6 |
| D 18 | 5.35 | 5.85 | 6.3 | 6.45 | 6.7 | 9.45 | 8.1 | 5.5 | 5.6 | 5.6 |
| D 19 | 5.15 | 10.4 | 12.25 | 10.4 | 10.5 | 11.7 | 14.2 | 11.25 | 7.85 | 10.05 |
| D 20 | 5.4 | 5.65 | 6.55 | 7.05 | 5.5 | 9 | 7.5 | 7.05 | 6.45 | 5.5 |
| D 21 | 3 | 3.45 | 5.45 | 4 | 6.7 | 6.35 | 6 | 5 | 3.15 | 5.05 |
| D 22 | 4.15 | 4.1 | 4.05 | 6.55 | 4.4 | 4.85 | 4.55 | 3.7 | 4.75 | 4.1 |
| D 23 | 4.5 | 7.5 | 3.95 | 10.7 | 8.9 | 11.3 | 6 | 4 | 6.8 | 4.4 |
| D 24 | 9.25 | 19.5 | 16.65 | 19.9 | 16.85 | 21.75 | 17 | 18 | 17.5 | 18.3 |
| D 25 | 353.5 | 311.7 | 278.05 | 487 | 377.75 | 375.85 | 472 | 402 | 380.3 | 380.6 |
| D 26 | 4.15 | 5.8 | 5.05 | 5.8 | 4.85 | 6.2 | 5 | 2 | 6.85 | 7.1 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.172144 | 0.211268 | 0.175274 | 0.186228 | 0.165884 | 0.29108 | 0.179969 | 0.158059 | 0.175274 | 0.162754 |
| D 2 | 0.007194 | 0.014388 | 0.014388 | 0.021583 | 0.014388 | 0.043165 | 0.035971 | 0.021583 | 0.014388 | 0.05036 |
| D 3 | 0.017699 | 0.044248 | 0.017699 | 0.026549 | 0.035398 | 0.035398 | 0.044248 | 0.026549 | 0.00885 | 0.026549 |
| D 4 | 0.035088 | 0.035088 | 0.052632 | 0.070175 | 0.052632 | 0.070175 | 0.035088 | 0.061404 | 0.061404 | 0.04386 |
| D 5 | 0.166667 | 0 | 0.166667 | 0 | 0 | 0.166667 | 0 | 0.166667 | 0 | 0.166667 |
| D 6 | 0 | 0 | 0.035714 | 0 | 0 | 0.035714 | 0.035714 | 0 | 0 | 0 |
| D 7 | 0.086207 | 0.155172 | 0.189655 | 0.155172 | 0.103448 | 0.12069 | 0.172414 | 0.137931 | 0.12069 | 0.155172 |
| D 8 | 0.057143 | 0.057143 | 0.042857 | 0.057143 | 0.028571 | 0.014286 | 0.014286 | 0.042857 | 0.028571 | 0.028571 |
| D 9 | 0.170732 | 0.195122 | 0.121951 | 0.121951 | 0.04878 | 0.170732 | 0.195122 | 0.195122 | 0.04878 | 0.219512 |
| D 10 | 0.103448 | 0.137931 | 0.137931 | 0.068966 | 0.103448 | 0.241379 | 0 | 0.103448 | 0.137931 | 0.172414 |
| D 11 | 0.396694 | 0.504132 | 0.487603 | 0.413223 | 0.446281 | 0.421488 | 0.446281 | 0.495868 | 0.454545 | 0.471074 |
| D 12 | 0.16 | 0.16 | 0.2 | 0.24 | 0.2 | 0.28 | 0.12 | 0.2 | 0.16 | 0.12 |
| D 13 | 0.1 | 0.15 | 0.15 | 0.05 | 0.1 | 0.15 | 0.05 | 0.05 | 0.05 | 0.1 |
| D 14 | 0.0625 | 0.1875 | 0.1875 | 0.125 | 0 | 0.3125 | 0.125 | 0.0625 | 0.1875 | 0.1875 |
| D 15 | 0 | 0.044248 | 0.035398 | 0.044248 | 0.026549 | 0.017699 | 0.00885 | 0.026549 | 0.044248 | 0.026549 |
| D 16 | 0.173913 | 0.217391 | 0.217391 | 0.086957 | 0.173913 | 0.130435 | 0.217391 | 0.086957 | 0.086957 | 0.217391 |
| D 17 | 0.035398 | 0.017699 | 0.035398 | 0.044248 | 0.026549 | 0.061947 | 0.00885 | 0.017699 | 0.00885 | 0.00885 |
| D 18 | 0.074074 | 0.111111 | 0.092593 | 0.12963 | 0.148148 | 0.185185 | 0.12963 | 0.12963 | 0.111111 | 0.092593 |
| D 19 | 0.113208 | 0.226415 | 0.132075 | 0.188679 | 0.301887 | 0.301887 | 0.264151 | 0.188679 | 0.169811 | 0.245283 |
| D 20 | 0.084746 | 0.186441 | 0.135593 | 0.067797 | 0.186441 | 0.169492 | 0.101695 | 0.169492 | 0.118644 | 0.135593 |
| D 21 | 0.202614 | 0.24183 | 0.235294 | 0.196078 | 0.196078 | 0.261438 | 0.156863 | 0.202614 | 0.202614 | 0.176471 |
| D 22 | 0.215517 | 0.25 | 0.258621 | 0.284483 | 0.25 | 0.293103 | 0.25 | 0.224138 | 0.25 | 0.215517 |
| D 23 | 0.224681 | 0.251915 | 0.250213 | 0.261277 | 0.248511 | 0.258723 | 0.250213 | 0.26383 | 0.257872 | 0.241702 |
| D 24 | 0.054795 | 0.068493 | 0.013699 | 0.013699 | 0.013699 | 0.123288 | 0.013699 | 0.027397 | 0.013699 | 0.013699 |
| D 25 | 0.258278 | 0.225166 | 0.15894 | 0.211921 | 0.145695 | 0.231788 | 0.205298 | 0.205298 | 0.192053 | 0.231788 |
| D 26 | 0.101695 | 0.186441 | 0.186441 | 0.152542 | 0.152542 | 0.186441 | 0.135593 | 0.135593 | 0.084746 | 0.101695 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.004694 | 0.012759 | 0.005573 | 0.004725 | 0.007239 | 0.025584 | 0.005869 | 0.005105 | 0.00566 | 0.00771 |
| D 2 | 0 | 0.003672 | 0.003672 | 0 | 0 | 0.004429 | 0 | 0 | 0 | 0.001609 |
| D 3 | 0.004161 | 0.009523 | 0.006718 | 0.003932 | 0.004868 | 0.004517 | 0.003632 | 0.005196 | 0 | 0.00567 |
| D 4 | 0 | 0.0036 | 0.008478 | 0 | 0 | 0.008285 | 0 | 0 | 0.001961 | 0.001961 |
| D 5 | 0.08507 | 0 | 0.068399 | 0 | 0 | 0.037268 | 0 | 5.7E-17 | 0 | 5.7E-17 |
| D 6 | 0 | 0 | 0.013084 | 0 | 0 | 0.007986 | 0 | 0 | 0 | 0 |
| D 7 | 0.003855 | 8.54E-17 | 0.015294 | 0.015698 | 0.003855 | 0.008666 | 0.00766 | 0.012999 | 7.12E-17 | 0.015294 |
| D 8 | 0.011234 | 0.006389 | 0.008546 | 0.004397 | 0.007859 | 0.006347 | 0 | 0.006389 | 0.00718 | 0.007292 |
| D 9 | 0.025611 | 0.016972 | 0.023206 | 0.015577 | 0.017472 | 0.020019 | 0.015577 | 0.016361 | 0.013418 | 0.033456 |
| D 10 | 0.017689 | 0.017332 | 0.020246 | 0.015319 | 0.023132 | 0.039554 | 0 | 0.007711 | 0.014151 | 0.020246 |
| D 11 | 0.006231 | 0.005672 | 0.009327 | 0.004218 | 0.005295 | 0.006823 | 5.7E-17 | 0.001848 | 0.003672 | 0.004998 |
| D 12 | 0.020417 | 0.024192 | 0.042252 | 0.016416 | 0.029019 | 0.024192 | 0.008944 | 0.008944 | 0.018353 | 0.020926 |
| D 13 | 0.01118 | 0.01539 | 0.036635 | 0 | 0.01118 | 0.01539 | 0 | 0 | 0 | 0.01118 |
| D 14 | 0.013975 | 0.031414 | 0.027766 | 0 | 0 | 0.038474 | 0 | 0.013975 | 0 | 0.044887 |
| D 15 | 0 | 0.004517 | 0.006356 | 0.005742 | 0.005936 | 0.006023 | 0 | 0.003242 | 0.00567 | 0.00454 |
| D 16 | 5.7E-17 | 0.017843 | 0.009722 | 0.009722 | 0.017843 | 0.019316 | 5.7E-17 | 4.27E-17 | 4.27E-17 | 0.009722 |
| D 17 | 0.007192 | 0.001979 | 0.004448 | 0.008172 | 0.00454 | 0.008554 | 0.003632 | 0 | 0.001979 | 0.004517 |
| D 18 | 0.009062 | 0.012423 | 0.0076 | 5.7E-17 | 0.009062 | 0.0152 | 0.0057 | 0.009062 | 0.004141 | 0.012166 |
| D 19 | 0 | 0.009483 | 0.006912 | 0.011411 | 0.01295 | 0.020555 | 0.008871 | 0.007743 | 0.014225 | 0.01295 |
| D 20 | 0.006956 | 0.008651 | 0.01263 | 0.005217 | 0.012867 | 0.013329 | 0 | 0.007969 | 0.008294 | 5.7E-17 |
| D 21 | 0.002012 | 0 | 0.005237 | 8.54E-17 | 0.004384 | 0.011276 | 5.7E-17 | 8.54E-17 | 8.54E-17 | 8.54E-17 |
| D 22 | 0.014151 | 0.004763 | 0.005062 | 0.008518 | 0.013089 | 0.014061 | 0.007861 | 0.004333 | 0 | 0.00383 |
| D 23 | 0.002864 | 0.004181 | 0.005335 | 0.00455 | 0.00447 | 0.007509 | 0.004923 | 0.00507 | 0.003888 | 0.004183 |
| D 24 | 0.012472 | 0.011988 | 0.006086 | 0.005018 | 0.006704 | 0.027924 | 0.004216 | 0.004216 | 0.004216 | 0.003063 |
| D 25 | 0.010186 | 0.006039 | 0.007717 | 0.006887 | 0.007865 | 0.008452 | 0.005644 | 0.00423 | 0.006115 | 0.00909 |
| D 26 | 0.00753 | 0.006209 | 0.00379 | 0.008294 | 0 | 0.018321 | 5.7E-17 | 0.00753 | 0 | 0 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 16.95 | 15.45 | 15.3 | 22.25 | 17.8 | 17.9 | 23.05 | 22 | 18.65 | 18.8 |
| D 2 | 3 | 5.8 | 4.65 | 8 | 5.8 | 4.9 | 5.3 | 4.2 | 5.85 | 4.35 |
| D 3 | 6.65 | 11.35 | 7.4 | 16.85 | 11.8 | 15.65 | 17.7 | 18 | 14 | 12.75 |
| D 4 | 5.05 | 6.2 | 5.55 | 6.25 | 5.8 | 4.95 | 6 | 5.35 | 5.85 | 6.05 |
| D 5 | 11.05 | 27.3 | 24.5 | 36.65 | 28.2 | 25.1 | 35.45 | 27.55 | 26.65 | 28.3 |
| D 6 | 2 | 4.45 | 3.35 | 3.55 | 3.95 | 3.5 | 5.3 | 2.7 | 2.5 | 3.9 |
| D 7 | 3.2 | 5.6 | 6.3 | 6.05 | 5.65 | 6.6 | 7.05 | 7.65 | 7.35 | 3.5 |
| D 8 | 10.6 | 12.8 | 13.45 | 20.95 | 13.1 | 24.1 | 20 | 18.05 | 15.1 | 16.35 |
| D 9 | 22.45 | 24 | 11.9 | 36.4 | 26.4 | 30.4 | 36.05 | 35.3 | 27.15 | 26.2 |
| D 10 | 6.7 | 8.25 | 9.6 | 11.5 | 8.4 | 8.05 | 11.95 | 10.9 | 8.2 | 8.7 |
| D 11 | 46.25 | 41.05 | 8.7 | 63.95 | 46.8 | 49.1 | 61.15 | 57.35 | 44.9 | 43.3 |
| D 12 | 122.25 | 130.05 | 44.4 | 191.8 | 148.8 | 152.9 | 196.75 | 196.15 | 154.65 | 139.95 |
| D 13 | 8.3 | 10 | 12 | 11.35 | 11.05 | 12.75 | 12.45 | 10.65 | 9.05 | 9.1 |
| D 14 | 3.3 | 6.35 | 9.4 | 9.9 | 11 | 9.35 | 10.2 | 10.75 | 9.05 | 6.45 |
| D 15 | 7.2 | 10.05 | 6.1 | 16.25 | 10.75 | 17.7 | 18.15 | 16.15 | 11.25 | 14.85 |
| D 16 | 2 | 3.05 | 4.15 | 4.05 | 4.9 | 4.5 | 6.05 | 6.1 | 5.45 | 3.85 |
| D 17 | 8.35 | 11.3 | 9.55 | 15.9 | 13.25 | 14.3 | 18.55 | 17.45 | 13.55 | 12.85 |
| D 18 | 6.9 | 6.6 | 6.65 | 9.25 | 8.95 | 7.6 | 7.8 | 6.5 | 6.05 | 6.85 |
| D 19 | 3.5 | 8.45 | 8.05 | 12.7 | 10.4 | 10.65 | 11.75 | 11.85 | 8.5 | 11.95 |
| D 20 | 5.7 | 7.45 | 8.15 | 8.6 | 6.5 | 8.45 | 7.75 | 7.9 | 6.65 | 6.7 |
| D 21 | 4.7 | 2.5 | 3.45 | 5.6 | 5 | 4.9 | 5 | 4.75 | 4 | 4.5 |
| D 22 | 3 | 3.6 | 4.75 | 2.75 | 5.2 | 4.6 | 3.4 | 3.15 | 3.35 | 5.7 |
| D 23 | 5.85 | 5.85 | 4.15 | 7.25 | 4.95 | 8.8 | 7.65 | 6.5 | 4.3 | 4.25 |
| D 24 | 15.05 | 17.5 | 18 | 22.95 | 18.8 | 18.6 | 23.85 | 22.7 | 19.05 | 21.7 |
| D 25 | 339.6 | 318.8 | 114.3 | 480.75 | 370.7 | 369.65 | 476.65 | 487 | 369.75 | 366.2 |
| D 26 | 4.85 | 6.3 | 2.85 | 4.85 | 4.75 | 4.65 | 3.8 | 4.55 | 3 | 6.6 |
References
- Cheng, X. A Comprehensive Study of Feature Selection Techniques in Machine Learning Models. Insights Comput. Signals Syst. 2024, 1, 65–78. [Google Scholar] [CrossRef]
- Lamsaf, A.; Carrilho, R.; Neves, J. C.; Proença, H. Causality, machine learning, and feature selection: a survey. Sensors 2025, 25, 2373. [Google Scholar] [CrossRef]
- Hosseinzadeh, M.; Ali, U.; Ali, S.; Abbaszadi, R.; Gharehchopogh, F. S.; Khoshvaght, P.; Porntaveetus, T.; Lansky, J. Improving phishing email detection performance through deep learning with adaptive optimization. Sci. Rep. 2025, 15, 36724. [Google Scholar] [CrossRef]
- Hosseinzadeh, M.; Tanveer, J.; Rahmani, A. M.; Baptista, M. L.; Abbaszadi, R.; Gharehchopogh, F. S.; Porntaveetus, T.; Lee, S. W. A Comprehensive Survey of Hybrid Whale Optimization Algorithm with Long-Short Term Memory: Applications, Improvements, and Future Perspective. Arch. Comput. Methods Eng. 2025, 1–42. [Google Scholar] [CrossRef]
- Khan, M. A. Special Issue “Algorithms for Feature Selection (2nd Edition)”. Algorithms 2025, 18, 16. [Google Scholar] [CrossRef]
- Sayed, G. I.; Hassanien, A. E.; Azar, A. T. Feature selection via a novel chaotic crow search algorithm. Neural Comput Appl. 2017, 171–188. [Google Scholar] [CrossRef]
- Albashish, D.; Hammouri, A. I.; Braik, M.; Atwan, J.; Sahran, S. Binary biogeography-based optimization based SVM-RFE for feature selection. Appl. Soft Comput. 2021, 101, 107026. [Google Scholar] [CrossRef]
- Tawhid, M. A.; Ibrahim, A. M. Hybrid Binary Particle Swarm Optimization and Flower Pollination Algorithm Based on Rough Set Approach for Feature Selection Problem. Nat.-Inspir. Comput. Data Min. Mach. Learn. 2020, 249–273. [Google Scholar]
- Sharifi, T.; Mirsalim, M.; Gharehchopogh, F. S.; Mirjalili, S. Cultural history optimization algorithm: a new human-inspired metaheuristic algorithm for engineering optimization problems. Neural Comput. Appl. 2025, 37, 21009–21068. [Google Scholar] [CrossRef]
- Ibrahim, A. M.; Tawhid, M. A.; Ward, R. A Binary Water Wave Optimization for Feature Selection. Int. J. Approx. Reason. 2020, 120, 74–91. [Google Scholar] [CrossRef]
- Zorarpacı, E.; Özel, S. A. A hybrid approach of differential evolution and artificial bee colony for feature selection. Expert Syst. With Appl. 2016, 62, 91–103. [Google Scholar] [CrossRef]
- Pashaei, E.; Pashaei, E. An efficient binary chimp optimization algorithm for feature selection in biomedical data classification. Neural Comput. Appl. 2022, 34, 6427–6451. [Google Scholar] [CrossRef]
- Dehghani, M.; Trojovská, E.; Zuščák, T. A new human-inspired metaheuristic algorithm for solving optimization problems based on mimicking sewing training. Sci. Rep. 2022, 12, 17387. [Google Scholar] [CrossRef]
- Haq, A. U.; Li, J.; Memon, M. H.; others. Heart Disease Prediction System Using Model of Machine Learning and Sequential Backward Selection Algorithm for Features Selection. 2019 IEEE 5th International Conference for Convergence in Technology (I2CT), 2019; pp. 1–4. [Google Scholar]
- Mohamad, M.; Selamat, A.; Krejcar, O.; Crespo, R. G.; Herrera-Viedma, E.; Fujita, H. Enhancing big data feature selection using a hybrid correlation-based feature selection. Electronics 2021, 10, 2984. [Google Scholar] [CrossRef]
- Witten, I. H.; Frank, E.; Hall, M. A.; Pal, C. J.; Foulds, J. Data Mining: Practical Machine Learning Tools and Techniques; Morgan Kaufmann, 2025. [Google Scholar]
- Duque, J.; Godinho, A.; Moreira, J.; Vasconcelos, J. Data Science with Data Mining and Machine Learning A design science research approach. Procedia Comput. Sci. 2024, 237, 245–252. [Google Scholar] [CrossRef]
- Epstein, E.; Nallapareddy, N.; Ray, S. On the relationship between feature selection metrics and accuracy. Entropy 2023, 25. [Google Scholar] [CrossRef]
- Posch, K.; Arbeiter, M.; Truden, C.; Pleschberger, M.; Pilz, J. Feature Selection Using Nearest Neighbor Gaussian Processes. Mathematics 2026, 14, 476. [Google Scholar] [CrossRef]
- Tawhid, M. A.; Ibrahim, A. M. Feature selection based on rough set approach, wrapper approach, and binary whale optimization algorithm. Int. J. Mach. Learn. Cybern. 2020, 11, 573–602. [Google Scholar] [CrossRef]
- Hashemi, A.; Dowlatshahi, M. B. Exploring Ant Colony Optimization for Feature Selection: A Comprehensive Review. Appl. Ant. Colony Optim. Its Var. 2024, 101–121. [Google Scholar]
- Pethe, Y. S.; Gourisaria, M. K.; Singh, P. K.; Das, H. FSBOA: Feature Selection Using Bat Optimization Algorithm for Software Fault Detection. Discov. Internet Things 2024, 4, 1–18. [Google Scholar] [CrossRef]
- Sekhar, L. C.; Sabu, M. K. Feature Selection Using Artificial Bee Colony and Discernibility Matrix in Rough Set Theory—A Hybrid Approach. Lect. Notes Netw. Syst. 2024, 834, 105–112. [Google Scholar]
- Xue, B.; Zhang, M.; Browne, W. N. Particle swarm optimisation for feature selection in classification: Novel initialisation and updating mechanisms. Appl. Soft Comput. 2014, 18, 261–276. [Google Scholar] [CrossRef]
- Simon, D. A Probabilistic Analysis of a Simplified Biogeography-Based Optimization Algorithm. Evol. Comput. 2011, 19, 167–188. [Google Scholar] [CrossRef]
- Goldberg, D. E. Genetic Algorithms in Search, Optimization and Machine Learning; Addison-Wesley, 1989. [Google Scholar]
- Zheng, L.; Diao, R.; Shen, Q. Efficient feature selection using a self-adjusting harmony search algorithm. 2013 13th UK Workshop on Computational Intelligence (UKCI), 2013; pp. 167–174. [Google Scholar]
- Latiffi, M. I. A.; Yaakub, M. R.; Ahmad, I. S. Flower Pollination Algorithm for Feature Selection in Tweets Sentiment Analysis. Int. J. Adv. Comput. Sci. Appl. 2022, 13, 429–435. [Google Scholar] [CrossRef]
- Kamel, S. R.; Yaghoubzadeh, R. Feature selection using grasshopper optimization algorithm in diagnosis of diabetes disease. Inform. Med. Unlocked 2021, 26, 100707. [Google Scholar] [CrossRef]
- Mafarja, M. M.; Eleyan, D.; Jaber, I.; Hammouri, A.; Mirjalili, S. Binary Dragonfly Algorithm for Feature Selection. 2017 International Conference on New Trends in Computing Sciences (ICTCS), 2017; pp. 12–17. [Google Scholar]
- Diaaeldin, I. M.; Hasanien, H. M.; Qais, M. H.; others. Multi scenario chaotic transient search optimization algorithm for global optimization technique. Sci. Rep. 2025, 15, 4284. [Google Scholar] [CrossRef]
- Alnaish, Z. a. H.; Algamal, Z. Y. Improving binary crow search algorithm for feature selection. J. Intell. Syst. 2023, 32, 45–62. [Google Scholar] [CrossRef]
- Eluri, U. R.; Devarakonda, R. Chaotic Dwarf Mongoose Optimization Algorithm for feature selection. Sci. Rep. 2023, 13, 50959. [Google Scholar]
- Alwakeel, A. S.; El-Rifaie, A. M.; Moustafa, G.; Shaheen, A. M. Newton Raphson based optimizer for optimal integration of FAS and RIS in wireless systems. Results Eng. 2025, 25, 103822. [Google Scholar] [CrossRef]
- Cheng, S.; Yin, J.; Liu, T. Multi Strategy Improvement of Newton-Raphson-Based Optimizer and Engineering Application. 2024 6th International Conference on Machine Learning, Big Data and Business Intelligence (MLBDBI); 2024, pp. 219–227.
- Ravichandran, S.; Manoharan, P.; Jangir, P. Newton–Raphson–Based Optimizer: A New Population–Based Metaheuristic Algorithm for Continuous Optimization Problems. Eng. Appl. Artif. Intell. 2024, 128, 107532. [Google Scholar]
- Li, Y.; Liu, B.; Chai, X.; Guo, F.; Li, Y.; Fu, D. Research on Shallow Water Depth Remote Sensing Based on the Improvement of the Newton–Raphson Optimizer. Water 2025, 17, 552. [Google Scholar] [CrossRef]
- Yousef, A.; Siam, A. I.; Barakat, S. I.; Mostafa, R. R. A Novel Newton Raphson Based Optimizer for Tomato Leaf Image Segmentation. Mansoura J. Comput. Inf. Sci. 2025, 20, 23–30. [Google Scholar]
- Lennard-Jones, J. E. On the Determination of Molecular Fields. Proc. R. Soc. A 1924, 106, 463–477. [Google Scholar]
- Hamming, R. W. Error detecting and error correcting codes. Bell Syst. Tech. J. 1950, 29, 147–160. [Google Scholar] [CrossRef]
- Xu, M.; Song, Q.; Xi, M.; Zhou, Z. Binary arithmetic optimization algorithm for feature selection. Soft Comput 2023, 11395–11429. [Google Scholar] [CrossRef] [PubMed]
- Mirjalili, S.; Mirjalili, S. M.; Yang, X. S. Binary bat algorithm. Neural Comput. Appl. 2014, 25, 663–681. [Google Scholar] [CrossRef]
- Rodrigues, D.; Yang, X. S.; de Souza, A. N.; Papa, J. P. Binary flower pollination algorithm and its application to feature selection. Stud. Comput. Intell. 2015, 85–100. [Google Scholar]
- Mirjalili, S.; Lewis, A. S-shaped versus V-shaped transfer functions for binary Particle Swarm Optimization. Swarm Evol. Comput. 2013, 9, 1–14. [Google Scholar] [CrossRef]
- Too, J.; Abdullah, A. R. Binary atom search optimization approaches for feature selection. Connect Sci. 2020, 32, 49–73. [Google Scholar] [CrossRef]
- Akinola, O. A.; Agushaka, J. O.; Ezugwu, A. E. Binary dwarf mongoose optimizer for solving high-dimensional feature selection problems. PLoS ONE 2022, 17, e0274850. [Google Scholar] [CrossRef]
- Anka, F.; Gharehchopogh, F. S.; Tejani, G. G.; Mousavirad, S. J. Advances in Mountain Gazelle Optimizer: A Comprehensive Study on its Classification and Applications. Int. J. Comput. Intell. Syst. 2025, 18, 247. [Google Scholar] [CrossRef]
- Hosseinzadeh, M.; Tanveer, J.; Rahmani, A. M.; Abbaszadi, R.; Gharehchopogh, F. S.; Porntaveetus, T.; Lee, S. W. A Comprehensive Survey Inspired by Elephant Optimization Algorithms: Comprehensive Analysis, Scrutinizing Analysis, and Future Research Directions. Arch. Comput. Methods Eng. 2025, 1–48. [Google Scholar] [CrossRef]
- Zhou, H.; Ren, D.; Xia, H.; Fan, M.; Yang, X.; Huang, H. An Attention-Based Interaction-Aware Spatio-Temporal Graph Neural Network for Trajectory Prediction. International Conference on Neural Information Processing, 2020; pp. 38–45. [Google Scholar]
- Fan, M.; Zhang, X.; Hu, J.; Gu, N.; Tao, D. Adaptive data structure regularized multiclass discriminative feature selection. IEEE Trans. Neural Netw. Learn. Syst. 2021, 33, 5859–5872. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Liu, W.; Wang, Q.; Zhang, S.; Hong, Z.; Xu, S. Reffacenet: Reference-based face image generation from line art drawings. Neurocomputing 2022, 488, 154–167. [Google Scholar] [CrossRef]
- Chaos-enhanced metaheuristics: classification, comparison, and convergence analysis. In Complex & Intelligent Systems; Limane, A., Zitouni, F., Harous, S., Eds.; 2025; Volume 11, p. 177. [Google Scholar]
- Authors. Multi scenario chaotic transient search optimization algorithm for global optimization technique. Scientific Reports, 2025.
- Wei, B.; Yang, S.; Zha, W.; Deng, L.; Huang, J.; Su, X.; Wang, F. Particle swarm optimization algorithm based on comprehensive scoring framework for high-dimensional feature selection. Swarm Evol. Comput. 2025, 95, 101915. [Google Scholar] [CrossRef]
- El maloufy, A.; Bencherqui, A.; Tahiri, M. A.; El Ghouate, N.; Karmouni, H.; Sayyouri, M.; Askar, S. S.; Abouhawwash, M. Chaos-enhanced white shark optimization algorithms CWSO for global optimization. Alex. Eng. J. 2025, 122, 465–483. [Google Scholar] [CrossRef]
- Ihsan, A.; Sag, T. Binary Puma Optimizer: A Novel Approach for Solving 0-1 Knapsack Problems and the Uncapacitated Facility Location Problem. Appl. Sci. 2025, 15, 9955. [Google Scholar] [CrossRef]
- Crawford, B.; Soto, R.; Caballero, H.; Astorga, G.; Cisternas-Caneo, F.; Solís-Piñones, F.; Giachetti, G. An Experimental Study of Transfer Functions and Binarization Strategies in Binary Arithmetic Optimization Algorithms for the Set Covering Problem. Mathematics 2025, 13, 3129. [Google Scholar] [CrossRef]
- Abdelrazek, M.; Abd Elaziz, M.; El-Baz, A. H. Chaotic Dwarf Mongoose Optimization Algorithm for feature selection. Sci. Rep. 2024, 14, 701. [Google Scholar] [CrossRef]
- Ramakrishnan, A.; Ramalingam, R.; Ramalingam, P.; Ravi, V.; Alahmadi, T. J.; Maidin, S. S. A novel chaotic binary butterfly optimization algorithm based feature selection model for classification of autism spectrum disorder. Int. J. Appl. Math. Comput. Sci. 2024, 34, 647–660. [Google Scholar] [CrossRef]
- Li, M.; Luo, Q.; Zhou, Y. Binary Grasshopper Optimization Algorithm with Time-Varying Gaussian Transfer Functions for Feature Selection. Biomimetics 2024, 9, 187. [Google Scholar] [CrossRef] [PubMed]
- Mehrabi, N.; Haeri Boroujeni, S. P.; Pashaei, E. An Efficient High-Dimensional Gene Selection Approach Based on Binary Horse Herd Optimization Algorithm for Biological Data Classification. Iran. J. Comput. Sci. 2024, 7, 1–17. [Google Scholar] [CrossRef]
- Ayeche, F.; Alti, A. Novel binary walrus optimization algorithms BWaOA and BWaOA-C with crossover operator for feature selection in high-dimensional data. Discov. Comput. 2025, 28, 234. [Google Scholar] [CrossRef]
- Nadimi-Shahraki, M. H.; Asghari Varzaneh, Z.; Zamani, H.; Mirjalili, S. Binary starling murmuration optimizer algorithm to select effective features from medical data. Appl. Sci. 2022, 13, 564. [Google Scholar] [CrossRef]
- Crawford, B.; López Cortés, B.; Cisternas-Caneo, F.; Gómez-Pulido, J. M.; Olivares, R.; Soto, R.; Barrera-Garcia, J.; Brante-Aguilera, C.; Giachetti, G. New Binary Reptile Search Algorithms for Binary Optimization Problems. Biomimetics 2025, 10, 653. [Google Scholar] [CrossRef]
- AbouOmar, M. S.; El Ferik, S. Multi-objective Newton-Raphson-based optimizer for fractional-order control of PEM fuel cells. Results Eng. 2025, 25, 104152. [Google Scholar] [CrossRef]
- Zelinka, I.; Diep, Q. B.; Snášel, V.; Das, S.; Innocenti, G.; Tesi, A.; Schoen, F.; Kuznetsov, N. V. Impact of chaotic dynamics on the performance of metaheuristic optimization algorithms: An experimental analysis. Inf. Sci. 2022, 587, 692–719. [Google Scholar] [CrossRef]
- Limane, A.; Zitouni, F.; Harous, S.; Lakbichi, R.; Ferhat, A.; Almazyad, A. S.; Jangir, P.; Mohamed, A. W. Chaos-enhanced metaheuristics: Classification, comparison, and convergence analysis. Complex Intell. Syst. 2025, 11. [Google Scholar] [CrossRef]
- Zhang, K.; Liu, Y.; Mei, F.; Sun, G.; Jin, J. Improved Binary Golden Jackal Optimization with Chaotic Tent Map and Cosine Similarity for Feature Selection. Entropy 2023, 25, 1128. [Google Scholar] [CrossRef]
- Zhao, W.; Wang, L.; Zhang, Z. A novel atom search optimization for dispersion coefficient estimation in groundwater. Future Gener. Comput. Syst. 2019, 91, 601–610. [Google Scholar] [CrossRef]
- El-Shorbagy, M. A.; Bouaouda, A.; Abualigah, L.; Hashim, F. A. Atom Search Optimization: a comprehensive review of its variants, applications, and future directions. PeerJ Comput. Sci. 2025, 11, e2722. [Google Scholar] [CrossRef] [PubMed]
- Devlin, S. M.; Kudenko, D. Dynamic potential-based reward shaping. 11th International Conference on Autonomous Agents and Multiagent Systems (AAMAS 2012), 2012; pp. 433–440. [Google Scholar]
- Beheshti, Z. BMNABC: Binary Multi-Neighborhood Artificial Bee Colony for High-Dimensional Discrete Optimization Problems. Cybern. Syst. 2018, 49, 452–474. [Google Scholar] [CrossRef]
- Asghari, A.; Zeinalabedinmalekmian, M.; Azgomi, H.; Alimoradi, M.; Ghaziantafrishi, S. Farmer ants optimization algorithm: a novel metaheuristic for solving discrete optimization problems. Information 2025, 16, 207. [Google Scholar] [CrossRef]
- Agrawal, P.; Ganesh, T.; Oliva, D.; Mohamed, A. W. S-shaped and v-shaped gaining-sharing knowledge-based algorithm for feature selection. Appl. Intell. 2022, 52, 81–112. [Google Scholar] [CrossRef]
- Beheshti, Z. UTF: Upgrade Transfer Function for Binary Meta-Heuristic Algorithms. Appl. Soft Comput. 2021, 106, 107346. [Google Scholar] [CrossRef]
- Too, J.; Abdullah, A. R.; Saad, N. M.; Ali, N. M. Feature selection based on binary tree growth algorithm for the classification of myoelectric signals. Machines 2018, 6, 72. [Google Scholar] [CrossRef]
- Dua, D.; Graff, C. UCI Machine Learning Repository. 2017. [Google Scholar]
- Jabbar, M. A.; Deekshatulu, B. L.; Chandra, P. A comprehensive study on decision tree classifiers for predicting heart disease. Mater. Today Proc. 2021, 45, 4968–4973. [Google Scholar]
- Saadatfar, H.; Khosravi, S.; Joloudari, J. H.; Mosavi, A.; Shamshirband, S. A New K-Nearest Neighbors Classifier for Big Data Based on Efficient Data Pruning. Mathematics 2020, 8, 1–18. [Google Scholar] [CrossRef]
- Hassaballah, M.; Muhammad, G.; Alabrah, A.; Al-Mutib, K.; Alsulaiman, M. Adaptive K-NN Metric Classification Based on Improved Kepler Optimization Algorithm. J. Supercomput. 2025, 66. [Google Scholar]
- Dokeroglu, T.; Kucukyilmaz, T. Multi-objective Harris Hawk metaheuristic algorithms for the diagnosis of Parkinson’s disease. Expert Syst. With Appl. 2025, 270, 126503. [Google Scholar] [CrossRef]
- Patil, A.; Sherekar, S. Classification Based on Decision Tree Algorithm for Machine Learning. Int. J. Sci. Res. Eng. Manag. (IJSREM) 2021, 5, 1–5. [Google Scholar]
- Singh Kushwah, J.; Kumar, A.; Patel, S.; Soni, R.; Gawande, A.; Gupta, S. Comparative study of regressor and classifier with decision tree using modern tools. Mater. Today Proc. 2022, 56, 3571–3576. [Google Scholar] [CrossRef]
- Yang, F. J. An Implementation of Naive Bayes Classifier. 2018 International Conference on Computational Science and Computational Intelligence (CSCI), 2018; pp. 301–306. [Google Scholar]
- Mortazavi, R.; Mortazavi, S.; Troncoso, A. Wrapper-based feature selection using regression trees to predict intrinsic viscosity of polymer. Eng. With Comput. 2022, 38, 2553–2565. [Google Scholar] [CrossRef]
- Derrac, J.; García, S.; Molina, D.; Herrera, F. A practical tutorial on the use of nonparametric statistical tests as a methodology for comparing evolutionary and swarm intelligence algorithms. Swarm Evol. Comput. 2011, 1, 3–18. [Google Scholar] [CrossRef]
- Karaboga, D.; Basturk, B. A powerful and efficient algorithm for numerical function optimization: artificial bee colony (ABC) algorithm. J. Glob. Optim. 2007, 39, 459–471. [Google Scholar] [CrossRef]
- Storn, R.; Price, K. Differential evolution–a simple and efficient heuristic for global optimization over continuous spaces. J. Glob. Optim. 1997, 11, 341–359. [Google Scholar] [CrossRef]
- Zhao, Z.; Morstatter, F.; Sharma, S.; Alelyani, S.; Anand, A.; Liu, H. Advancing Feature Selection Research; Arizona State University, ASU Feature Selection Repository Report, 2010; Volume 32, pp. 1–28. [Google Scholar]
| Algorithm | Dataset(s) | Class. | Mechanism | Chaos | Main Notes | Limitations |
|---|---|---|---|---|---|---|
| BPO [56] | Benchmark (Knapsack) |
N/A | Sigmoid and probabilistic |
No | Strong exploitation via puma hunting. |
Limited validation on feature selection datasets. |
| BAOA [57] | SCP datasets |
N/A | S-shaped and V-shaped TFs |
No | Analysis of transfer functions impact. |
Not validated on medical datasets. |
| CDMO [58] | UCI datasets |
k-NN | Chaotic map thresholding |
Yes | Improves search over standard DMO. |
Sensitivity to initial chaotic parameters. |
| CBBOA [59] | ASD / classification |
NB, k-NN |
Chaos-based transfer |
Yes | Enhances classification accuracy. |
Risk of local optima in complex data. |
| BGOA [60] | UCI, DEAP |
k-NN | Gaussian TF |
No | Strong global search and fast convergence. |
Needs careful tuning of parameters. |
| BHOA [61] | Microarray datasets |
SVM | X-shape TF |
No | Hybrid MRMR improves gene selection. |
Dependency on the filter-based stage. |
| BWaOA [62] | High-dim UCI |
k-NN | Crossover update |
No | Improves convergence quality and speed. |
Increased computational overhead. |
| BSMO [63] | Medical datasets |
k-NN, SVM |
S-shaped transfer |
No | Models collective bird behavior. |
Premature convergence risk; needs tuning. |
| BRSA [64] | Benchmark, UCI |
k-NN, SVM |
S/V-shaped TFs |
No | Strong exploration and exploitation. |
Performance may degrade in high-dim spaces. |
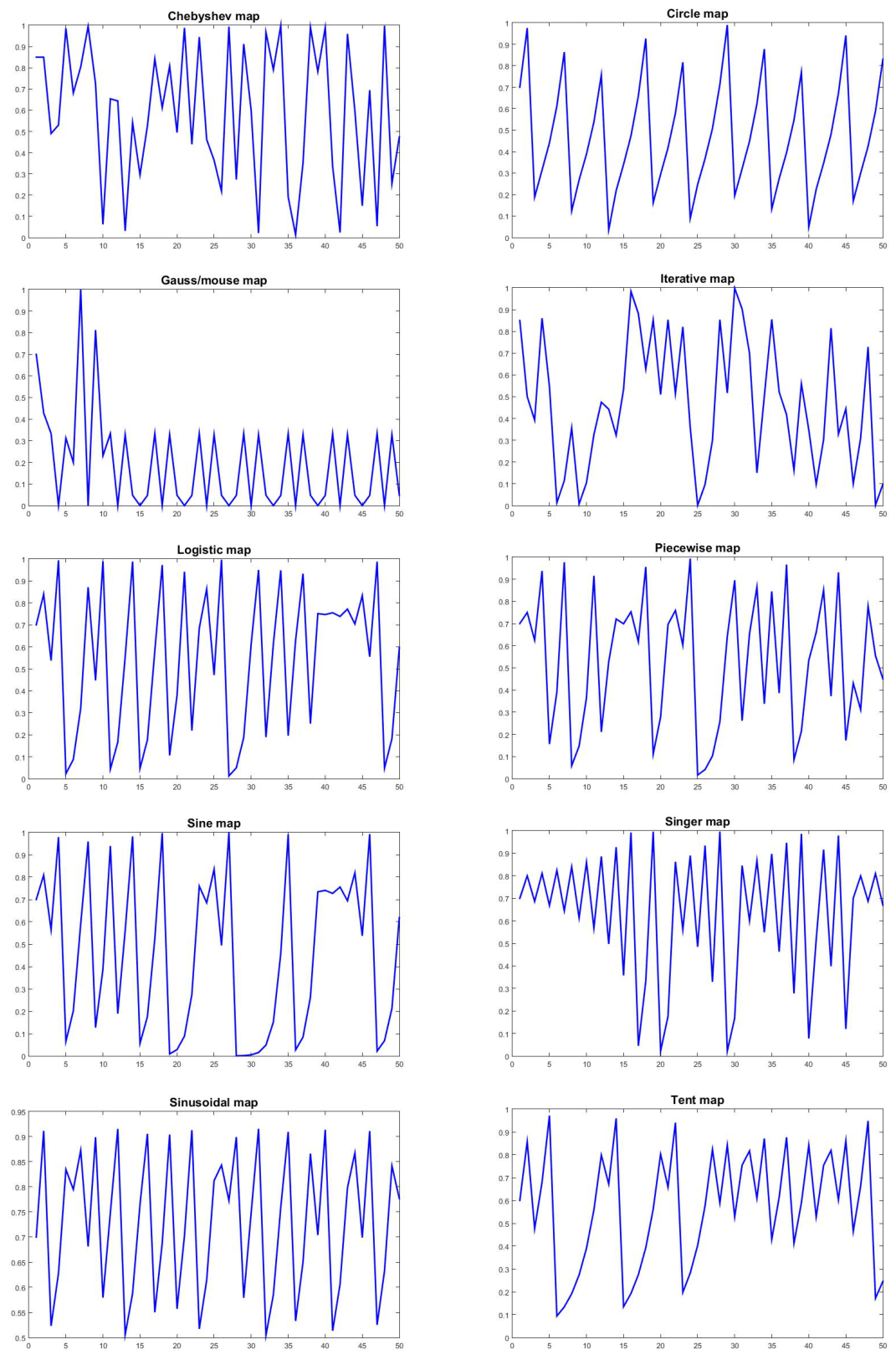
| Maps No. | Map name | Math formula | Range |
|---|---|---|---|
| Map1 | Chebyshev | ||
| Map2 | Circle | ||
| Map3 | Gauss/Mouse | ||
| Map4 | Iterative | ||
| Map5 | Logistic | ||
| Map6 | Piecewise | ||
| Map7 | Sine | ||
| Map8 | Singer | ||
| Map9 | Sinusoidal | ||
| Map10 | Tent |
| Algorithm | Parameter | Value |
|---|---|---|
| 0]*BAOA[41] | 0.99 | |
| 0.01 | ||
| the maximum values of MOP | 5 | |
| the minimum values of MOP | 0.2 | |
| 0]*BASO[45] | Depth weight, | 50 |
| Multiplier weight, | 0.2 | |
| 0]*BFPA [43] | Switch Probablity, P | 0.8 |
| Levy coefficient, | 1.5 | |
| 1]*BBA [42] | Maximum frequency, Fmax | 2 |
| Minimum frequency, Fmin | 0 | |
| Two constants, and | 0.9 | |
| 1]*BCCSA[32] | probability of awareness (AP) | 0.2 |
| flight length , | [1 , 1.8] | |
| 0]*BPSO [44] | Acceleration coefficients, C1 and C2 | 2 |
| Inertia weight, W | 0.1 | |
| Maximum Inertia weight, W | 0.9 | |
| Minimum Inertia weight, W | 0.4 | |
| 0]*BDA [46] | Crossover rate, CR | 1 |
| 0]*BDE | 32 | |
| Constant factor F | [0,2] | |
| Crossover constant CR | [0,1] | |
| Global_minimum | 1 | |
| VTR | 1.05 | |
| 0]*BABC | Acceleration coefficient | [-1,1] |
| Dataset No. | Dataset Name | No. of samples | No. of features |
|---|---|---|---|
| D1 | Chess | 3196 | 36 |
| D2 | Wisconsin | 699 | 9 |
| D3 | Breast | 569 | 30 |
| D4 | Olive | 572 | 8 |
| D5 | Lung Cancer | 32 | 56 |
| D6 | Diabetes | 144 | 7 |
| D7 | Heart | 294 | 13 |
| D8 | Ionosphere | 351 | 34 |
| D9 | Sonar | 208 | 60 |
| D10 | Lymphography | 148 | 18 |
| D11 | Hillvalley | 606 | 100 |
| D12 | LSVT | 126 | 310 |
| D13 | Zoo | 101 | 16 |
| D14 | Hepatitis | 80 | 19 |
| D15 | Diagnostic | 569 | 30 |
| D16 | Coimbra | 116 | 9 |
| D17 | BreastEw | 568 | 30 |
| D18 | HeartEw | 568 | 30 |
| D19 | SPECT | 267 | 22 |
| D20 | Diabetes | 768 | 8 |
| D21 | Cleveland | 297 | 13 |
| D22 | ILPD | 583 | 10 |
| D23 | Parkinsons | 5875 | 19 |
| D24 | Dermatology | 366 | 34 |
| D25 | Pd Speech | 756 | 753 |
| D26 | Heart Failure Clinical | 299 | 12 |
| Dataset | Algorithms | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BNRBO | Map1 | Map2 | Map3 | Map4 | Map5 | Map6 | Map7 | Map8 | Map9 | Map10 | ||
| Chess | Mean | 0.0326 | 0.0405 | 0.0408 | 0.0373 | 0.0357 | 0.0430 | 0.0463 | 0.0376 | 0.0418 | 0.0555 | 0.0342 |
| Std | 0.0059 | 0.0069 | 0.0064 | 0.0085 | 0.0074 | 0.0088 | 0.0081 | 0.0051 | 0.0051 | 0.0047 | 0.0070 | |
| Lung Cancer | Mean | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.1250 | 0.0833 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 |
| Std | 0.0143 | 0.0275 | 0.0164 | 0.0175 | 0.0277 | 0.0124 | 0.0198 | 0.0182 | 0.0222 | 0.0150 | 0.0123 | |
| Diabetes | Mean | 0.0714 | 0.0714 | 0.0000 | 0.0357 | 0.0357 | 0.0357 | 0.0357 | 0.0000 | 0.0000 | 0.0357 | 0.0000 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | |
| Sonar | Mean | 0.0207 | 0.0805 | 0.0573 | 0.0427 | 0.0817 | 0.0354 | 0.0524 | 0.0524 | 0.0671 | 0.0537 | 0.0146 |
| Std | 0.0134 | 0.0150 | 0.0142 | 0.0156 | 0.0157 | 0.0168 | 0.0160 | 0.0121 | 0.0145 | 0.0101 | 0.0089 | |
| Hillvalley | Mean | 0.3826 | 0.3103 | 0.4029 | 0.4223 | 0.3413 | 0.3587 | 0.3260 | 0.3715 | 0.3694 | 0.3831 | 0.3223 |
| Std | 0.0179 | 0.0325 | 0.0255 | 0.0799 | 0.0000 | 0.0398 | 0.0000 | 0.0325 | 0.0470 | 0.0440 | 0.0147 | |
| LSVT | Mean | 0.3160 | 0.2260 | 0.2700 | 0.2760 | 0.2000 | 0.1840 | 0.2400 | 0.3460 | 0.2280 | 0.2620 | 0.3140 |
| Std | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0740 | 0.0855 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | |
| Pd Speech | Mean | 0.2248 | 0.2500 | 0.2179 | 0.2159 | 0.1950 | 0.2056 | 0.1990 | 0.2136 | 0.2182 | 0.1878 | 0.2252 |
| Std | 0.0391 | 0.0236 | 0.0436 | 0.0404 | 0.0479 | 0.0329 | 0.0070 | 0.0457 | 0.0586 | 0.0049 | 0.0000 | |
| Parkinsons | Mean | 0.0068 | 0.0221 | 0.0184 | 0.0111 | 0.0106 | 0.0179 | 0.0077 | 0.0034 | 0.0162 | 0.0151 | 0.0060 |
| Std | 0.0000 | 0.0000 | 0.0004 | 0.0000 | 0.0004 | 0.0000 | 0.0002 | 0.0002 | 0.0000 | 0.0006 | 0.0000 | |
| Dermatology | Mean | 0.0000 | 0.0000 | 0.0021 | 0.0041 | 0.0027 | 0.0000 | 0.0137 | 0.0000 | 0.0000 | 0.0000 | 0.0000 |
| Std | 0.0000 | 0.0000 | 0.0050 | 0.0064 | 0.0056 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | |
| Pd Speech | Mean | 0.2248 | 0.2500 | 0.2179 | 0.2159 | 0.1950 | 0.2056 | 0.1990 | 0.2136 | 0.2182 | 0.1878 | 0.2252 |
| Std | 0.0391 | 0.0236 | 0.0436 | 0.0404 | 0.0479 | 0.0329 | 0.0070 | 0.0457 | 0.0586 | 0.0049 | 0.0000 | |
| Dataset | BNRBO | BCNRBO | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Map1 | Map2 | Map3 | Map4 | Map5 | Map6 | Map7 | Map8 | Map9 | Map10 | ||
| Chess | 3.1 | 6.2 | 6.475 | 4.95 | 4.05 | 6.95 | 8.2 | 5.175 | 6.775 | 10.55 | 3.575 |
| Lung Cancer | 5.375 | 5.375 | 5.375 | 5.375 | 9.5 | 8.125 | 5.375 | 5.375 | 5.375 | 5.375 | 5.375 |
| Diabetes | 10.5 | 10.5 | 2.5 | 7 | 7 | 7 | 7 | 2.5 | 2.5 | 7 | 2.5 |
| Sonar | 2.5 | 8.95 | 6.975 | 5.15 | 9.175 | 4.325 | 6.2 | 6.35 | 7.95 | 6.475 | 1.95 |
| Hillvalley | 8.225 | 1.5 | 9.725 | 10.825 | 3.85 | 5.6 | 2.725 | 6.825 | 6.575 | 7.85 | 2.3 |
| LSVT | 9.35 | 4.225 | 6.75 | 6.375 | 2.375 | 2.15 | 4.925 | 10.325 | 4.275 | 6.125 | 9.125 |
| Diagnostic | 7.025 | 8.1 | 11 | 1.9 | 7.475 | 9.975 | 1.1 | 4.95 | 7.025 | 3.175 | 4.275 |
| Parkinsons | 3 | 11 | 9.825 | 5.75 | 5.25 | 9.175 | 4 | 1 | 7.95 | 7.05 | 2 |
| Dermatology | 5.175 | 5.175 | 6 | 6.825 | 6.275 | 5.175 | 10.675 | 5.175 | 5.175 | 5.175 | 5.175 |
| Pd Speech | 7.225 | 9.175 | 6.6 | 5.9 | 4.225 | 5.1 | 5.25 | 6.15 | 5.75 | 3.5 | 7.125 |
| Sum. | 61.475 | 70.2 | 71.225 | 60.05 | 59.175 | 63.575 | 55.45 | 53.825 | 59.35 | 62.275 | 43.4 |
| Avg. | 6.148 | 7.020 | 7.123 | 6.005 | 5.918 | 6.358 | 5.545 | 5.383 | 5.935 | 6.228 | 4.340 |
| BCNRBO | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Dataset | BNRBO | Map1 | Map2 | Map3 | Map4 | Map5 | Map6 | Map7 | Map8 | Map9 | ||||||||||||||||||||
| value | R | value | R | value | R | value | R | value | R | value | R | value | R | value | R | value | R | value | R | |||||||||||
| Chess | 0.37118 | 0 | 0.03179 | 1 | 0.00373 | 1 | 0.13751 | 0 | 0.33755 | 0 | 0.0013 | 1 | 0.00029 | 1 | 0.04219 | 1 | 0.00297 | 1 | 0.00013 | 1 | ||||||||||
| Lung Cancer | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 6.1E-05 | 1 | 0.00195 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | ||||||||||
| Diabetes | 7.7E-06 | 1 | 7.7E-06 | 1 | 1 | 0 | 7.7E-06 | 1 | 7.7E-06 | 1 | 7.7E-06 | 1 | 7.7E-06 | 1 | 1 | 0 | 1 | 0 | 7.7E-06 | 1 | ||||||||||
| Sonar | 0.25781 | 0 | 9.9E-05 | 1 | 0.00018 | 1 | 0.00018 | 1 | 0.00012 | 1 | 0.00012 | 1 | 0.00026 | 1 | 0.00016 | 1 | 0.0001 | 1 | 0.00016 | 1 | ||||||||||
| Hillvalley | 8.5E-05 | 1 | 0.01344 | -1 | 8.3E-05 | 1 | 7.9E-05 | 1 | 0.00194 | 1 | 0.00013 | 1 | 0.2913 | 0 | 7.9E-05 | 1 | 8.1E-05 | 1 | 8.1E-05 | 1 | ||||||||||
| LSVT | 1 | 0 | 3.7E-05 | -1 | 0.00011 | -1 | 0.00866 | -1 | 2.3E-05 | -1 | 5.7E-05 | -1 | 2.3E-05 | -1 | 0.00452 | 1 | 0.00018 | -1 | 0.00012 | 1 | ||||||||||
| Diagnostic | 7.7E-06 | 1 | 5.4E-05 | 1 | 7.7E-06 | 1 | 2.9E-05 | -1 | 2.9E-05 | 1 | 1.2E-05 | 1 | 7.7E-06 | -1 | 0.0625 | 0 | 7.7E-06 | 1 | 6.3E-05 | -1 | ||||||||||
| Parkinsons | 7.7E-06 | 1 | 7.7E-06 | 1 | 4.7E-05 | 1 | 7.7E-06 | 1 | 5.4E-05 | 1 | 7.7E-06 | 1 | 1.2E-05 | 1 | 1.2E-05 | -1 | 7.7E-06 | 1 | 6.1E-05 | 1 | ||||||||||
| Dermatology | 1 | 0 | 1 | 0 | 0.25 | 0 | 0.03125 | 1 | 0.125 | 0 | 1 | 0 | 7.7E-06 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | ||||||||||
| Pd Speech | 0.83688 | 0 | 0.00093 | 1 | 1 | 0 | 0.33046 | 0 | 0.01471 | -1 | 0.10488 | 0 | 6.3E-05 | -1 | 0.95493 | 0 | 0.89558 | 0 | 5.8E-05 | -1 | ||||||||||
| Won | 4 | 6 | 5 | 5 | 6 | 7 | 5 | 4 | 5 | 6 | ||||||||||||||||||||
| Loss | 0 | 2 | 1 | 2 | 2 | 1 | 3 | 1 | 1 | 2 | ||||||||||||||||||||
| Equal | 6 | 2 | 4 | 3 | 2 | 2 | 2 | 5 | 4 | 2 | ||||||||||||||||||||
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.025039 | 0.051643 | 0.032864 | 0.046948 | 0.025039 | 0.059468 | 0.040689 | 0.039124 | 0.035994 | 0.025039 |
| D 2 | 0.007194 | 0.035971 | 0.014388 | 0.007194 | 0.014388 | 0.007194 | 0.035971 | 0.007194 | 0.007194 | 0.007194 |
| D 3 | 0.035398 | 0.026549 | 0.026549 | 0.053097 | 0.053097 | 0.035398 | 0.044248 | 0.035398 | 0.035398 | 0.017699 |
| D 4 | 0.087719 | 0.087719 | 0.061404 | 0.12281 | 0.087719 | 0.061404 | 0.10526 | 0.087719 | 0.087719 | 0.070175 |
| D 5 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.16667 | 0 | 0 |
| D 6 | 0 | 0.035714 | 0.035714 | 0.035714 | 0.035714 | 0 | 0 | 0 | 0 | 0.071429 |
| D 7 | 0.13793 | 0.051724 | 0.10345 | 0.068966 | 0.12069 | 0.13793 | 0.051724 | 0.15517 | 0.13793 | 0.12069 |
| D 8 | 0.028571 | 0.014286 | 0.071429 | 0.1 | 0.042857 | 0.12857 | 0.071429 | 0.057143 | 0.042857 | 0.014286 |
| D 9 | 0 | 0.02439 | 0.073171 | 0.073171 | 0 | 0.097561 | 0.097561 | 0.04878 | 0.02439 | 0.02439 |
| D 10 | 0.10345 | 0.10345 | 0.068966 | 0.13793 | 0.068966 | 0.068966 | 0.034483 | 0.10345 | 0.034483 | 0.068966 |
| D 11 | 0.29752 | 0.36364 | 0.35537 | 0.35537 | 0.38017 | 0.40496 | 0.3719 | 0.33058 | 0.3719 | 0.39669 |
| D 12 | 0.28 | 0.2 | 0.16 | 0.28 | 0.16 | 0.28 | 0.4 | 0.2 | 0.4 | 0.2 |
| D 13 | 0.05 | 0 | 0.05 | 0 | 0 | 0 | 0 | 0 | 0 | 0.05 |
| D 14 | 0.1875 | 0.125 | 0.125 | 0.125 | 0 | 0.0625 | 0.125 | 0.125 | 0.1875 | 0.125 |
| D 15 | 0.026549 | 0.044248 | 0.026549 | 0.044248 | 0.00885 | 0.053097 | 0.035398 | 0.017699 | 0.044248 | 0.035398 |
| D 16 | 0.086957 | 0.13043 | 0.17391 | 0.13043 | 0 | 0.043478 | 0.086957 | 0.086957 | 0.17391 | 0.13043 |
| D 17 | 0.026549 | 0.035398 | 0.017699 | 0.035398 | 0.026549 | 0.044248 | 0.026549 | 0.044248 | 0.035398 | 0.035398 |
| D 18 | 0.092593 | 0.074074 | 0.074074 | 0.092593 | 0.11111 | 0.14815 | 0.12963 | 0.092593 | 0.11111 | 0.11111 |
| D 19 | 0.16981 | 0.11321 | 0.15094 | 0.16981 | 0.13208 | 0.22642 | 0.16981 | 0.13208 | 0.18868 | 0.09434 |
| D 20 | 0.084746 | 0.10169 | 0.15254 | 0.15254 | 0.084746 | 0.15254 | 0.084746 | 0.067797 | 0.15254 | 0.067797 |
| D 21 | 0.22222 | 0.23529 | 0.19608 | 0.22876 | 0.21569 | 0.20915 | 0.22222 | 0.24183 | 0.20915 | 0.20915 |
| D 22 | 0.23276 | 0.21552 | 0.24138 | 0.25862 | 0.19828 | 0.25 | 0.24138 | 0.2069 | 0.25862 | 0.24138 |
| D 23 | 0.005957 | 0.021277 | 0.014468 | 0.024681 | 0.012766 | 0.01617 | 0.017021 | 0.020426 | 0.014468 | 0.019574 |
| D 24 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| D 25 | 0.22517 | 0.17219 | 0.15894 | 0.25166 | 0.15894 | 0.22517 | 0.23179 | 0.24503 | 0.17881 | 0.1457 |
| D 26 | 0.11864 | 0.13559 | 0.13559 | 0.18644 | 0.20339 | 0.10169 | 0.13559 | 0.13559 | 0.11864 | 0.13559 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.034194 | 0.064319 | 0.052739 | 0.057199 | 0.036385 | 0.063928 | 0.054773 | 0.046479 | 0.041628 | 0.034351 |
| D 2 | 0.007554 | 0.035971 | 0.016547 | 0.007194 | 0.020144 | 0.014748 | 0.035971 | 0.007194 | 0.007194 | 0.008633 |
| D 3 | 0.044248 | 0.026991 | 0.050442 | 0.059735 | 0.069912 | 0.047788 | 0.056637 | 0.043363 | 0.038496 | 0.017699 |
| D 4 | 0.087719 | 0.088158 | 0.069737 | 0.12281 | 0.089035 | 0.069298 | 0.10526 | 0.087719 | 0.087719 | 0.070175 |
| D 5 | 0 | 0 | 0.10833 | 0.15 | 0 | 0 | 0 | 0.16667 | 0 | 0 |
| D 6 | 0 | 0.035714 | 0.035714 | 0.035714 | 0.035714 | 0 | 0 | 0 | 0 | 0.071429 |
| D 7 | 0.14052 | 0.063793 | 0.11034 | 0.073276 | 0.12069 | 0.18879 | 0.061207 | 0.1569 | 0.14828 | 0.13017 |
| D 8 | 0.057857 | 0.029286 | 0.086429 | 0.105 | 0.077857 | 0.15714 | 0.081429 | 0.086429 | 0.054286 | 0.032857 |
| D 9 | 0.014634 | 0.064634 | 0.12195 | 0.10366 | 0.029268 | 0.13049 | 0.15366 | 0.080488 | 0.030488 | 0.045122 |
| D 10 | 0.11034 | 0.12586 | 0.096552 | 0.1569 | 0.10345 | 0.12759 | 0.058621 | 0.10345 | 0.034483 | 0.096552 |
| D 11 | 0.32231 | 0.38058 | 0.41157 | 0.36157 | 0.40496 | 0.43512 | 0.38223 | 0.35496 | 0.39215 | 0.42562 |
| D 12 | 0.314 | 0.264 | 0.202 | 0.308 | 0.274 | 0.288 | 0.456 | 0.2 | 0.446 | 0.206 |
| D 13 | 0.05 | 0.0275 | 0.05 | 0.0075 | 0 | 0.0275 | 0 | 0 | 0 | 0.055 |
| D 14 | 0.19375 | 0.125 | 0.125 | 0.15 | 0.14375 | 0.1125 | 0.125 | 0.20938 | 0.1875 | 0.125 |
| D 15 | 0.026549 | 0.04646 | 0.05354 | 0.044248 | 0.032301 | 0.071239 | 0.05 | 0.017699 | 0.050885 | 0.049115 |
| D 16 | 0.086957 | 0.15217 | 0.18478 | 0.13043 | 0 | 0.10217 | 0.086957 | 0.086957 | 0.17391 | 0.13043 |
| D 17 | 0.026549 | 0.04292 | 0.034956 | 0.043363 | 0.026991 | 0.050442 | 0.033186 | 0.052655 | 0.035398 | 0.035398 |
| D 18 | 0.1 | 0.098148 | 0.074074 | 0.10648 | 0.13704 | 0.22315 | 0.12963 | 0.10741 | 0.11481 | 0.12315 |
| D 19 | 0.20566 | 0.15283 | 0.18208 | 0.21038 | 0.17358 | 0.28113 | 0.19906 | 0.16792 | 0.2 | 0.12736 |
| D 20 | 0.085593 | 0.13898 | 0.17458 | 0.15424 | 0.10339 | 0.21186 | 0.11102 | 0.09322 | 0.15508 | 0.085593 |
| D 21 | 0.22222 | 0.23529 | 0.19935 | 0.22908 | 0.2232 | 0.2317 | 0.22222 | 0.24183 | 0.20915 | 0.20915 |
| D 22 | 0.23276 | 0.21595 | 0.24914 | 0.26078 | 0.21552 | 0.25345 | 0.25819 | 0.20733 | 0.25862 | 0.24138 |
| D 23 | 0.005957 | 0.022553 | 0.014468 | 0.025404 | 0.016979 | 0.021106 | 0.017872 | 0.020426 | 0.014468 | 0.019574 |
| D 24 | 0 | 0 | 0 | 0 | 0 | 0.012329 | 0 | 0 | 0 | 0 |
| D 25 | 0.22517 | 0.2 | 0.19503 | 0.25166 | 0.22053 | 0.2298 | 0.23742 | 0.25066 | 0.22947 | 0.20695 |
| D 26 | 0.11864 | 0.13559 | 0.13729 | 0.18644 | 0.20339 | 0.12034 | 0.14746 | 0.13559 | 0.11864 | 0.13559 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 2.175 | 9.05 | 6.55 | 7.275 | 2.6 | 9.3 | 6.925 | 5.125 | 3.7 | 2.3 |
| D 2 | 2.55 | 9.5 | 5.925 | 2.425 | 6.975 | 5.325 | 9.5 | 7.375 | 2.425 | 3 |
| D 3 | 5.5 | 2.1 | 6.6 | 8.375 | 9.825 | 6.15 | 7.575 | 3.625 | 4.25 | 1 |
| D 4 | 6.3 | 6.4 | 2.95 | 10 | 6.625 | 2.975 | 9 | 1.275 | 6.3 | 3.175 |
| D 5 | 4.25 | 4.25 | 7.3 | 8.725 | 4.25 | 4.25 | 4.25 | 9.225 | 4.25 | 4.25 |
| D 6 | 3 | 7.5 | 7.5 | 7.5 | 7.5 | 3 | 3 | 3 | 3 | 10 |
| D 7 | 7.05 | 1.875 | 4.375 | 2.475 | 5.05 | 9.875 | 1.65 | 8.725 | 7.875 | 6.05 |
| D 8 | 3.8 | 1.575 | 6.7 | 8.725 | 5.925 | 10 | 6.25 | 6.775 | 3.475 | 1.775 |
| D 9 | 1.625 | 5 | 8.225 | 7.25 | 2.575 | 8.525 | 9.575 | 5.85 | 2.675 | 3.7 |
| D 10 | 6.225 | 7.725 | 5.025 | 9.5 | 5.475 | 7.275 | 2.15 | 5.55 | 1.15 | 4.925 |
| D 11 | 1 | 4.725 | 7.75 | 2.85 | 7.075 | 9.525 | 4.925 | 2.35 | 6 | 8.8 |
| D 12 | 7.025 | 4.85 | 2.3 | 6.75 | 4.55 | 5.775 | 9.575 | 2.35 | 9.25 | 2.575 |
| D 13 | 8.325 | 6.1 | 8.325 | 4.1 | 3.375 | 6.1 | 3.375 | 3.375 | 3.375 | 8.55 |
| D 14 | 8.525 | 3.725 | 3.725 | 5.6 | 5.65 | 3.15 | 3.725 | 8.85 | 8.325 | 3.725 |
| D 15 | 2.35 | 5.825 | 6.525 | 5.175 | 3.625 | 9.65 | 7.1 | 1.275 | 7.15 | 6.325 |
| D 16 | 3.425 | 7.75 | 9.25 | 6.4 | 1 | 4.85 | 3.425 | 3.425 | 9.075 | 6.4 |
| D 17 | 1.95 | 7 | 5.15 | 7.2 | 2.075 | 8.95 | 3.925 | 9.5 | 4.625 | 4.625 |
| D 18 | 3.55 | 3.425 | 1.075 | 4.525 | 7.525 | 9.875 | 7.95 | 4.75 | 5.8 | 6.525 |
| D 19 | 7.2 | 2.675 | 5.05 | 7.575 | 4.375 | 9.925 | 6.525 | 3.65 | 6.65 | 1.375 |
| D 20 | 2.125 | 6.25 | 8.825 | 7.45 | 3.375 | 9.8 | 4.35 | 2.95 | 7.45 | 2.425 |
| D 21 | 5.5 | 8.4 | 1.3 | 7.25 | 5.65 | 6.825 | 5.5 | 9.575 | 2.5 | 2.5 |
| D 22 | 3.775 | 2.575 | 6.6 | 8.85 | 2.25 | 7.275 | 8.475 | 1.475 | 8.4 | 5.325 |
| D 23 | 1 | 8.575 | 3.05 | 9.925 | 4.9 | 7.475 | 4.675 | 6.725 | 3.05 | 5.625 |
| D 24 | 5.175 | 5.175 | 5.175 | 5.175 | 5.175 | 8.425 | 5.175 | 5.175 | 5.175 | 5.175 |
| D 25 | 4.45 | 2.075 | 2.3 | 9.425 | 4.75 | 5.75 | 7.25 | 9.275 | 6.2 | 3.525 |
| D 26 | 2.05 | 5.3 | 5.55 | 9 | 10 | 3.225 | 7.225 | 5.3 | 2.05 | 5.3 |
| summation | 109.900 | 139.400 | 143.100 | 179.500 | 132.150 | 183.250 | 153.050 | 136.525 | 134.175 | 118.950 |
| Average | 4.227 | 5.362 | 5.504 | 6.904 | 5.083 | 7.048 | 5.887 | 5.251 | 5.161 | 4.575 |
| Dataset | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | |||||||||
| D 1 | 8.81E-05 | 1 | 8.73E-05 | 1 | 8.7E-05 | 1 | 0.421649 | 0 | 8.73E-05 | 1 | 0.000154 | 1 | 0.000231 | 1 | 0.00088 | 1 | 0.982563 | 0 | ||||||||
| D 2 | 1.19E-05 | 1 | 4.26E-05 | 1 | 1 | 0 | 3.62E-05 | 1 | 0.000105 | 1 | 1.19E-05 | 1 | 1.19E-05 | 1 | 1 | 0 | 0.375 | 0 | ||||||||
| D 3 | 9.43E-05 | -1 | 0.011963 | 1 | 0.000243 | 1 | 9.43E-05 | 1 | 0.391479 | 0 | 0.004354 | 1 | 0.000977 | 1 | 0.004395 | 1 | 6.2E-05 | -1 | ||||||||
| D 4 | 1 | 0 | 0.00029 | -1 | 7.74E-06 | 1 | 0.25 | 0 | 1.71E-05 | -1 | 7.74E-06 | 1 | 7.74E-06 | -1 | 1 | 0 | 7.74E-06 | -1 | ||||||||
| D 5 | 1 | 0 | 0.000488 | 1 | 2.21E-05 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 7.74E-06 | 1 | 1 | 0 | 1 | 0 | ||||||||
| D 6 | 7.74E-06 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 7.74E-06 | 1 | ||||||||
| D 7 | 6.03E-05 | -1 | 6.69E-05 | -1 | 5.57E-05 | -1 | 2.3E-05 | -1 | 0.000105 | 1 | 6.39E-05 | -1 | 3.38E-05 | 1 | 0.011719 | 1 | 0.001953 | 1 | ||||||||
| D 8 | 0.00025 | -1 | 0.000392 | 1 | 7.77E-05 | 1 | 0.001496 | 1 | 8.38E-05 | 1 | 0.000407 | 1 | 0.000181 | 1 | 0.434082 | 0 | 0.00027 | -1 | ||||||||
| D 9 | 0.000118 | 1 | 7.68E-05 | 1 | 6.19E-05 | 1 | 0.003906 | 1 | 6.26E-05 | 1 | 6.75E-05 | 1 | 7.19E-05 | 1 | 0.000977 | 1 | 0.000327 | 1 | ||||||||
| D 10 | 0.011719 | 1 | 0.217529 | 0 | 0.000142 | 1 | 0.125 | 0 | 0.010742 | 1 | 5.62E-05 | -1 | 0.125 | 0 | 2.93E-05 | -1 | 0.041016 | 1 | ||||||||
| D 11 | 7.75E-05 | 1 | 8.4E-05 | 1 | 7.12E-05 | 1 | 8.56E-05 | 1 | 8.4E-05 | 1 | 7.51E-05 | 1 | 7.88E-05 | 1 | 7.85E-05 | 1 | 8.29E-05 | 1 | ||||||||
| D 12 | 0.000194 | -1 | 7.39E-05 | -1 | 0.375 | 0 | 0.330354 | 0 | 0.000244 | 1 | 4.26E-05 | 1 | 2.3E-05 | -1 | 6.06E-05 | 1 | 2.97E-05 | -1 | ||||||||
| D 13 | 0.003906 | 1 | 1 | 0 | 3.74E-05 | -1 | 7.74E-06 | -1 | 0.003906 | 1 | 7.74E-06 | -1 | 7.74E-06 | -1 | 7.74E-06 | -1 | 0.5 | 0 | ||||||||
| D 14 | 1.71E-05 | -1 | 1.71E-05 | -1 | 0.000488 | 1 | 0.000488 | 1 | 4.15E-05 | -1 | 1.71E-05 | -1 | 0.125 | 0 | 0.5 | 0 | 1.71E-05 | -1 | ||||||||
| D 15 | 3.56E-05 | 1 | 0.000179 | 1 | 7.74E-06 | 1 | 0.189651 | 0 | 7.5E-05 | 1 | 4.26E-05 | 1 | 7.74E-06 | -1 | 3.56E-05 | 1 | 5.6E-05 | 1 | ||||||||
| D 16 | 5.4E-05 | 1 | 2.31E-05 | 1 | 7.74E-06 | 1 | 7.74E-06 | -1 | 0.75493 | 0 | 1 | 0 | 1 | 0 | 7.74E-06 | 1 | 7.74E-06 | 1 | ||||||||
| D 17 | 4.94E-05 | 1 | 0.002827 | 1 | 1.71E-05 | 1 | 1 | 0 | 4.15E-05 | 1 | 6.1E-05 | 1 | 1.19E-05 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | ||||||||
| D 18 | 0.923828 | 0 | 5.06E-05 | -1 | 0.09375 | 0 | 0.000183 | 1 | 8.63E-05 | 1 | 5.06E-05 | 1 | 0.080078 | 0 | 0.000244 | 1 | 6.1E-05 | 1 | ||||||||
| D 19 | 0.000112 | -1 | 0.00621 | -1 | 0.484375 | 0 | 0.001468 | -1 | 0.000123 | 1 | 0.334473 | 0 | 0.00011 | -1 | 0.206328 | 0 | 8.01E-05 | -1 | ||||||||
| D 20 | 6.72E-05 | 1 | 7.19E-05 | 1 | 2.31E-05 | 1 | 0.001953 | 1 | 7.32E-05 | 1 | 0.000157 | 1 | 0.011719 | 1 | 1.71E-05 | 1 | 1 | 0 | ||||||||
| D 21 | 7.74E-06 | 1 | 3.56E-05 | -1 | 1.19E-05 | 1 | 0.826823 | 0 | 0.093832 | 0 | 1 | 0 | 7.74E-06 | 1 | 7.74E-06 | -1 | 7.74E-06 | -1 | ||||||||
| D 22 | 1.19E-05 | -1 | 6.06E-05 | 1 | 3.56E-05 | 1 | 0.000488 | 1 | 4.32E-05 | 1 | 6.75E-05 | 1 | 1.19E-05 | -1 | 7.74E-06 | 1 | 7.74E-06 | 1 | ||||||||
| D 23 | 7.45E-05 | 1 | 7.74E-06 | 1 | 2.3E-05 | 1 | 5.31E-05 | 1 | 7.28E-05 | 1 | 2.31E-05 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | ||||||||
| D 24 | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 0.000244 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | ||||||||
| D 25 | 8.4E-05 | -1 | 0.000467 | -1 | 7.74E-06 | 1 | 0.546875 | 0 | 0.000122 | 1 | 6.39E-05 | 1 | 2.3E-05 | 1 | 0.062374 | 0 | 0.013672 | 1 | ||||||||
| D 26 | 7.74E-06 | 1 | 1.71E-05 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 0.672214 | 0 | 4.15E-05 | 1 | 7.74E-06 | 1 | 1 | 0 | 7.74E-06 | 1 | ||||||||
| Won | 14 | 15 | 19 | 12 | 18 | 16 | 14 | 13 | 13 | |||||||||||||||||
| Loss | 8 | 8 | 2 | 4 | 2 | 4 | 6 | 3 | 7 | |||||||||||||||||
| Equal | 4 | 3 | 5 | 10 | 6 | 6 | 6 | 10 | 6 | |||||||||||||||||
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.004695 | 0.023474 | 0.00939 | 0.004695 | 0.004695 | 0.017214 | 0.007825 | 0.00626 | 0.00939 | 0.00939 |
| D 2 | 0.007194 | 0.007194 | 0.028777 | 0.028777 | 0.035971 | 0.021583 | 0.028777 | 0.014388 | 0.028777 | 0.014388 |
| D 3 | 0 | 0.044248 | 0.00885 | 0.00885 | 0.00885 | 0.017699 | 0.026549 | 0.00885 | 0.026549 | 0.026549 |
| D 4 | 0.078947 | 0.061404 | 0.070175 | 0.04386 | 0.078947 | 0.035088 | 0.061404 | 0.026316 | 0.052632 | 0.070175 |
| D 5 | 0.166667 | 0 | 0 | 0.166667 | 0 | 0 | 0 | 0 | 0 | 0.166667 |
| D 6 | 0 | 0.107143 | 0.035714 | 0.035714 | 0.035714 | 0 | 0.035714 | 0.035714 | 0.071429 | 0.035714 |
| D 7 | 0.103448 | 0.137931 | 0.086207 | 0.12069 | 0.103448 | 0.103448 | 0.103448 | 0.137931 | 0.137931 | 0.086207 |
| D 8 | 0 | 0.028571 | 0.014286 | 0.014286 | 0.028571 | 0.042857 | 0.028571 | 0.014286 | 0.042857 | 0.014286 |
| D 9 | 0.02439 | 0.04878 | 0.04878 | 0.097561 | 0.02439 | 0.146341 | 0.121951 | 0.097561 | 0.02439 | 0 |
| D 10 | 0.103448 | 0.068966 | 0.103448 | 0.034483 | 0 | 0.103448 | 0.103448 | 0.034483 | 0.068966 | 0.103448 |
| D 11 | 0.272727 | 0.31405 | 0.305785 | 0.289256 | 0.214876 | 0.371901 | 0.305785 | 0.272727 | 0.31405 | 0.247934 |
| D 12 | 0.04 | 0.04 | 0.04 | 0.08 | 0.04 | 0.12 | 0.04 | 0.04 | 0.04 | 0.08 |
| D 13 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.05 |
| D 14 | 0 | 0.125 | 0.125 | 0.0625 | 0.125 | 0.0625 | 0 | 0.125 | 0.125 | 0.0625 |
| D 15 | 0.00885 | 0.035398 | 0.017699 | 0.017699 | 0 | 0 | 0.026549 | 0.035398 | 0.00885 | 0.017699 |
| D 16 | 0.173913 | 0.086957 | 0.086957 | 0.086957 | 0.173913 | 0.217391 | 0.130435 | 0.173913 | 0 | 0.173913 |
| D 17 | 0.00885 | 0.00885 | 0.026549 | 0.017699 | 0.017699 | 0.044248 | 0.026549 | 0.00885 | 0 | 0 |
| D 18 | 0.12963 | 0.111111 | 0.111111 | 0.148148 | 0.092593 | 0.148148 | 0.092593 | 0.12963 | 0.055556 | 0.074074 |
| D 19 | 0.207547 | 0.132075 | 0.150943 | 0.169811 | 0.132075 | 0.226415 | 0.150943 | 0.113208 | 0.150943 | 0.150943 |
| D 20 | 0.084746 | 0.101695 | 0.101695 | 0.101695 | 0.135593 | 0.135593 | 0.118644 | 0.118644 | 0.135593 | 0.084746 |
| D 21 | 0.281046 | 0.267974 | 0.228758 | 0.235294 | 0.235294 | 0.228758 | 0.24183 | 0.215686 | 0.24183 | 0.24183 |
| D 22 | 0.224138 | 0.224138 | 0.241379 | 0.25 | 0.258621 | 0.224138 | 0.206897 | 0.224138 | 0.241379 | 0.241379 |
| D 23 | 0.01617 | 0.04 | 0.006809 | 0.040851 | 0.012766 | 0.059574 | 0.04766 | 0.036596 | 0.026383 | 0.015319 |
| D 24 | 0 | 0.013699 | 0.013699 | 0.013699 | 0 | 0.013699 | 0 | 0.013699 | 0.027397 | 0 |
| D 25 | 0.05298 | 0.099338 | 0.086093 | 0.072848 | 0.046358 | 0.139073 | 0.099338 | 0.13245 | 0.066225 | 0.059603 |
| D 26 | 0.118644 | 0.152542 | 0.101695 | 0.084746 | 0.118644 | 0.118644 | 0.118644 | 0.050847 | 0.067797 | 0.118644 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.00626 | 0.035681 | 0.011581 | 0.00759 | 0.004851 | 0.017214 | 0.012207 | 0.007981 | 0.012911 | 0.012676 |
| D 2 | 0.007194 | 0.007554 | 0.033813 | 0.028777 | 0.042806 | 0.034532 | 0.029137 | 0.014388 | 0.029137 | 0.014388 |
| D 3 | 0.003097 | 0.05177 | 0.019027 | 0.021681 | 0.015929 | 0.030973 | 0.034956 | 0.015487 | 0.031416 | 0.038938 |
| D 4 | 0.078947 | 0.065789 | 0.077632 | 0.04386 | 0.079386 | 0.059211 | 0.061404 | 0.026316 | 0.053509 | 0.070175 |
| D 5 | 0.2 | 0.141667 | 0.141667 | 0.166667 | 0.15 | 0.141667 | 0 | 0.141667 | 0 | 0.166667 |
| D 6 | 0 | 0.107143 | 0.05 | 0.035714 | 0.035714 | 0.001786 | 0.035714 | 0.035714 | 0.071429 | 0.035714 |
| D 7 | 0.114655 | 0.155172 | 0.086207 | 0.152586 | 0.114655 | 0.113793 | 0.10431 | 0.137931 | 0.137931 | 0.091379 |
| D 8 | 0.019286 | 0.046429 | 0.027143 | 0.027857 | 0.032857 | 0.054286 | 0.029286 | 0.027857 | 0.067143 | 0.015714 |
| D 9 | 0.052439 | 0.069512 | 0.084146 | 0.126829 | 0.057317 | 0.192683 | 0.143902 | 0.117073 | 0.063415 | 0.045122 |
| D 10 | 0.113793 | 0.106897 | 0.103448 | 0.037931 | 0.027586 | 0.155172 | 0.103448 | 0.034483 | 0.124138 | 0.105172 |
| D 11 | 0.301653 | 0.34876 | 0.327686 | 0.320661 | 0.263636 | 0.392149 | 0.328512 | 0.301653 | 0.339256 | 0.281818 |
| D 12 | 0.05 | 0.07 | 0.108 | 0.082 | 0.04 | 0.18 | 0.042 | 0.04 | 0.11 | 0.106 |
| D 13 | 0 | 0 | 0.01 | 0.0025 | 0.02 | 0.0375 | 0.005 | 0 | 0 | 0.05 |
| D 14 | 0.034375 | 0.134375 | 0.175 | 0.103125 | 0.178125 | 0.175 | 0.109375 | 0.125 | 0.18125 | 0.09375 |
| D 15 | 0.010177 | 0.038938 | 0.031416 | 0.025664 | 0.016372 | 0.019912 | 0.029204 | 0.035841 | 0.023451 | 0.022566 |
| D 16 | 0.173913 | 0.086957 | 0.086957 | 0.086957 | 0.184783 | 0.271739 | 0.130435 | 0.180435 | 0 | 0.173913 |
| D 17 | 0.015044 | 0.023009 | 0.033186 | 0.019469 | 0.025664 | 0.052655 | 0.031858 | 0.017257 | 0.008407 | 0 |
| D 18 | 0.14537 | 0.131481 | 0.116667 | 0.168519 | 0.092593 | 0.173148 | 0.092593 | 0.138889 | 0.055556 | 0.082407 |
| D 19 | 0.220755 | 0.15566 | 0.184906 | 0.2 | 0.136792 | 0.256604 | 0.181132 | 0.150943 | 0.187736 | 0.174528 |
| D 20 | 0.101695 | 0.116949 | 0.118644 | 0.114407 | 0.138136 | 0.170339 | 0.118644 | 0.121186 | 0.138136 | 0.098305 |
| D 21 | 0.281046 | 0.269935 | 0.231699 | 0.235294 | 0.236601 | 0.251634 | 0.24183 | 0.215686 | 0.242157 | 0.242157 |
| D 22 | 0.225 | 0.228879 | 0.246983 | 0.25 | 0.269397 | 0.278017 | 0.211638 | 0.227155 | 0.243534 | 0.242241 |
| D 23 | 0.018851 | 0.06383 | 0.018468 | 0.050298 | 0.058468 | 0.086255 | 0.059404 | 0.058681 | 0.040596 | 0.025064 |
| D 24 | 0 | 0.013699 | 0.017808 | 0.015753 | 0.000685 | 0.028767 | 0.013014 | 0.013699 | 0.039726 | 0.003425 |
| D 25 | 0.067219 | 0.110596 | 0.111589 | 0.09404 | 0.068543 | 0.168874 | 0.119205 | 0.142053 | 0.074503 | 0.080795 |
| D 26 | 0.118644 | 0.152542 | 0.104237 | 0.090678 | 0.120339 | 0.160169 | 0.118644 | 0.055085 | 0.067797 | 0.118644 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 2.45 | 10 | 5.95 | 3.275 | 1.375 | 8.925 | 6.325 | 3.4 | 6.65 | 6.65 |
| D 2 | 1.475 | 1.575 | 7.575 | 6.425 | 9.7 | 8.175 | 6.55 | 3.475 | 6.575 | 3.475 |
| D 3 | 1.15 | 9.875 | 3.85 | 4.575 | 3.125 | 6.725 | 7.65 | 3.025 | 6.7 | 8.325 |
| D 4 | 8.975 | 5.65 | 8.45 | 2.175 | 9.05 | 4.85 | 4.725 | 1 | 3.425 | 6.7 |
| D 5 | 7.5 | 6 | 6.025 | 6.75 | 6.25 | 6.025 | 1.85 | 6 | 1.85 | 6.75 |
| D 6 | 1.475 | 10 | 6.7 | 5.275 | 5.275 | 1.65 | 5.275 | 5.275 | 8.8 | 5.275 |
| D 7 | 4.8 | 9.475 | 1.35 | 9.225 | 4.8 | 4.7 | 3.625 | 7.475 | 7.475 | 2.075 |
| D 8 | 2.925 | 7.8 | 4.3 | 4.6 | 5.525 | 8.775 | 4.925 | 4.625 | 9.625 | 1.9 |
| D 9 | 2.775 | 4.15 | 5.35 | 7.8 | 3.175 | 9.875 | 8.625 | 7.15 | 3.675 | 2.425 |
| D 10 | 6.95 | 6.375 | 6 | 2.175 | 1.775 | 9.525 | 6 | 2.05 | 8.025 | 6.125 |
| D 11 | 3.6 | 8.4 | 6.4 | 5.475 | 1.5 | 10 | 6.45 | 3.375 | 7.775 | 2.025 |
| D 12 | 3.375 | 5.15 | 7.6 | 6.125 | 2.575 | 9.875 | 2.75 | 2.575 | 7.375 | 7.6 |
| D 13 | 4.25 | 4.25 | 5.25 | 4.5 | 6.25 | 8 | 4.75 | 4.25 | 4.25 | 9.25 |
| D 14 | 1.35 | 5.5 | 7.7 | 3.875 | 8.025 | 7.65 | 4.25 | 4.925 | 8.325 | 3.4 |
| D 15 | 1.4 | 9.175 | 7.3 | 5.5 | 2.875 | 3.925 | 6.55 | 8.675 | 4.95 | 4.65 |
| D 16 | 7.3 | 3 | 3 | 3 | 7.825 | 9.9 | 5 | 7.675 | 1 | 7.3 |
| D 17 | 3.875 | 6 | 8.075 | 5.1 | 6.325 | 9.975 | 7.975 | 4.425 | 2.225 | 1.025 |
| D 18 | 7.575 | 6.35 | 5.425 | 9.275 | 3.275 | 9.425 | 3.275 | 6.95 | 1 | 2.45 |
| D 19 | 8.825 | 3.05 | 5.725 | 7.125 | 1.6 | 9.95 | 5.375 | 2.55 | 5.95 | 4.85 |
| D 20 | 2.225 | 4.8 | 5.1 | 4.275 | 8.25 | 9.85 | 5.025 | 5.4 | 8.25 | 1.825 |
| D 21 | 9.95 | 9.05 | 2.55 | 3.375 | 3.6 | 7.025 | 6.1 | 1 | 6.175 | 6.175 |
| D 22 | 2.575 | 3.35 | 6.625 | 7.625 | 9.2 | 9.35 | 1.375 | 2.95 | 6.125 | 5.825 |
| D 23 | 2.45 | 7.25 | 2.15 | 5.4 | 7.225 | 9.825 | 7.125 | 6.875 | 4.075 | 2.625 |
| D 24 | 1.875 | 5.725 | 6.625 | 6.175 | 2.05 | 8.6 | 5.5 | 5.725 | 9.775 | 2.95 |
| D 25 | 1.9 | 6.6 | 6.7 | 4.875 | 1.925 | 9.95 | 7.575 | 9.05 | 2.825 | 3.6 |
| D 26 | 6.35 | 9.375 | 4.1 | 3.525 | 6.575 | 9.375 | 6.35 | 1.125 | 1.875 | 6.35 |
| Summation | 109.350 | 167.925 | 145.875 | 137.500 | 129.125 | 211.900 | 140.975 | 121.000 | 144.750 | 121.600 |
| Average | 4.206 | 6.459 | 5.611 | 5.288 | 4.966 | 8.150 | 5.422 | 4.654 | 5.567 | 4.677 |
| Dataset | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | |||||||||
| D 1 | 8.76E-05 | 1 | 0.000184 | 1 | 0.080139 | 0 | 0.022461 | 1 | 6.59E-05 | 1 | 0.000188 | 1 | 0.0183 | 1 | 0.000128 | 1 | 0.000107 | 1 | ||||||||
| D 2 | 1 | 0 | 4.67E-05 | 1 | 7.74E-06 | 1 | 6.86E-05 | 1 | 5.02E-05 | 1 | 1.19E-05 | 1 | 7.74E-06 | 1 | 1.19E-05 | 1 | 7.74E-06 | 1 | ||||||||
| D 3 | 6.91E-05 | 1 | 4.26E-05 | 1 | 9.84E-05 | 1 | 0.000117 | 1 | 7.03E-05 | 1 | 4.83E-05 | 1 | 0.000171 | 1 | 7.5E-05 | 1 | 7.5E-05 | 1 | ||||||||
| D 4 | 7.93E-05 | -1 | 0.581055 | 0 | 7.74E-06 | -1 | 1 | 0 | 0.000176 | -1 | 7.74E-06 | -1 | 7.74E-06 | -1 | 1.19E-05 | -1 | 7.74E-06 | -1 | ||||||||
| D 5 | 0.015625 | 1 | 0.03125 | 1 | 0.125 | 0 | 0.03125 | 1 | 0.03125 | 1 | 2.93E-05 | -1 | 0.015625 | 1 | 2.93E-05 | -1 | 0.125 | 0 | ||||||||
| D 6 | 7.74E-06 | 1 | 5.06E-05 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 1 | 0 | 7.74E-06 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | ||||||||
| D 7 | 6.07E-05 | 1 | 4.67E-05 | -1 | 5.47E-05 | 1 | 1 | 0 | 0.818546 | 0 | 0.000488 | 1 | 4.67E-05 | 1 | 4.67E-05 | 1 | 0.000185 | -1 | ||||||||
| D 8 | 0.000177 | 1 | 0.159058 | 0 | 0.016357 | 1 | 0.000122 | 1 | 8.12E-05 | 1 | 0.000488 | 1 | 0.001953 | 1 | 7.71E-05 | 1 | 0.039063 | 1 | ||||||||
| D 9 | 0.002197 | 1 | 6.1E-05 | 1 | 7.19E-05 | 1 | 0.283203 | 0 | 8.29E-05 | 1 | 7.14E-05 | 1 | 8.12E-05 | 1 | 0.072205 | 0 | 0.256348 | 0 | ||||||||
| D 10 | 0.289063 | 0 | 0.03125 | 1 | 4.26E-05 | -1 | 7.7E-05 | -1 | 0.000768 | 1 | 0.03125 | 1 | 4.15E-05 | -1 | 0.122308 | 0 | 0.0625 | 0 | ||||||||
| D 11 | 0.000125 | 1 | 9.99E-05 | 1 | 6.1E-05 | 1 | 0.000279 | -1 | 8.07E-05 | 1 | 0.000181 | 1 | 0.874756 | 0 | 0.000123 | 1 | 0.001309 | -1 | ||||||||
| D 12 | 0.116333 | 0 | 0.000216 | 1 | 0.009311 | 1 | 0.25 | 0 | 0.0001 | 1 | 0.375 | 0 | 0.25 | 0 | 0.000439 | 1 | 0.000299 | 1 | ||||||||
| D 13 | 1 | 0 | 0.125 | 0 | 1 | 0 | 0.007813 | 1 | 6.1E-05 | 1 | 0.5 | 0 | 1 | 0 | 1 | 0 | 7.74E-06 | 1 | ||||||||
| D 14 | 5.79E-05 | 1 | 6.56E-05 | 1 | 6.1E-05 | 1 | 6.53E-05 | 1 | 5.06E-05 | 1 | 0.000329 | 1 | 5.31E-05 | 1 | 4.67E-05 | 1 | 6.1E-05 | 1 | ||||||||
| D 15 | 6.2E-05 | 1 | 6.03E-05 | 1 | 6.53E-05 | 1 | 0.000465 | 1 | 0.001099 | 1 | 4.94E-05 | 1 | 2.96E-05 | 1 | 0.000131 | 1 | 5.06E-05 | 1 | ||||||||
| D 16 | 7.74E-06 | -1 | 7.74E-06 | -1 | 7.74E-06 | -1 | 0.0625 | 0 | 5.62E-05 | 1 | 7.74E-06 | -1 | 0.25 | 0 | 7.74E-06 | -1 | 1 | 0 | ||||||||
| D 17 | 0.000244 | 1 | 0.000115 | 1 | 0.027344 | 1 | 0.000732 | 1 | 7.19E-05 | 1 | 5.57E-05 | 1 | 0.226563 | 0 | 0.000488 | 1 | 6.12E-05 | -1 | ||||||||
| D 18 | 0.010075 | -1 | 5.62E-05 | -1 | 0.000192 | 1 | 5.48E-05 | -1 | 0.000438 | 1 | 5.48E-05 | -1 | 0.143066 | 0 | 5.48E-05 | -1 | 6.6E-05 | -1 | ||||||||
| D 19 | 7.48E-05 | -1 | 0.000167 | -1 | 0.000545 | -1 | 6.4E-05 | -1 | 0.000108 | 1 | 7.02E-05 | -1 | 7.18E-05 | -1 | 0.000101 | -1 | 7.65E-05 | -1 | ||||||||
| D 20 | 0.000488 | 1 | 0.000217 | 1 | 0.002686 | 1 | 4.39E-05 | 1 | 7.94E-05 | 1 | 4.78E-05 | 1 | 8E-05 | 1 | 4.39E-05 | 1 | 0.1875 | 0 | ||||||||
| D 21 | 4.78E-05 | -1 | 5.31E-05 | -1 | 7.74E-06 | -1 | 1.19E-05 | -1 | 7.67E-05 | -1 | 7.74E-06 | -1 | 7.74E-06 | -1 | 1.19E-05 | -1 | 1.19E-05 | -1 | ||||||||
| D 22 | 0.044922 | 1 | 6.2E-05 | 1 | 1.71E-05 | 1 | 7.06E-05 | 1 | 0.000122 | 1 | 0.000218 | -1 | 0.125 | 0 | 3.62E-05 | 1 | 2.97E-05 | 1 | ||||||||
| D 23 | 8.83E-05 | 1 | 0.24283 | 0 | 8.72E-05 | 1 | 0.000383 | 1 | 8.82E-05 | 1 | 8.77E-05 | 1 | 8.79E-05 | 1 | 0.000103 | 1 | 0.05419 | 0 | ||||||||
| D 24 | 7.74E-06 | 1 | 4.15E-05 | 1 | 2.3E-05 | 1 | 1 | 0 | 5.57E-05 | 1 | 1.31E-05 | 1 | 7.74E-06 | 1 | 1.71E-05 | 1 | 0.0625 | 0 | ||||||||
| D 25 | 8.31E-05 | 1 | 0.000129 | 1 | 8.49E-05 | 1 | 0.958643 | 0 | 8.75E-05 | 1 | 8.5E-05 | 1 | 7.95E-05 | 1 | 0.016987 | 1 | 0.001889 | 1 | ||||||||
| D 26 | 7.74E-06 | 1 | 3.74E-05 | -1 | 5.7E-05 | -1 | 0.5 | 0 | 0.000174 | 1 | 1 | 0 | 3.56E-05 | -1 | 7.74E-06 | -1 | 1 | 0 | ||||||||
| Won | 17 | 16 | 17 | 13 | 22 | 16 | 14 | 16 | 11 | |||||||||||||||||
| Loss | 5 | 6 | 6 | 5 | 2 | 7 | 5 | 7 | 7 | |||||||||||||||||
| Equal | 4 | 4 | 3 | 8 | 2 | 3 | 7 | 3 | 8 | |||||||||||||||||
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.156495 | 0.167449 | 0.156495 | 0.170579 | 0.14241 | 0.203443 | 0.161189 | 0.14241 | 0.156495 | 0.137715 |
| D 2 | 0.007194 | 0.007194 | 0.007194 | 0.021583 | 0.014388 | 0.028777 | 0.035971 | 0.021583 | 0.014388 | 0.043165 |
| D 3 | 0.00885 | 0.00885 | 0 | 0.017699 | 0.017699 | 0.017699 | 0.035398 | 0.00885 | 0.00885 | 0.00885 |
| D 4 | 0.035088 | 0.026316 | 0.035088 | 0.070175 | 0.052632 | 0.04386 | 0.035088 | 0.061404 | 0.052632 | 0.035088 |
| D 5 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.166667 | 0 | 0.166667 |
| D 6 | 0 | 0 | 0 | 0 | 0 | 0 | 0.035714 | 0 | 0 | 0 |
| D 7 | 0.068966 | 0.155172 | 0.137931 | 0.103448 | 0.086207 | 0.103448 | 0.155172 | 0.103448 | 0.12069 | 0.103448 |
| D 8 | 0.014286 | 0.028571 | 0.014286 | 0.042857 | 0 | 0 | 0.014286 | 0.014286 | 0.014286 | 0 |
| D 9 | 0.073171 | 0.146341 | 0.02439 | 0.073171 | 0 | 0.097561 | 0.146341 | 0.121951 | 0 | 0.097561 |
| D 10 | 0.068966 | 0.103448 | 0.068966 | 0.034483 | 0.034483 | 0.068966 | 0 | 0.068966 | 0.103448 | 0.103448 |
| D 11 | 0.371901 | 0.487603 | 0.454545 | 0.404959 | 0.429752 | 0.396694 | 0.446281 | 0.487603 | 0.446281 | 0.454545 |
| D 12 | 0.12 | 0.08 | 0.04 | 0.2 | 0.08 | 0.2 | 0.08 | 0.16 | 0.08 | 0.04 |
| D 13 | 0.05 | 0.1 | 0.05 | 0.05 | 0.05 | 0.1 | 0.05 | 0.05 | 0.05 | 0.05 |
| D 14 | 0 | 0.125 | 0.125 | 0.125 | 0 | 0.1875 | 0.125 | 0 | 0.1875 | 0.0625 |
| D 15 | 0 | 0.026549 | 0.00885 | 0.026549 | 0.00885 | 0 | 0.00885 | 0.017699 | 0.026549 | 0.017699 |
| D 16 | 0.173913 | 0.173913 | 0.173913 | 0.043478 | 0.130435 | 0.086957 | 0.217391 | 0.086957 | 0.086957 | 0.173913 |
| D 17 | 0.00885 | 0.00885 | 0.026549 | 0.00885 | 0.017699 | 0.035398 | 0 | 0.017699 | 0 | 0 |
| D 18 | 0.055556 | 0.074074 | 0.074074 | 0.12963 | 0.111111 | 0.12963 | 0.111111 | 0.111111 | 0.092593 | 0.055556 |
| D 19 | 0.113208 | 0.207547 | 0.113208 | 0.132075 | 0.245283 | 0.226415 | 0.245283 | 0.169811 | 0.113208 | 0.207547 |
| D 20 | 0.067797 | 0.169492 | 0.101695 | 0.050847 | 0.135593 | 0.118644 | 0.101695 | 0.152542 | 0.084746 | 0.135593 |
| D 21 | 0.196078 | 0.24183 | 0.215686 | 0.196078 | 0.176471 | 0.215686 | 0.156863 | 0.202614 | 0.202614 | 0.176471 |
| D 22 | 0.181034 | 0.232759 | 0.241379 | 0.258621 | 0.206897 | 0.25 | 0.224138 | 0.215517 | 0.25 | 0.206897 |
| D 23 | 0.215319 | 0.237447 | 0.231489 | 0.241702 | 0.234894 | 0.225532 | 0.233191 | 0.244255 | 0.245106 | 0.228936 |
| D 24 | 0.013699 | 0.027397 | 0 | 0 | 0 | 0.027397 | 0 | 0.013699 | 0 | 0 |
| D 25 | 0.211921 | 0.198675 | 0.125828 | 0.192053 | 0.119205 | 0.205298 | 0.18543 | 0.192053 | 0.165563 | 0.198675 |
| D 26 | 0.084746 | 0.169492 | 0.169492 | 0.135593 | 0.152542 | 0.135593 | 0.135593 | 0.118644 | 0.084746 | 0.101695 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 0.163615 | 0.191628 | 0.164241 | 0.178091 | 0.150782 | 0.237715 | 0.170892 | 0.149139 | 0.166432 | 0.151487 |
| D 2 | 0.007194 | 0.010432 | 0.011151 | 0.021583 | 0.014388 | 0.034532 | 0.035971 | 0.021583 | 0.014388 | 0.043525 |
| D 3 | 0.015044 | 0.026549 | 0.009292 | 0.024336 | 0.019912 | 0.026991 | 0.042478 | 0.023451 | 0.00885 | 0.018584 |
| D 4 | 0.035088 | 0.02807 | 0.041667 | 0.070175 | 0.052632 | 0.057456 | 0.035088 | 0.061404 | 0.05307 | 0.035526 |
| D 5 | 0.075 | 0 | 0.133333 | 0 | 0 | 0.008333 | 0 | 0.166667 | 0 | 0.166667 |
| D 6 | 0 | 0 | 0.005357 | 0 | 0 | 0.001786 | 0.035714 | 0 | 0 | 0 |
| D 7 | 0.069828 | 0.155172 | 0.164655 | 0.133621 | 0.087069 | 0.110345 | 0.168103 | 0.113793 | 0.12069 | 0.111207 |
| D 8 | 0.039286 | 0.044286 | 0.022857 | 0.055714 | 0.017857 | 0.010714 | 0.014286 | 0.03 | 0.022857 | 0.013571 |
| D 9 | 0.135366 | 0.17561 | 0.078049 | 0.103659 | 0.018293 | 0.136585 | 0.164634 | 0.162195 | 0.042683 | 0.152439 |
| D 10 | 0.086207 | 0.124138 | 0.091379 | 0.043103 | 0.056897 | 0.155172 | 0 | 0.101724 | 0.131034 | 0.115517 |
| D 11 | 0.385124 | 0.499587 | 0.468595 | 0.409504 | 0.438843 | 0.408678 | 0.446281 | 0.495455 | 0.448347 | 0.467355 |
| D 12 | 0.138 | 0.122 | 0.128 | 0.232 | 0.12 | 0.242 | 0.118 | 0.198 | 0.12 | 0.088 |
| D 13 | 0.0525 | 0.105 | 0.065 | 0.05 | 0.0525 | 0.145 | 0.05 | 0.05 | 0.05 | 0.0525 |
| D 14 | 0.059375 | 0.15 | 0.171875 | 0.125 | 0 | 0.2625 | 0.125 | 0.059375 | 0.1875 | 0.13125 |
| D 15 | 0 | 0.034956 | 0.018584 | 0.035398 | 0.014602 | 0.012389 | 0.00885 | 0.019027 | 0.034513 | 0.022124 |
| D 16 | 0.173913 | 0.182609 | 0.176087 | 0.045652 | 0.13913 | 0.119565 | 0.217391 | 0.086957 | 0.086957 | 0.176087 |
| D 17 | 0.020796 | 0.009292 | 0.030088 | 0.032743 | 0.022124 | 0.050885 | 0.00708 | 0.017699 | 0.000442 | 0.004867 |
| D 18 | 0.062037 | 0.099074 | 0.077778 | 0.12963 | 0.113889 | 0.155556 | 0.112963 | 0.117593 | 0.093519 | 0.068519 |
| D 19 | 0.113208 | 0.215094 | 0.129245 | 0.170755 | 0.265094 | 0.261321 | 0.258491 | 0.184906 | 0.143396 | 0.227358 |
| D 20 | 0.081356 | 0.177119 | 0.107627 | 0.066102 | 0.15339 | 0.148305 | 0.101695 | 0.157627 | 0.087288 | 0.135593 |
| D 21 | 0.196732 | 0.24183 | 0.217647 | 0.196078 | 0.177451 | 0.227778 | 0.156863 | 0.202614 | 0.202614 | 0.176471 |
| D 22 | 0.187931 | 0.242241 | 0.246983 | 0.270259 | 0.216379 | 0.272845 | 0.233621 | 0.218966 | 0.25 | 0.209052 |
| D 23 | 0.218553 | 0.244383 | 0.236043 | 0.254936 | 0.238894 | 0.24834 | 0.243191 | 0.255574 | 0.250511 | 0.232383 |
| D 24 | 0.023973 | 0.05274 | 0.003425 | 0.002055 | 0.008904 | 0.076027 | 0.012329 | 0.026027 | 0.00137 | 0.000685 |
| D 25 | 0.231457 | 0.211258 | 0.138411 | 0.204305 | 0.135099 | 0.218874 | 0.192715 | 0.19702 | 0.176821 | 0.212583 |
| D 26 | 0.097458 | 0.172034 | 0.170339 | 0.14661 | 0.152542 | 0.157627 | 0.135593 | 0.131356 | 0.084746 | 0.101695 |
| Dataset | BCNRBO | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA |
|---|---|---|---|---|---|---|---|---|---|---|
| D 1 | 4.85 | 8.8 | 5.1 | 7.875 | 2.3 | 9.9 | 6.425 | 1.725 | 5.525 | 2.5 |
| D 2 | 1.5 | 2.625 | 2.875 | 6.5 | 4 | 8.45 | 8.6 | 6.5 | 4 | 9.95 |
| D 3 | 3.6 | 6.9 | 2.4 | 6.725 | 5 | 7.45 | 9.925 | 6.45 | 1.775 | 4.775 |
| D 4 | 3.175 | 1.375 | 4.55 | 9.95 | 6.575 | 7.65 | 3.175 | 8.6 | 6.675 | 3.275 |
| D 5 | 6.1 | 3.85 | 7.85 | 3.85 | 3.85 | 4.1 | 3.85 | 8.85 | 3.85 | 8.85 |
| D 6 | 4.9 | 4.9 | 5.65 | 4.9 | 4.9 | 5.15 | 9.9 | 4.9 | 4.9 | 4.9 |
| D 7 | 1.025 | 8.225 | 9.075 | 6.525 | 2.025 | 4.325 | 9.4 | 4.575 | 5.55 | 4.275 |
| D 8 | 7.675 | 8.6 | 5 | 9.825 | 3.925 | 2.375 | 3.025 | 6.475 | 5.05 | 3.05 |
| D 9 | 6.05 | 8.875 | 3.15 | 4.25 | 1.175 | 6.075 | 8.2 | 8.025 | 1.95 | 7.25 |
| D 10 | 4.9 | 7.925 | 5.3 | 2.475 | 3.075 | 9 | 1 | 6.025 | 8.375 | 6.925 |
| D 11 | 1 | 9.75 | 7.55 | 2.525 | 4.2 | 2.475 | 5.25 | 9.25 | 5.575 | 7.425 |
| D 12 | 5.325 | 4.3 | 4.5 | 9.3 | 4.075 | 9.45 | 3.975 | 8.05 | 4.1 | 1.925 |
| D 13 | 4.575 | 8.875 | 5.2 | 4.35 | 4.575 | 9.8 | 4.35 | 4.35 | 4.35 | 4.575 |
| D 14 | 2.575 | 6.575 | 7.55 | 5.375 | 1.05 | 9.875 | 5.375 | 2.575 | 8.325 | 5.725 |
| D 15 | 1.05 | 8.85 | 5.325 | 8.925 | 4.175 | 3.65 | 2.6 | 5.425 | 8.8 | 6.2 |
| D 16 | 7.25 | 7.775 | 7.375 | 1.075 | 5.075 | 4.025 | 9.85 | 2.6 | 2.6 | 7.375 |
| D 17 | 6.225 | 3.475 | 8.05 | 8.275 | 6.675 | 9.85 | 2.95 | 5.675 | 1.4 | 2.425 |
| D 18 | 1.5 | 5 | 2.725 | 8.75 | 6.775 | 9.875 | 6.7 | 7.3 | 4.3 | 2.075 |
| D 19 | 1.125 | 6.325 | 2.1 | 4.1 | 9.05 | 8.9 | 8.825 | 4.825 | 2.85 | 6.9 |
| D 20 | 2.275 | 9.75 | 4.675 | 1.075 | 7.975 | 7.575 | 4.325 | 8.325 | 2.8 | 6.225 |
| D 21 | 4.625 | 9.925 | 8.25 | 4.425 | 2.575 | 8.825 | 1 | 6.45 | 6.45 | 2.475 |
| D 22 | 1.325 | 6.1 | 6.95 | 9.325 | 3.15 | 9.55 | 5.15 | 3.675 | 7.55 | 2.225 |
| D 23 | 1 | 5.675 | 3.625 | 8.95 | 4.025 | 6.875 | 5.45 | 9 | 7.775 | 2.625 |
| D 24 | 6.95 | 9.2 | 3.325 | 2.925 | 4.625 | 9.625 | 5.4 | 7.55 | 2.775 | 2.625 |
| D 25 | 9.85 | 7.275 | 1.6 | 6.25 | 1.4 | 8.425 | 4.325 | 5.125 | 3.025 | 7.725 |
| D 26 | 2.25 | 9.3 | 9.175 | 6.3 | 7.125 | 7.55 | 4.925 | 4.625 | 1.125 | 2.625 |
| summation | 102.675 | 180.225 | 138.925 | 154.8 | 113.35 | 190.8 | 143.95 | 156.925 | 121.45 | 126.9 |
| Average | 3.949038 | 6.931731 | 5.343269 | 5.953846 | 4.359615 | 7.338462 | 5.536538 | 6.035577 | 4.671154 | 4.880769 |
| Dataset | BAOA | jBASO | BFPA | BBA | BCCSA | BDE | BABC | BPSO | BDA | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | P Value | R | |||||||||
| D 1 | 8.79E-05 | 1 | 0.428794 | 0 | 8.66E-05 | 1 | 0.000391 | -1 | 8.81E-05 | 1 | 0.001323 | 1 | 0.000128 | -1 | 0.082975 | 0 | 0.000129 | -1 | ||||||||
| D 2 | 0.003906 | 1 | 0.000977 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 5.47E-05 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 1.19E-05 | 1 | ||||||||
| D 3 | 0.000769 | 1 | 0.01416 | 1 | 6.32E-05 | 1 | 0.007813 | 1 | 0.000113 | 1 | 6.18E-05 | 1 | 0.000829 | 1 | 0.000122 | 1 | 0.055664 | 0 | ||||||||
| D 4 | 6.33E-05 | -1 | 0.007813 | 1 | 7.74E-06 | 1 | 7.74E-06 | 1 | 6.26E-05 | 1 | 1 | 0 | 7.74E-06 | 1 | 1.19E-05 | 1 | 1 | 0 | ||||||||
| D 5 | 0.003906 | 1 | 0.039063 | 1 | 0.003906 | 1 | 0.003906 | 1 | 0.021484 | 1 | 0.003906 | 1 | 0.000977 | 1 | 0.003906 | 1 | 0.000977 | 1 | ||||||||
| D 6 | 1 | 0 | 0.25 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 7.74E-06 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | ||||||||
| D 7 | 1.19E-05 | 1 | 5.69E-05 | 1 | 7.4E-05 | 1 | 2.01E-05 | 1 | 5.62E-05 | 1 | 4.26E-05 | 1 | 6.18E-05 | 1 | 1.19E-05 | 1 | 4.39E-05 | 1 | ||||||||
| D 8 | 0.15625 | 0 | 0.001404 | 1 | 0.000156 | 1 | 0.000225 | -1 | 9.95E-05 | -1 | 9.7E-05 | -1 | 0.046647 | -1 | 0.001022 | -1 | 0.000189 | -1 | ||||||||
| D 9 | 0.000363 | 1 | 0.000179 | -1 | 0.000488 | 1 | 7.95E-05 | -1 | 1 | 0 | 0.001651 | 1 | 0.000854 | 1 | 7.12E-05 | -1 | 0.131165 | 0 | ||||||||
| D 10 | 0.000242 | 1 | 0.3125 | 0 | 0.00022 | -1 | 0.00077 | -1 | 0.000145 | 1 | 5.4E-05 | -1 | 0.011719 | 1 | 0.000144 | 1 | 0.000977 | 1 | ||||||||
| D 11 | 7.78E-05 | 1 | 8.27E-05 | 1 | 7.24E-05 | 1 | 6.74E-05 | 1 | 7.59E-05 | 1 | 5.6E-05 | 1 | 6.19E-05 | 1 | 6.36E-05 | 1 | 6.74E-05 | 1 | ||||||||
| D 12 | 0.072266 | 0 | 0.154408 | 0 | 6.36E-05 | 1 | 0.017578 | 1 | 6.59E-05 | 1 | 0.001953 | 1 | 7.93E-05 | 1 | 0.019531 | 1 | 0.000194 | -1 | ||||||||
| D 13 | 2.86E-05 | 1 | 0.25 | 0 | 1 | 0 | 1 | 0 | 2.31E-05 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | ||||||||
| D 14 | 5.3E-05 | 1 | 4.26E-05 | 1 | 1.19E-05 | 1 | 1.31E-05 | -1 | 5.6E-05 | 1 | 1.19E-05 | 1 | 1 | 0 | 1.19E-05 | 1 | 0.00029 | 1 | ||||||||
| D 15 | 3.65E-05 | 1 | 5.05E-05 | 1 | 5.61E-05 | 1 | 6.2E-05 | 1 | 0.000124 | 1 | 7.74E-06 | 1 | 2.3E-05 | 1 | 5.57E-05 | 1 | 5.4E-05 | 1 | ||||||||
| D 16 | 0.125 | 0 | 1 | 0 | 1.19E-05 | -1 | 6.33E-05 | -1 | 3.56E-05 | -1 | 7.74E-06 | 1 | 7.74E-06 | -1 | 7.74E-06 | -1 | 1 | 0 | ||||||||
| D 17 | 0.0002 | -1 | 0.000122 | 1 | 0.002116 | 1 | 0.613281 | 0 | 0.000113 | 1 | 0.000151 | -1 | 0.113281 | 0 | 5.68E-05 | -1 | 0.000101 | -1 | ||||||||
| D 18 | 0.000114 | 1 | 6.1E-05 | 1 | 4.67E-05 | 1 | 5.47E-05 | 1 | 7.11E-05 | 1 | 5.87E-05 | 1 | 5.61E-05 | 1 | 5.3E-05 | 1 | 0.091797 | 0 | ||||||||
| D 19 | 5.06E-05 | 1 | 3.74E-05 | 1 | 2.96E-05 | 1 | 4.37E-05 | 1 | 7.25E-05 | 1 | 4.15E-05 | 1 | 2.93E-05 | 1 | 0.00011 | 1 | 6.12E-05 | 1 | ||||||||
| D 20 | 6.36E-05 | 1 | 5.66E-05 | 1 | 8.02E-05 | -1 | 6.33E-05 | 1 | 7.5E-05 | 1 | 2.93E-05 | 1 | 5.62E-05 | 1 | 0.03125 | 1 | 2.93E-05 | 1 | ||||||||
| D 21 | 1.71E-05 | 1 | 3.68E-05 | 1 | 0.5 | 0 | 2.86E-05 | -1 | 8.14E-05 | 1 | 1.71E-05 | -1 | 2.21E-05 | 1 | 2.21E-05 | 1 | 1.71E-05 | -1 | ||||||||
| D 22 | 6.32E-05 | 1 | 6.6E-05 | 1 | 7.67E-05 | 1 | 0.000111 | 1 | 8.46E-05 | 1 | 7.6E-05 | 1 | 0.000136 | 1 | 2.93E-05 | 1 | 0.000219 | 1 | ||||||||
| D 23 | 8.72E-05 | 1 | 8.66E-05 | 1 | 8.62E-05 | 1 | 8.57E-05 | 1 | 8.77E-05 | 1 | 8.63E-05 | 1 | 8.79E-05 | 1 | 8.68E-05 | 1 | 8.45E-05 | 1 | ||||||||
| D 24 | 0.000167 | 1 | 0.000383 | -1 | 0.000106 | -1 | 0.000488 | 1 | 0.000182 | 1 | 0.000488 | 1 | 0.831055 | 0 | 9.89E-05 | -1 | 0.000102 | -1 | ||||||||
| D 25 | 0.000172 | -1 | 8.26E-05 | -1 | 7.87E-05 | -1 | 8.29E-05 | -1 | 0.000149 | -1 | 8.26E-05 | -1 | 8.2E-05 | -1 | 8.13E-05 | -1 | 0.000173 | -1 | ||||||||
| D 26 | 5.06E-05 | 1 | 4.15E-05 | 1 | 6.71E-05 | 1 | 3.56E-05 | 1 | 8.01E-05 | 1 | 3.56E-05 | 1 | 2.97E-05 | 1 | 6.1E-05 | 1 | 0.0625 | 0 | ||||||||
| Won | 19 | 17 | 18 | 15 | 21 | 19 | 17 | 17 | 11 | |||||||||||||||||
| Loss | 3 | 3 | 5 | 8 | 3 | 5 | 4 | 6 | 7 | |||||||||||||||||
| Equal | 4 | 6 | 3 | 3 | 2 | 2 | 5 | 3 | 8 | |||||||||||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




